Abstract
Poultry has been suggested as an important source for extended-spectrum beta-lactamase (ESBL)-producing bacteria that can lead to difficult-to treat infections in humans. Therefore, this study aims to determine the frequency, the genetics, and antimicrobial resistance profiles of ESBL-producing Escherichia coli in domestic free-range poultry in Agogo, Ghana. The study was set up and piloted from January 2019 until June 2019. Between June and December 2019, fecal samples (N = 144) were collected from free-roaming chickens from domestic farms in the regions of Sukuumu, Bontodiase, and Freetown and cultured on ESBL screening agar. Strain identification and antibiotic susceptibility were performed using the VITEK 2 compact system. ESBL-producing E. coli were confirmed using the double disk synergy test. Molecular characterization of ESBL-associated genes (bla TEM, bla SHV, and bla CTX-M) were performed using conventional polymerase chain reaction (PCR) and further sequencing of obtained PCR amplicons. The result showed that 56.2% (n/N = 81/144) of collected fecal samples were positive for ESBL-producing E. coli. Majority of the isolates showed resistance to tetracycline (93.8%, n/N = 76/81) and trimethoprim-sulfamethoxazole (66.7, n/N = 54/81), whereas resistance to carbapenems was not found. The majority of ESBL-producing E. coli carried the bla CTX-M genes, with bla CTX-M-15 being the dominant (95.1%, n/N = 77/81) genotype. In this study, we report high frequencies of ESBL-producing E. coli in smallholder free-range poultry representing a potential source of infection, highlighting the need for control of antibiotic use and animal hygiene/sanitation measures, both important from a One Health perspective.
Introduction
The rapid increase of resistance to commonly used antibiotics in human medicine and animal husbandry has become one of the leading global health concerns (Rousham et al, 2018). Antimicrobial resistance (AMR)-related infections have become the leading cause of deaths worldwide with 1.2 million annually (Murray et al, 2022). Without preventive measures in place, by 2050, the annual deaths due to infections with AMR bacteria are expected to reach 10 million globally (O'Neill, 2014).
In sub-Saharan Africa, AMR infections, including infections caused by extended-spectrum beta-lactamase (ESBL)-producing Escherichia coli, are of major concern. ESBL enzymes can hydrolyze beta-lactam antibiotics, including third-generation cephalosporins (Pitout and Laupland, 2008), which are among the most commonly prescribed antibiotics around the world. Among the several variants of ESBL genes, the most frequent and clinically relevant genes are bla TEM, bla SHV, and bla CTX-M, with bla CTX-M being the most dominant worldwide (Castanheira et al, 2021).
ESBL-producing bacteria are often linked to concomitant resistance to other classes of antibiotic agents leading to multidrug resistance (MDR) (WHO, 2020). Therefore, there are global ongoing efforts to address this issue and the issue of AMR as a whole (Mendelson and Matsoso, 2015; O'Neill, 2016). These pathogens can be acquired by direct contact with infected animals or by consumption of contaminated meat products. In particular, the consumption of poultry meat has proven to be a major source of ESBL-producing bacteria according to a global study (Lazarus et al, 2015). Moreover, a recent study from Ghana indicated poultry as a potential source for ESBL-producing bacteria (Falgenhauer et al, 2019), which can lead to difficult-to-treat infections in humans (Haruki et al, 2018; Lee et al, 2018).
Despite that, poultry is the most popular among all the meats and is largely consumed in all regions of Ghana. The majority (79%) of these poultry meats are imported from other countries as the domestic producers are failing to sustain the high production costs (Zamani et al, 2021). In the quest to meet this demand and to compete with imported birds, poultry keepers employ antimicrobials as growth promoter, and to prevent and treat infections (Paintsil et al, 2021; Prestinaci et al, 2015). Subsequently, this contributes to the emergence and spread of AMR bacteria in the farm animals (Nhung et al, 2017).
