Abstract
PulseNet International (PNI) is a global network of 88 countries who work together through their regional and national public health laboratories to track foodborne disease around the world. The vision of PNI is to implement globally standardized surveillance using whole genome sequencing (WGS) for real-time identification and subtyping of foodborne pathogens to strengthen preparedness and response and lower the burden of disease. Several countries in North America and Europe have experienced significant benefits in disease mitigation after implementing WGS. To broaden the routine use of WGS around the world, challenges and barriers must be overcome. We conducted this study to determine the challenges and barriers countries are encountering in their attempts to implement WGS and to identify how PNI can provide support to improve and become a better integrated system overall. A survey was designed with a set of qualitative questions to capture the status, challenges, barriers, and successes of countries in the implementation of WGS and was administered to laboratories in Africa, Asia-Pacific, Latin America and the Caribbean, and Middle East. One-third of respondents do not use WGS, and only 8% reported using WGS for routine, real-time surveillance. The main barriers for implementation of WGS were lack of funding, gaps in expertise, and training, especially for data analysis and interpretation. Features of an ideal system to facilitate implementation and global surveillance were identified as an all-in-one software that is free, accessible, standardized and validated. This survey highlights the minimal use of WGS for foodborne disease surveillance outside the United States, Canada, and Europe to date. Although funding remains a major barrier to WGS-based surveillance, critical gaps in expertise and availability of tools must be overcome. Opportunities to seek sustainable funding, provide training, and identify solutions for a globally standardized surveillance platform will accelerate implementation of WGS worldwide.
Introduction
Background
PulseNet International (PNI) is a global network of national, regional, and subregional laboratories and laboratory networks of 88 countries that span Africa, Asia-Pacific, Canada, Europe, Latin America and the Caribbean, Middle East, and the United States. PNI has implemented standardized genotyping methods around the world, pioneered sharing of subtyping data for surveillance and outbreak response, and enabled global data sharing through capacity building and training activities (Swaminathan et al., 2006; Nadon et al., 2013).
Standardized subtyping methods pulsed-field gel electrophoresis and multilocus variable-number tandem-repeat analysis, in use for many years, were ultimately replaced beginning in the mid-2010s when evidence emerged that whole genome sequencing (WGS) was far superior in performance (e.g., sensitivity, specificity, and the ability to resolve clusters).
The benefits of WGS were estimated to be significant enough to warrant the substantial upheaval that its implementation would cause. The high resolution and epidemiological concordance of WGS, plus the potential for unambiguous nomenclature amenable to global surveillance compelled a new vision for PNI: to implement WGS for foodborne disease surveillance worldwide (Nadon et al., 2017).
Benefits of WGS for foodborne disease surveillance and outbreak response
Several countries in North America, Europe, and elsewhere have already experienced significant benefits in outbreak detection, response, and disease mitigation after implementing WGS. In the United States, WGS for foodborne disease surveillance is viewed as paradigm shifting “precision public health”; the beginning of a new era where pathogen genomics is anticipated to provide improvements for all areas of infectious diseases (Armstrong et al., 2019; Kubota et al., 2019).
Implementation of WGS has led to numerous benefits compared with molecular subtyping. For example, WGS (combined with epidemiological and food safety traceback data) led to an increase in the proportion of listeriosis outbreaks that could be solved (Jackson et al., 2016). In Europe, WGS is a key part of the European Centre for Disease Prevention and Control Public Health Microbiology Strategy for 2018–22, and by 2017 had been implemented in 18 countries of the European Union/European Economic Area (ECDC, 2018, 2019).
In Europe, WGS has enabled linking of sporadic cases with food products, improved outbreak investigations and source attribution and identified transmission pathways and antimicrobial resistance (ECDC, 2019). In Canada, using WGS for routine surveillance led to increases in outbreak resolution, epidemiological concordance, and led to the detection and linkage of multiple Salmonella Enteritidis outbreaks to food products that rapidly led to changes to national food safety policy and a subsequent decrease in disease incidence (Rumore et al., 2018; Morton et al., 2019). Laboratories in Australia, New Zealand, China, and Latin America and the Caribbean have described similar benefits (Kwong et al., 2016; Baker et al., 2017; Ford et al., 2018; Zhang et al., 2020).
The benefits of WGS have also been realized in smaller countries, where the improved performance plus the all-in-one workflow and reduction in labor-intensive steps of WGS are appealing to small countries (Nouws et al., 2020). In South Africa, the Centre for Enteric Diseases was in the early stages of WGS implementation when the technology was thrust into prominence and lauded for its critical role in resolving the world's largest listeriosis outbreak in 2017 (Smith et al., 2019). A national surveillance system using WGS was then implemented in South Africa, demonstrating the feasibility of WGS use in a middle-income country (Thomas et al., 2020).
