Abstract
Three recent studies report the first biochemical reconstitution of DNA double-strand break (DSB) formation by SPO11, the topoisomerase-derived transesterase that initiates meiotic recombination in sexually reproducing organisms. A central conclusion of these studies is that SPO11 is sufficient to catalyze DSBs in vitro, but cleavage is limited by the poor propensity of SPO11 to dimerize, thereby providing an effective mechanism to prevent uncontrolled breaks. The studies yield new insights into the mechanism of DNA DSB formation and raise new questions regarding the functions of SPO11 partners, the impact of the DNA substrate, the coordination between cleavage events, and the reversibility of the reaction.
SPO11 Initiates Meiotic Recombination
Twenty-eight years ago, SPO11 was identified as the enzyme that catalyzes the formation of DNA double-strand breaks (DSBs) at the onset of meiosis (Bergerat et al., 1997; Keeney et al., 1997). DSB formation by SPO11 is an evolutionarily conserved mechanism that triggers homologous recombination and consequently promotes parental chromosome pairing and genetic diversity through allelic exchanges (Yadav and Claeys Bouuaert, 2021). However, these programmed breaks are inherently risky and must be tightly regulated to avoid untimely or excessive DNA damages. Conjointly with two other research groups, we have now reported the first successful in vitro reconstitution of SPO11-dependent DNA cleavage (Oger and Claeys Bouuaert, 2025; Tang et al., 2025; Zheng et al., 2025), paving the way to decipher the molecular mechanism and regulation of meiotic DSB formation.
SPO11 Is a Broken Pair of Scissors
SPO11 evolved from the DNA-cleavage subunit of an ancestral type IIB topoisomerase, Topo VI (Bergerat et al., 1997; Robert et al., 2016b). Like its evolutionary precursor, SPO11 cleaves DNA through a nucleophilic attack from a catalytic tyrosine that forms a covalent link with the DNA backbone (Keeney and Kleckner, 1995; Liu et al., 1995). To catalyze DSB formation, each DNA strand is cleaved by an active site formed at the interface between two SPO11 subunits. One subunit contributes the catalytic tyrosine and the other subunit coordinates metal ions necessary for the reaction (Diaz et al., 2002; Nichols et al., 1999) (Fig. 1A).
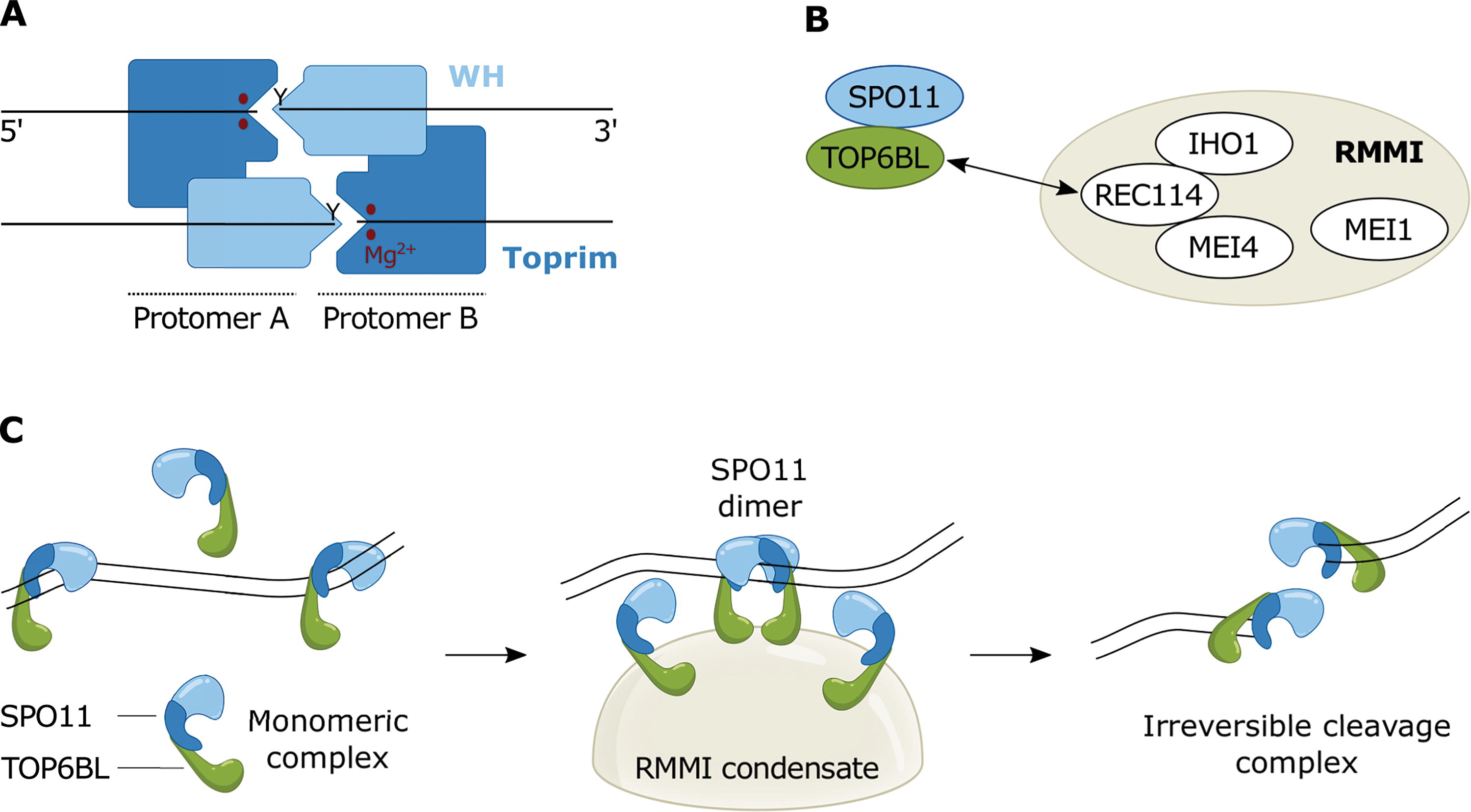
Model of SPO11 activation in mouse.
