Abstract
The endomembrane system plays an important role during cellular adaptation of the plants with the extracellular environment. The small GTP-binding protein Rab7 located at the vacuolar membrane regulates the vesicle fusion with the vacuole and thereby helps in recycling of the molecules. This is the first report on isolation and characterization of AlRab7 gene from the halophyte plant, Aeluropus that extrudes NaCl through salt glands and grows luxuriantly throughout the year at the Gujarat coast, India. The AlRab7 encodes a protein with 206 amino acids, and a highly conserved effector-binding domain and four nucleotide-binding domains. The in silico analysis predicts the presence of the prenylation site for Rab geranylgeranyltransferase 2 and the Rab escort protein site. The C-terminal two cysteine residues in –XCC sequence are present for membrane attachment. Transcript expression of the AlRab7 gene was differentially regulated by different environmental stimuli such as dehydration, salinity, and hormone abscisic acid (ABA). The recombinant Escherichia coli cells showed improved growth in Luria Bertani medium supplemented with NaCl, KCl, mannitol, ABA, and indole-3-acetic acid. A novel Rab7 interacting partner AlRabring7 was identified by yeast two-hybrid screening.
Introduction
P
The plant endomembrane system consists of several biochemically distinct membrane-bound organelles that are responsible for the biosynthesis of proteins, polysaccharides, and lipids. In addition to its role in the biogenesis of the cell wall, plasma membrane, and vacuole, the endomembrane system contributes to the control of development and toward response to biotic and abiotic stresses (Surpin and Raikhel, 2004). The relationship between abiotic stress signaling and endocytic traffic is not understood (Vieria et al., 1996). However, its involvement in abiotic stress regulation has been indicated by NtSyr1, an important component of vesicle trafficking in eukaryotes, which is abscisic acid (ABA)-dependent (Leyman et al., 1999).
Vesicular trafficking of soluble and membrane proteins begins with the budding of transport vesicles from donor membranes and ends with their fusion to target organelles. A single round in the transport process (from vesicle budding to its fusion) basically involves two classes of Ras-like, small GTPases known as SAR/ARF, which act on the budding vesicle and Rab, which function in targeting and/or tethering transport vesicles to acceptor compartments. The plant Rab GTPases constitute the largest of the RAS-related super families of small GTPases (Yang, 2002; Nielsen et al., 2008). These small GTPases show conservation throughout eukaryotes and has expanded in the more complex animal and plant systems. The Rab GTPase was one of the first proteins, identified as Ypt in Saccharomyces cerevisiae and found to be involved in the regulation of vesicular trafficking (Gallwitz et al., 1983). The term “RAB” protein was coined as it was identified in rat brain cDNA libraries (Touchot et al., 1987). The 60 Rab proteins have been described in mammals and 57 in the Arabidopsis genome (Pereira-Leal and Seabra, 2001), whereas, only seven in the fission yeast (S. pombe; Lazar et al., 1997). The Rab family of Arabidopsis are grouped into eight clades, A–H (Bischoff et al., 1999; Pereira-Leal and Seabra, 2001; Rutherford and Moore, 2002; Vernoud et al., 2003); these are related to yeast and animal Rabs in terms of homology: Rab1, Rab2, Rab5, Rab6, Rab7, Rab8, Rab11, and Rab18; however, Rab2 and Rab11 are not present in yeast (Lazar et al., 1997). Rabs are localized to the cytoplasmic face of organelles and involved in vesicle formation, transportation, and docking during exocytic and endocytic pathways in eukaryotic cells (Chavrier and Goud, 1999; Pereira-Leal and Seabra, 2001). The role of Rabs is characterized in phytochrome signaling, pathogen response, and in guard cell differentiation. Interestingly, some of the Rab genes are found to be responsive to environmental cues like drought (O'Mahony and Oliver, 1999; Nahm et al., 2003), aluminum stress (Mikami et al., 1997), ABA application (Nicolás et al., 1998), salt treatment (Bolte et al., 2000), and to elicitors (Legendre et al., 1993; Kieffer et al., 1997; Xing et al., 1997). However, their basic function is not explored yet. The Rab GTPases are important components of endocytic network in plant cells. Among different Rab proteins, Rab7-related proteins regulate the fusion of membrane with lysosomes and vacuoles in mammalian and yeast cells, which is the final step of the endocytic pathway. In plants the Rab7 proteins are involved in transporting vesicles to vacuole and in vacuole biogenesis and also play role in abiotic stress, biotic stress, and hormone signaling (Agarwal et al., 2009). The Arabidopsis genome encodes eight Rab7-related proteins, seven of which appear to be located on the vacuolar membrane and regulate the vesicle fusion with the vacuole (Saito et al., 2002). OsRab7 showed its localization at vacuoles, suggesting its role in transporting vesicles to vacuole and involvement in vacuole biogenesis (Nahm et al., 2003). The transgenic plants overexpressing AtRab7 and PgRab7 showed increased tolerance to salt and osmotic stresses (Mazel et al., 2004; Agarwal et al., 2008). Arabidopsis transgenics with AtRab7 showed accelerated endocytosis in roots, leaves, and protoplast that could be due to faster vesicle trafficking at the end point, that is, endosome–vacuole fusion (Mazel et al., 2004).
