Abstract
Laryngeal squamous cell carcinoma (LSCC) is the most common and prevalent malignant tumor in head and neck squamous cell carcinoma. Recent studies have shown that circular RNAs (circRNAs) play a vital role in cancer development, but their specific role in the tumorigenesis and development of LSCC remains unclear. We selected five pairs of LSCC tumor tissues and paracancerous tissues for RNA sequencing. Reverse transcription-quantitative PCR (RT-qPCR), Sanger sequencing, and fluorescence in situ hybridization were utilized to study the expression, localization, and clinical significance of circTRIO in LSCC tissues, and TU212 and TU686 cell lines. Furthermore, cell counting Kit-8, colony-forming assay, Transwell, and flow cytometry assays were assessed to illustrate the crucial role played by the circTRIO in proliferation, colony-forming ability, migration, and apoptosis in LSCC cells. Finally, the molecule's role as a microRNA (miRNA) sponge was analyzed. In the results, we screened out a promising upregulated novel circRNA-circTRIO in LSCC tumor tissues compared with paracancerous tissues using RNA sequencing. Then, we used qPCR to evaluate the expression of circTRIO in 20 additional paired LSCC tissues and two cells, and the results showed that circTRIO was highly expressed in LSCC and that this high expression was closely related to the malignant progression of LSCC. Furthermore, we examined circTRIO expression in the Gene Expression Omnibus data sets GSE142083 and GSE27020, and circTRIO expression was considerably higher in tumor tissues compared with the adjacent tissues. Kaplan–Meier survival analysis showed that the expression of circTRIO was associated with worse disease-free survival. The Gene Set Enrichment Analysis biological pathway evaluation results demonstrated that circTRIO was mainly enriched in cancer pathways. Moreover, we confirmed that silencing circTRIOs can help to significantly inhibit LSCC cell proliferation and migration while triggering apoptosis. Upregulated circTRIO expression levels may play an important role in the tumorigenesis and development of LSCC.
Introduction
Over 12,000
Several different noncoding RNAs, such as long noncoding RNAs (lncRNAs), microRNAs (miRNAs), and circular RNAs (circRNAs), have recently been demonstrated to be related to the pathophysiology of LSCC (Carron et al., 2021; Gao et al., 2020; Li et al., 2020b; Lv et al., 2023; Wu et al., 2020). Research on circRNAs has historically been less extensive in favor of the more directly involved miRNAs—yet increasing evidence points to the critical role of circRNAs in the development of LSCC. CircRNAs are covalently closed, single-stranded RNA molecules without the free 5′ ends, 3′ ends, or poly-A tails (Meng et al., 2017). As per its name, CircRNA has a unique ring structure, leading to major stability and other notable qualities, including heterogeneity, high evolutionary conservation, and tissue specificity (Kristensen et al., 2019).
Via various mechanisms, CircRNA affects a broad range of biological functions, including the regulation of gene transcription and translation, protein interactions, and miRNA sponges (Lei et al., 2020; Shi et al., 2020; Zang et al., 2020; Zhang et al., 2020). Because CircRNA, particularly its dysregulation, plays a significant role in several diseases and tissues, research on this noncoding RNA could lead to new diagnostic biomarkers and therapeutic targets for numerous diseases, including cancers.
Regarding LSCC, several potential circRNA biomarkers have been identified to be dysregulated and linked to tumorigenesis, including hsa_circ_0036722, circCORO1C, hsa_circ_100290, and hsa_circ_0067934 (Chu, 2020; Guo et al., 2020; Wang et al., 2020; Wu et al., 2020). Most of these circRNAs function as miRNA molecular sponges, competitively binding to the corresponding miRNAs, which in turn antagonize the repressive effects of miRNAs on downstream target genes. For example, hsa_circ_0042823 promotes cancer progression by modulating miR-877-5p and FOXM1 in LSCC (Wu et al., 2021b). While these studies have provided some insight as to the vital role played by circRNA in the LSCC development, such mechanisms are so complicated that more research must be conducted to understand the role of circRNA in LSCC.
In this study, we identified the circTRIO molecule as a potential novel diagnostic biomarker and therapeutic target for LSCC. We initially performed RNA sequencing and constructed circRNA expression profiles for five pairs of malignant LSCC tumors and their surrounding tissues. The differentially expressed circRNAs (DECs) were then investigated, and circTRIO was shown to be one of the top three aberrant circRNAs in LSCC samples. This study's experimental focus is therefore to explore the molecular mechanism and function of circTRIO. The expression of circTRIO was first validated in 20 pairs of LSCC tissues and LSCC cell lines. Then, the role of CircTRIO in the proliferation, migration, and apoptosis of LSCC cells was assessed. Our findings imply that artificially silencing circTRIO can inhibit LSCC cell proliferation and migration while promoting apoptosis, which can provide crucial clues for the development of novel diagnostic and therapeutic targets in LSCC patients.
Materials and Methods
Patients and clinical information
Clinical LSCC and paracancerous tissue samples were taken from patients who underwent surgery in Beijing Tongren Hospital, Capital Medical University (Beijing, China) between 2019 and 2021. The inclusion criteria were as follows: (1) patients who were only diagnosed with LSCC, which excluded other malignancies; (2) patients who did not receive any treatment (chemotherapy, immunotherapy, or Traditional Chinese Medicine) before surgery; and (3) samples of LSCC tumors (cancerous) and adjacent normal tissues 1–3 cm away (paracancerous) that were taken during surgery. These tissues were immediately preserved in liquid nitrogen and paired together for analysis.
The World Health Organization (WHO) Classification of Tumors system was used to determine the histological types of LSCC, and the TNM (Tumor, Node, Metastasis) stage was determined using the criteria of the American Joint Committee on Cancer (AJCC, 8th edition). Of the 25 paired LSCC and paracancerous tissues, 5 were used for RNA sequencing, and 20 were used for quantitative PCR (qPCR) analysis (Supplementary Table S1). The study protocol was reviewed and approved by the Ethics Committee of Beijing Tongren Hospital (Approval No. 20190215).
