Abstract
Shiga toxin–producing Escherichia coli (STEC) are an important cause of bacterial enteric infection. STEC strains cause serious human gastrointestinal disease, which may result in life-threatening complications such as hemolytic uremic syndrome. They have the potential to impact public health due to diagnostic challenges of identifying non-O157 strains in the clinical laboratory. The Wadsworth Center (WC), the public health laboratory of the New York State Department of Health, has isolated and identified non-O157 STEC for decades. A shift from initially available enzyme immunoassay testing to culture-independent diagnostic tests (CIDTs) has increased the uptake of testing at clinical microbiology laboratories. This testing change has resulted in an increased number of specimen submissions to WC. During a 12-year period between 2011 and 2022, WC received 5037 broths and/or stool specimens for STEC confirmation from clinical microbiology laboratories. Of these, 3992 were positive for Shiga toxin genes (stx1 and/or stx2) by real-time PCR. Furthermore, culture methods were utilized to isolate, identify, and characterize 2925 STEC from these primary specimens. Notably, WC observed a >200% increase in the number of STEC specimens received in 2021–2022 compared with 2011–2012 and an 18% increase in the number of non-O157 STEC identified using the same methodologies. During the past decade, the WC testing algorithm has been updated to manage the increase in specimens received, while also navigating the novel COVID-19 pandemic, which took priority over other testing for a period of time. This report summarizes updated methods for confirmation, surveillance, and outbreak detection of STEC and describes findings that may be related to our algorithm updates and the increased use of CIDTs, which is starting to elucidate the true incidence of non-O157 STEC.
Introduction
Shiga toxin–producing Escherichia coli (STEC) infections are a major cause of bacterial foodborne illness (Scallan et al., 2011). Illness presents with a range of symptoms from mild diarrhea to severe bloody diarrhea and hemolytic uremic syndrome (HUS) (Nataro et al., 2007). STEC produce Shiga toxins but can occur as normal commensal flora in certain animals, specifically cattle. STEC infections are typically acquired in humans by the consumption of raw or undercooked beef or from food and water contaminated with feces of cattle or other ruminant animals (Hunt, 2010). However, other animal hosts, including migratory birds, may serve as reservoirs of STEC (Amézquita-López et al., 2018).
Culture isolation of STEC from stool specimens is critical for outbreak investigations and tracing patterns of disease transmission (Hunt, 2010). Historically, standard culture methods for the detection of O157 STEC have not identified non-O157 STEC; therefore, the number of infections has likely been underestimated until recent years, with the addition of Chromagar specific media, enzyme immunoassay (EIA), and culture-independent diagnostic tests (CIDTs) (Nataro et al., 2007).
STEC identification is further complicated by the fact that there are more than 150 serogroups of E. coli known to carry Shiga toxin genes, the most common of which are STEC serogroups O157, O26, O103, O111, O121, O145, and O45 (Ludwig et al., 2020). The Wadsworth Center (WC) has been identifying and characterizing STEC with molecular methods since 2002 and continuing to improve methods overtime expanding detection of the most recent gene variants as of 2008. These include gene variants for Shiga toxins 1, 1c, and 1d and Shiga toxins 2, 2c, 2d, 2e, and 2g. In addition, real-time PCR assays targeted to detect for the most common serogroups have also been employed (Mingle et al., 2012).
In this study, we describe a 12-year period of isolation, identification, and serotyping of STEC in our laboratory utilizing culture-based methods and a panel of laboratory-developed test (LDT) real-time PCR assays. Samples were submitted to us from local laboratories throughout the state from ill patients presenting with diarrhea. We detail the methodology used and the results we obtained by updating our testing algorithm to improve efficacy, optimize recovery, and streamline processes.
Materials and Methods
Stool enrichment and nucleic acid extraction
A portion of the submitted broth or stool was diluted 1:10 with an in-house prepared E. coli broth (120 g of tryptone, 30.6 g of lactose, 24 g of potassium hydrogen phosphate, 9 g of potassium dihydrogen phosphate, 30 g of sodium chloride in 6 L of Millipore H2O; pH 6.9) and incubated for 18–24 h aerobically at 37°C. A 1 mL aliquot of enrichment broth was then centrifuged for 2 min at >13,000 rpm in a microcentrifuge. The supernatant was discarded, and the pellet was resuspended in 1 mL of 10 × PCR buffer (Alpha Teknova, Inc.) and diluted to 1 × with molecular grade H2O. These steps were repeated for a total of two washes, and the sample was then heat lysed at 95°C for 20 min. Isolates submitted to WC were plated directly onto Sorbitol MacConkey Agar (SMAC) and incubated for 18–24 h aerobically at 37°C. A single colony isolate was placed in 300 μL 1 × PCR buffer and heat lysed at 95°C for 20 min.