Till date the Rab7 genes have been isolated only from glycophytes; however, this gene has not been characterized from any halophytes. Halophytes have a unique genetic make-up allowing it to thrive under harsh climatic conditions. Aeluropus lagopoides (Linn.) Trin. Ex Thw. (poaceae) is a salt-secreting rhizomatous perennial halophyte growing throughout the year as a forage plant. Its foliage shows little variation in salt content despite a three-fold increase in soil salinity in summer (Joshi and Bhoite, 1988); this is possibly due to salt secretion through leaf glands. The halophytes carry salt glands that shift ions from mesophyll tissues to leaf surfaces, from where it is either washed or blown away. Barhoumi et al. (2008) reported that the increasing salinity increased the number and size of vacuoles in the gland cells and also increased the density and size of the intricate membrane system of the basal gland cell in Aeluropus. Therefore, it becomes meaningful to isolate and characterize AlRab7 gene, involved in vesicle trafficking from A. lagopoides, growing luxuriantly on the coasts of Gujarat, India and would contribute toward understanding the function of plant Rab family proteins. Furthermore, we identify AlRabring7 as its interacting partner for the first time in the plant system.
Materials and Methods
Plant growth and stress treatments
Aeluropus lagopoides plants were collected from the saline marshy areas near CSIR-CSMCRI salt farms. Nodal cuttings with two to three pair of leaves were placed in holes made on thermocol sheets in 1/2 strength Hoagland medium (Hoagland and Arnon, 1950). The plants were maintained in a growth chamber with dark/light (30–35 μmol m−2 s−1 spectral flux photon of photosynthetically active radiations) cycle of 16/8 h at 25°C. Two week later, when the roots were formed the plants were subjected to different stress treatments in 1/2 strength Hoagland medium; 250 mM NaCl and 20 μM ABA for 6, 12, 24, and 48 h and dehydration (plants were removed from medium and wrapped in tissue paper and kept at room temperature) for 1, 2, and 3 h. Another set of rooted plantlets were maintained under control conditions. Three sets of plants were used for each treatment and leaf tissue was collected and stored at −80°C until use.
Cloning of AlRab7 cDNA
Total RNA was isolated from the control and stress-treated plants by GITC method (Chomczynski and Sacchi, 1987). About 2.5 μg of total RNA was used for first strand cDNA synthesis using manufacturer's protocol (Thermo Scientific). The degenerate, oligo dT, and 3′ RACE primers used are as follows: Rab7-2F 5′-AAGACSTCSCTGATGAACCAR-3′, Rab7-3F 5′-AGTCTTGGTGTBGCATTYTA-3′ and PAOligodT 5′-CAGACGAGAGTGTGGAGGACTGCTGCTGGTGTAGCT(17)-3′, PAR1 5′-CAGACGAGAGTGTGGAGG-3′, PAR2 5′-GACTGCTGCTGGTGTAGC-3′. Using the above primers, the fragment obtained was cloned in pJET 1.2 vector (MBI Fermentas) and sequenced. After confirmation of the sequence by BLAST search, the 5′ RACE were conducted. The clone was made full length by 5′ RACE (Invitrogen) with following primers AlGSP1 5′-TCTCAGAGACCACGCGGCTGTTGC-3′, AlGSP2 5′-ATCTACATC CACTTTGTTTCCCAG-3′, AlGSP3 5′-GCCAGTTGTTTAAGTTGTCAAATGA-3′. The PCR was carried out using cDNA as template, 150 ng of each primer, 200 μM dNTPs, and 2.5 U Taq DNA polymerase in a 50 μL reaction volume. The PCR conditions were 94°C, 5 min for 1 cycle; 94°C, 1 min, 55°C, 1 min, and 72°C, 1 min for 35 cycles; and last 72°C, 7 min for 1 cycle. The clone was sequenced at Macrogen. The sequence was analyzed by the BLAST program provided by NCBI (National Center of Biotechnology Information,
Transcript analysis of AlRab7 under different stress treatments by reverse transcription–polymerase chain reaction
Total RNA was isolated from control and stress-treated Aeluropus plantlets and first strand cDNA synthesis was carried out using manufacturer's protocol (Thermo Scientific). The cDNA was used as template for reverse transcription–polymerase chain reaction (RT-PCR) using AlRab7 primers (forward 5′-AGTCTTGGTGTGGCATTCTA-3′ and reverse 5′-GAACCATGGAGACCCGTCAA-3′), and the reference actin gene (GW796822) primers (forward 5′-TACGAAGGGTTTACGCTTCCT-3′and reverse 5′-TCTCCAACTCCTCCTCGTAAT-3′). Real-time was performed in a CFX detection system (Bio-Rad) with 1× Sso Advanced SYBR green supermix (Bio-Rad) using 75 ng of each gene-specific and actin primers with following PCR conditions: 94°C, 2 min for 1 cycle; 94°C, 30 s, 55°C, 30 s, and 72°C, 30 s for 40 cycles; and 72°C, 7 s for 1 cycle. At the end of the PCR cycles, the products were subjected to melt curve analysis to verify the specificity of PCR amplification. Three independent experiments were performed with three replicates. Relative normalized expression was determined using Livak method (Livak and Schmittgen, 2001). The mean and standard deviation (SD) was calculated using Microsoft Excel.
Cloning of AlRab7 into an Escherichia coli expression vector and purification of the recombinant protein
Based on the complete cDNA sequence, two primers were designed with the flanking restriction sites of BamHI in forward primer 5′-CGCGGATCCATGGCCGCGCGCCGCCGT-3′ and XhoI site in the reverse primer 5′-CCGCTCGAGTTAACAGCAGCCGGATGA-3′. Using these primers the complete coding sequence for the AlRab7 was PCR amplified (150 ng of each primer, 200 μM dNTPs, 2.5 U Taq DNA polymerase in a 50 μL reaction, with 94°C, 5 min for 1 cycle, 94°C, 1 min, 55°C, 1 min, and 72°C, 1 min for 30 cycles; and 72°C, 7 min for 1 cycle) using Aeluropus cDNA clone as template. The amplified product was gel purified, digested, and cloned in pET 28a expression vector (Novagen). The recombinant plasmid was transformed in BL21 (DE3) star Escherichia coli strain and grown in Luria Bertani (LB) medium at 37°C to OD600 0.5. AlRab7 protein was induced with 1 mM isopropyl beta-D-thiogalactoside (IPTG).