RNA-seq and circRNA screening
Five pairs of cancerous and matched paracancerous tissues were selected for whole-transcriptome high-throughput sequencing (RNA-seq) with the purpose of quantifying RNA expression. Sequencing was carried out by Novogene (Beijing, China) utilizing the standard protocol of a HiSeq 4000 (Illumina, San Diego, CA) instrument. Total RNA was extracted from the samples and used for cDNA library construction after ribosomal RNA (rRNA) depletion. CIRI (Gao et al., 2015) and Find_circ (Memczak et al., 2013) were used to classify circRNAs, and expression levels were assessed using fragments per kilobase per million mapped reads. DECs were identified using the edgeR package, with a subset of these circRNAs being annotated using the circBase database. p-Values were adjusted as q (padj) values by the Benjamini and Hochberg method (Storey, 2003); DECs were sorted by the following threshold: log2-fold change >1.5 (or <−1.5), with a q (padj)-value <0.05.
Gene Expression Omnibus data analysis and Gene Set Enrichment Analysis
Transcriptome sequencing data and clinical features of LSCC were downloaded from the Gene Expression Omnibus (GEO) database. In particular, the LSCC data set consisting of GSE142083 and GSE27020 was utilized. Patients were classified into high-(circTRIOhi) and low-expression (circTRIOlow) groups at a median cutoff of circTRIO expression to calculate the log2-fold change using R package “limma” implemented procedures. Moreover, the following three gene lists were obtained from the Molecular Signatures Database (MSigDB): h.all.v2022.1.Hs.symbols.gmt, c2.cp.kegg.v2022.1.Hs.symbols.gmt, and
Cell culture, siRNAs, and cell transfection
The human oral epithelial cell line and human LSCC cell lines (TU686 and TU212) were cultured in DMEM/high glucose (HyClone) supplemented with 10% fetal bovine serum (FBS) (#04-001-1ACS, BI). All cell lines were maintained at 37°C and 5% CO2. After cell confluence reached 80%, 0.25% trypsin was used to digest cells for passaging. SiRNAs targeting circTRIO (si-circTRIO#1; si-circTRIO#2; si-circTRIO#3) as well as negative control siRNAs (si-NC) were synthesized by Guangzhou Geneseed Biotech (Guangzhou, China). SiRNAs used in this study are shown in Supplementary Table S2. After transfection with Lipofectamine 3000 (Invitrogen) in accordance with the manufacturer's protocols, cells were incubated at 37°C and 5% CO2 for 48 h.
Real-time quantitative reverse transcription polymerase chain reaction
Twenty additional cancerous–paracancerous tissue pairs were collected and verified with real-time quantitative reverse transcription polymerase chain reaction (qRT-PCR). Using the TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.), total RNA was isolated from tumors and the surrounding nontumor tissues. RNA integrity and quality were measured using 1% agarose gel electrophoresis and spectrophotometry, respectively. The A260/A280 ratios were 1.8–2.2, and the absorbance at 260 nm (A260)/A230 ratios had to be >1.7 for the RNA samples to be regarded as qualified. RNA was then reverse transcribed using the Transcriptor First Strand cDNA Synthesis Kit (Roche). Quantitative qPCR was performed using PowerUp™ SYBR® Green Master Mix (Thermo Fisher) on a CFX96 real-time PCR detection (CFX96; Bio-Rad) system. Specific primer sequences used can be found in Supplementary Table S3. The 2(−△△CT) method was used to calculate the relative expression levels. Samples were analyzed in triplicate.
Fluorescence in situ hybridization
To visualize the subcellular location of circTRIO in LSCC cells, fluorescence in situ hybridization (FISH) was performed using a FISH kit (GenePharma, Shanghai, China), according to the manufacturer's instructions. After prehybridization at 75°C for 10 min, the cells were hybridized with Cy3-labeled circ-TRIO probes at 37°C for 30 min. Nuclei were stained with DAPI and images were then acquired on a fluorescence microscope (Leica Microsystems, Germany).
Cell counting kit-8
CCK8 was used to assess cell proliferation viability. Forty eight hours post-transfection, tumor cells (1 × 103) were digested and seeded into 96-well plates for incubation at 37°C with 5% CO2 (each well in quintuplicate). A mixture of 100 μL DMEM/high glucose and CCK8 (Dojindo, Kumamoto, Japan) (9:1) was added and refreshed at regular intervals of 0, 24, 48, 72, and 96 h. Absorbance was measured at 450 nm using the Multiskan Go microplate reader (Thermo scientific).
Colony formation assay
Transfected cells (1 × 103/well) were transferred to six-well plates and maintained for 14 days. Subsequently, the cells were washed with PBS once, fixed with 4% paraformaldehyde for 20 min, and then stained with 0.1% crystal violet solution at room temperature for 20 min.
Transwell migration assays
LSCC cells were assessed by a Transwell cell migration assay with an 8-μm-pore polycarbonate membrane (Corning Life Sciences, Corning, NY). An FBS-free medium containing 1 × 105 LSCC cells was seeded into the top chamber. After a 24–48-h incubation period, the cells permeating into the inferior chamber (containing 600 μL of DMEM and 10% FBS) were fixed, stained, imaged, and quantified.
Flow cytometric apoptosis analysis by annexin-propidium iodide staining
The annexinV-FITC apoptosis detection kit (Beyotime, China) was used in accordance with the manufacturer's protocols to detect cellular apoptosis. After transfection, the LSCC cells were digested by trypsin (without EDTA) and carefully washed with PBS for three times. After centrifugation at 1000 g for 5 min, the cell pellets were resuspended in 195 μL of annexin V binding buffer and gently mixed with an additional 5 μL of annexin V binding buffer and 10 μL of propidium iodide (PI). The suspension was then incubated in the dark for 10 min at room temperature. The Guava Easy Cyte System (Millipore) was used to measure the apoptotic rate.
Prediction of miRNA interaction
The targets of circTRIO were predicted by bioinformatic analysis via the online Circular RNA Interactome, circbank, and StarBase v2.0 database.
The The Cancer Genome Atlas (TCGA) LSCC data set including miRNA expression data and clinical information of 112 LSCC patients was downloaded. Differential analysis was performed using the miRNA expression data. The optimal prognostic cutoff was determined using the R package “survminer,” providing a value of a cutoff that corresponds to the most significant relationship with overall survival.