Challenges with implementation
The four main areas of challenges for WGS implementation, as experienced by early adopters, include organizational (i.e., legislative/regulatory barriers, economic drivers, and funding), cultural (cross-sector collaboration, language barriers, and resistance to change), technical (data storage and computing power), and scientific (sequence generation and bioinformatics) (WHO, 2018).
An economic analysis of WGS implementation for Salmonella found that WGS costs 1.2–4.3 times more than traditional molecular methods, but that the resulting benefits would offset this difference even if WGS leads to only a relatively small number of cases prevented (Alleweldt et al., 2021). Indeed, the Africa Centres for Disease Control and Prevention recently launched a US$100-million Pathogen Genomics Initiative to integrate pathogen genomics and bioinformatics for improved disease control and prevention in Africa (Makoni, 2020).
Despite these examples of economic benefits and advantages, many countries still struggle. Although sustainable funding may be considered a universal root cause of slow implementation, the paradigm shift that WGS represents may present a more nuanced set of challenges to be overcome. Although WGS is also being planned for implementation in disease surveillance in many countries for many pathogens, the global COVID-19 pandemic has vastly and rapidly expanded the critical importance of genomics to the prevention and control of infectious diseases (Cardona-Ospina et al., 2021). These advances owing to COVID-19 may provide an opportunity for other infectious disease areas to leverage the progress for targeting genomics-based modernizations across the board.
Purpose
To broaden the routine use of WGS for foodborne disease surveillance and response worldwide, challenges and barriers must be identified. As a network, PNI is positioned to provide guidance and advocacy for capacity building and international surveillance; however, additional intelligence on where to best focus activities is needed. The purpose of this study was to ascertain the status of WGS implementation for foodborne disease surveillance and outbreak response by laboratories in low and middle-income countries (LMICs) participating in PNI regional networks, and to identify the specific challenges and barriers to implementing WGS.
Methods
Survey design
A qualitative survey, comprising multiple choice and open-ended questions, was designed to capture status, challenges, barriers, and plans of laboratories in WGS implementation during the 2019 calendar year (Supplementary Table S3).
Sampling plan and data collection
A total of 54 PNI-participating countries with laboratories in Africa, Asia-Pacific, Latin America and the Caribbean, and Middle East were invited to complete the survey. The survey was administered online via Google Forms (Alphabet, Inc., USA) by invitation only. There were no exclusion criteria. Institutional Review was not required for this technical survey with no human data.
Quantitative and qualitative analysis
Survey respondents were deidentified (names and institutes anonymized). Answers were converted into key messages and stratified by country and region in Excel (Microsoft, USA) following general grounded theory methodology (Corbin and Strauss, 2008). Initially, responses were coded to categories, then re-examined to identify concepts, and then assessed for trends, themes, and deviations. Multiple choice questions and coded qualitative responses were analyzed and visualized using the ggmap and ggplot2 package in RStudio (Kahle and Wickham, 2013; Wickham, 2016; RStudio Team, 2020). Questions with no responses were coded as missing values and were excluded from the analysis for that question; thus, response rates were tabulated for each question individually.
Results and Discussion
The survey represents a cross-section of laboratories in LMICs. Forty-one institutions from 33 of the 54 countries completed the survey: a response rate of 61% (country level). The survey was administered in early 2020, that is, at the beginning of the global COVID-19 pandemic. Many, if not all, the targeted laboratories were involved to various degrees in the initial pandemic response, and this could have impacted the response rate. The responding laboratories spanned all regions of PNI: Africa (42%), Asia-Pacific (15%), Latin America and the Caribbean (20%), and Middle East (25%) (Fig. 1). The majority of laboratories (66%) were national reference laboratories, including those from national public health, agriculture, and food safety authorities. The remaining laboratories were from the academic and research sector (24%) or could not be categorized (10%).