Functional analysis of E. coli cells transformed with AlRab7 under different stresses
Spot and liquid culture assay
The effect of 500 mM NaCl, 500 mM KCl, 400 mM mannitol, 20 μM ABA, and 20 μM indole-3-acetic acid (IAA) was studied on the growth of E. coli cells containing recombinant plasmid and vector plasmid with the help of spot and liquid culture assay. E. coli cells were grown in LB medium to an OD600 0.6. Thereafter, 1 mM IPTG was added and cells were grown for 12 h at 37°C, and then diluted to 10−2, 10−3, and 10−4. Ten microliters from each dilution was spotted on LB basal plates, and on plates supplemented with 500 mM NaCl, 500 mM KCl, 400 mM mannitol, 20 μM ABA, and 20 μM IAA. The experiment was repeated thrice independently. For liquid assay, E. coli cells were grown as for spot assay, diluted at OD600 to 0.6. The 400 μl cells were inoculated in 10 mL LB basal medium and medium containing 500 mM NaCl, 500 mM KCl, 400 mM mannitol, 20 μM ABA, and 20 μM IAA. The bacterial suspension was allowed to grow at 37°C at 160 rpm and harvested at every 2 h upto 12 h for growth measurements at OD600. The experiment was repeated thrice independently; mean and SD were calculated by Microsoft Excel) with p≤0.05.
Yeast two-hybrid screening
For yeast two-hybrid assay the cDNA library was prepared using manufacture's protocol (Clonetech). The AlRab7 cDNA was cloned in the BamHI/PstI sites of the pGBKT7 vector. The interaction of Rab7 and cDNA library was carried out for 16 h at 30°C and plated on SD−Ade/−His/−Leu/−Trp medium. The colonies were screened using pGADT7 vector specific primers. DNA was isolated from the PCR-positive colonies and sequenced at Macrogen. The self-activation was carried out to overrule the interaction with vectors.
Results
Cloning and sequence analysis of AlRab7 cDNA from Aeluropus lagopoides
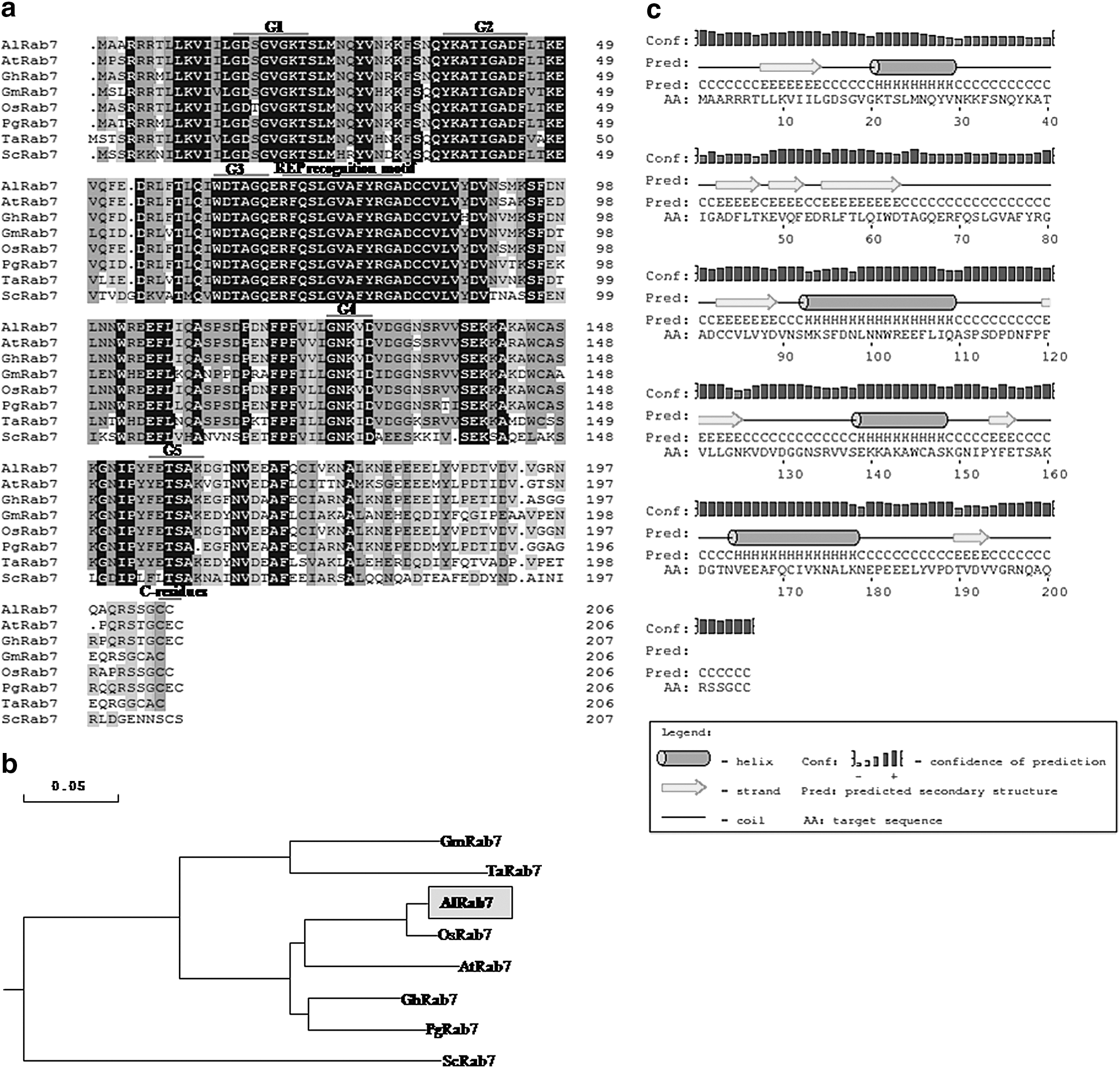
The full-length clone of AlRab7 was isolated using degenerate primer approach. The 3′ RACE using degenerate primers (AlRab7-3F) gave an amplicon of 555 bp. After confirming its sequence with NCBI BLAST search, 5′ RACE was conducted and 358 bp amplicon was obtained. The full-length cDNA (NCBI acc. no. KM023774) comprised of an open reading frame of 621 bp, with 41 bp 5′-untranslated and 147 bp 3′-untranslated region. The AlRab7 encodes 206 amino acids polypeptide with a