Abstract
Renal ischemia/reperfusion injury (IRI) is a main risk factor for the occurrence of delayed graft function or primary graft nonfunction of kidney transplantation. However, it lacks ideal molecular markers for indicating IRI in kidney transplantation. The present study is to explore novel candidate genes involved in renal IRI. Experimental renal IRI mouse models were constructed, and the differentially expressed genes were screened using a microarray assay. Gene Ontology and Kyoto Encyclopedia of Genes and Genomes enrichment analysis were performed. The expression of genes was detected using real-time qPCR assay. Western blotting and immunohistochemistry staining assays were performed for protein determination. We identified that renal IRI induced the upregulation of SPRR2F, SPRR1A, MMP-10, and long noncoding RNA (lncRNA) Malat1 in kidney tissues for 479.3-, 4.98-, 238.1-, and 3.79-fold, respectively. The expression of miR-139-5p in kidney tissues of IRI-treated mice was decreased to 40.4% compared with the sham-operated mice. These genes are associated with keratinocyte differentiation, regeneration and repair of kidney tissues, extracellular matrix degradation and remodeling, inflammation, and cell proliferation in renal IRI. Identification of novel biomarkers involved in renal IRI may provide evidences for the diagnosis and treatment of renal IRI.
Introduction
Kidney transplantation is an effective therapy for most of the end-stage renal disease patients. As the extending criteria of deceased kidney donors, the rates of delayed graft function (DGF) or primary graft nonfunction (PGN) are increasing. The occurrence of DGF and PGN can be caused by many aspects, such as ischemia/reperfusion injury (IRI), immunological rejection, primary diseases of the recipients, and the quality of the graft. However, there is no effective treatment for DGF and PGN after kidney transplantation.
IRI is one of the most important but unavoidable risk factors for causing the impairment of graft function. Renal IRI triggers the generation of reactivate oxygen species, causing oxidative injury, acute inflammation and eventually necrosis, and apoptosis of renal tubular epithelial cells (Gu et al., 2018; Situmorang and Sheerin, 2018). These responses can evoke the anti-inflammatory and repair processes to initiate the recovery of the graft function (Sanchez-Nino et al., 2016; Pressly and Park, 2017). On the contrary, many factors released during the IRI, such as transforming growth factor-β, matrix metalloproteinases (MMPs), and altered extracellular matrix (ECM) components, can induce epithelial–mesenchymal transition, ECM remodeling, and interstitial fibrosis (Jain et al., 2000; Danobeitia et al., 2017; Kim et al., 2017; Liu et al., 2018), which hinder the recovery of complete function. Therefore, IRI is a complex and multistep pathological process that the mechanism has not been fully clarified yet. Identifying novel targets of renal IRI is crucial for exploring the diagnosis and treatment of DGF or PGN after kidney transplantation. In the present study, we screened for differentially expressed genes in renal IRI based on an experimental IRI model.
Materials and Methods
Animal studies
All the studies related to animals were approved by the Beijing Friendship Hospital Animal Care and Use Committee (approval no.: 2018-P2-167-01). The renal IRI experiments were performed as previously described (Hesketh et al., 2014). In brief, 8-week-old male C57/BL6 mice were anesthetized through intraperitoneal injection of pentobarbital sodium (0.05 mg/g body weight). The mice were randomly divided into injury and sham-treated groups. For IRI treatment, the mice were subjected to 45 min of ischemia by bilaterally clamping the renal artery. After 24 h of reperfusion, the mice were euthanized and the kidney samples were immediately frozen at −80°C or fixed in 4% paraformaldehyde for further analysis.
Microarray assay
Total RNA from the kidney samples was extracted using TRIzol reagent (Thermo Fisher Scientific, Schuylerville, NY). The differentially expressed genes were detected using an Affymetrix Mouse Genome 430 2.0 microarray (Thermo Fisher Scientific) from three mice of renal IRI and three sham-treated mice. The experimental procedures for microarray were performed at the Shanghai Biotechnology Corporation (Shanghai, China). The differential expression genes fulfill at least twofold change and p < 0.05 were considered for further analysis. Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis were performed.