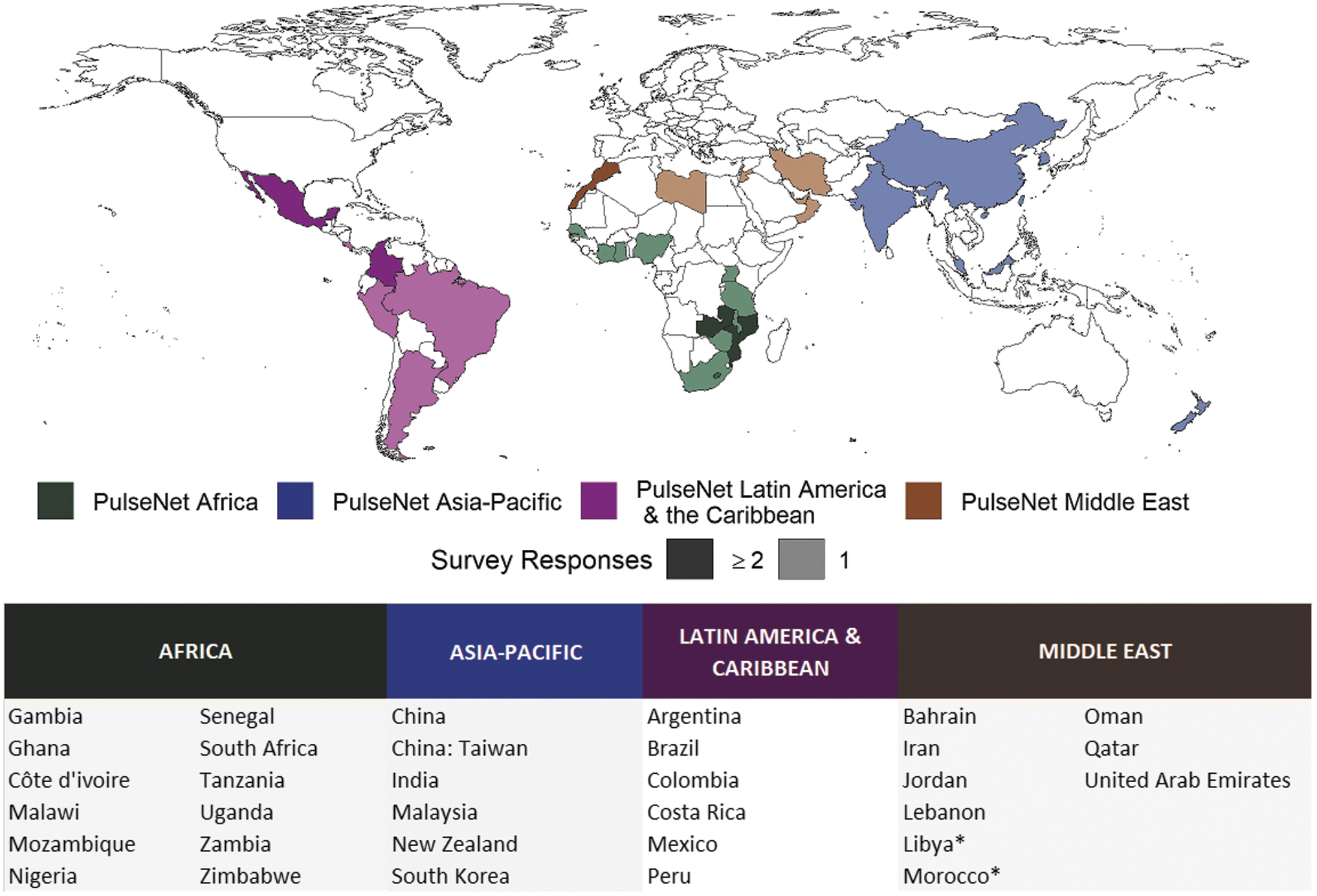
Map of PNI participating countries that responded to the survey (n = 41). Colors represent the PNI regions and the darkening of shades indicates countries with ≥2 survey responses. The country breakdown of responses is given below the map. *Countries are identified as belonging to regions as per the PNI networks, found at
WGS implementation is underway, but use in surveillance is highly limited. Only 8% of the laboratories use WGS for routine foodborne disease surveillance (Fig. 2A). A larger proportion (20%) use WGS for outbreak investigations after they are identified by other means (e.g., molecular subtyping), and 28% use WGS for research and pilot studies only.
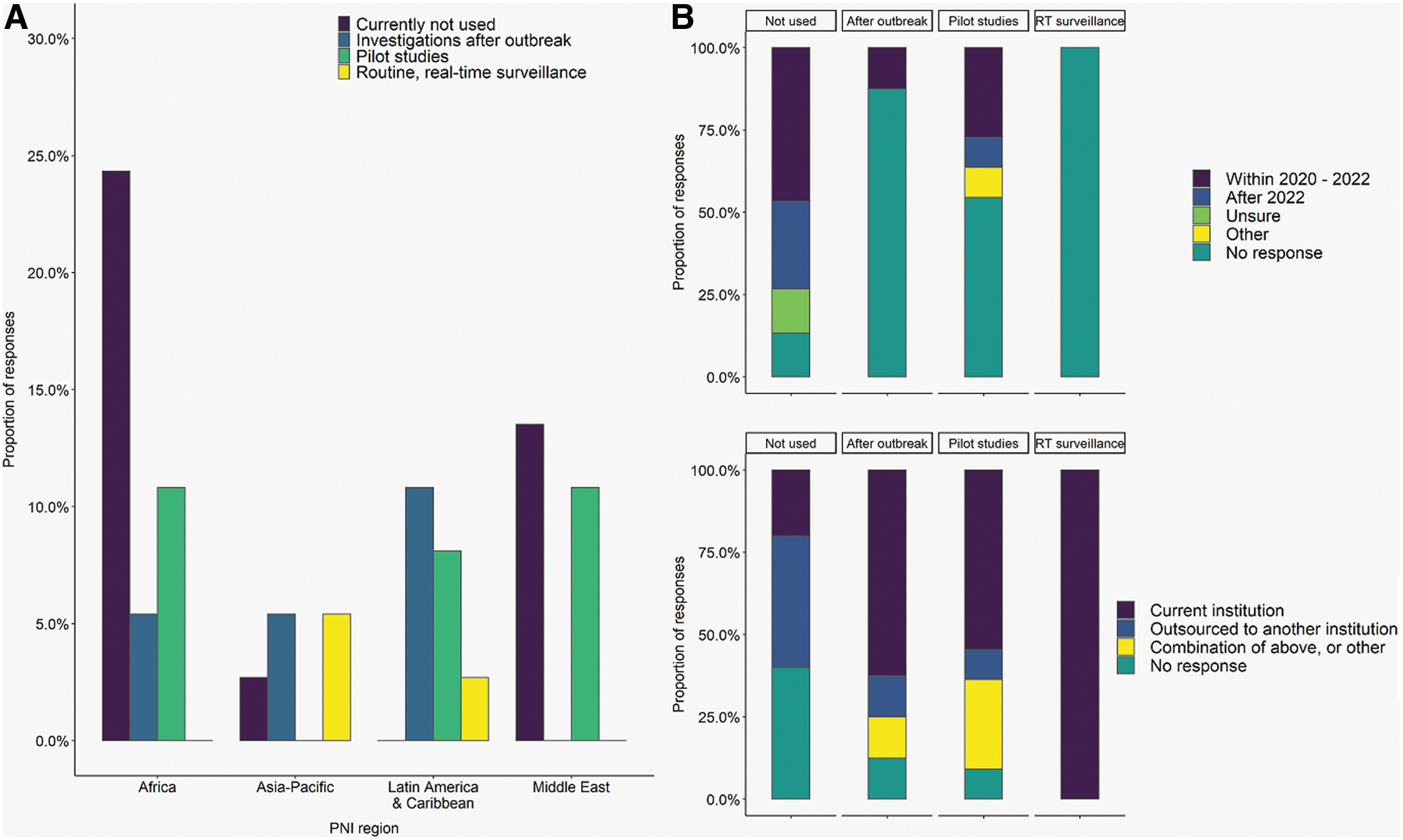
Characterization of WGS use for foodborne disease surveillance based on PNI region
Of the laboratories that currently do not have WGS implemented, 40% reported that they outsource sequencing to another institution (e.g., commercial partner, academic institution, or government laboratory), but have plans to implement WGS in-house around or after 2022 (47% and 27%, respectively) (Fig. 2B). Twenty percent of laboratories do not currently use WGS for foodborne disease surveillance, although sequencing is performed on-site for other purposes. The majority of laboratories that use WGS for investigations after an outbreak (63%) or for pilot studies (55%) perform sequencing at their own institution. Of these, 13% and 27%, respectively, plan to implement WGS at their own institution by 2022. All the laboratories that use WGS for routine, real-time surveillance perform these sequencing activities at their own institution.
In 2019 only 5% of laboratories sequenced over 1000 isolates (data not shown). Although 66% of laboratories sequenced 0–100 isolates that year, the remaining laboratories sequenced between 100 and 1000 isolates. The majority of sequencing was performed on priority foodborne pathogens: Salmonella (75%), Escherichia coli/Shigella (45%), Vibrio (20%), Campylobacter (15%), and Listeria (10%).
The capacity for generating sequences is generally higher than the capacity for analyzing or interpreting the data. Sixty-one percent of respondents indicated their capacities for utilizing, optimizing, and troubleshooting wet lab protocols for WGS were good to excellent (Fig. 3A). Approximately 70% of laboratories reported using Illumina instrumentation, with MiSeq being the most common model used (data not shown). Other platforms reported included Oxford Nanopore, Ion Torrent, and Pacific Biosciences. Capacity for computation and bioinformatics was found to be generally low. Local high-performance computing is available in 28% of laboratories, the remaining laboratories rely on laptop computers (28%), external hard drives (14%), cloud-based resources (11%), onsite servers (11%), or a combination of the above (6%) for computing and storage (Fig. 3C).