calculated molecular weight of 23.05 kDa and the predicted pI of 5.38. The conserved domain search of AlRab7 amino acid sequence conducted in NCBI predicted its high sequence homology to small GTP-binding protein Rab7, a member of Ras super family (Fig. 1a). Amino acid alignment and phylogenetic analysis of AlRab7 revealed its high homology with OsRab7 (97%), AtRab7 (89%) and PgRab7 (84%, Fig. 1b). Multiple sequence alignment of AlRab7 with homologous sequences showed that it possesses functionally important motifs peculiar to small GTP-binding proteins. The effector-binding domain G2 and four nucleotide-binding domains (G1, G3, G4, and G5) responsible for nucleotide binding and hydrolysis are highly conserved in AlRab7 (Fig. 1a). The G1 (P-loop; GDSGVGKT) is involved during Mg+2 and phosphate binding, G3 (Switch II; WDTAGQ) where DTAG interacts with γ-phosphate of GTP, G4 (GNKVD) where NKVD is the guanine specificity region, G5 (ETSAK) where ETSA interacts with the D residue in the NKVD. The effector region G2 (Switch I; YKATIGADF) contains information for individual function, and its TIGADF motif interacts with specific GTPase-activating proteins (GAPs; Haubruck et al., 1990; Zerial and Stenmark, 1993). The PrePS identified LKNEPEEELYVPDTVDVVGRNQAQRSSGCC (177–206 aa) as the prenylation site for Rab geranylgeranyltransferase 2 (RGGT2) and RFQSLGVAFYRGA (69–81 aa) as the REP (Rab escort protein) site. The sequence motif YRG (78–80 aa) characteristic of Ypt/Rab subfamily is also conserved, although its function is not known. Furthermore, the AlRab7 exhibited C-terminal sequence of CC residues, which is required for membrane attachment. The secondary structure of AlRab7 amino acid sequence was deduced by PSIPRED protein structure prediction server (ExPASy tools), which showed the presence of four α helices and eight β sheet strands (Fig. 1c).
Differential expression analysis of AlRab7 gene under different stress conditions
The transcript study of AlRab7 was carried out by RT-PCR in A. lagopoides plantlets subjected to 250 mM NaCl, dehydration and 20 μM ABA treatments (Fig. 2). In dehydration treatment the transcript showed ∼1.6-fold higher expression at 1 h as compared to control (Fig. 2a). The transcript level decreased slightly at 2 h and then again increased at 3 h. There was no significant increase in AlRab7 transcript accumulation after the imposition of 250 mM NaCl stress from 6 to 48 h (Fig. 2b). The transcript was down regulated with 20 μM ABA at 6 h treatment, however, got upregulated at 12 h (Fig. 2c). The transcript levels decreased gradually after 24 h which continued up to 48 h of stress.