Statistical analysis
Statistical analysis was performed using GraphPad Prism 8.0 software (La Jolla). A two-tailed Student's t-test was used to compare the two groups. Correlations were analyzed by Pearson's correlation, and disease-free survival analysis was performed by Kaplan–Meier plots using the log-rank test. Results are presented as mean ± standard deviation, and p-values <0.05 were considered statistically significant.
Results
CircRNA expression profiling and screening for DECs
To elucidate the expression of circRNAs, RNA-sequencing was performed between tumor tissues and paracancerous tissues from five LSCC patients. A total of 9449 circRNA transcripts were identified from the sequencing data in total, where 5460 and 3989 were up- and downregulated, respectively, in this cohort. We identified 21 circRNAs, which were most up- or downregulated in tumor tissues, using the criterion of log2-fold change >1.5 (or <−1.5), with a q (padj)-value <0.05, with 14 circRNAs significantly upregulated and 7 circRNAs significantly downregulated (Fig. 1A). We then constructed a heatmap of the top 20 DECs in tumor and paracancerous tissues (Fig. 1B), in which the 7 upregulated circRNAs were chosen for further investigation. Reverse transcription-quantitative PCR (RT-qPCR) results revealed that hsa_circ_0005260 (circTRIO) was the most considerably elevated circRNA both in LSCC TU212 and TU686 cell lines (Supplementary Fig. S1).
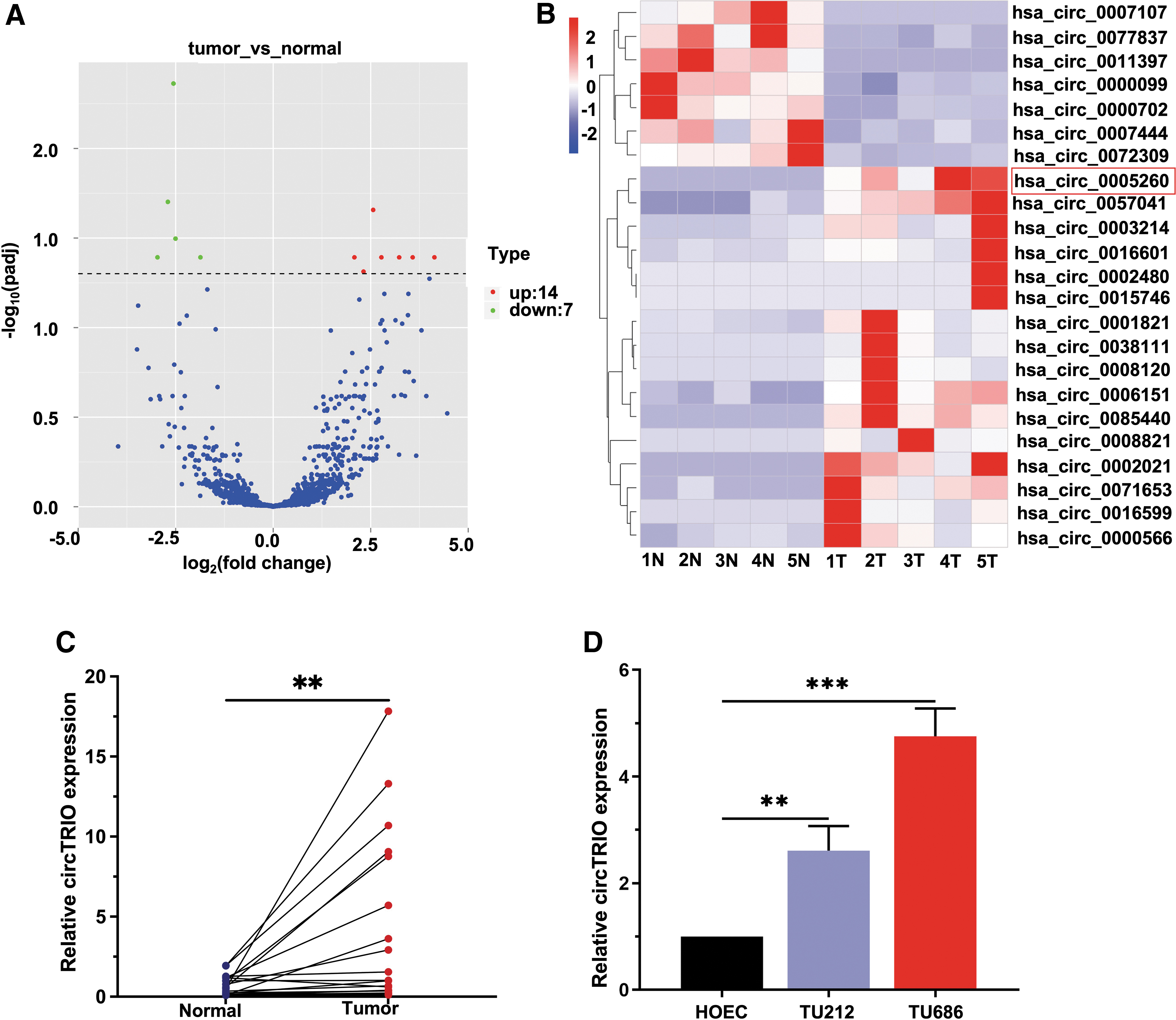
RNA sequencing of five pairs of LSCC and matched paracancerous tissues to screen differentially expressed circRNAs.
Next, the expression of seven upregulated circRNAs in addition to the LSCC tissues was determined and the results indicated that compared with paracancerous tissues, the expression of circTRIO was prominently elevated in LSCC tumor tissues (Fig. 1C, D, and Supplementary Fig. S2). Hence, we focused on this circTRIO in this study.
CircTRIO was upregulated in LSCC and correlated with clinicopathological factors
To investigate the correlation between circTRIO levels and LSCC, we analyzed the correlation between the expression of circTRIO and clinic characteristics in 20 pairs of tumor and paracancerous tissues. The expression level of circTRIO was significantly related to T stage, lymphatic metastasis, and clinical stage. Patients in the advanced stage and those with cervical lymph node metastasis had high expression levels of circTRIO (Table 1).
Correlation Between circTRIO Expression and Clinicopathological Characteristics in 20 Patients with Laryngeal Squamous Cell Carcinoma
p < 0.05, ** p < 0.01.
SD, standard deviation.