Although unable to meet the high demands due to economic reasons, poultry farming in Ghana is still one of the predominant animal businesses with an estimated number of 74.5 million birds produced per year (NEA, 2019). The majority of which are reared intensively on commercial farms, whereas the remaining birds are kept on smallholder farm level, freely roaming within communities. Extensive use of antimicrobials has been reported in both these types of farms in Ghana (Paintsil et al, 2021), and it has resulted in the emergence of ESBL-producing bacteria, particularly E. coli, in commercial poultry farms (Aworh et al, 2020; Eibach et al, 2018; Falgenhauer et al, 2019). However, to our knowledge, such bacteriological studies have not been conducted on free-range chickens reared on smallholder farms in Ghana. Nevertheless, due to the animals' proximity to humans and their significance in the food chain, such studies are essential to identify possible reservoirs of ESBL-carrying pathogens, which can be a source of transmission within communities.
Therefore, this study aimed at determining the frequency of ESBL-producing E. coli, their genetic characteristics, and their antibiotic-resistant profiles among smallholder farm free-range chicken in Agogo, a town located in the Ashanti region of Ghana.
Methods
Study site and sample collection
Fecal samples from chickens were collected from six smallholder farms in three different communities in Agogo in the Asante Akyem North municipality within the Ashanti region of Ghana (Fig. 1). Farm selection was based on agreement of farmers to take part in this study (nonprobability sampling). Sampling included backyard poultry farms and was subject to the willingness of the owners/farmers to participate in the study. Sample collection took place between June and December 2019. Each freshly voided fecal sample was collected from a single chicken.
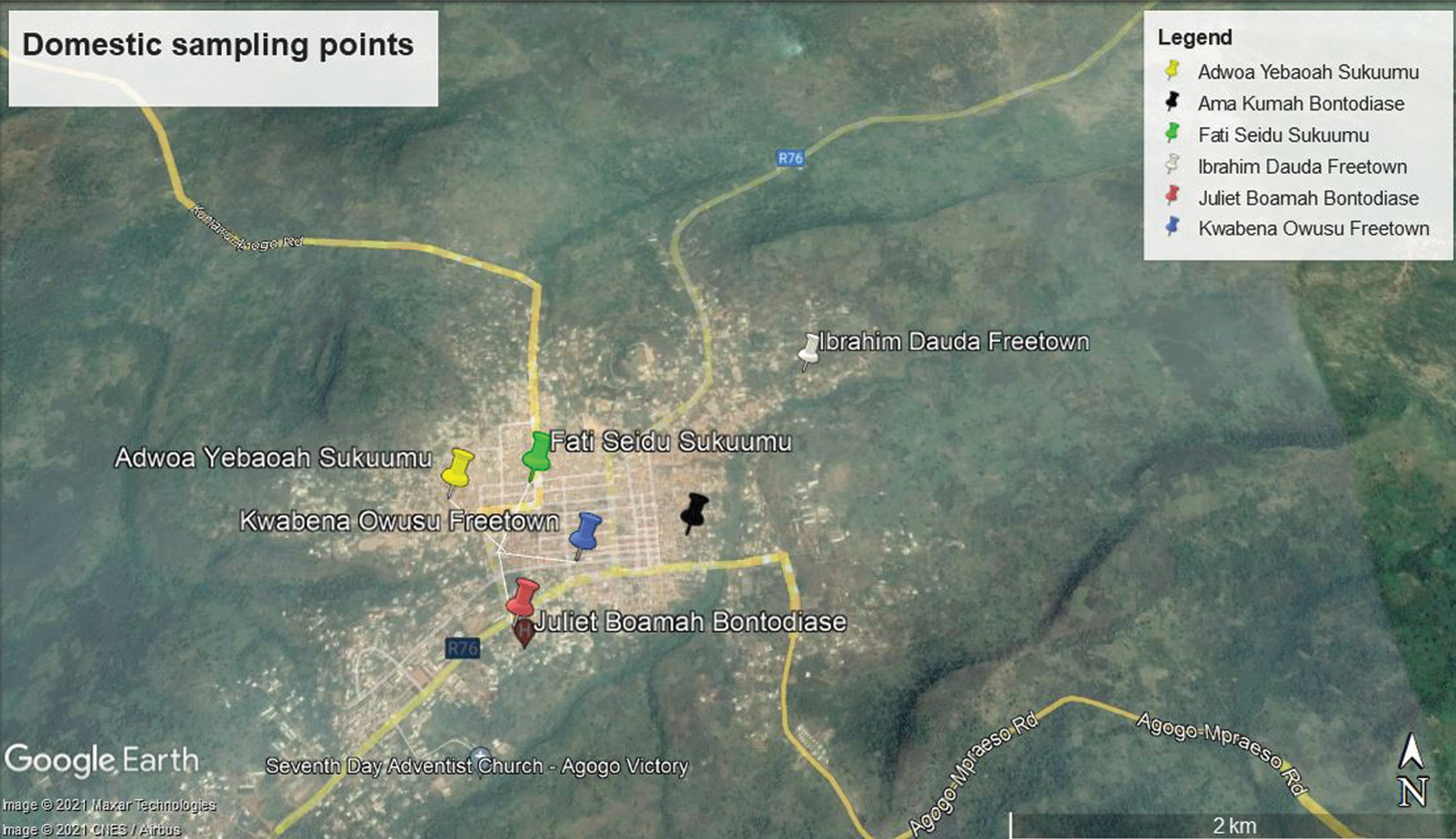
Geographical location of domestic (household) farms in Agogo, Ghana.
They were collected from smallholder farms where chickens were kept in the backyard. Each smallholder farm possessed ∼25 chickens and was visited once during the study period. About 1 g of the fresh stool per individual dropping was collected using a sterile spoon and placed in a dry container. Only fecal samples that had minimal contact with the surface of the ground were taken. Samples were transported at 2–8°C in a cool box to the microbiology laboratory at the Kumasi Centre for Collaborative Research in Tropical Medicine (KCCR) within 2–4 h after sampling for further analyses.