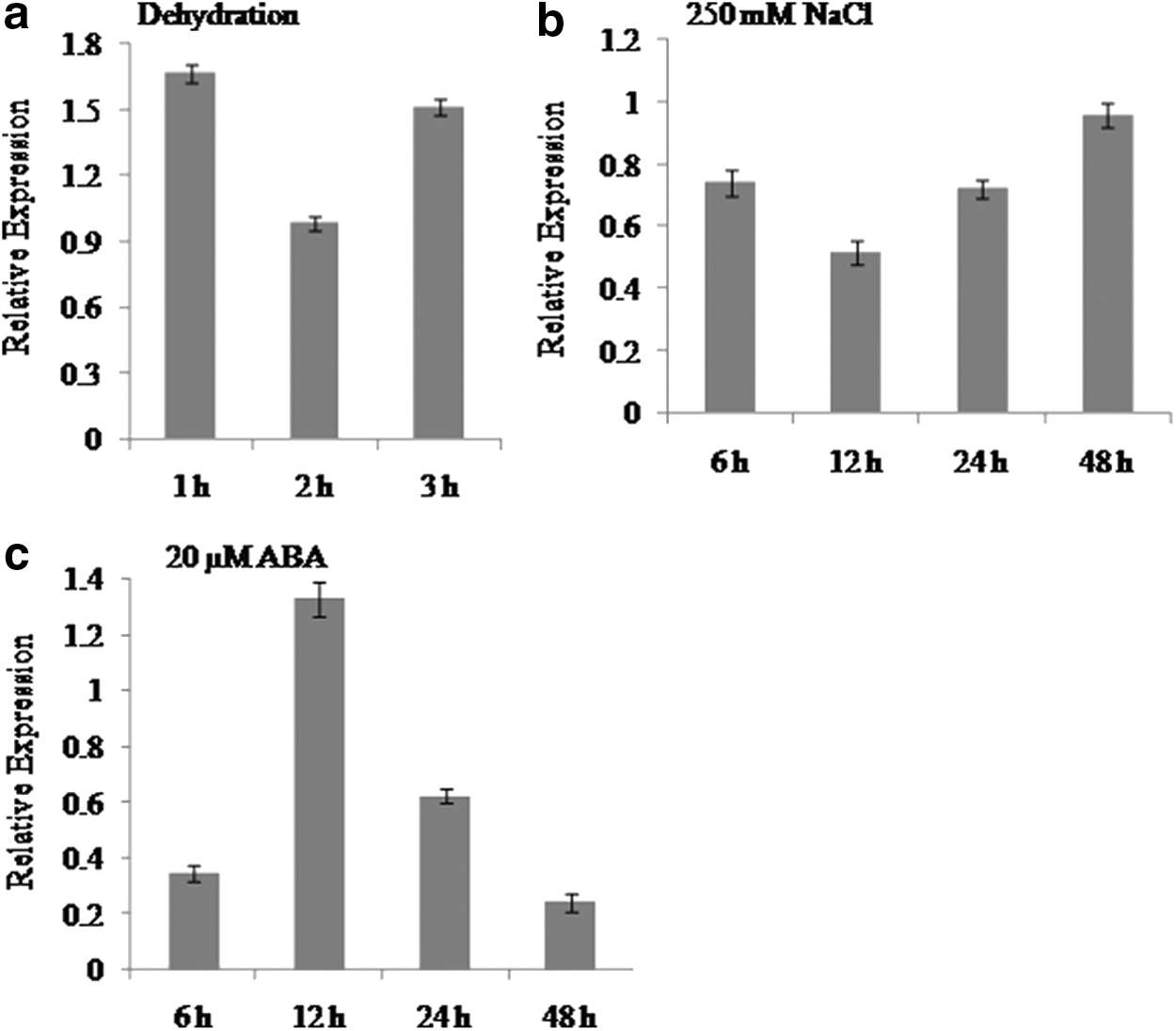
Relative expression of AlRab7 gene in A. lagopoides plantlets exposed to
Expression of AlRab7 protein promotes the growth of Escherichia coli cells under stress conditions
The 6X His-Tag fused AlRab7 recombinant protein (23.05 kDa) was induced with 1 mM IPTG for different time periods (Fig. 3). To study the role of AlRab7 in conferring stress tolerance to E. coli, the growth of the recombinant E. coli cells harboring AlRab7 (recombinant cells) and vector pET28a (control cells) was monitored qualitatively (spot assay, Fig. 4) and quantitatively (liquid assay, Fig. 5). The spot and liquid assay was performed on LB basal medium and medium supplemented with 500 mM NaCl, 500 mM KCl, 400 mM mannitol, 20 μM ABA, and 20 μM IAA. The recombinant and control cells followed similar growth pattern on LB basal medium (Figs. 4a and 5a). The recombinant cells displayed a better growth and increase in number of colonies on medium containing NaCl, mannitol, and ABA (Fig. 4b, d, e), however, with KCl and IAA the number of recombinant colonies were similar to control cells (Fig. 4c, f). In liquid culture assay the ionic stress proved very toxic for both recombinant and control cells (Fig. 5b, c). The AlRab7 recombinant cells grew at similar rate as that of vector alone in mannitol up to 8 h and thereafter the growth of recombinant cells increased (Fig. 5d). In presence of ABA and IAA, the recombinant E. coli cells and vector alone growth was almost similar until 6 h of inoculation and thereafter the growth of recombinant cells showed progressive increase as compared to vector alone (Fig. 5e, f).
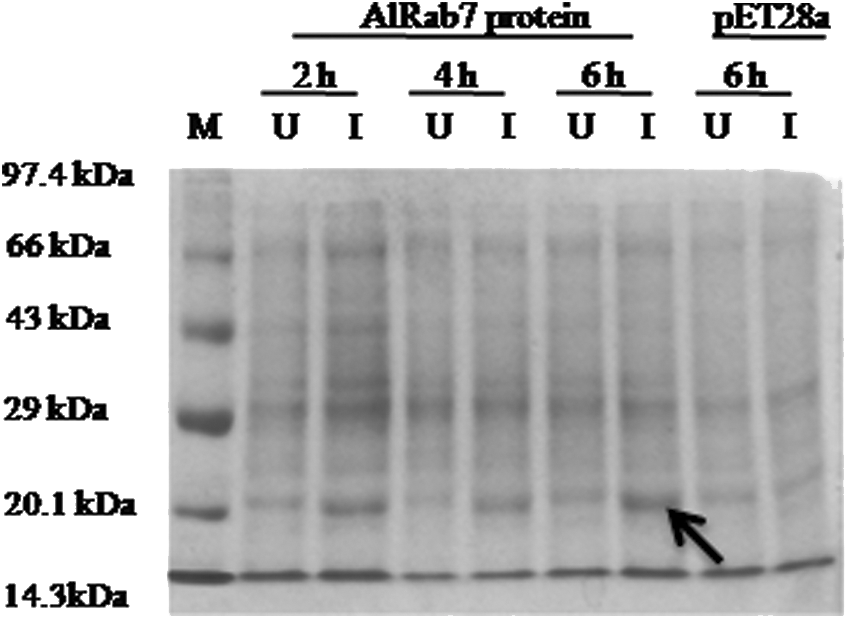
Protein expression study of induced and uninduced AlRab7 recombinant protein in Escherichia coli BL21 (DE3) cells. M marker, lane 1 uninduced protein 2 h, lane 2 induced protein 2 h, lane 3 uninduced protein 4 h, lane 4 induced protein 4 h, lane 5 uninduced protein 6 h, lane 6 induced protein 6 h (shown by arrow), lanes 7 and 8 uninduced and induced vector alone after 6 h, respectively.