To further verify the expression of circTRIO in LSCC in other databases, we analyzed the GEO data sets GSE142083 (53 LSCC tissues paired with 53 adjacent normal mucosa tissues) and GSE27020 (training set: 59 samples; validation set: 50 samples). We found that the expression of circTRIO was significantly upregulated in tumor tissues compared with the matched surrounding tissues (Fig. 2A), this was consistent with the RNA sequencing and RT-qPCR analysis results. Kaplan–Meier survival analysis showed that patients with higher expression levels of circTRIO were associated with worse disease-free survival in these two LSCC cohorts (Fig. 2B). In addition, these LSCC patients also displayed an association between poor differentiation and high expression of circTRIO (Fig. 2C).
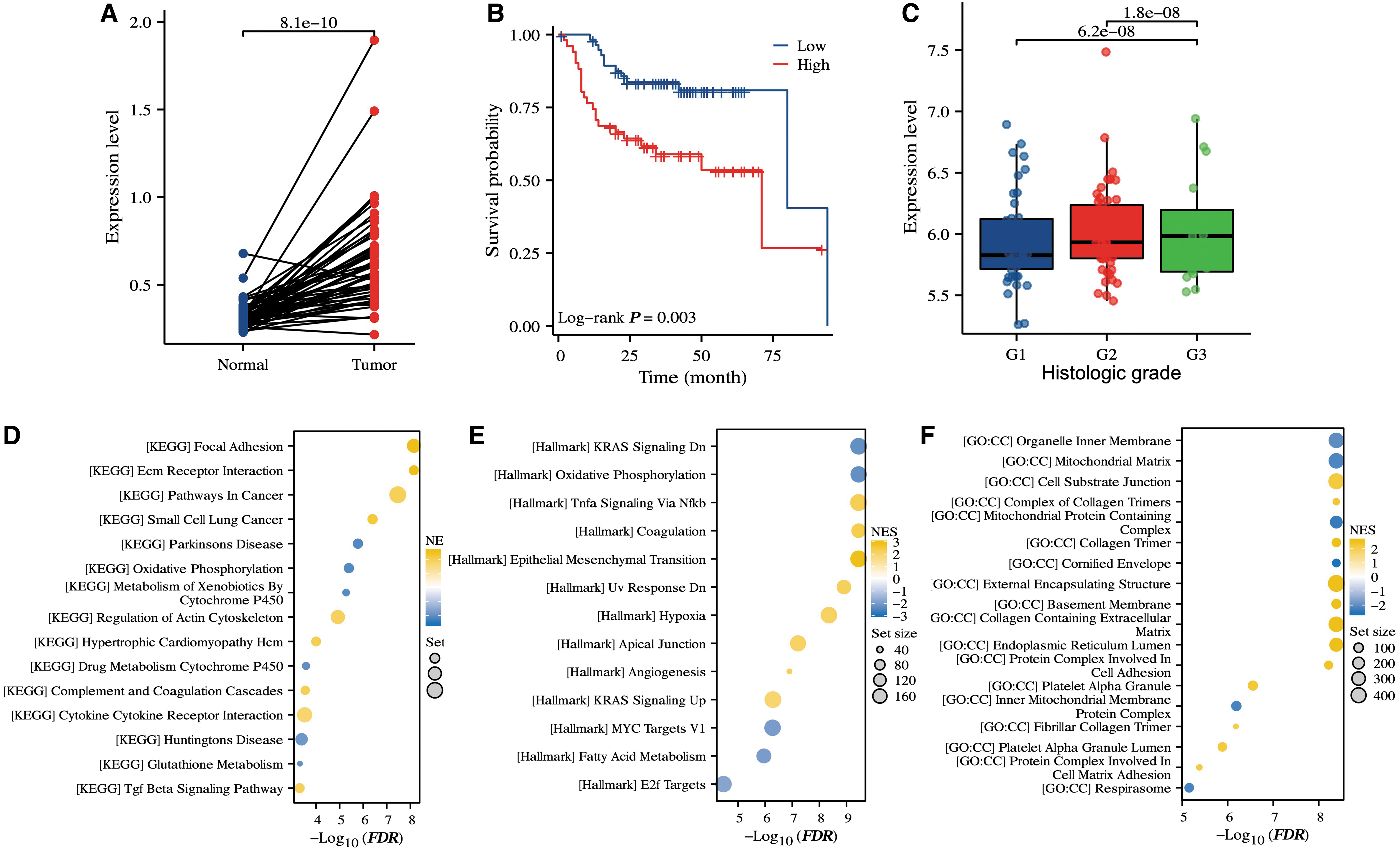
CircTRIO was upregulated in LSCC and correlated with clinicopathological factors.
To further understand the potential roles of circTRIO, we performed GSEA to evaluate the enriched biological signaling in circTRIOhi versus circTRIOlow tumors. The results demonstrated that increased expression of circTRIO was associated with cancer progression (i.e., MYC1), and metastasis (i.e., epithelial–mesenchymal transition) (Fig. 2D–F).
Genomic information of circTRIO
In this study, circTRIO (hsa_circ_0005260, chr5:14,316,621–14,336,836) was estimated as the third highly upregulated circRNA in LSCC, with a log2-fold change of 5.97 as reported by RNA-sequencing analysis. Following that, we validated the DECs with additional LSCC tissues and cells, and qPCR results confirmed that circTRIO was highly differentially expressed in LSCC. Via browsing the GRCh37/hg19 human reference genome and the circBase database, we pinpointed circTRIO's formation through the back-splicing three exons (exons 9, 10, and 11) of the triple functional domain (TRIO) gene (Fig. 3A). As a rho guanine nucleotide exchange factor, TRIO has been demonstrated to be closely related to the cytoskeleton dynamics and actin remodeling—its role in regulating cell migration and cytoskeleton reorganization has also been extensively studied (Peng et al., 2010). Increased m6A modification of TRIO mRNA and downregulation of TRIO are also closely associated with the process of myocardial ischemia/reperfusion injury (Li et al., 2022).