Real-time PCR
Shiga toxin genes (stx1 and stx2) and the E. coli O157-specific rfb gene were detected using an in-house developed LDT (New York State [NYS] Clinical Laboratory Evaluation Program approved) real-time PCR assay (StxO157PCR). All three targets were detected in a multiplex assay for enrichment broths that also include an inhibition control. To detect the most current gene variants, these were redesigned in 2008 (Mingle et al., 2012). A separate assay just targeting the stx1 and stx2 genes (StxPCR) was used for single isolate and colony pool testing, in which individual colonies were combined into a single tube to create pools and screened. Primer and probe sequences are listed in Table 1. These were individually diluted to 45 μM and then combined respectively, if necessary.
Primer and Probe Sequences of Real-Time PCR Assays
Integrated DNA Technologies, Coralville, IA, USA.
Applied Biosystems, Foster City, CA, USA.
Plasmid containing the same primer binding sites as stx2.
Between 2011 and 2020, enrichment broths for real-time PCRs were prepared and run as previously described (Mingle et al., 2012). However, in 2021, samples were spiked with an inhibition control directly, and the assay was validated with 1 × PerfeCTa qPCR ToughMix (Quanta Biosciences, Gaithersburg, MD). A 25 μL final volume was used, and each reaction contained 450 nM of O157 rfb primers, 900 nM of each Stx1 and Stx2 primer or primer mix, 250 nM of each probe or probe mix, 4 μL of sterile H2O, and 5 μL of DNA template. Standard thermocycling conditions were as follows: 1 cycle at 95°C for 3 min, 40 cycles at 95°C for 15 s, and 60°C for 45 s. Thermocycling, fluorescence data collection, and data analysis were performed with the ABI 7500 Fast Real-Time PCR system according to the manufacturer's instructions, with the passive reference dye, turned off. This change has helped streamline our testing by allowing multiple assays to be run and analyzed on the same optical plate under the same cycling conditions, which are shorter in duration.
In addition, our laboratory has a two-step multiplex real-time PCR assay for detection of the top six non-O157 STEC serogroups isolated from humans. Surveillance data since 2003 have shown that STEC serogroups O26, O103, O111, O45, O121, and O145 are consistently the most detected serogroups other than O157 (Beutin et al., 2009; Ludwig et al., 2020). The assay consists of a three-plex real-time PCR targeting the wzx gene of STEC O26 and STEC O103 and the wbd1 gene of STEC O111, followed by a three-plex real-time PCR targeting the wzy gene of STEC O45, STEC O121, and STEC O145 (Mingle et al., 2012). This assay is currently performed on all positive screened broths that are O157 negative and any O157-negative Shiga toxin–positive isolates. PCRs were prepared using 1 × PerfeCTa Multiplex qPCR Supermix (Quanta Biosciences). Each reaction contained 900 nM of each primer, 250 nM of each probe, 3.75 μL of sterile H2O, and 5 μL of DNA template. Primer and probe sequences used were described previously (Mingle et al., 2012).
Pulsed-field gel electrophoresis and next-generation sequencing isolate characterization
Until March 2019, pulsed-field gel electrophoresis was performed on all confirmed STEC isolates. These methods were outlined by the PulseNet national surveillance system (Ribot et al., 2006). Tiff images were analyzed and uploaded to the PulseNet national database using BioNumerics (Applied Maths, Sint-Martens-Latem, Belgium). Beginning in March 2019, a next-generation sequencing method (NGS) method, whole-genome sequencing (WGS), and bioinformatic analysis were performed on every STEC isolate using methods previously described (Baker et al., 2023). Sequence files were analyzed and uploaded to both the PulseNet national database and the National Center for Biotechnology Information Sequence Read Archive using BioNumerics.
Algorithm for testing Shiga toxin EIA- and CIDT-positive specimens
Our current laboratory algorithm for Shiga toxin confirmation and identification of EIA- or CIDT-positive stools and/or broths from submitting clinical laboratories is shown in Figure 1. Before updating our testing methods, stools and/or broths were plated directly onto SMAC culture media along with enrichment in E. coli broth on each day of receipt. Individual colonies were initially tested alongside the stool enrichment broth by real-time PCR. However, we determined that only a small percentage of our enrichment broths with a Ct (cycle threshold) >30 resulted in recovery of the STEC organism (Table 2).