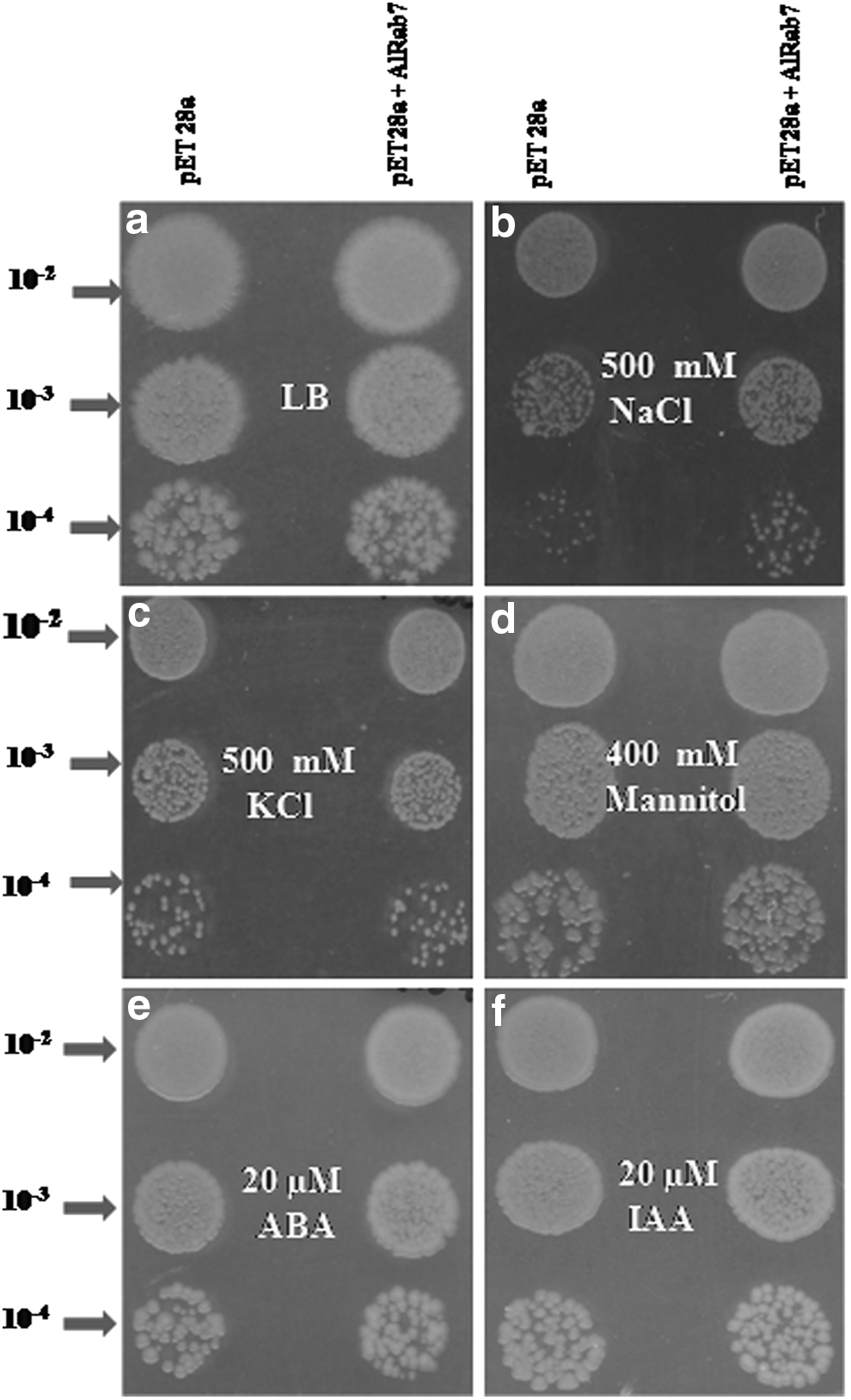
Spot assay of BL/pET28a and BL/AlRab7-pET28a on the Luria Bertani (LB) plates
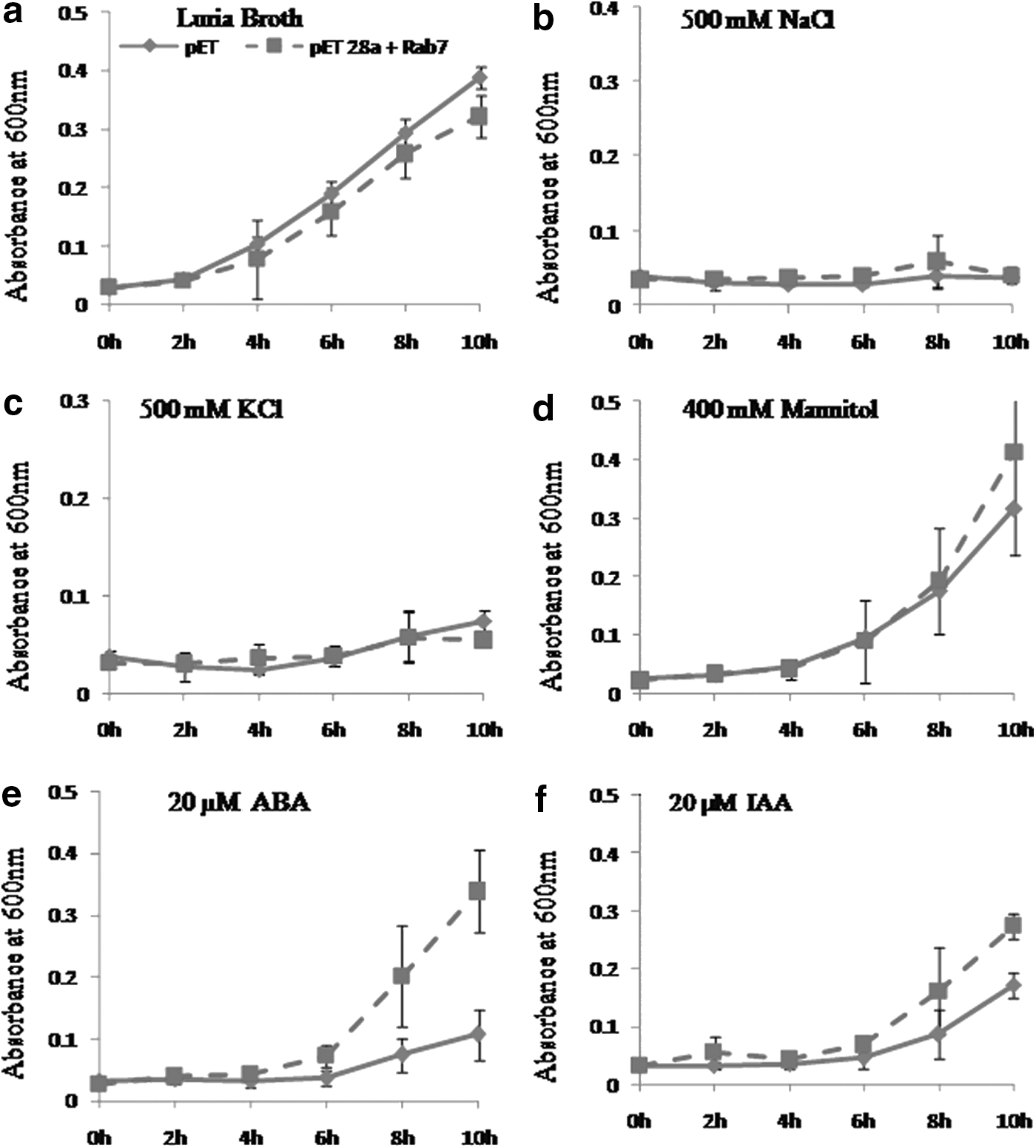
Growth study of E. coli BL21 (DE3) cells with only pET28a and pET28a-AlRab7 in liquid LB medium
AlRab7 interacts with AlRabring7
The full-length AlRab7 cDNA did not show self-activation with pGADT7 vector, therefore the native gene was used as bait for identification of Rab7 target proteins. Approximately, 2.4×105 transformants were screened and 30 positive clones were identified. Two similar clones (AlRabring7, acc no JZ712387) of 197 bp showed 85% and 66% identity with