RNA extraction and real-time qPCR analysis
Total RNA from kidney samples was extracted with the miRNeasy Mini Kit (Qiagen, Hilden, Germany). For quantitative detection of mRNA or lncRNA, the complementary DNAs (cDNAs) were synthesized with a cDNA Synthesis Kit (Takara, Dalian, China). The real-time qPCR was performed by using the following specific primers: MMP-10, 5′-TCCAGGAATTGAGCCACAAG-3′ and 5′-GCATTGGGGTCAAACTCGAAC-3′; SPRR1A, 5′-GAGAACCTGCTCTTCTCTGAGT-3′ and 5′-CAAGGTTCCTGGGGTGGTG-3′; SPRR2F, 5′-GGAGAACCTGATCCTGAGGC-3′ and 5′-CCCTGTCCTGAAGACTGCTG-3′; Malat1, 5′-TGCAGTGTGCCAATGTTTCG-3′and 5′-GGCCAGCTGCAAACATTCAA-3′. GAPDH was used as an internal control, and the primers are: 5′-GGCAAATTCAACGGCACAGT-3′ and 5′-AGATGGTGATGGGCTTCCC-3′. For the detection of mature form of miRNA-139-5p, the specific primer containing a stem-loop structure (5′-CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGACTGGAGA-3′) was used for reverse transcription, and the specific primers were used for amplification: 5′-ACACTCCAGCTGGGTCTACAGTGCACGTGTC-3′ and 5′-TGGTGTCGTGGAGTCG-3′. The ribosomal U6 was used as an internal control for normalization: 5′-CTCGCTTCGGCAGCACA-3′ and 5′-AACGCTTCACGAATTTGCGT-3′.
Immunochemistry staining
Kidney tissues fixed in 4% paraformaldehyde were dehydrated and embedded in the paraffin. The samples were cut into 4 μm thick sections. The sections were then dewaxed and rehydrated. The antigen was repaired by a heating process, and the endogenous peroxidases were blocked with 3% H2O2 for 20 min at room temperature. After washed with phosphate-buffered saline (PBS) for three times, the sections were incubated with rabbit anti-MMP-10 primary antibody (BBI; Sangon Biotech, Shanghai, China) at 4°C overnight. After that, the sections were washed three times with PBS and incubated in an HRP/Fab polymer-conjugated secondary antibody (ZSGB-BIO, Beijing, China) for 30 min at room temperature. The sections were then washed for five times with PBS and were developed using a diaminobenzidine reagent. The nuclei were counter stained with hematoxylin and visualized with a microscopy.
Protein extraction and western blotting
Total protein from tissue samples were extracted with RIPA lysis buffer containing protease and phosphatase inhibitors. The lysates were centrifuged with 12,000 round per minute at 4°C for 10 min. The supernatants were collected and quantified using bicinchoninic acid (BCA) assay (Thermo Fisher Scientific). The protein samples were normalized and diluted at a final concentration of 5 μg/μL. For electrophoretic separation, a total of 50 μg proteins from each sample were loaded onto a sodium dodecyl sulphate (SDS) polyacrylamide gel. After the electrophoresis, the proteins on the SDS polyacrylamide gel were blotted onto a nitrocellulose membrane and then blocked with 5% nonfat milk at room temperature for 1 h. After that, the membrane was incubated with specific antibodies: rabbit anti-MMP-10 (BBI; Sangon Biotech) and mouse anti-β-actin (Santa Cruz, Dallas, TX). The membrane was washed with TBS-T and incubated with HRP-conjugated goat anti-rabbit or mouse secondary antibodies (ZSGB-BIO) for 1 h at room temperature. After washing with TBS-T for four times, the specific bands were developed using Super Signal West Femto Maximum Sensitivity Substrate (Pierce, Rockford, IL) and scanned using Quantity One software V 4.6.2 (Bio-Rad, Hercules, CA). The expression of β-actin was used as an internal control.
Statistical analysis
All the statistical calculations were performed with SPSS 19.0 software (SPSS, Inc., Chicago, IL). A two-tailed Student's t-test was used for analyzing the differences between two groups. p < 0.05 was considered as statistically significant.