Using purified mouse SPO11, we developed a plasmid DNA cleavage assay that recapitulates all the hallmarks of meiotic DSB formation. We showed that SPO11 produces DSBs with two-nucleotide 5′ overhangs and remains covalently bound to the 5′ DNA ends. Mutating the catalytic tyrosine (Y138F) or the metal-ion binding pocket (E224A) completely abolished DNA cleavage. However, mixing the two inactive mutants led to single-strand-break products, indicating that the formation of heterodimers restores one functional active site per complex (Oger and Claeys Bouuaert, 2025; Zheng et al., 2025). This formally establishes that SPO11 dimerizes to cleave two DNA strands using two hybrid active sites.
Mouse SPO11 behaves mostly as an inactive monomer in solution (Oger and Claeys Bouuaert, 2025; Zheng et al., 2025). This monomeric stoichiometry had also been observed for S. cerevisiae and C. elegans Spo11/SPO-11 (Claeys Bouuaert et al., 2021b; Yeh et al., 2017), but since no cleavage activity had been reported for these proteins, it remained unclear whether the monomeric state was a feature or an artifact. Our current data strongly indicate that this is indeed an essential and conserved property of SPO11.
In vitro, cleavage requires a high protein-to-DNA ratio, which is best explained by promoting the encounter of SPO11 subunits on DNA (Oger and Claeys Bouuaert, 2025). Consistently, we found that cleavage could be promoted by adding an excess of inactive SPO11 to stimulate dimerization. Furthermore, Zheng and colleagues demonstrated that artificial dimerization of SPO11 stimulates DNA cleavage in vitro (Zheng et al., 2025). Based on our model that dimerization controls SPO11 activity, we expect that artificial dimerization of SPO11 will also promote DNA cleavage in vivo, independently of any additional cofactors.
What Is the Function of SPO11’s Partners?
SPO11 universally relies on a constellation of accessory factors to catalyze DSB formation in vivo (Lam and Keeney, 2014; Wang and Copenhaver, 2018; Yadav and Claeys Bouuaert, 2021). In mice, for instance, SPO11 requires the topoisomerase-derived B-like subunit, TOP6BL, and at least four meiosis-specific proteins, REC114, MEI4, MEI1, and IHO1 (RMMI) (Fig. 1B) (Kumar et al., 2010; Kumar et al., 2018; Libby et al., 2003; Robert et al., 2016a; Stanzione et al., 2016). Hence, we were surprised to discover that mouse SPO11 catalyzes DSBs without any of its partners (Oger and Claeys Bouuaert, 2025). What, then, is the function of all these accessory proteins in vivo?