RNA-binding protein of Oryza sativa and ring finger protein 141 of Aegilops tauschii, respectively. Further, Rab7 and this clone was confirmed to overrule the self-activation with pGADT7 and pGBKT7 vectors respectively (Fig. 6), confirming that Rab7 interacts with Rabring7 protein.
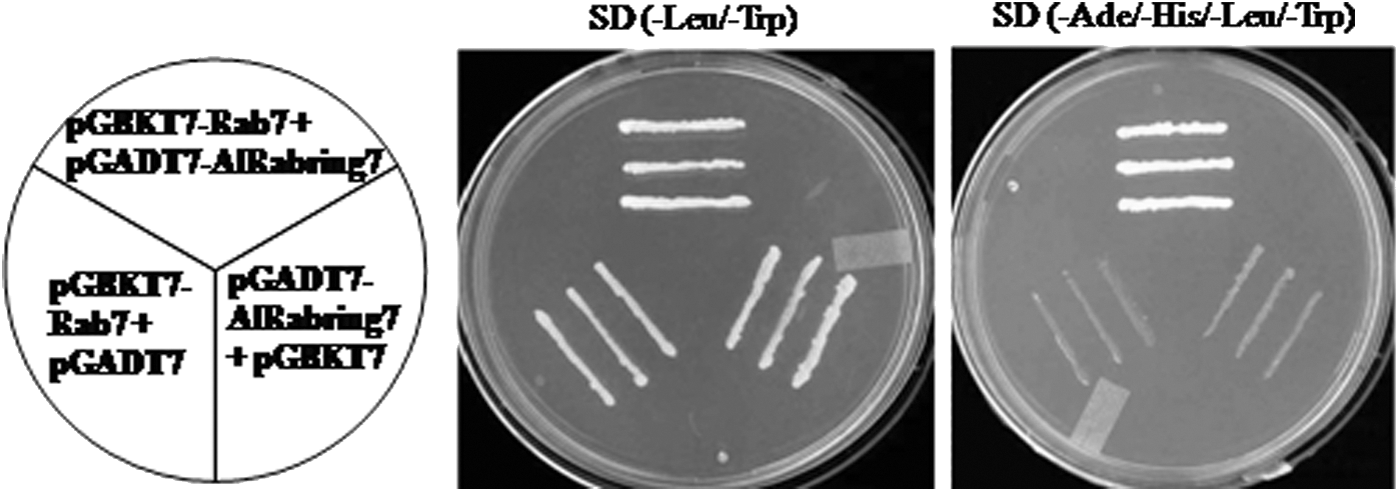
Yeast two-hybrid assay showing the interaction of AlRab7 and AlRabring7. The pGBKT7-Rab7+pGADT7-AlRabring7 shows the growth on both SD-2 (–Leu/–Trp) and SD-4 (–Ade/–His/–Leu/–Trp) medium, whereas the pGBKT7-Rab7+pGADT7 and pGADT7-AlRabring7+pGBKT7 do not show the growth on SD-4 (–Ade/–His/–Leu/–Trp) medium.