Species identification and antibiotic susceptibility testing
On arrival at the laboratory, samples were cultured on two MacConkey agar plates supplemented with 1 mg/L ceftazidime and 1 mg/L cefotaxime, respectively. Plates were incubated at 35–37°C for 18–24 h in normal atmosphere. All morphological different lactose fermenting colonies (not exceeding three colonies) were subsequently subcultured on Columbia blood agar (Oxoid). Isolate identification and antibiotic susceptibility were done using the VITEK 2 compact system (bioMérieux, Marcy L'Etoile, France).
Confirmed E. coli were classified as S (Susceptible, standard dosing regimen), I (Susceptible, increased exposure), or R (Resistant) to commonly used antibiotics according to the European Committee on Antimicrobial Susceptibility Testing (EUCAST) guidelines (EUCAST guidelines, version 10.0, 2020;
Production of ESBLs was confirmed by the combined double-disk test with ceftazidime and cefotaxime alone, and in combination with clavulanic acid (Becton, Dickinson Company, Sparks, MD) as defined by EUCAST (EUCAST guidelines, version 10.0, 2020). Strains confirmed as ESBL-producing E. coli were saved in microbanks (Pro-lab Diagnostics, Richmond Hill, ON, Canada) and stored at −80°C for further analysis. Quality control was performed using the E. coli ATCC 25922 and a bla CTX-M-15 positive E. coli. Results of the quality control strains were within acceptable ranges for controls throughout this study.
Genotyping of ESBL genes by polymerase chain reaction
DNA extraction for E. coli was performed using the boiling method as described by Tellevik et al (2016). DNA was subjected to molecular characterization by polymerase chain reaction (PCR) for the detection of the resistance genes bla
TEM (temoneira), bla
SHV (sulfhydryl variable enzyme), and bla
CTX-M (cefotaximase-Munich), using known primers as described elsewhere (Campos et al, 2014). To further differentiate the bla
CTX-M genes, specific target primers (Table 1) were used for PCR amplification and sequencing of these amplicons (Campos et al, 2014). The PCR amplicons were sent to Microsynth Seqlab (Göttingen, Germany) for Sanger sequencing. The resulting sequences were aligned and mutations were identified by comparison with known sequences using CLC Sequence Viewer 8.0 (
Primers for Extended-Spectrum Beta-Lactamase Genes
Statistical analysis
Descriptive statistics were applied and continuous categorical variations were shown as frequencies with percentages. All analyses were done using R language for statistical computing, version 4.0.2.