SPO11 forms with TOP6BL a stable 1:1 complex of similar activity to SPO11 alone, suggesting that TOP6BL has no direct role in dimerization or cleavage (Oger and Claeys Bouuaert, 2025). Previous work showed that the yeast homologs of the mouse RMMI proteins undergo DNA-dependent condensation in vitro (Claeys Bouuaert et al., 2021a). Despite extreme sequence divergence, mouse REC114-MEI4 and IHO1 complexes share structural and functional properties with their yeast counterparts, including condensation (Daccache et al., 2023; Gao et al., 2020; Laroussi et al., 2023; Liu et al., 2023). Mouse REC114 was also shown to bind TOP6BL directly (Nore et al., 2022). Based on these insights, we suggest that RMMI form condensates along meiotic chromosomes and recruit SPO11 via TOP6BL, thereby locally increasing SPO11 concentration to allow dimerization and cleavage (Fig. 1C) (Oger and Claeys Bouuaert, 2025; Zheng et al., 2025). In addition, it is possible that SPO11 anchoring at the surface of condensates imposes an orientation favorable for dimerization.
This model suggests that TOP6BL and RMMI can be bypassed at high SPO11 concentration, explaining why they are dispensable in vitro. Since REC114-MEI4 has a 2:1 stoichiometry (Claeys Bouuaert et al., 2021a; Daccache et al., 2023; Laroussi et al., 2023), REC114-MEI4 may also serve as a bridging partner to promote SPO11 dimer formation. However, a simple bridging model fails to account for the formation of bright RMMI foci (Kumar et al., 2015; Kumar et al., 2018; Stanzione et al., 2016) and the frequent catalysis of closely spaced double-DSBs (Johnson et al., 2021; Prieler et al., 2021). Whether SPO11 dimerizes in the context of a stoichiometric complex or by concentration in a nonstoichiometric condensate, the expectation is that RMMI will stimulate DNA cleavage at low SPO11 concentrations in vitro.
SPO11 Creates Single-Strand Nicks
In addition to the expected double-strand cleavage products, the in vitro assay revealed that SPO11 often cleaves a single DNA strand (Oger and Claeys Bouuaert, 2025; Zheng et al., 2025). Hence, the two cleavage events are not strictly coordinated.
Similar observations were previously reported for type II topoisomerases. For instance, half of the cleavage complexes formed by human Topo IIα are single-stranded (Deweese and Osheroff, 2009). Nevertheless, it has been demonstrated that the first nick increases cleavage of the opposite strand up to 20-fold (Deweese and Osheroff, 2009; Zechiedrich et al., 1989). This imperfect coordination is mediated by DNA bending, since cleavage of the first strand increases the flexibility of DNA and, as a result, enhances the probability of second-strand cleavage by maintaining the DNA substrate in a cleavage-competent bent conformation (Lee et al., 2012). Future studies will clarify whether SPO11 has inherited a similar inter-protomer coordination mechanism.
This observation that SPO11 frequently cleaves a single DNA strand raises the question of whether nicks accumulate during meiosis. Previous attempts to address this failed to detect single-strand nicked products in yeast (de Massy et al., 1995; Liu et al., 1995; Xu and Kleckner, 1995). Perhaps in vivo, the presence of partners reinforces the coordination of cleavage on both DNA strand, for instance by stabilizing DNA bending or preventing the complex from falling apart during catalysis. It is also possible that nicked products are efficiently resealed during DNA isolation, hampering their detection.
SPO11 Relaxes Supercoiled Plasmids
In conditions that accumulated single-strand nicks, for example, when SPO11 dimers only contained one functional active site, we observed that supercoiled plasmids were converted into partially relaxed plasmid topoisomers (Oger and Claeys Bouuaert, 2025). A similar observation was reported for the SPO11–TOP6BL complex (Zheng et al., 2025).
Despite its relationship with Topo VI, a plasmid relaxation activity was not expected for SPO11 because it is not a bona fide type II topoisomerase. Indeed, this activity was independent of ATP and occurred in the absence of DSB intermediate and hence cannot function by a strand passage mechanism. Instead, we proposed that SPO11 relaxation occurs by a swiveling mechanism, which may stem from the weak dimer interface of SPO11. Indeed, dissociation of the dimer after first-strand cleavage would lead to the rotation of the DNA double helix around the intact phosphodiester bond. Upon restoration of the dimer, the cleavage reaction can be reversed, resulting in the release of a partially relaxed, covalently closed plasmid (Fig. 2A). An alternative is that swiveling results from the dissociation of one SPO11 subunit from DNA, but this is less likely given that the dimer interface is weaker than the DNA-binding surface.