Results
Screening for differential expression genes in renal IRI
Previously, we found 2218 differentially expressed genes, including 1103 upregulated genes and 1115 downregulated genes by using microarray assay (Fig. 1a; Hu et al., 2019). In the present study, GO and KEGG enrichment analyses were performed, and the top 30 enrichments were listed, respectively (Fig. 1b, c, complete data see Supplementary Tables S1 and S2). We found that many essential pathways were associated with renal IRI, such as tumor necrosis factor signaling pathway, MAPK signaling pathway, Hif-1 signaling pathway, ECM-receptor interaction, focal adhesion, keratinocyte development, Toll-like receptor signaling, IκB/NF-κB signaling pathway, positive regulation of pri-miRNA transcription, and metanephric nephron tubule development.
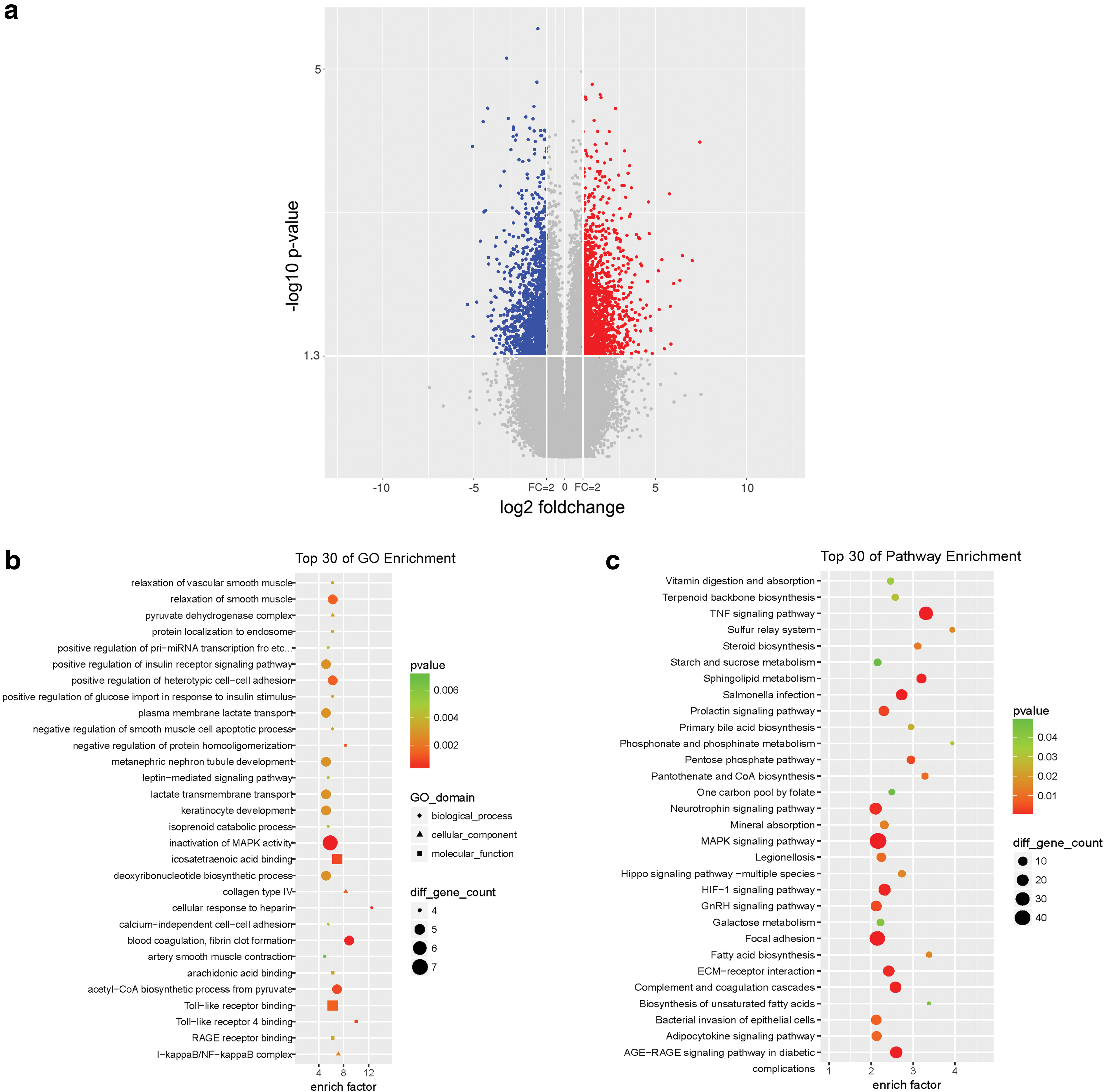
Screening for differentially expressed genes in renal IRI.