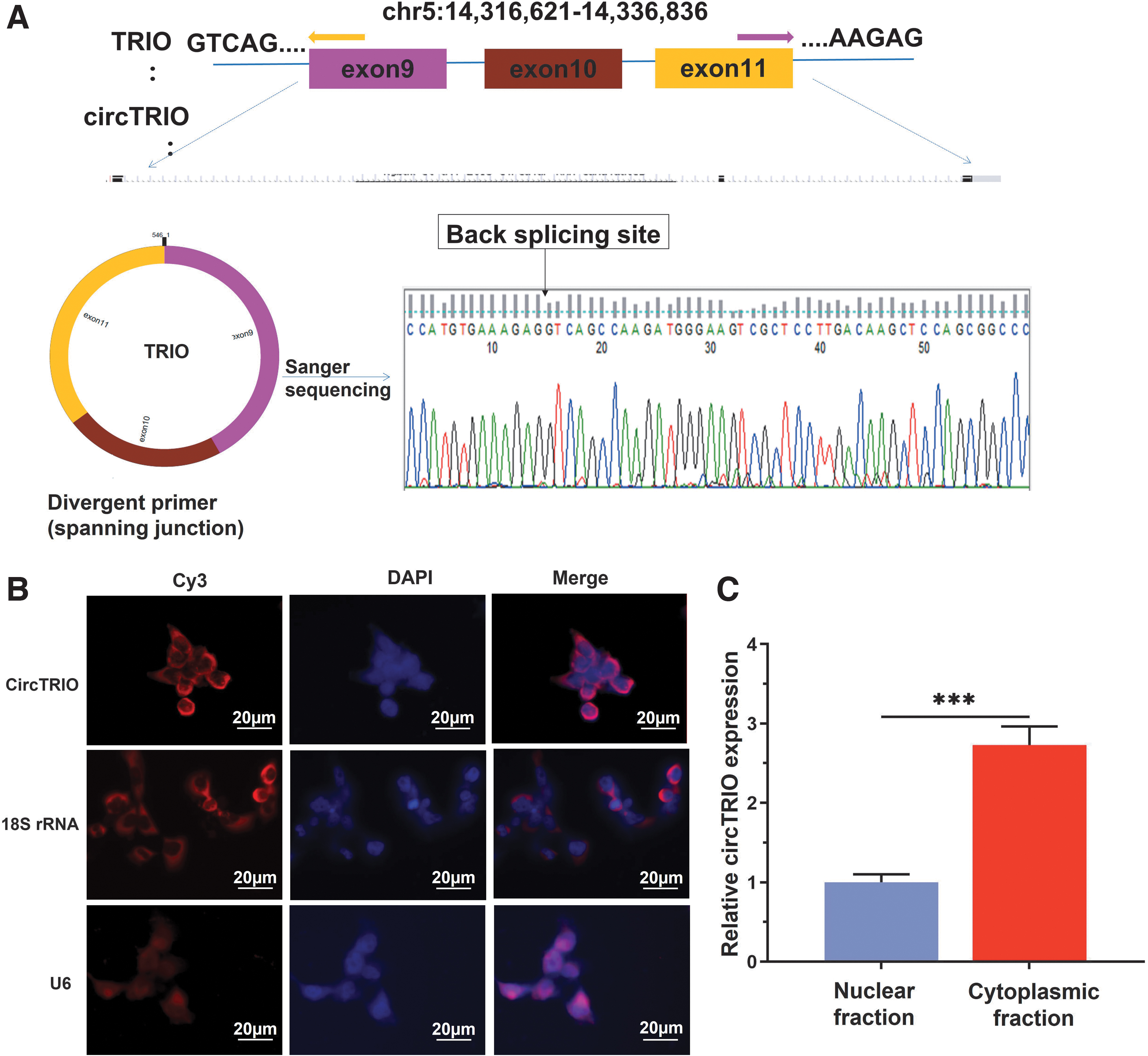
Genomic composition and origin of circTRIO.
To further confirm the prevalence of circTRIO in LSCC, we amplified circTRIO in LSCC cell lines and conducted Sanger sequencing. The result confirmed that the head-to-tail splicing in the reverse-transcription (RT)-PCR product was in conformity to the expected length and location of circTRIO, as shown in Figure 3A.
We subsequently performed FISH to investigate the intracellular location of circTRIO in LSCC cell lines, as the function of circRNA is typically related to its localization in cells. The results revealed that circTRIO was mainly localized in the cytoplasm of LSCC cells (Fig. 3B). Next, using the Cytoplasmic and Nuclear RNA Purification Kit (Norgen Biotek, Canada), cytoplasmic and nucleoplasmic RNA was isolated. Reverse transcription and qPCR were conducted to compare the expression of circTRIO in these two subdivisions. The results of qPCR further revealed that circTRIO was concentrated mainly in the cell's cytoplasm as opposed to its nuclei (Fig. 3C).
Downregulation of circTRIO inhibits both proliferation and colony-forming ability
To explore the biological functions of circTRIO in vitro, three small interfering RNAs (siRNAs) targeting the junction sites of circTRIO were designed to target the circTRIO junction sites and silence its expression in TU212 and TU686 cell lines. These siRNAs significantly decreased circTRIO expression as expected while having no effect on TRIO, its liner isoform (Fig. 4A, B). In addition, we introduced a negative control siRNA (si-NC) that did not target circTRIO. CircTRIO#3 was our silencing siRNA of choice for the subsequent knockdown experiments due to its high inhibitory efficiency. Finally, we tested for cell counts and colony formation with CCK8 and colony formation assays, respectively. The CCK8 assay showed that circTRIO knockdown markedly suppresses cell proliferation ability in TU212 and TU686 cell lines (Fig. 4C, D). Colony formation assay indicated that the silencing of circTRIO significantly inhibits the colony-forming ability of TU212 and TU686 (Fig. 4E).