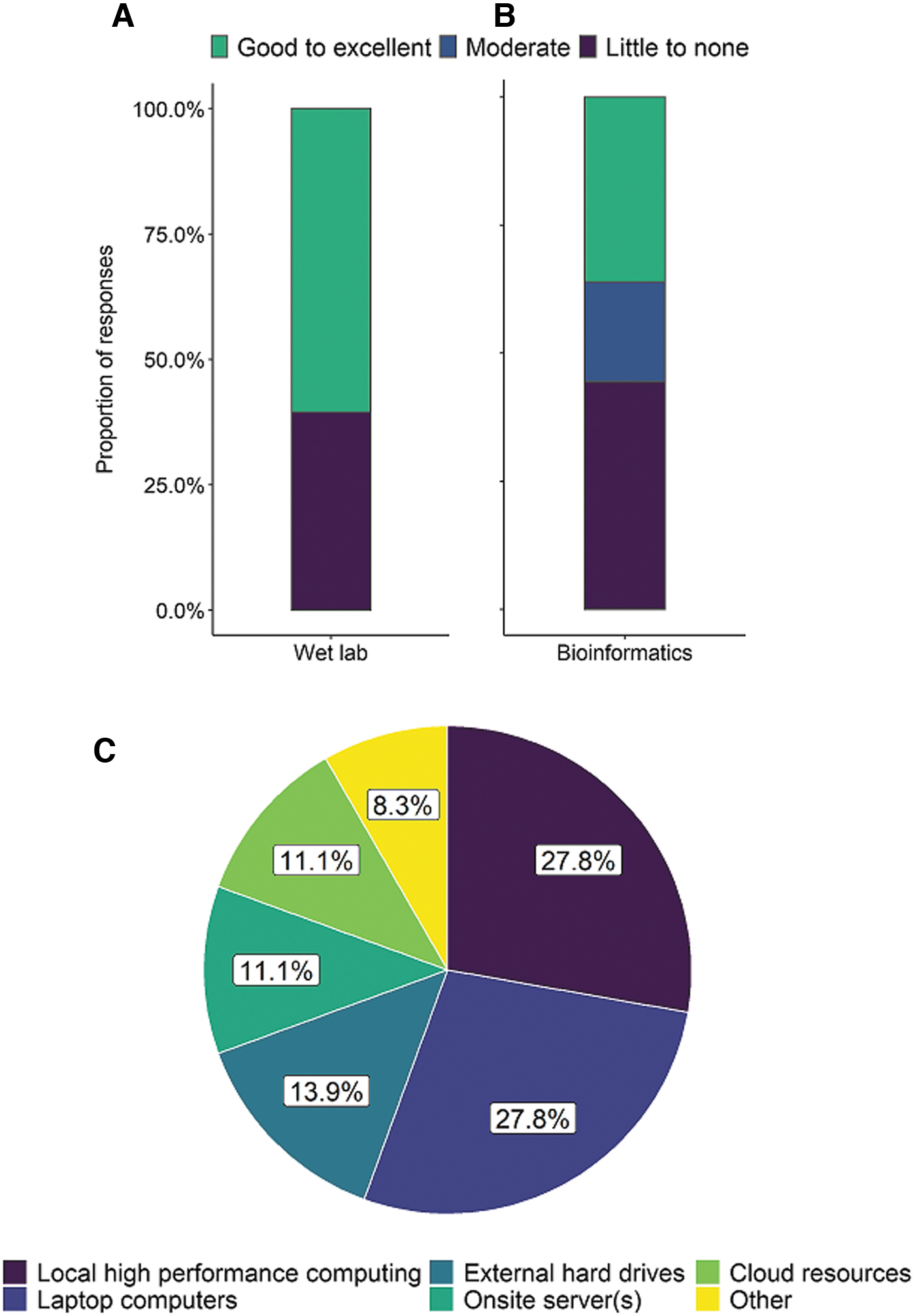
Characterization of responding laboratories capacity to optimize and troubleshoot whole genome sequencing wet lab protocols
Forty-four percent of responses characterized their laboratories' capacity and expertise to utilize, develop, optimize, and troubleshoot bioinformatics analysis protocols for WGS data as little or none (Fig. 3B). Core- or whole-genome MLST was identified as the in silico subtyping method used for determining allelic differences in 50% of laboratories, followed by single nucleotide polymorphism/single nucleotide variant (SNP/SNV)-based approaches (25%; data not shown). At least 75% of the laboratories that use WGS also use in silico phenotype prediction tools for antimicrobial resistance, virulence factors, and serotype.
The majority of laboratories (61%) do not have established guidelines for interpreting WGS data (e.g., number of allele or SNV/SNP differences for outbreak detection), whereas 21% rely on interpretation criteria developed by others or found in the literature (consulting with bioinformaticians, epidemiologists, and external partners as needed). Only 9% of laboratories have established guidelines, and another 9% interpret results on a case-by-case basis.
End-user knowledge for effective use of WGS data is low. The primary users of WGS data for outbreak detection and surveillance are epidemiologists (60%), ministry of health or equivalent authorities (40%), researchers/academicians (29%), and biologists (i.e., microbiologists, bacteriologists, and geneticists; 23%; Fig. 4A). Only 33% of laboratories characterized the level of knowledge and ability to utilize WGS data for public health decision-making as good to excellent (Fig. 4B). The remaining laboratories indicated their capacity to do so was moderate (19%) or minimal/nonexistent (48%).