Discussion
The Rab7 proteins regulate fusion of membrane with lysosomes and vacuoles in mammalian and yeast cells, in the last step of endocytosis (Saito et al., 2002; Ueda and Nakano, 2002). Rab7 is a small GTP-binding protein involved in intracellular vesicle trafficking from late endosome to the vacuole. There are eight Rab7-related proteins encoded in the Arabidopsis genome, seven of which appear to be located on the vacuolar membrane and regulate the vesicle fusion with the vacuole (Saito et al., 2002). The OsRab7 showed localization at vacuoles suggesting that OsRab7 transports vesicles to vacuole and has involvement in vacuole biogenesis (Nahm et al., 2003). Similarly, Arabidopsis transgenics overexpressing AtRab7 showed accelerated endocytosis in roots, leaves, and protoplast that could be due to faster vesicle trafficking at the end point (endosome–vacuole fusion, Mazel et al., 2004). Interestingly, we have earlier shown that PgRab7 transgenic tobacco showed enhanced alkaline-phosphatase (ALP) activity, because of higher rate of ALP transport to the vacuole, highlighting the involvement of Rab7 gene in vacuolar transport in plants (Agarwal et al., 2008). In yeasts, membrane-bound enzyme ALP is transported to the vacuole via late endosome-like compartment for its maturation; furthermore, the ypt7 null mutant is characterized by highly fragmented vacuoles and differential defects in vacuolar protein transport and maturation (Wichmann et al., 1992).
The AlRab7 protein showed highest homology to OsRab7, they show similar C-terminal two cysteine residues (CC). The C-terminal is preceded by a hyper-variable domain, which directs the trafficking and association of Rab7 proteins with their specific targets (Nahm et al., 2003). The C-terminal sequence is essential for post-translational modification (prenylation), necessary for the localization of these proteins to appropriate membranes of different subcellular compartments (Chavrier et al., 1990). However, to date no isoprenylation activity has been reported in plants (Terryn et al., 1993; Ma, 1994). PrePs predict the presence of prenylation site for Rab geranylgeranyltransferase and REP site. Rab geranylgeranylation first requires the recognition of a Rab GTPase protein by its REP, the complex of Rab and REP is then recognized by the lipid attaching RGGT2. The RGG2 anchors to the last two cysteines residues of a suitable sequence being –XCC, –CXC, and –CCX. Subsequent carboxymethylation has only been reported for –CXC but not –XCC motifs (Konstantinopoulos et al., 2007)
Rab7 from rice and Pennisetum have been shown to play an important role in abiotic stress. The AlRab7 transcript showed upregulation in response to dehydration, similar response was reported for the PgRab7 and OsRab7 genes. In salt stress, the AlRab7 transcript was not upregulated similar to the OsRab7 transcript, however, the PgRab7 showed significant increased transcript under salt stress (Nahm et al., 2003; Agarwal et al., 2008). The ABA treatment showed increased AlRab7 transcript at 12 h with 20 μM ABA, however, a significant increase in OsRab7 was observed with 100 μM ABA (Nahm et al., 2003). The weak transcript expression of AlRab7 gene by dehydration, stress hormone ABA, and no induction during salt treatment indicate that its transcript is not significantly regulated by stress; however, its protein when overexpressed in-vivo shows enhanced ionic and osmotic tolerance (Agarwal et al., 2009). The AlRab7 recombinant E. coli cells showed enhanced growth under different stress conditions. In spot assay, the recombinant cells showed increased number of colonies with NaCl compared with control cells; however, in liquid assay not much difference was observed, probably the ionic stress perceived in broth was higher compared with semi-solid medium. Similarly, the OsRab7 gene in E. coli increased their resistance to heat, cold, and salt stress (Peng et al., 2011). The recombinant cells also showed enhanced growth with IAA. In E. coli 50 genes (1.1%) exhibited significantly different expression profiles on IAA supplemented medium (Bianco et al., 2006a). Some of these genes encoded for proteins involved in cell envelope biogenesis and in cellular responses under stress conditions, including GTP-binding protein chain elongation factor EF-G (Bianco et al., 2006b). Previously, we have observed that the PgRab7 gene showed increased transcript accumulation with IAA treatment (Agarwal et al., 2008). The PgRab7 overexpressed tobacco transgenics showed elongated roots on high auxin concentration, whereas the WT line showed shorter root length (Agarwal et al., unpublished data). This may be because the Rab7 gene might be involved in auxin signaling.
The protein-interacting partners of Rab7 are not reported from the plants. AlRabring7 was identified as a target protein for AlRab7 using Y2H for the first time in the plant system. Earlier this gene was identified as an interacting partner from a murine immature B cell cDNA library (Mizuno et al., 2003). The AlRabring7 shows homology to RING-finger (Really Interesting New Gene) domain, a specialized type of Zn-finger of 40–60 residues that binds two atoms of zinc. The Rabring7 from mice is predominantly found in cytoplasm and gets localized to late endosome and regulates vesicle traffic (Mizuno et al., 2003). AlRabring7 might also be involved along with AlRab7 during vesicle transport. Further studies towards characterization of AlRabring7 will facilitate in deciphering its function during endocytosis.
Footnotes
Acknowledgments
CSIR-CSMCRI Communication No. 115 as provided by BDIM. The financial assistance from CSIR (OLP 0067 and CSC 0203) is greatly acknowledged. P.S. and J.K. are thankful to CSIR for Senior Research Fellowship and Junior Research Fellowship, respectively. P.S. and J.K. are thankful to AcSIR for enrollment in PhD.
Disclosure Statement
No competing financial interests exist.