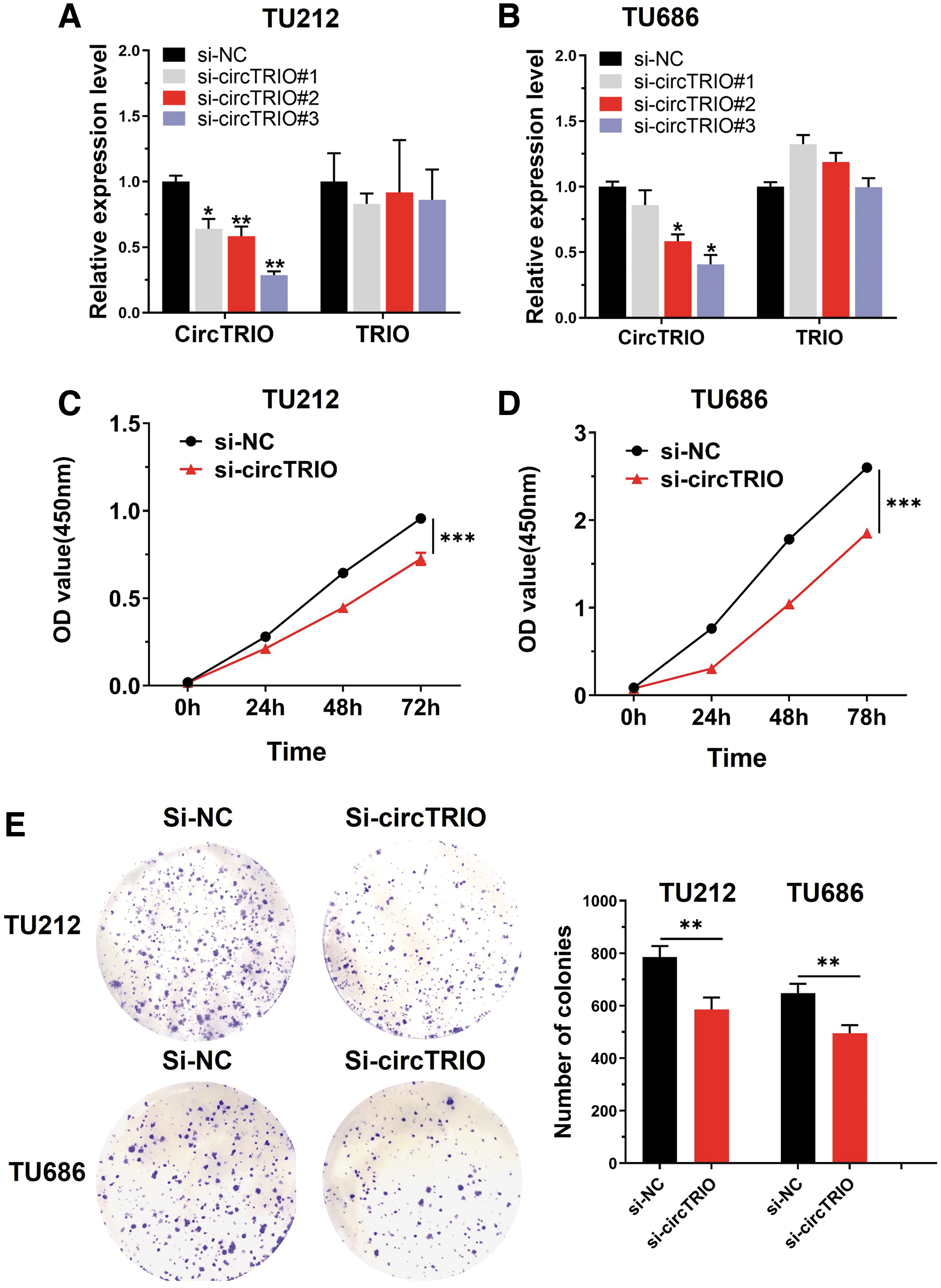
CircTRIO knockdown inhibits cell proliferation and cell colony forming ability.
Downregulation of circTRIO inhibits migration and promotes apoptosis of LSCC cells
We subjected post-transfection, TU212 and TU686 cells to a Transwell migration assay, illustrating that migration capabilities were suppressed in the si-circTRIO group compared with the si-NC group (Fig. 5A). Moreover, flow cytometry analysis showed that more apoptotic cells are presented in the si-circTRIO group of both LSCC cell lines, suggesting that the knockdown of circTRIO induced apoptosis (Fig. 5B). Collectively, our data indicate that silencing of circTRIO might moderate LSCC progression.
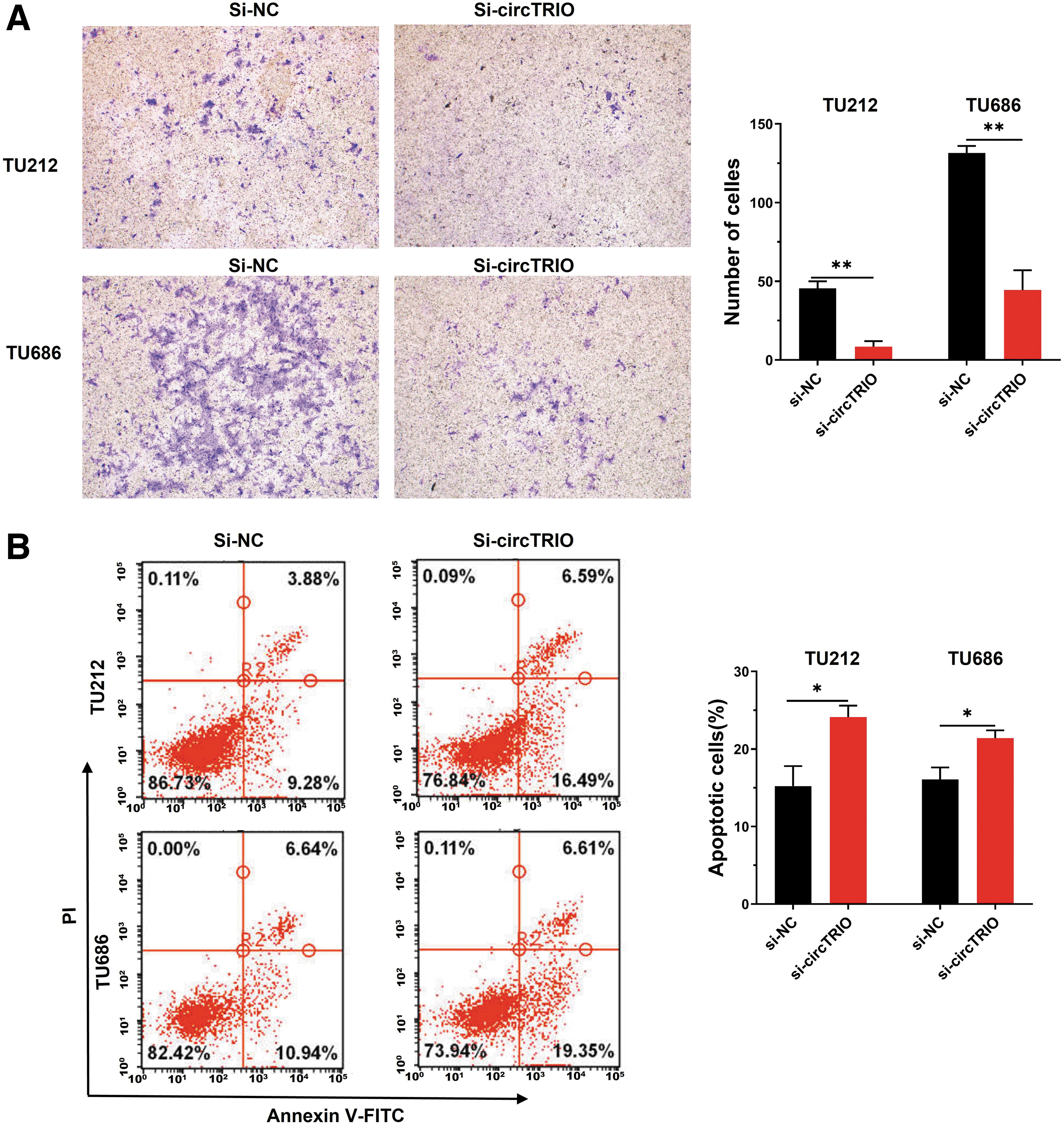
CircTRIO knockdown inhibits migration and promotes apoptosis.