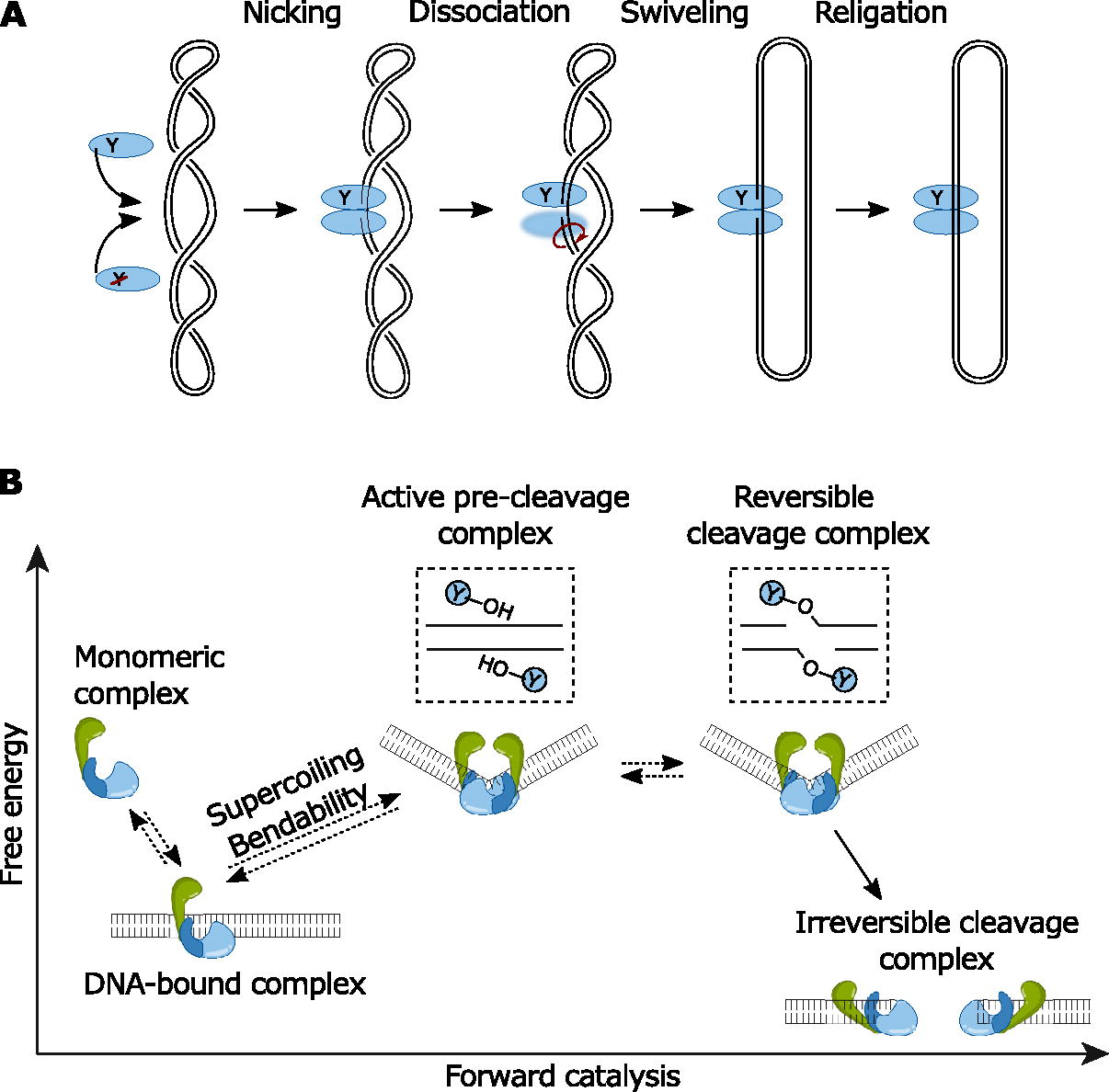
Model of DNA relaxation by SPO11 and proposed thermodynamic representation of SPO11-TOP6BL catalysis.
A role for SPO11 in DNA relaxation was not anticipated and is not currently backed by any biological evidence. In vivo, we envision SPO11 to be tethered at the surface of condensates, which may anchor both SPO11 protomers and hence prevent swiveling during catalysis. However, more detailed analyses may be required, since this putative relaxation activity could manifest only in specific conditions.