SPRR2F and SPRR1A is upregulated in renal IRI
Keratinocytes are known to participate in IRI, and the keratinocyte-derived chemokine is a known biomarker for indicating early renal injury before the increase of serum creatinine (Molls et al., 2006). Data from GO enrichment analysis (Fig. 1b) also supported that genes-related keratinocytes might be potential biomarkers of IRI. By analyzing these differentially expressed genes, we found that the expression of SPRR2F and SPRR1A, annotated as “keratinocytes differentiation,” was significantly upregulated in the kidney tissues of IRI mice. The results from real-time qPCR consistently verified that the expression of SPRR2F in kidneys of IRI mice was 479.3-fold higher than the sham-operated mice (Fig. 2a). Likewise, the expression of SPRR1A increased up to 4.98-fold of the sham-treated group (Fig. 2b). Our data indicated that the SPRR2F and SPRR1A might be involved in IRI.
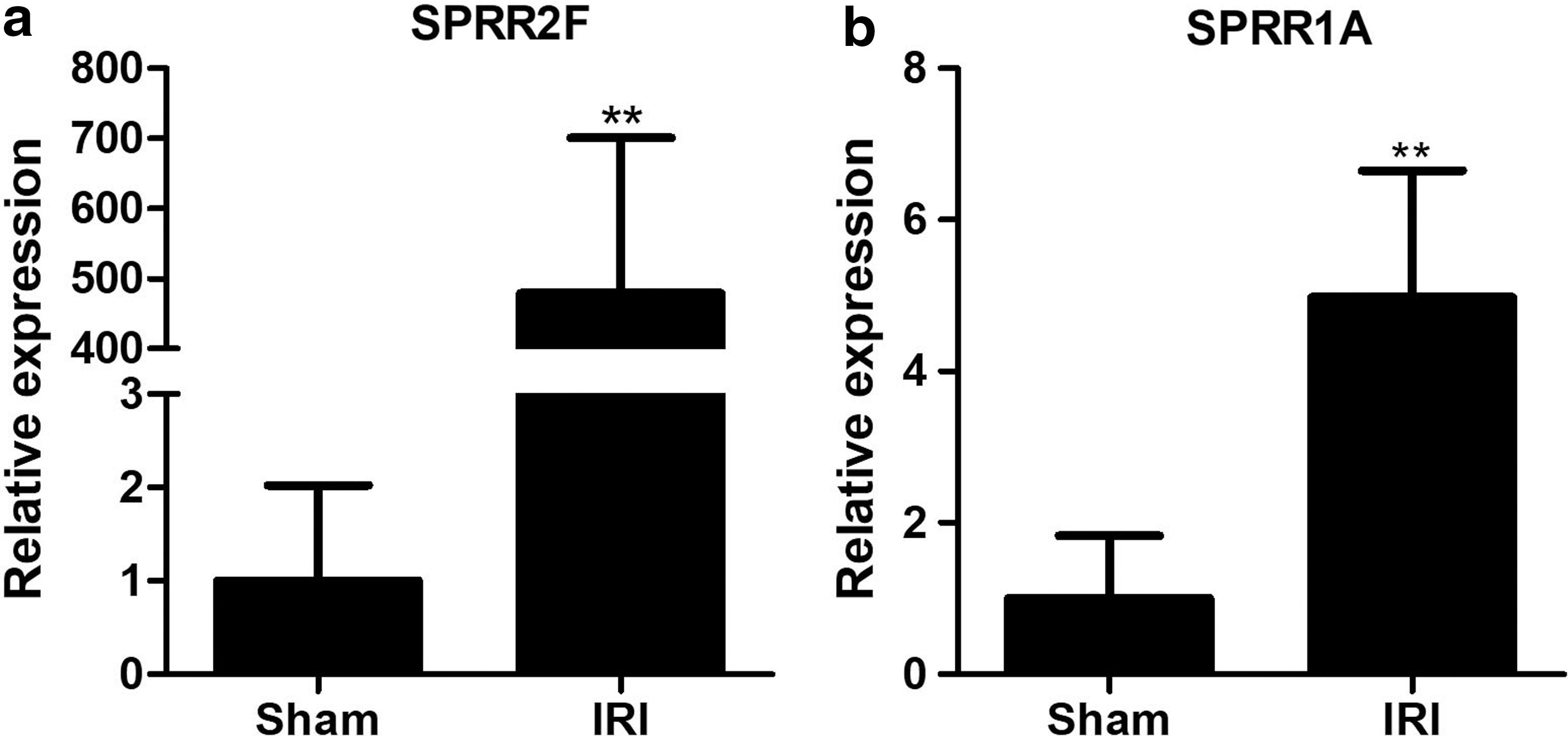
SPRR2F and SPRR1A are upregulated in IRI.