Results
In total, 144 fecal samples were collected from 6 smallholder farms in 3 communities in Agogo (Fig. 1) and bacteriologically investigated. Single ESBL E. coli was isolated from each sample. Of these 144 isolates, 81 (56.3%, n/N = 81/144) were phenotypically positive for ESBL-producing E. coli. Frequencies detected within each of the communities were 76.1% (n/N = 35/46) in Sukuumu, 52.9% (n/N = 37/70) in Freetown, and 32.1% (n/N = 9/28) in Bontodiase (Table 2).
Distribution of Extended-Spectrum Beta-Lactamase Isolates and Its Genotype Across the Sampling Sites
Out of total isolates from Sukuumu.
Out of total isolates from Freetown.
Out of total isolates from Bontodiase.
Out of total isolates tested in this study (N = 144).
ESBL, extended-spectrum beta-lactamase.
Genotype identification of ESBL-producing E. coli
Genotype characterization of the investigated ESBL-producing E. coli isolates identified two different beta-lactamase-encoding genes (bla CTX-M and bla TEM) out of the three genes tested (bla CTX-M, bla SHV, and bla TEM) (Table 2). The majority of isolates were positive for bla CTX-M-15 (95.1%, n/N = 77/81), whereas only a minority carried bla TEM-1b (2.5%, n/N = 2/81) and bla CTX-M-15/bla TEM-1b (1.2%, n/N = 1/81). None of the ESBL E. coli isolates carried the bla SHV gene. However, in one of the phenotypically confirmed ESBL E. coli isolates, none of the tested genes (bla CTX-M, bla SHV, and bla TEM) were identified.
In terms of distribution of the ESBL isolates and its genotype across the sample sites, higher percentage of ESBL-positive isolates were observed in Sukuumu (76.1%), followed by Freetown (52.9%) and the least in Bontodiase (9%). Similarly, the majority of bla CTX-M-15 positive isolates was observed in Sukuumu (74%) followed by Freetown (35%) and Bontodiase (28.5%). Whereas only one bla TEM-1b positive isolate was observed in Sukuumu (2.2%) and Bontodiase (3.6%) but not in Freetown. One isolate of ESBL E. coli carrying both bla CTX-M and bla TEM genes and an isolate with an unidentified gene were observed only in Freetown.
Antibiotic susceptibility of ESBL-producing E. coli
All E. coli isolates of this study were sensitive to meropenem, imipenem, ertapenem, and tigecycline (100%, 81/81) (Fig. 2). Apart from 100% resistance to cephalosporins shown by the E. coli isolates, the highest resistance was observed for tetracycline (93.8%, n/N = 76/81), followed by trimethoprim-sulfamethoxazole (66.7%, n/N = 54/81), and ciprofloxacin (35.8%, n/N = 29/81) as shown in Figure 2. Among these strains, 30.8% (n/N = 25/81) were MDR with 19.75% (n/N = 16/81) being resistant to three (i.e., fluoroquinolone, tetracycline, and sulfonamides) and 11.1% (n/N = 9/81) to four classes of antibiotics (i.e., fluoroquinolone, tetracycline, aminoglycoside, and sulfonamides). No difference in resistance profile was observed within the farms. Antibiotic usage on these farms was not assessed. The result of the E. coli ATCC 25922 and a bla CTX-M-15 positive E. coli used as internal quality control fell within the acceptable quality ranges defined by EUCAST.