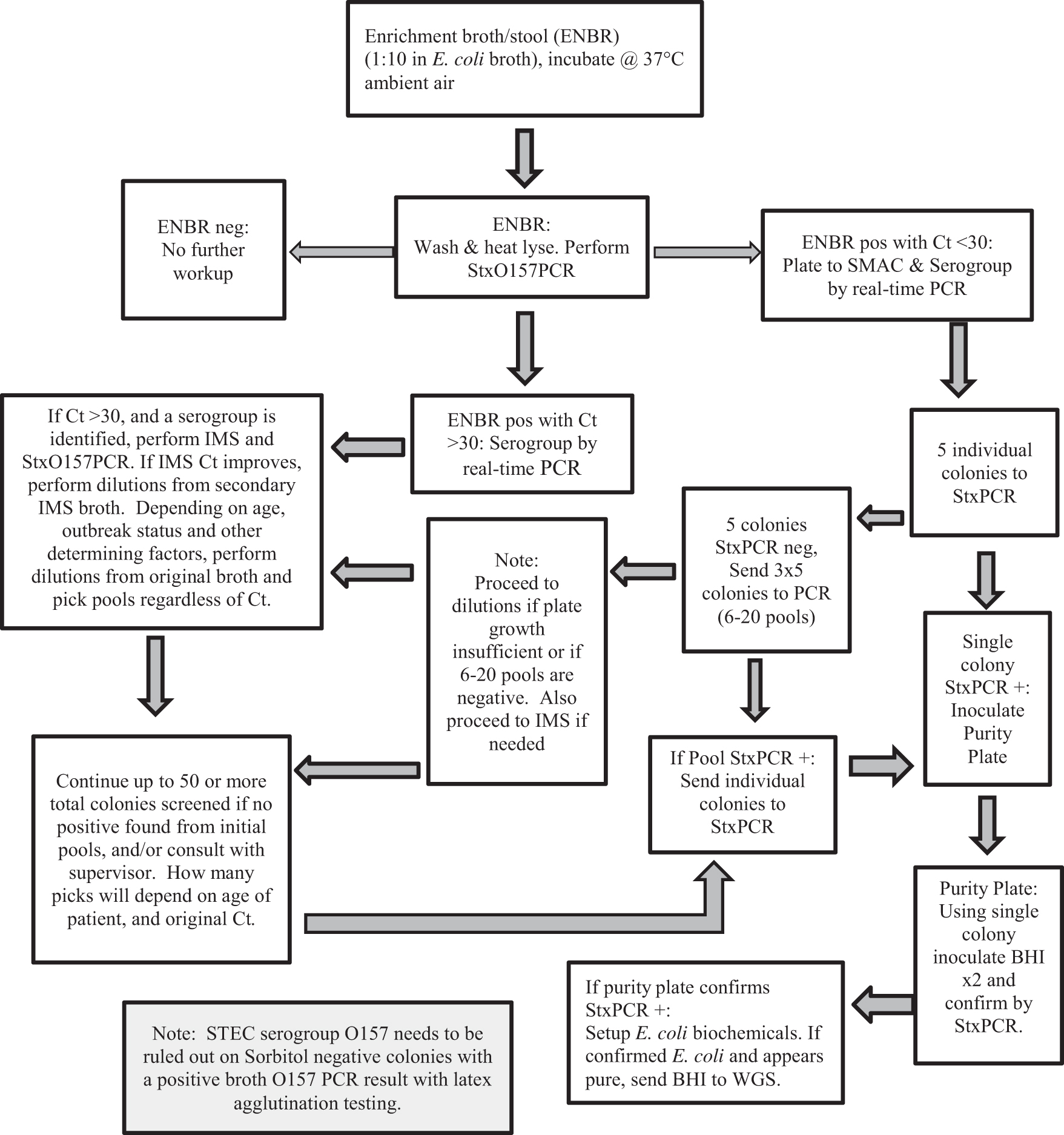
Wadsworth Center algorithm for isolating STEC from EIA/CIDT-positive enrichment broths and stools. BHI, brain–heart infusion agar; CIDT, culture-independent diagnostic test; Ct, cycle threshold; EIA, enzyme immunoassay; ENBR, enrichment broth; SMAC, Sorbitol MacConkey Agar; StxO157PCR, Shiga toxin and Escherichia coli O157 real-time PCR; StxPCR, Shiga toxin real-time PCR; IMS, immunomagnetic bead separation; STEC, Shiga toxin–producing Escherichia coli; WGS, whole-genome sequencing.