The expression of MMP-10 is activated in renal IRI
ECM degradation and renal remodeling are important pathological changes of IRI hindering the recovery of kidney. During the acute phase, breakdown of normal structures of ECM aggravates kidney injury, whereas the interstitial fibrosis impairs kidney function in the chronic phase. From the data of the differentially expressed genes of IRI, we found that MMP-10, an enzyme mediating the degradation of ECM, was significantly upregulated by IRI treatment. By using real-time qPCR, we observed that MMP-10 was highly expressed in all the mice undergone IRI operation (Fig. 3a). In contrast, MMP-10 is undetectable in most of the mice of the sham-operated group. The products from real-time qPCR were visualized through an ultraviolet imager (Fig. 3b).
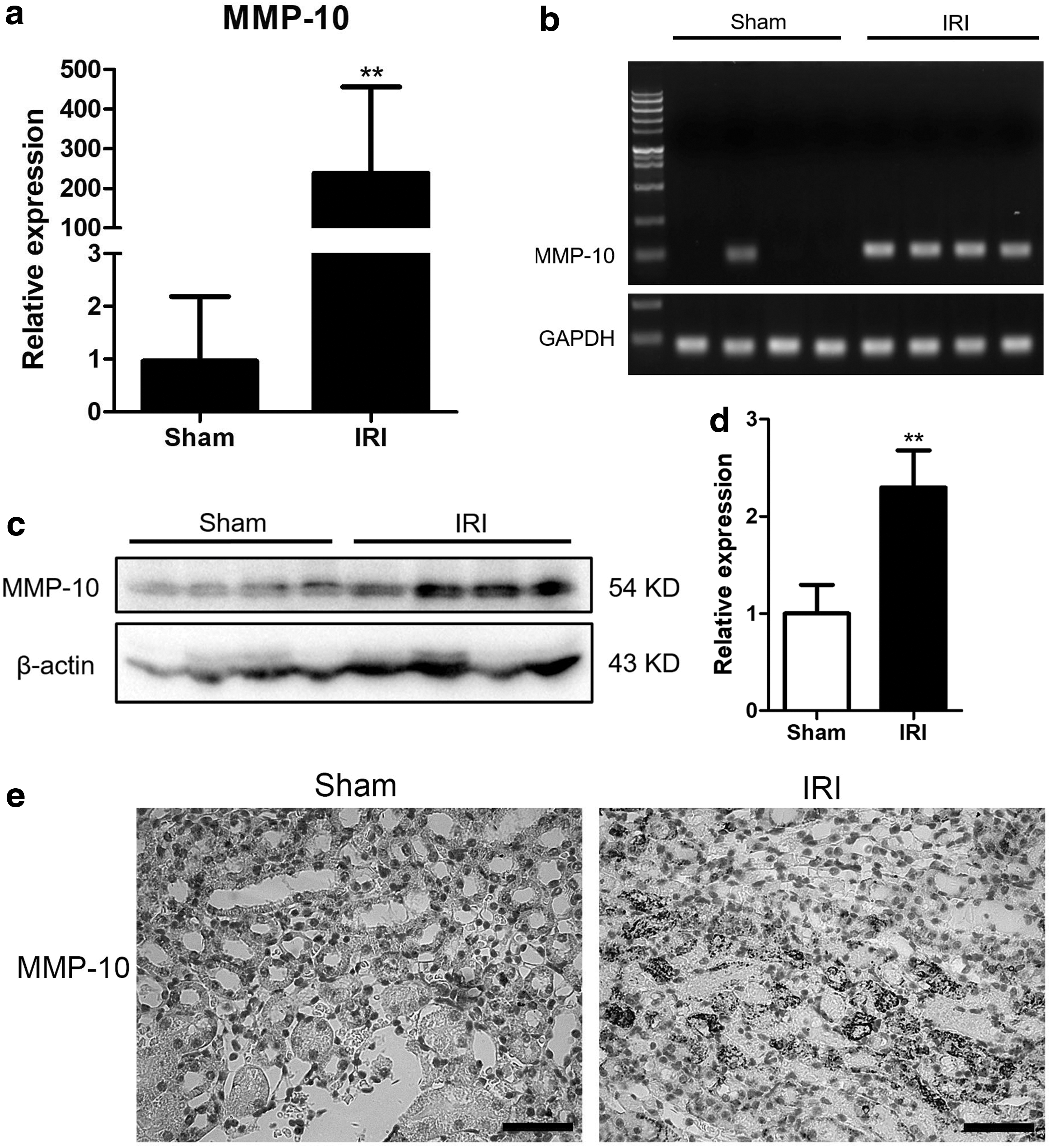
MMP-10 is induced by renal IRI.
We then analyzed the expression of MMP-10 at protein levels, using western blotting. We found that MMP-10 was indeed upregulated in kidneys of IRI-treated mice (Fig. 3c, d). Interestingly, MMP-10 was presented in kidney tissues of sham-operated mice, which is not paralleled with the results of its mRNA level. Consistently, data from immunohistochemistry staining indicated that MMP-10 was strongly stained in the injury area of the kidney, but weakly stained in kidneys of sham-operated mice (Fig. 3e). Our data suggested that MMP-10 might be a potential molecular candidate participating in renal IRI.
The expression of Malat1 and miR-139-5p in renal IRI
Noncoding RNAs, including miRNAs, lncRNAs, and circular RNAs (circRNAs), play important roles in many biological processes. Our data of microarray screened that the lncRNA Malat1 was upregulated in IRI. The real-time qPCR verified that Malat1 in kidneys of IRI mice was increased to 3.79-fold compared with the sham-operated mice (Fig. 4a). In addition, we detected the expression of miR-139-5p, which is not included in the microarray data but closely associated with cell proliferation. IRI treatment repressed the renal miR-139-5p expression to 40.4% of the sham-operated mice (Fig. 4b). These data demonstrated that both noncoding RNAs Malat1 and miR-139-5p were involved in IRI.