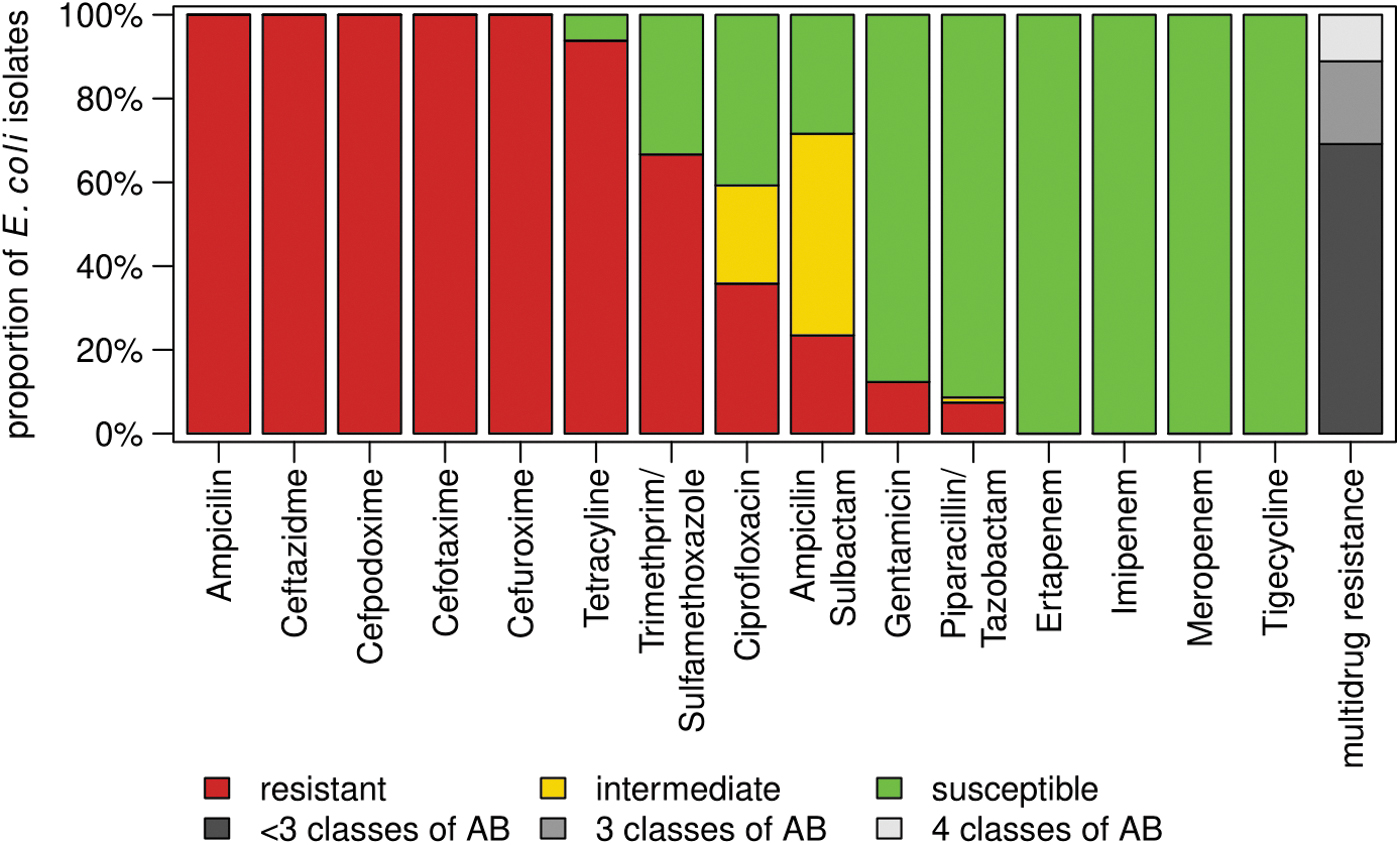
Antibiotic resistance among ESBL-producing Escherichia coli isolated from smallholder farm poultry. AB, antibiotics; ESBL, extended-spectrum beta-lactamase.