Number of Shiga Toxin–Producing Escherichia coli Identifications by Successful Isolation from Culture Colony Subculture Compared Across Cycle Threshold Values, 2017–2020
In 2020, we implemented an initial molecular screen of the enrichment broths to detect the presence of stx1 and stx2 and O157 DNA by real-time PCR. If real-time PCR was negative for stx1/stx2 genes, regardless of the presence of O157 DNA, no further workup was performed. If the Ct was >30 and no serogroup was determined, no further workup was performed unless the patient was <20 years of age. A dilution series was performed from the original broth, and pools of colonies were tested by StxPCR. If the Ct was >30, and the enrichment broth was positive for O157 DNA, or one of the six most prevalent non-O157 STEC serogroups, we would proceed to immunomagnetic bead separation (IMS) of the enrichment broth for that serogroup.
The magnetic beads are coated with an antibody to a particular serogroup. When combined with the enrichment broth, these bind to specific antigens from the target serogroup. Organisms are attached through antibody/antigen reactions. The beads were then removed from the broth using a magnet and wash method and were cultured to allow for detection and isolation of the suspect organism. Real-time PCR was run to verify the improved Ct value score. This indicated that the IMS bead procedure had worked properly. If the Ct did not improve with IMS, no further workup was performed. The exception to this was if the patient was <20 years of age. If the Ct of the secondary IMS broth was stronger, dilutions were performed from the IMS broth and pools of colonies were tested by StxPCR.
If the Ct was <30, a portion of the primary enrichment broth dilution was directly plated onto an SMAC plate, and 5 individual colonies or up to 50–100 individual colonies in pools were screened from growth on this plate by StxPCR. A Shiga toxin–positive colony was then plated to a new SMAC plate to confirm purity and Shiga toxin presence before initiating WGS. The Shiga toxin–positive colony selected was identified biochemically as E. coli using triple sugar iron, urea, indole, citrate, and sorbitol fermentation. If the Shiga toxin–positive isolate was sorbitol negative, and O157 PCR positive on the initial primary enrichment broth screen, O157 antigen testing was performed using the O157 latex agglutination assay (Oxoid, Basingstoke, Hants, UK). If the Shiga toxin–positive isolate was sorbitol positive or sorbitol negative, and O157 negative on the initial enrichment broth screen, real-time PCR for the top six non-O157 serogroups was performed on the isolate.
Results
From 2011 to 2022, 5037 primary stool and/or broth specimens were received and tested by WC (Table 3). The number of specimens received has steadily increased each year. The exception to this was a slight decrease from 2016 to 2017 and a significant decrease from 2019 to 2020, most likely due to the COVID-19 pandemic. STEC was isolated from 58% (2925/5037) of specimens. Culture-negative, Shiga toxin–positive specimens represented 21% (1067/5037) of the total specimens received and tested by real-time PCR. Furthermore, 21% (1045/5037) of specimens were determined to be STEC negative by StxO157PCR.
Summary of EIA/CIDT-Positive Broth and Stool Testing, 2011–2022
STEC are Shiga toxin–producing Escherichia coli isolates for which a serogroup was determined to be serogroup undetermined, or results were not available from Wadsworth Center or Centers for Disease Control and Prevention at the time of our report.
STEC serogroup identifications listed from 4 to 1 over the 12-year period: O109, O166, O119, O21, O22, O23, O66, O153, O136, O158, O28/O42:H25, O17/O44/O77, O93, O178, O110, O179, O61:H16, O88:H25, O126, O182, O1, O38, O123/O186:H11, O100:H20, O100:H30, O105:H34, O108, O54, O78, O96, O143, O43, O183, O185, O32, O168:H8, O13/O135, O100:/O154:H25, O163, O106, O65:H2, O68, O123, O124, O132:H27, O172, O6, O7, O15:H27, O24, O29:H45, O39, O46, O50, O52:H10, O87:H2, O142:H16, O148, O176:H4, O188, O3, O27:H30, O32/O8:H21, O104, O123/O186:H10, O153/O178:H7, O10:H19, O171:H2.
StxPCR positive only are broths/stools that were positive for stx1 or stx2 by real-time PCR, but for which no Shiga toxin–positive isolate was obtained.
Negative are broths/stools that could not be confirmed by real-time PCR or culture as STEC. Serogroup O157 determined by latex agglutination; serogroups O103, O26, O111, O45, O145, O121 determined by real-time PCR. Remaining STEC serogroups identified by WGS and BioNumerics analysis.