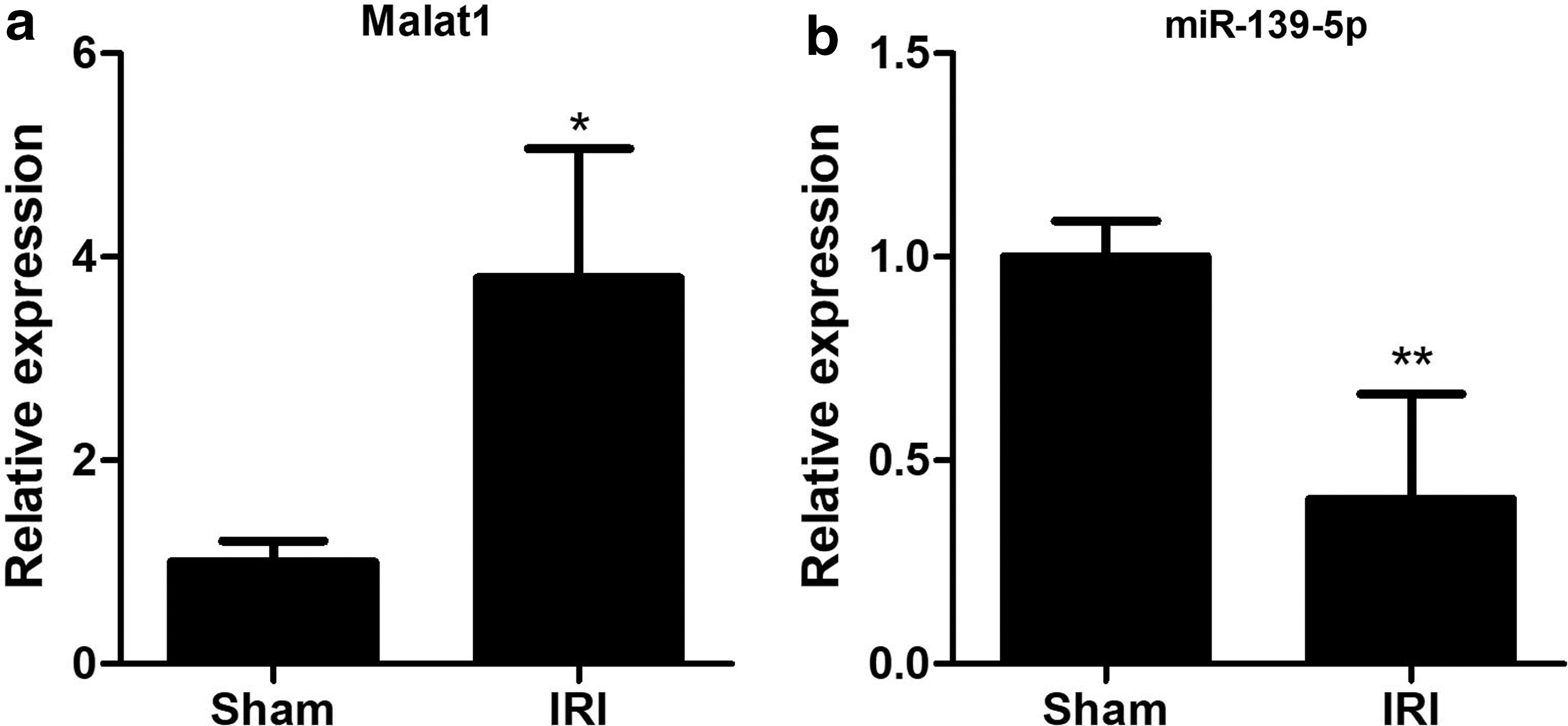
The expression of lncRNA Malat1 and miR-139-5p in kidneys of IRI.
Discussion
Renal IRI is a main risk factor for the occurrence of DGF and PGN in kidney transplantation. IRI is a multistep pathological process and is currently not fully clarified. Therefore, it lacks an ideal biomarker for the diagnosis and treatment. Recipients undergo IRI of the graft frequently requires a redialysis, causing heavy burden both physically and mentally. Therefore, identification of novel molecular biomarkers is important for the diagnosis and treatment of DGF and PGN. Previously, we screened differentially expressed genes in experimental IRI models through a microarray assay. In the present study, we used these data to perform enrichment analysis and identified that SPRR2F, SPRR1A, MMP-10, and Malat1were significantly altered in our experimental models, indicating that these genes may be involved in renal IRI.
SPRR2F and SPRR1A belong to the small proline-rich protein family, which are both important markers of keratinocyte terminal differentiation (Sark et al., 1998; Chen et al., 2010). SPRR2F is highly expressed in uterus and ovary (Song et al., 1999; Cabral et al., 2001) and briefly expressed during nephrogenesis (Schwab et al., 2003). We observed that SPRR2F was seldom expressed in normal kidney tissues, but could be strongly induced by IRI treatment. The function of SPRR2F remains far from understanding. In contrast, SPRR1A, another member of SPRR family, is well profiled and is known to be associated with many diseases, such as breast cancer and diffuse large B cell lymphoma (Zhang et al., 2014; Chen et al., 2015). SPRR1A is a gene related with tissue regeneration (Gaudet et al., 2016; Gey et al., 2016; Danaher et al., 2018) and can be controlled by ATF3 (Danaher et al., 2018). In the present study, we found an increased expression of SPRR1A in kidneys of IRI-treated mice. Therefore, we speculate that SPRR2F and SPRR1A are biomarkers of kidney repair and regeneration after IRI, which may reflect the recovery of the kidney function. However, the detailed role of SPRR2F and SPRR1A in renal IRI remains to be investigated.