Discussion
Free-range smallholder farm chicken plays a significant role in enhancing food security in rural Africa (Melesse, 2014). However, the unregulated use of antibiotics in these farms (Paintsil et al, 2021) may result in the spread of drug-resistant bacteria, particularly ESBL-producing E. coli, in the community (Carattoli, 2008). Moreover, several studies have highlighted the role of ESBL-producing E. coli in severe human infections (Haruki et al, 2018; Melzer and Petersen, 2007).
The considerably high rate of ESBL-producing E. coli in this study (Table 2) exceeds what was found in commercial poultry farming in Ghana (Falgenhauer et al, 2019). This is not surprising, as Paintsil et al (2021) demonstrated that 43% of smallholder farmers from the same study area used antibiotics in poultry farming, which in turn increases the risk of higher ESBL acquisition in the normal gut flora of the animal (Carattoli, 2008). In general, not exclusively at smallholder farm level, frequencies of ESBL-producing bacteria found in chickens seem high on the African and Asian continents.
For example, a review by Alonso et al (2017) has reported 42.0% and 55.5% ESBL-producing E. coli among chicken feces from Tunisia and Algeria, respectively, as well as 61.6% from chicken meat in Egypt. In Asia, a recent study reported ESBL-producing E. coli among chicken (broilers) from backyard farming in India (11.7%) (Chowdhury et al, 2022) and 80% among poultry in subsistent farming in Nepal (Subramanya et al, 2021). The high prevalence of ESBL-producing E. coli is alarming and should be monitored closely, especially in developing countries to control foodborne diseases in humans.
Generally, in resource-limited countries, inadequate sanitation aside from the overuse of antibiotics is likely to play significant roles in the selection and transmission of antibiotic resistance (Aslam et al, 2018). This is particularly true for areas where free-range chickens (in smallholder farms) are raised in communities with close contact to their owners. In sub-Saharan African countries, the potential transmission risk of antimicrobial-resistant bacteria from animals or animal products to humans has been demonstrated before (Falgenhauer et al, 2019).
The resistance profile of E. coli isolates shown in this study (Fig. 2), that is, high level of resistance to beta-lactam antibiotics, the degree of resistance seen for other classes of antibiotics such as tetracycline and trimethoprim-sulfamethoxazole as well as the much lower levels of resistance seen for gentamicin and piperacillin is in line with what was found in other studies around the globe (Bengtsson-Palme et al, 2018; Braykov et al, 2016; Jaja et al, 2020). Drug resistance seen in this study is possibly a reflection on the overall usage and availability of antibiotics in Ghana (Nkansa et al, 2020; Turkson, 2008).
Fluoroquinolones, such as ciprofloxacin, are important drugs in Ghana for the treatment of infections in humans (D'Arcy et al, 2021; Yevutsey et al, 2017). The level of ciprofloxacin resistance (35.8%, n/N = 29/81) (Fig. 2) seen in our sample is similar to that reported in a study on poultry from Nigeria (Aworh et al, 2020) but lower than what was reported in an earlier study in Ghana on poultry meat, where 59% of the strains were resistant to ciprofloxacin (Eibach et al, 2018). The difference in resistance seen might be attributed to the fact that ciprofloxacin usage in smallholder farms is less common in this study area (Paintsil et al, 2021).
Among the most frequent and clinically relevant ESBL genes, bla CTX-M positive E. coli are the most common genotype found in clinical isolates from humans and animals (Alonso et al, 2017; Castanheira et al, 2021). This correlates to our study as the bla CTX-M genes were found in 95% of the isolates. However, some studies suggest otherwise, for example, a study in Nigeria reported bla SHV as the predominant gene followed by bla TEM and bla CTX-M (Mohammed et al, 2016). Another study in Bangladesh reported bla TEM to be predominant among the three genes (Yesmin et al, 2014). These findings suggest that the predominance of these genes is subjected to regional variations.
