Abstract
Background:
The aim of this study was to investigate the molecular status of gingival mesenchymal stem cells (GMSCs) and to elucidate their role in regenerative medicine. We performed gene expression analysis for pluripotency genes in detail using low density array.
Methodology:
Methodology involved the in vitro expansion of GMSCs using specified media with growth factors. They were characterized by trilineage differentiation into osteocytes, adipocytes, and chondrocytes that were verified by staining. RNA was extracted from the confluent monolayer followed by cDNA preparation and real-time PCR. A low density array microfluidic card was run, which had 92 genes specific for pluripotency and stemness profiling.
Results:
The GMSCs demonstrated rapid proliferation and their initial differentiation into osteocytes, adipocytes, and chondrocytes were confirmed by Alizarin Red S, Oil Red O, and Alcian blue staining, respectively. Moreover, our results of TaqMan™ Array Human Stem Cell Pluripotency Panel found several novel pluripotent genes in GMSCs, other than the previously reported SOX2, NANOG, OCT4, and LIN28. Our study also differentiates the pluripotent markers in GMSCs and pluripotent stem cells.
Conclusion:
The available data on GMSCs show that they can produce multiple cell lineages, and to explore their expression further, we performed real-time PCR with 92 pluripotent genes. As a result, mRNA expression of many pluripotent genes can be seen that were unreported before and they could be further investigated to explore the role of GMSCs in regenerative medicine.

Introduction
Anatomically, gingiva covers the alveolar bone present around the teeth and protects the delicate structures such as vessels and nerves. Histologically, gingiva has two main components; gingival epithelium followed by connective tissue called lamina propria that is firmly attached to the underlying bone. The gingival epithelium is thick and resistant to abrasion and external injuries. The epithelial cells have a basal layer that consists of progenitor cells and maturing cells. The progenitor cells have stem cell-like properties as they provide new cells and the maturing cells migrate to upper layers of epithelium and form the keratinized oral mucosa. 1 On the other hand, lamina propria supports and provides structural framework to the oral epithelium. It predominantly contains collagen fibers that are surrounded by blood vessels, extracellular matrix, lymphatics, nerves, and numerous types of cells. 2
Gingival cells are frequently injured because of their anatomical location in the oral cavity and these injured sites are ideal environment for bacterial invasion. 3 The external adverse stimuli produce gingival inflammation and initiate protective response of immunity that further starts recruitment of cells. Many studies have established that gingival mesenchymal stem cells (GMSCs) help in bone regeneration and their healing capacity is comparable with bone marrow-mesenchymal stem cells. 4
One of the biggest differences in oral and dermal wound healing is that scar tissue is not formed in the oral cavity. This happens because of better blood circulation, rapid wound healing, superior intrinsic properties of cells such as keratinocytes and gingival fibroblasts, and the mineral environment produced by saliva and crevicular fluid. Therefore, these characteristic features produce fast and scarless repaired tissue in gingiva and oral cavity. 5
Dental tissues in the oral cavity are regarded as rich sources of mesenchymal stem cells, which can be obtained easily. However, some limitations include scarce tissue available in dental pulp stem cells, stem cells from deciduous teeth, and dental follicle stem cell. Likewise, stem cells from apical papilla and periodontal ligament stem cell may require extraction for tissue isolation. The gingival tissue is abundant and a better source as GMSCs reside in latent phase in lamina propria.6,7
The higher proliferative index and stable morphological feature make GMSCs ideal for isolation. They have multilineage potential and the molecular expression of GMSCs includes mesenchymal stem cell (MSC) markers such as CD44, CD90, and CD73. 6 Reportedly, in long-term cultures, GMSCs display fewer senescence features. 8 Recent studies have reported that these cells exert immune modulatory effects on both adaptive and innate immunity through secretory vesicles and cell to cell signal transmission. 9 Furthermore, they also express pluripotent markers such as octamer-binding transcription factor 4 (OCT4), SRY-Box transcription factor 2 (SOX2), and gene encoding for homeobox protein NANOG (Nanog). 10
In a pluripotent state, a stem cell can divide into ectoderm, mesoderm, and endoderm. The pluripotent markers are crucial as they maintain cell growth and differentiation at the time of repair and regeneration.11,12 The somatic cells can also be converted into induced pluripotent stem cells by force induction of OCT4, SOX2, and cellular myelocytomatosis oncogene (c-Myc).13,14
GMSCs have been shown to differentiate into many cell types that include mesodermal, ectodermal, and endodermal lineages. For instance, they can be transformed into chondrocytes, 15 keratinocytes, 16 neural cells, 17 and endothelial cells.18,19 This displays that GMSCs are unique kinds of MSCs that have certain properties of pluripotency. The expression of OCT4, NANOG, SOX2, and LIN28 (Lin-28 homolog A) has already been established in previously published literature. Therefore, we tried to investigate the expression of pluripotent genes in GMSCs in order to find more target genes that can be studied in detail and the role of GMSCs in regenerative medicine can be further explored.