Most of the MMPs are secreted enzymes mediating ECM degradation. MMP-10 (also known as stromelysin 2, SL-2) belongs to the stromelysins subgroup of MMPs, catalyzing substrates include collagen IV, laminin, fibronectin, and elastin. MMP-10 was reported to be associated with atherosclerosis (Purroy et al., 2018), intracerebral hemorrhage (Howe et al., 2018), pulmonary hypertension (Avouac et al., 2017), and cancers (Miyata et al., 2007; Du et al., 2014; Kadeh et al., 2016; Juchniewicz et al., 2017). Recently, Guvercin et al. reported that the increase of serum MMP-10 is related to acute kidney injury in nondiabetic geriatric patients (Guvercin et al., 2018). Our data demonstrated that the expression of MMP-10 in normal kidney tissues is very weak, but could be strongly activated by IRI treatment, supporting that MMP-10 is a potential candidate involved in renal IRI. Interestingly, we observed that the expression of MMP-10 at mRNA level was not paralleled with its protein level in normal kidney tissues. Because mature form of MMP-10 enzyme can be secreted into the circulation, we postulate that the circulatory MMP-10 in the plasma may be recruited to the kidney, causing the present of MMP-10 protein in the kidney tissues. Due to the stromelysin activity, the upregulation of MMP-10 in tissues results in the degradation and remodeling of the ECM (Avouac et al., 2017), further causing intestinal fibrosis and thus hindering the recovery of kidney function (Sokai et al., 2015). Our data indeed showed a significant accumulation of MMP-10 in the injured area of the kidney, indicating that tissue MMP-10 may be a potential biomarker to reflect the severity of tissue injury. We speculate that MMP-10 may be a potential therapeutic target for acute kidney injury.
From the data of the differentially expressed genes, we found that lncRNA Malat1 was upregulated in kidneys of IRI-treated mice, which has been reported from other groups (Kölling et al., 2018; Tian et al., 2018). However, these reports remain controversy. Tian et al. reported that the increase of Malat1 inhibits hypoxia-induced inflammation (Tian et al., 2018), whereas Kölling, et al. speculated Malat1 as dispensable (Kölling et al., 2018). Our data support that Malat1 participates in IRI, but its role remains to be clarified. In addition to the high throughput data, we identified that miR-139-5p was decreased in renal IRI. miR-139-5p was reported to inhibit tumorigenesis and reverses drug resistance in many cancers (Wang et al., 2017; Jiang et al., 2018; Zhang et al., 2018). Targets of miR-139-5p, such as c-Jun (Jiang et al., 2018; Su et al., 2018), PI3K/Akt (Maoa et al., 2015; Catanzaro et al., 2018), IGF-1/IGF-1R (Xu et al., 2015), VEGF, and Wnt/β-catenin (Long et al., 2017; Xie and Shen, 2018), are potent stimulators of cell proliferation, some of which are important pathways promoting injury repair (Zhou et al., 2016; Guise and Chade, 2018; Xie et al., 2018). Therefore, we postulate that the downregulation of miR-139-5p may reflect an activation of cell proliferation, promoting the repair and regeneration of injured kidney.
Taken together, our data indicated that the expression of SPRR2F, SPRR1A, MMP-10, Malat1, and miR-139-5p in kidney was altered by IRI, which may serve as potential biomarkers for renal IRI. These findings may also provide evidences for exploring novel targets for the prevention and treatment of DGF and PGN after kidney transplantation.
Footnotes
Acknowledgments
This study was supported by the National Natural Science Foundation of China (No. 81602214 and 81500236) and Beijing Municipal Administration of Hospitals Clinical Medicine Development of Special Funding Support (ZYLX201604).
Disclosure Statement
No competing financial interests exist.
Supplementary Material
Supplementary Table S1
Supplementary Table S2
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