STEC, Shiga toxin–producing Escherichia coli; WGS, whole-genome sequencing.
The most frequently detected STEC serogroup from primary specimens was O103, which represented 20% (595/2925) of the specimens with STEC isolation. The different STEC serogroups isolated and identified from primary stool and/or broth specimens from 2011 to 2022 are shown in Table 3. STEC serogroups O103, O26, O111, O121, O45, and O145 remained the top identified, along with O157, which included O157:H7. It is notable that compared with the previous 6-year study, some of the percentages of STEC isolation and identifications have markedly changed. In the 2005–2010 period, the number of STEC serogroups >1% of the total was in order O157 (27.7%), O103 (16.4%), O26 (13.4%), O111 (9.5%), O45 (8.0%), O121 (3.4%), O145 (2.2%), and O165 (1.9%).
In the 2011–2022 period, the number of STEC serogroups >1% of the total was in order O103 (20.3%), O157 (14.0%), O26 (12.4%), O111 (10.4%), O121 (4.5%), O45 (3.7%), O145 (2.2%), O91 (1.6%), O118/O151:H2 (1.5%), and O8 (1.4%). Rare STEC serogroup identification (<5 times over the 12-year period) comprised 4.8% (140/2925) of the STEC isolated. The number of STEC serogroups for which a serogroup was undetermined, or had no serogroup identified by the WC or Centers for Disease Control and Prevention (CDC), was 8.1% (237/2925) for the 2011–2022 period.
Table 4 displays the total number of STEC that were identified over the 12-year time frame. A total of 3559 isolates were Shiga toxin–positive, either originally submitted as a primary stool/broth specimen or as a culture isolate. Overall, only 27% (965/3559) of these were O157 STEC, whereas 73% were non-O157 STEC. Of the 634 STEC-positive isolates that were originally submitted as culture isolates to our laboratory, 555 (88%) were O157 positive. The other 79 samples (12%) were identified as non-O157 E. coli. From primary specimens, only 14% (410/2925) were isolated and identified as O157 STEC, whereas 86% (2515/2925) were non-O157 STEC (Table 4).
Summary of Shiga Toxin–Producing Escherichia coli Identified and Characterized by the Wadsworth Center, 2011–2022
Discussion
STEC causes an estimated 265,000 illnesses each year in the United States (Bryan et al., 2015). The increased sensitivity of testing methods used in the clinical laboratory combined with confirmatory methods at WC has contributed not only to an increase in the number of specimens we receive but also to an increase in STEC-confirmed positive cases and non-O157 serogroup identifications. Clinical laboratories use a combination of culture, immunoassays for STEC-specific antigens, and molecular testing to detect STEC. CIDTs can detect stx1 and/or stx2 genes; however, for epidemiological and outbreak purposes, culture isolation and identification are necessary (Bryan et al., 2015).
CIDTs in the clinical laboratory are increasing in use; however, foodborne illness surveillance, specifically for STEC, has traditionally relied on confirmatory culture methodology to identify outbreaks through molecular subtyping and analysis (Imdad et al., 2018). Although some direct NGS methods have been described (Singh et al., 2019), having access to STEC isolates has been critical for public health surveillance. WC has continued to yield fully characterized STEC isolates to monitor trends and support foodborne investigations and outbreaks (Iwamoto et al., 2015).
It is important to note that during the 12-year time frame, a higher occurrence of non-O157 STEC (86% [2515/2925]) compared with O157 STEC (14% [410/2925]) was observed in the 2925 STEC confirmed positive stool/broth submissions (Table 3). Interestingly, during the previous 6-year period (Mingle et al., 2012), non-O157 was identified in 68% of the primary stool/broth submissions, whereas O157 STEC was only identified in 32%. This change represents an 18% increase in non-O157 STEC identifications and likely correlates with the additional use of CIDTs and serves to support the concern that non-O157 STEC have been and continue to be underreported (Kuehne et al., 2016). Increasing trends of non-O157 STEC in other regions and countries have also been observed, specifically in a study from Cairo between 2018 and 2019 in which the percentage of O157 STEC to non-O157 was 32% to 68%, respectively (Elshimy et al., 2023). Also of interest is the possible geographic factors associated with a higher risk of HUS associated with STEC, specifically serogroup O26 in France (Jones et al., 2023).