The miRNA interactome of circTRIO
The ceRNA hypothesis (Karreth and Pandolfi, 2013) suggests an inverse relationship between circRNA and miRNA expression; indeed, it has been widely reported that circRNA functions as an “miRNA sponge” in many cancers (Hansen et al., 2013; Kristensen et al., 2019; Li et al., 2020a). To ascertain whether circTRIO could sponge miRNAs in LSCC cancer cells, three databases [the Circular RNA Interactome, CircBank (Liu et al., 2019), and StarBase (Li et al., 2014)] were used to predict the circTRIO-miRNA binding sites. Twenty-seven miRNAs from Circular RNA Interactome, 37 miRNAs from CircBank, and 24 miRNAs from StarBase databases were identified (Fig. 6A). Distinguishing overlapping miRNA produced two candidates: miR-432-5p and miR-370-3p. Figure 6B shows the potential circRNA-miRNA network formed by circTRIO and the above two miRNAs for better visualization. According to the CircInteractome database, circTRIO (hsa_circ_0005260) can directly combine with miR-432-5p and miR-370-3p (Fig. 6C).
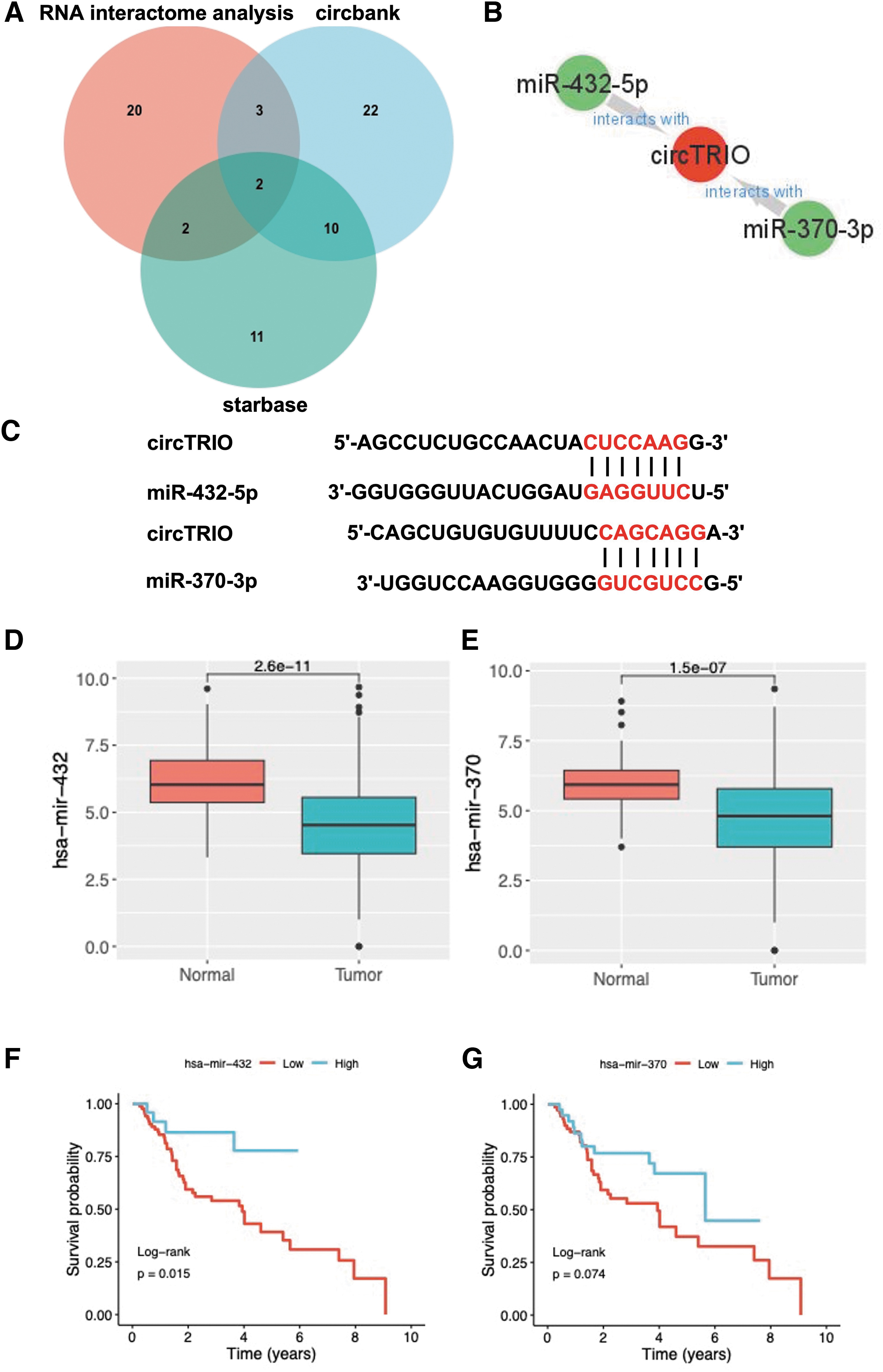
The miRNA interactome of circTRIO.