Among the bla CTX-M gene, bla CTX-M-15 is the most prevalent gene associated with human infections (Irrgang et al, 2017). This was also true for this study, where bla CTX-M-15 was found to be the dominant genotype among the ESBL-positive isolates (Table 2) and this is also in line with data from a previous study analyzing chicken meat in Ghana (Falgenhauer et al, 2019). Moreover, the dominance of bla CTX-M-15 has also been found in infected patients from two hospitals in Ghana (Mahazu et al, 2022). bla CTX-M-15 as predominant genotype among the ESBL-producing strains is of particular interest, as it has been associated with clinical E. coli strains to carry several virulence factors (Falgenhauer et al, 2021; Lau et al, 2008). bla CTX-M-15 is often found on incompatibility group FII plasmids (Coque et al, 2008), known as “epidemic resistance plasmids” (Carattoli, 2011) due to their tendency to acquire other resistance genes and their high potential to be transmitted through horizontal gene transfer (Cantón and Coque, 2006).
Notably, the coexistence of bla CTX-M-15 and bla TEM-1b observed in this study (Table 2) was considerably lower (1.2%) than what has already been reported in other parts of the world (Ramos et al, 2020). Nevertheless, continuous surveillance on bla CTX-M and other ESBL-positive isolates should be prioritized as they often confer resistance to multiple other antibiotic classes, leaving few or no other treatment options (Mellata, 2013). In support of this, a recent study reported a higher percentage of quinolone resistance genes in ESBL isolates than in non-ESBL isolates (FarajzadehSheikh et al, 2019). This could possibly explain the presence of MDR (30.8%, n/N = 25/81) in this study (Fig. 2), which is in line with those reported in rectal ESBL E. coli isolated from poultry in Tanzania (Kimera et al, 2021). However, other resistance genes were not screened for in this study.
Our study has limitations, which have to be considered when interpreting the results. Fecal samples were collected from only three communities in the study area and from six smallholder farms using a nonprobability sampling approach and the sample size was rather small. Thus, this study may not be representative for other regions of Ghana. Seasonality was not captured within this study as smallholder farms were visited only once within the 6 months study period.
Conclusion
The high prevalence of ESBL-producing E. coli in domestic farm chickens represent a potential transmission reservoir for human infections. Moreover, high resistance to locally used antibiotics poses a serious health threat to humans. These findings show that monitoring food hygiene and restricting antibiotic use must be reinforced in animal husbandry. This should be supported by the implementation of surveillance systems that monitor antimicrobial use and inform on emerging antibiotic-resistant bacterial strains. Instead, better hygiene measures among farmers could compensate for reduced usage of antibiotics. In addition, surveillance of imported meat should be a point of consideration for future studies. Moreover, further studies are required to discriminate between chromosomal and plasmid locations of the ESBL genes in E. coli to determine the potential of further AMR dissemination and the risk to human health.
Footnotes
Acknowledgments
We would like to thank Seth Ofori, Kwabena Oppong, Abdul Seidu Razak, Cynthia Adu Kyerewaa, and Charlotte Ama Tweneboa Adu of Kumasi Centre for Collaborative Research in Tropical Medicine (KCCR) and Doris Winter for technical assistance. We also thank the household chicken owners in Agogo.
Authors' Contributions
Conceptualization by C.W.A., L.A.O., and D.D. Data Curation by N.S. and C.W.A. Formal analysis by R.K., J.M., and N.A.K. Funding acquisition by L.A.O. and D.D. Methodology by C.W.A., L.A.O., D.D., and D.E. Project administration by R.O.P. and K.O.-D. Project administration by C.W.A. and E.K.P. Software by T.T. and J.M. Resources by D.D., R.O.P., J.M., and D.F. Supervision by L.A.O., D.D., and K.O.-D. Validation by C.W.A. and D.E. Visualization by C.W.A. and N.A.K. Writing—original draft preparation by C.W.A. Writing—review and editing by R.O.P., D.E., J.M., K.O.-D. L.A.O., D.D., T.T., E.K.P., N.A.K., R.K., N.S., and D.F.
Declarations
Disclosure Statement
No competing financial interests exist.
Funding Information
This research was funded by the German Research Foundation as part of the SASSA Project (DFG; project number 380545990). The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the article; or in the decision to publish the results.