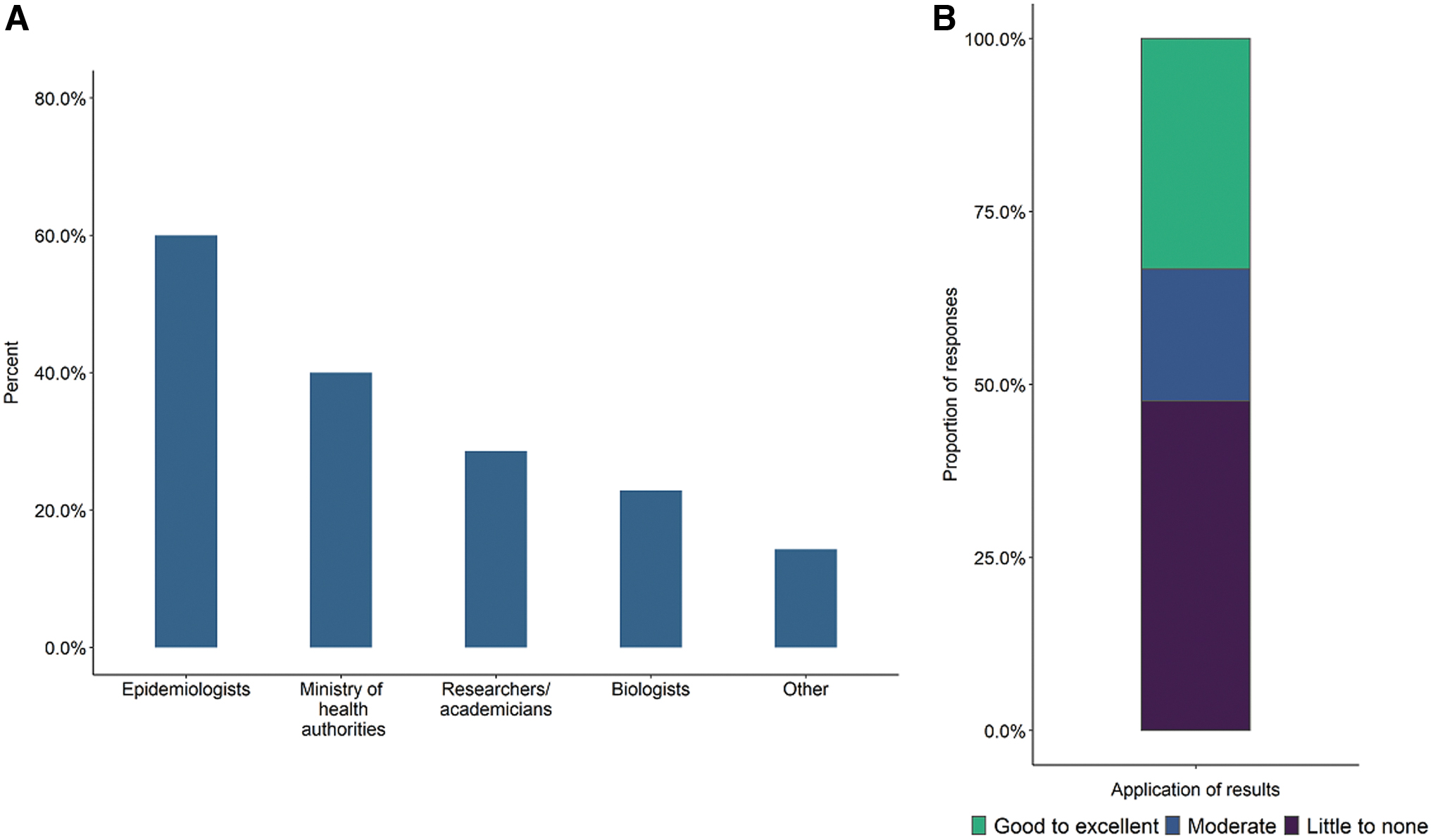
The end users of WGS data
The dissemination of WGS results is largely achieved through traditional mechanisms, and data sharing is highly limited. For cluster and outbreak detection, subtyping results are communicated to stakeholders through a variety of ways. Traditional methods, including excel spreadsheets (32%), hard copies (including fax; 24%), and in-person or telephone communication (18%), dominated over modern communication methods such as laboratory information management systems and internal bespoke websites (18%). A few laboratories disseminate WGS findings through formal written reports and manuscripts as their primary method of communication (9%; Fig. 5B).