To determine the expression and prognosis of miR-432 and miR-370 in LSCC, we used miRNA expression data and clinicopathological data from LSCC patients in the TCGA database, and the results are shown in Figure 6D–G. The expression of miR-432 and miR-370 was significantly downregulated in LSCC tissues compared with normal tissues (p < 0.001) (Fig. 6D, E). The prognostic values of these two miRNAs in LSCC were also assessed through the Kaplan–Meier analysis in Figure 6F and G. It was observed that both low expressions of miR-432 and miR-370 were related to shorter survival times, with significantly worse survival in the miR-432 low-expression group than in the high-expression group (p = 0.015, Fig. 6F).
Discussion
Driven by the improvements of high-throughput sequencing technology and data analysis technique, studies of circRNAs (and noncoding RNA research as a whole) have become increasingly crucial (Caiment et al., 2015; Qu et al., 2015). As a major subtype of laryngeal cancer, LSCC continues to be the leading cause of tumor-related deaths worldwide. The study of circRNAs in LSCC was first conducted in 2016 (Xuan et al., 2016). Using microarray biochips, the researchers analyzed DECs in four pairs of LSCC tissue and the surrounding normal tissue. Rising evidence supports circRNA's considerable potential in the diagnosis, therapy, and prognosis of cancer and other diseases. However, there have been only a few functional circRNAs discovered in LSCC so far (Wu et al., 2021a; Yang et al., 2022).
In our study, five pairs LSCC tissues and normal tissues have been sequenced and screened for the abnormal expression of circRNA, revealing hsa_circ_0005260 as the most significantly upregulated molecule in the LSCC tissues. Furthermore, we confirmed higher circTRIO expression in LSCC tissues and cells compared with adjacent noncancerous tissues and normal cells with the help of two open-access GEO data sets, 20 pairs of fresh LSCC tissues, and cultured LSCC cell lines. We then discovered that the abnormal expression of circTRIO is also associated with the histological grade, T stage, lymphatic metastasis, clinical stage, and prognosis of LSCC, concluding that circTRIO expression may be specific in patients with LSCC. Hsa_circ_0005260, derived from exons 9–11 of the TRIO gene, was named as circTRIO and identified as a potential oncogenic regulator of triple-negative breast cancer (TNBC) in 2022 (Wang et al., 2022a).
As a triple functional domain protein, TRIO has been implicated in numerous cancerous processes in recent years. For example, some studies have shown that TRIO can regulate the migration and invasion of colorectal cancer and cervical cancer (Hou et al., 2018; Sonoshita et al., 2015). Up to now, important roles of circTRIO have only been identified in the development of osteoarthritis and TNBC (Yang et al., 2022). Nevertheless, the expression and function of circTRIO in LSCC and other cancers remain unclear.
Several studies have already shown that circRNAs control the proliferation and migration of tumor cells via sponging miRNA. CircPARD3 stimulates cell growth in LSCC by interacting with miR-145-5p, while CircMYLK activates cyclin D1 by interacting with miR-195, both of which enhance tumor cell proliferation (Duan et al., 2019; Gao et al., 2020). In addition, the miR-145-5p/OTX1 regulatory axis can be used to decrease metastatic function following circMYOF knockdown (Wang et al., 2022b). Our study investigated the functional role of circTRIO in LSCC. After circTRIO knockdown with siRNA, CCK-8, colony-forming, Transwell migration, and flow cytometry assays showed that its knockdown restrained LSCC proliferation, colony-forming ability, and migration, coupled with promoted apoptosis—collectively suggesting circTRIO's pivotal role in LSCC progression. As cancer progression remains the main factor of an unsatisfactory outcome in cancer patients, uncovering the fundamental mechanisms of cancer proliferation and metastasis is crucial for cancer therapy.
As previously mentioned, circRNAs can act as a translation regulator as well as an miRNA sponge. To better understand the role and mechanism of circTRIO regulation, three databases were used to predict the binding site of circTRIO with miRNAs. MiR-432-5p and miR-370-3p were predicted in all three databases, showing their high affinity for circTRIO. Therefore, we suggest that miR-432-5p and miR-370-3p are the two miRNAs most likely to function as sponge miRNAs for circTRIO. According to the ceRNA hypothesis, miRNAs negatively regulate the expression of circRNAs. The result shows that miR-432 and miR-370 were both downregulated and correlated with the overall survival in LSCC. The difference in miR-432 was especially significant, unsurprisingly, as miR-432 has been reported to play a role in several cancers (Wang et al., 2022a).
In LSCC tissues, miR-432 was found to be downregulated, and its artificial knockdown increased cell proliferation and migration (Zhao et al., 2022). We believe that future research can benefit from exploring the role of circTRIO as an miRNA sponge.
Conclusions
In this study, we demonstrated that circTRIO expression was upregulated in LSCC tissues. Moreover, circTRIO was implicated in LSCC cell proliferation, migration, and apoptosis, suggesting the potential of circTRIO as a novel biomarker for LSCC. Meanwhile, circTRIO may assist in the development of targeted molecular therapy as an effective therapeutic target for patients with LSCC.
Footnotes
Authors' Contributions
J.W. and H.L.: Conceptualization and methodology; J.W. and S.H.: Data curation and writing—original draft preparation; J.W., D.W., Y.J., and W.S.: Visualization and investigation; H.L.: Supervision; J.W., A.H., and Z.Z.: Writing—reviewing and editing.
Data Availability
The data sets used and/or analyzed during the current study are available from the corresponding author upon request.
Ethics Approval and Consent to Participate
The present study was approved by the Ethics Committee of Beijing Tongren Hospital (Beijing, China; approval No. 20190215). All patients provided written informed consent and agreed to participate in the study.
Disclosure Statement
No competing financial interests exist.
Funding Information
The present study was supported by the Beijing Municipal Education Commission (grant no. KZ201910025033) and Beijing Municipal Administration of Hospitals Clinical Medicine Development of Special Funding (grant no. ZYLX201814).
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