Materials and Methods
Study setting and ethical consideration
The experiments were conducted in the Stem Cells and Regenerative Medicine laboratory, Dow Research Institute of Biotechnology and Biomedical Sciences (DRIBBS) at Dow University of Health Sciences (DUHS), Karachi. Ethical approval of the research project was granted by the Institutional Review Board (IRB), DUHS (IRB-2039/DUHS/Approval/2021).
Peripheral blood mononuclear cell isolation
For control group study, peripheral blood mononuclear cells (PBMCs) from healthy individuals were isolated following the standard density gradient centrifugation process. Briefly, blood was diluted with sterile phosphate-buffered saline solution in a 1:1 ratio. Ficoll-Paque was used as a density gradient medium. Diluted blood was layered on the top of the density gradient and the tube was centrifuged at 1,000 × g for 20 min at room temperature. The interphase layer containing PBMCs was carefully harvested and used in subsequent studies.
GMSC culturing
The cell line of GMSCs was established in DRIBBS by our research group. At that time, samples were taken from healthy tissue after taking consent from each patient. One vial of frozen cells that was stored in dimethyl sulfoxide frozen media at −80°C was thawed. Later on, the cells were centrifuged and their viability was counted by Vi-Cell Counter. Then they were plated in a six-well plate with 1,500 μl of complete Dulbecco’s modified eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS) and 1% antibiotic-antimycotic. Cells were allowed to expand at 37°C in the incubator. The culture medium was changed after every third day.
Characterization of GMSC chondrogenic differentiation
When GMSCs became 80%–90% confluent, they were trypsinized and centrifuged to obtain a cell pellet. The pellet was resuspended in complete media to make a cell suspension of 100 μl that contained 1 × 105 cells approximately. In each well of a six-well plate, 3–4 droplets of 20 μl were added and then the plate was placed in the incubator for 2 h. Afterward, 50 μl of chondrogenic differentiation media (Stempro Chondrogenesis Differentiation Kit, Cat no. A10071-01, Thermo Scientific-Life Tech, New York, United States) was added on each micro-mass of droplet. To prepare differentiation media, Stempro chondrogenesis kit instructions were followed.
Chondrocyte expansion
After 3–4 days the chondrocytes appeared and chondrocyte expansion media was added. Expansion media was changed in the six-well plate after every third day until the cells reached 70% confluency. The expansion media was formulated by following CGMTM Chondrocyte Growth BulletKitTM instructions (Cat no. CC-3216, Lonza, Walkersville, USA). In the basal media, we added human recombinant growth factor beta, insulin-like growth factor, transferrin, FBS, insulin, and gentamicin/amphotericin-B.
Staining of chondrocytes
Staining was accomplished using 1% Alcian blue (cat no. TMS-010-C, Merck Millipore, Canada) at the 10th day. The stain was prepared in 0.1 N glacial acetic acid. First, cells were washed with PBS (cat no. 14190144, Thermofisher, Paisley, UK) twice and then fixed with 4% formaldehyde solution. The cells were incubated for 30 min and washed again with PBS. Second, preformed 1% Alcian blue was added for 30 min and 0.1 N glacial acetic acid was used to wash cells. Then distilled water was placed to observe cells under the phase-contrast light microscope.
Chondrocytes real-time PCR
To further characterize chondrocytes and confirm their presence, we looked for the SRY-Box transcription factor 9 (SOX9) gene that plays key role in chondrocyte differentiation. For this reason, the RNA was extracted and cDNA were prepared from chondrocytes. We used preformed Taqman Gene expression assays for SOX9 and TBP (housekeeping gene) and Taqman FAST advanced master mix was used for the qPCR performed in Quant StudioTM 7 Flex Real-Time PCR System (Applied Biosystems) to verify the expresion of SOX9 in GMSCs at mRNA level.
Adipogenic and osteogenic differentiation
In a 24-well culture plate, GMSCs were plated with a count of 30,000–40,000 cells per well. Osteogenesis was induced by providing osteogenic differentiation medium [low glucose DMEM supplemented with 10% FBS, 0.1 μM dexamethasone, 10 mM b-glycerophosphate, and 0.2 mM ascorbic acid-2-phosphate] and adipogenesis was induced by providing adipogenic differentiation medium [low glucose DMEM supplemented with 10% FBS, 1 μM dexamethasone, 10 μg/ml insulin, and 100 μM indomethacine] to attached cells. The medium was changed every 3–4 days. After 21 days of differentiation, calcium deposition for osteogenic differentiation and lipid droplets deposition for adipogenic differentiation was observed by Alizarin Red S and Oil Red O staining, respectively.