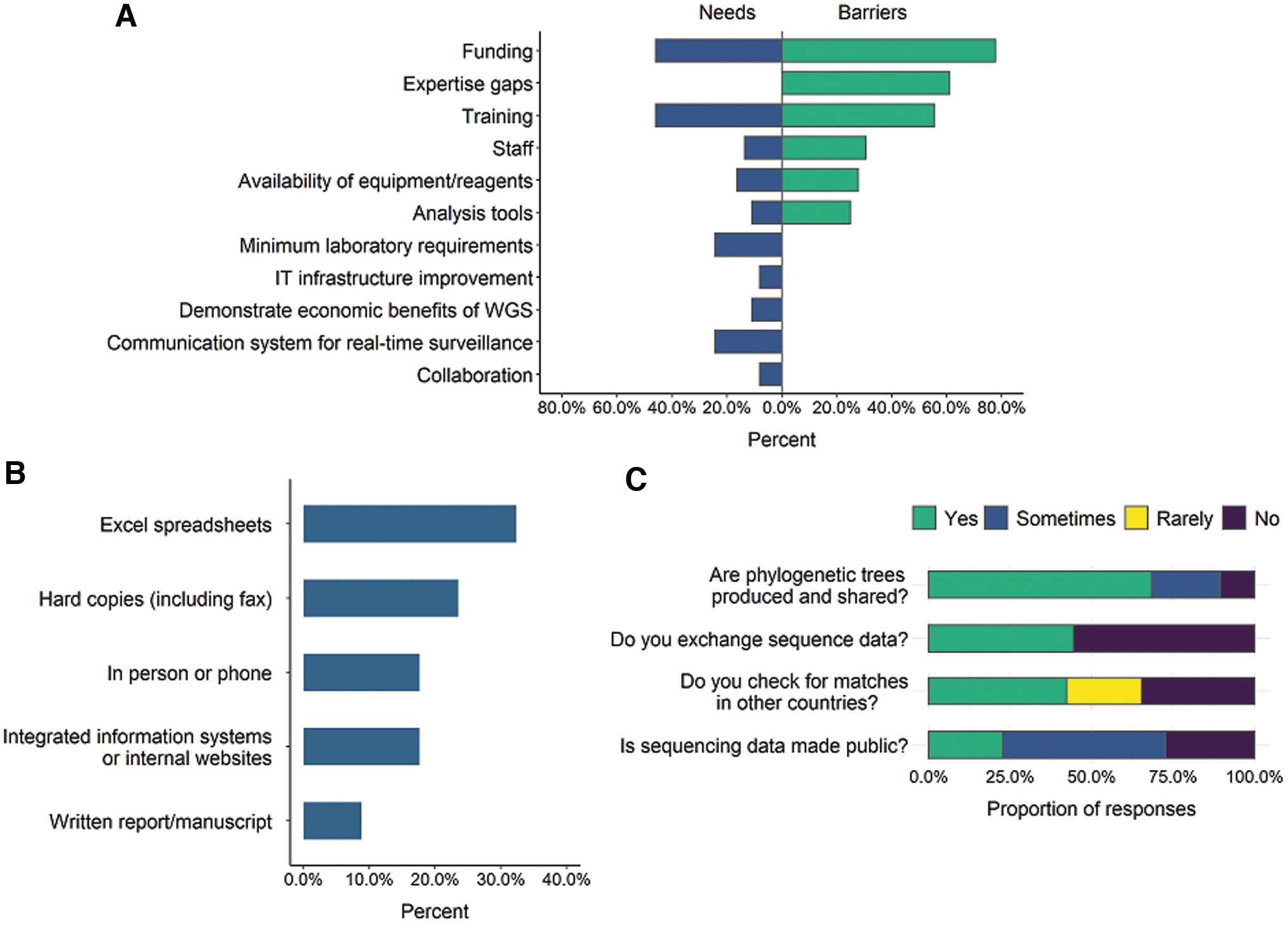
Although 68% of laboratories produce and share phylogenetic trees, the majority (56%) of laboratories do not exchange sequence data with external partners within their country. Forty-two percent of laboratories check for genetic similarity between their isolates and those in other countries on a routine basis. Otherwise, identifying “matches” in other countries is rarely done (23%) or not done at all (35%). Only half of responding laboratories make their sequencing data publicly available sometimes (Fig. 5C). Placing all sequencing data on public repositories is a routine practice for only 22% of laboratories.
Although 50% of laboratories reported not knowing reasoning behind this, 21% have not actually considered public release; their sole priority focused on national reporting only (data not shown). Lack of time and personnel to actually submit the WGS data to public archives (18%), not being permitted to share publicly because of legislation or policy restrictions (11%), and concerns of personal data protection and privacy (14%) were all reported as reasons for the lack of public release.
The primary challenges and barriers to WGS implementation are related to access to qualified personnel and data analysis and interpretation; however, there are additional needs that should be prioritized. Many of the biggest needs reflect the most frequently encountered barriers laboratories are experiencing. Lack of funding and training were identified as both the top barriers for WGS implementation (78% and 56%), and the biggest needs for achieving WGS within their institution (46% and 46%; Fig. 5A). Respondents emphasized that training is especially important for analysis and interpretation of WGS data, which would help fill the gaps in expertise (61%) that stand as a major barrier to utilizing WGS.