As indicated in Table 4, 58% of the O157 STEC identified were originally submitted as pure culture isolates, and 42% as stool/broth specimens, suggesting that the submitting laboratories are identifying O157 isolates independently and reducing the number of O157 STEC identified at WC. This finding is also supported by the change seen in the 2005–2010 testing period where 543 of 600 (90%) E. coli culture isolates submitted to the WC were identified as O157 STEC (Mingle et al., 2012).
Shiga toxin–positive specimens that were culture negative comprised 21% of our total specimens received. This may indicate a higher sensitivity of molecular methods compared with culture isolation. This could also suggest that Shiga toxin–positive organisms in the samples were not viable. Also, consistent with previous findings (Mingle et al., 2012) is the number of STEC for which a serogroup was undetermined or had no serogroup identified by the WC or CDC 8.6% (51/591) for the 2005–2010 period and 8.1% (237/2925) for the 2011–2022 period.
Of interest is the growing number of non-Shiga toxin–producing samples that were O157 positive, which accounted for a portion of our negative samples. In New York, clinical laboratories are required to submit isolates to the WC with a positive O157 finding; however, without a full virulence profile, the pathogenicity of these organisms is unknown. Additional evaluation will be needed to assess if this testing should continue and if these Shiga toxin–negative O157 strains are relevant to public health.
Over the years, WC has been confirming and serotyping STEC with several different algorithms that have been updated and adjusted to improve testing. We have found that a few factors determine isolate yield on a specimen. The most critical determinate is the Ct value of the enrichment broth that we test by real-time PCR. We have found that isolate recovery is inversely related to the Ct value, that is, the lower the Ct, the greater amount of organism DNA present. The success of culture isolation decreases with Cts >25 and is further reduced with Cts >30 (Table 2). In the future, we plan to take a closer look into the media used for specimen submissions to evaluate any impact on organism viability and isolation.
Overall, isolation and characterization of STEC is tedious and labor intensive, but important for public health. We have determined that using a Ct value cutoff helps predict successful isolation of STEC. The use of IMS beads can also improve the potential for isolation of STEC. A patient's age (<20 years) and clinical presentation is also considered as STEC associated-HUS accounts for 60% of case patients in children younger than 16 years (Bruyand et al., 2018), which can result in acute kidney injury, thrombocytopenia, and microangiopathic hemolytic anemia. We put additional time and effort into the isolation from these specimens because of the impact the STEC identification can have for the clinical case and determination of the source for an investigation. Approximately 10–15% of children will develop HUS following STEC infection (Walsh and Johnson, 2018).
In summary, utilization of our updated algorithm will continue to allow for the most effective process for isolating STEC in NYS. In addition, clinical laboratory practices are key for disease surveillance and patient care (Hoefer et al., 2011). With more sensitive CIDTs being utilized in recent years, an increase in specimen submissions and a marked increase in the percentage of non-O157 STEC have been observed, which has positive implications for public health. Our 12-year period of identification and serotyping of STEC utilizing culture-based methods and a panel of LDT real-time PCR assays has provided additional data to better understand STEC disease and may lead to effective and improved investigations and monitoring of trends in STEC disease epidemiology.
Footnotes
Acknowledgments
The authors would like to thank NYS Epidemiology, and Emerging Infections Program (EIP) and Epidemiology and Laboratory Capacity (ELC) staff members for their long-term support of the project. The authors also thank the Tissue Culture and Media Core, the Wadsworth Center Bacteriology Laboratory, as well as the Wadsworth Center Applied Genomic Technologies Core for performing WGS. Finally, the authors would also like to thank Marie-Claire Rowlinson, Nellie Dumas, Timothy Root, Tammy Quinlan, Ron Limberger, and many past and present members of the Wadsworth Center Bacteriology Laboratory for support of this project.
Authors' Contributions
K.A.M. and L.A.M.: Conceptualization. A.E.C., D.W., S.E.W, L.M.T., A.L.S., J.A.C., D.J.B., M.C.D., C.E.M., C.V., and D.T.W.: Methodology, data curation. A.E.C.: Writing—original draft preparation. A.E.C., D.W., S.E.W., and D.J.B.: Visualization, investigation. D.W. and L.A.M.: Supervision. A.E.C., S.E.W., K.A.M., and L.A.M.: Writing—reviewing and editing.
Disclosure Statement
No competing financial interests exist.
Funding Information
This work was in part supported by the CDC, National Center for Infectious Diseases, ELC for Infectious Disease Cooperative Agreement (grant NU50CK000423), the CDC EIP Cooperative Agreement (grant U50/CCU511190).