Reversibility of DNA Cleavage
A change in the linking number of a supercoiled plasmid implies that at least one strand of the DNA duplex has been opened and resealed. Therefore, the production of topoisomers indicates that SPO11 can both cleave and religate DNA. In the context of type II topoisomerases, “reversal” refers to the breakage of 5′-phosphotyrosyl bonds by free 3′-hydroxyl groups in the absence of strand transfer. For Topo II, this equilibrium between cleavage and ligation is in favor of reversal, which is thought to be important for genome integrity (Deweese et al., 2008).
Given that the transesterification reaction is isoenergetic, in contrast to a hydrolysis reaction, it has long been suspected that SPO11 cleavage would involve a reversible state (Keeney, 2008; Keeney et al., 1997). Possible evidence for reversal in vivo comes from previous analysis of Spo11 chromatin association in a yeast rad50S mutant (Prieler et al., 2005), which accumulates unrepaired breaks with Spo11 bound to the DNA ends. Spo11 immunoprecipitation revealed a crosslinking-independent DNA signal that accumulated before DSBs could be detected by Southern blot analysis. This association was sensitive to salt in the extraction buffer and required the catalytic tyrosine of Spo11, leading the authors to suggest that they captured a reversible cleavage intermediate (Prieler et al., 2005). This reversible intermediate may escape detection by Southern blot analysis, perhaps because the DNA extraction buffer promotes religation, a known phenomenon with Topo II (Liu et al., 1983; Osheroff and Zechiedrich, 1987).
So far, our in vitro results only support strand religation in the context of a single-strand nicked product. Presumably, the intact DNA strand helps position the free 3′ hydroxyl and the 5′-phosphotyrosyl bond in a configuration prone to reversal. The double-strand cleavage product also likely involves a reversible stage, but the extent to which reversal occurs remains unclear.
What, then, triggers the transition from a reversible to an irreversible break? A clue comes from comparing the cryo-EM structure of the yeast complex that mimics a post-cleavage state with models of a pre-cleavage complex, which suggested that cleavage is followed by a conformational transition that is incompatible with Spo11 dimerization (Yu et al., 2025). In this model, the cleavage complex is a high-energy state that relaxes into a thermodynamically more stable configuration only after both strands have been cleaved, causing an irreversible collapse of the dimer (Fig. 2B). This model also suggests that DNA topology may provide the energy required to drive the reaction forward.
The Role of DNA Bending in Catalysis
Our in vitro assays indicated that SPO11-dependent cleavage is affected by the sequence, supercoiling, and bendability of the DNA substrate (Oger and Claeys Bouuaert, 2025; Zheng et al., 2025). AlphaFold modeling of SPO11 complexes across several phyla suggested that the pre-cleavage complex bends DNA with an angle of approximately 100°. This is consistent with atomic force microscopy images of DNA-bound yeast and mouse SPO11 complexes and is conserved in Topo VI (Claeys Bouuaert et al., 2021b; Wendorff and Berger, 2018; Zheng et al., 2025).
These observations raise interesting questions regarding the function of DNA bending. Perhaps bending helps melt the double helix to facilitate cleavage. Perhaps it provides a mechanism to coordinate the two cleavage events by stabilizing the nicked intermediate. Another possibility is that DNA bending helps drive the cleavage reaction toward the formation of an irreversible break. Under this scenario, DNA bending energizes the pre-cleavage complex, making cleavage reversal energetically unfavorable. The requirement for DNA bending may participate in driving breaks near active promoters and help couple DSB timing with the buildup of topological stress during meiotic prophase I (Heldrich et al., 2020; Prieler et al., 2021).
Conclusions
The reconstitution of SPO11 activity in vitro has already brought key new insights into the mechanism and regulation of meiotic DSB formation, the impact of the target sequence and topology, and the reversibility of the cleavage reaction. Undoubtedly, this breakthrough will spur further in vitro studies that will lead to a better understanding of the initiation of meiotic recombination.
Our research also highlights new research questions to be addressed in vivo. A key prediction of our model is that artificial dimerization of SPO11 may bypass the requirement of SPO11 on accessory partners. Other questions of interest include the possibilities that meiotic cells accumulate single-strand nicks, that SPO11 may participate in DNA relaxation through swiveling activities, and that cleavage involves an elusive reversible state.
Footnotes
Acknowledgments
Research in the Claeys Bouuaert laboratory is supported by the European Research Council and the Fonds National de la Recherche Scientifique.
Authors’ Contributions
C.O. and C.C. wrote the article.
Disclosure Statement
No competing financial interests exist.