The staffing barrier (31%) aligns with the current need for more personnel to dedicate to sequencing activities (14%). Lack of available equipment and reagents was identified as a need (16%), and there is a major barrier to purchase these resources within some countries (28%). Analysis tools are still a major barrier (25%), and development of tools that are accessible, validated, and standardized was also specified as a priority (11%). Some needs, although they were not the most frequently identified, need attention: minimum laboratory requirements (e.g., basics in reagents, equipment, personnel, and Internet connection) were identified as needed in 24% of laboratories.
Additional high priority needs include a communication system for real-time surveillance (24%), enhanced collaboration between partner laboratories and PNI member countries (8%), and demonstrating the economic and other benefits (11%).
Priorities of PNI should facilitate global surveillance while continuing to build capacity via training and bespoke development support. With WGS comes universal, unambiguous, and comparable data generated by all and the opportunity to enable global foodborne disease surveillance. The ideal system to support global WGS-based foodborne disease surveillance should include an “all-in-one” solution for analyzing, visualizing, sharing, and storing WGS data (41%; Table 1). The ideal global surveillance system would also feature or make available properly validated and standardized pipelines (35%), with a common global database (35%), and that is accessible by all PNI-participating laboratories (14%).
Moving PulseNet International Forward
PNI, PulseNet International; WGS, whole genome sequencing.
Nearly half of laboratories (51%) believe that the priorities of PNI should focus on training, especially in WGS data analysis, and that having access to globally standardized and validated analysis tools and pipelines (31%) are key to progress toward global foodborne disease surveillance using WGS (Table 1). Moving forward, providing tailored support to countries that are struggling (29%) and help in coordinating funding opportunities (20%) are also relevant priorities. To better integrate activities around the world, laboratories indicated that PNI efforts should enable data sharing between participating laboratories without public release (23%), and provide guidance for increasing communication (17%) and collaboration (11%) between regions.
Conclusions
This study reveals that use of WGS for routine foodborne disease surveillance in the countries surveyed is low. Although funding remains a major barrier to WGS, there are critical gaps in expertise and availability of bioinformatics analysis that must be addressed. Our findings indicate that the implementation of WGS is complex and multifaceted. In addition to focusing on the funding and training aspects of capacity building, needs in areas outside the science per se have been illuminated.
Solutions for a globally standardized WGS-based surveillance platform and pipelines, accessible to all members with or without public sharing, are critically needed. These findings are in general agreement with an assessment conducted in Europe, where capacity building targeted at harmonizing operations and the interoperability of data were identified as needs (EFSA Panel on Biological Hazards et al., 2019).
Efforts to meet this need, that is, a secure mechanism for conducting country level and global surveillance, should be done in parallel to promoting the public release of data. To date, data-sharing efforts have been targeted toward the public release of sequences to enable global foodborne disease surveillance; indeed, this forms a core component of PNI's vision and must be continued (Nadon et al., 2017).
Many countries struggle to share sequence data with public repositories; the data available in NCBI's Pathogen Detection system (
The majority of S. enterica and E. coli/Shigella are from the United States (45% and 38%, respectively) and Europe (30% and 34%, respectively; accessed June 2021; Supplementary Table S2). There remains a critical need for cluster detection and outbreak response tools that bring the public health benefits of WGS to more countries regardless of their ability to share outside their jurisdiction. In addition, tools that can be used with all sequencing platforms are critical, because the COVID-19 response may have shifted the availability and use of instrumentation such as Oxford Nanopore. Since this survey was conducted, however, significant strides have been made in global sharing of pathogen sequence data via the COVID-19 pandemic, with ∼4 million SARS-CoV-2 sequences shared via the public-facing Global Initiative on Sharing All Influenza Data (as of October 2021).
With a spotlight now on the world's largest global genomic epidemiology effort to date, it is possible these advances will translate to other disease areas, including foodborne. In addition to designing capacity-building activities targeted at the needs identified in this study, the strategies used by laboratories that have implemented and sustained WGS-based surveillance could be harnessed and used as an implementation toolkit. The findings of this study will be useful in furthering our understanding of the forces at play in the design and implementation of WGS for foodborne disease surveillance.
Footnotes
Acknowledgments
The authors thank the members of the PNI steering committee for their thoughtful guidance and input on the study and article. The authors also thank all participants who completed the survey.
Disclosure Statement
No competing financial interests exist.
Funding Information
This publication was supported by Cooperative Agreement number NU60OE000104, funded by the Centers for Disease Control and Prevention. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the Centers for Disease Control and Prevention or the Department of Health and Human Services.
Supplementary Material
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