Pluripotent gene expression analysis by quantitative real-time PCR
Total RNA extraction
To isolate RNA from PBMCs (control group) and GMSCs (test group), the protocol of the RNA easy mini kit (Qiagen cat no. 741047, Austin, Texas) was followed and total elution volume of 30 μl was extracted. To quantify the prepared RNA, 2 μl of RNA was placed over Nanodrop (cat no. ND-LITE, Thermo Fisher Scientific). Next, the cDNA was prepared through Revert aid first strand cDNA synthesis kit (cat no. 4366596, Applied Biosystems, Thermo Fisher Scientific, Vilnius, Lithuania).
Preparation of TaqMan™ array human stem cell pluripotency panel
The pluripotent gene expression was analyzed by the TaqMan™ Array Human Stem Cell Pluripotency Panel (cat no. 4385344, Foster City, USA). The pluripotent markers and TaqMan® MGB Probes had been dried down in this low density array card. There were eight reservoirs in a card and each one was filled with 100 μl of prepared sample-specific PCR reaction mixture. To fill it properly, the card was centrifuged, sealed, and then positioned in real-time PCR. This array card has the ability to provide the data of four replicates simultaneously. It contained genetic markers that exhibited correlation of stemness and differentiation. It also carried genes that can identify undifferentiated stem cells and conserve pluripotency (Table 1).
Markers in TaqMan™ array human stem cell pluripotency panel with subgroup divisions
To enumerate the mRNA expression of pluripotent genes the most widely used 2 delta-delta CT method was used. Three housekeeping genes 18S rRNA, ACTB, and GAPDH were used to normalize the data with each gene (Table 2).
The pluripotent genetic markers with assay ID
Statistical analysis of data
For statistical analysis, one-way ANOVA and Tukey’s post hoc test were applied. The mean of groups was compared in this test and the p value < 0.05 was taken as statistically significant. The Graph pad prism software version 9.2 was used to investigate the fold change of pluripotent genes and to make the graphs with error bars.
Results
Morphologic assessment of GMSCs and chondrocytes in primary culture media
The GMSCs were plated and they depicted spindle or fibroblast-like shapes. Initially, we observed fewer cell attachments till day 2, and then after the 5th day, bunches of cell appeared that were allowed to expand until 70%–80% of confluency was reached. They had homogenous morphology and produced monolayer with each passage. The unstained micro-mass of chondrocytes showed rounded and small elongated cells that completely expand into elongated chondrocytes (Fig. 1).

Characterization of GMSCs by 1% Alcian blue staining of the micro-mass of chondrocytes
GMSCs droplets of 20 μl (micro-mass) were cultured with chondrocyte differentiation media and the rounded cells in the droplets appeared after 4–5 days. The cells in the micro-mass were allowed to expand, and approximately after 8 days, elongated cells were seen in the culture. After 12 days of chondrocyte expansion, they were stained with 1% Alcian blue. The cells displayed positive stain with Alcian blue, which stains acid mucopolysaccharides in a cell. This was easily evident under the phase-contrast microscope (Fig. 1).
Characterization of GMSCs by chondrocyte gene expression analysis—SOX9
The gene SOX9 is expressed in chondrocytes but not in GMSCs. The experiment was performed in triplicate and SOX9 (SRY-Box transcription factor 9) gene expression was checked in expanded and stained chondrocytes. To accomplish this step, RNA extracted from chondrocytes was measured by Nanodrop and it showed 39 ng/μl of RNA with 2.05 absorbance ratio. Figure 1 shows the positive expression of SOX9 in GMSCs differentiated chondrocytes.
Differentiation of GMSCs into osteocytes and adipocytes
Differentiation of GMSCs was confirmed through Alizarin Red (osteogenic) and Oil Red O (adipogenic) staining. GMSCs in adipogenic media showed intracellular lipid droplets, in osteogenic media, cells showed calcium deposits (Fig. 2). These results confirmed that the isolated cell population retained main characteristics of GMSCs.

Differentiation of GMSCs into osteocytes and adipocytes. Adipogenic differentiation was indicated by Oil Red O staining
Expression of pluripotent genes by Taqman™ array human stem cell pluripotency panel
The RNA isolated from PBMCs (Control group) and GMSCs (Test group) was of high quality and integrity, as the absorbance ratio (A260nm/A280nm ratio) on spectrophotometry was 2.00. Furthermore, cDNA was synthesized and PCR mixture was prepared to fill the reservoirs of TaqMan array card and then real-time PCR was performed. The data of genes obtained from qPCR were normalized by three housekeeping genes that were 18S ribosomal RNA (18S), glyceraldehyde-3-phosphatedehydrogenase (GAPDH), and beta-actin (ACTB) (Table 3).
Comparative method showing fold change after normalizing with three housekeeping genes (18S, ACTB, and GAPDH)
In our study, low density pluripotency array contributed in displaying the range of pluripotency genes present in GMSCs that have not been reported before (Table 4). We identified 32 pluripotent genes expressed in these cells with strong expression of teratocarcinoma-derived growth factor 1, interferon-induced transmembrane 1, interferon-induced transmembrane 2, choriogonadotropin subunit beta, etc. Moreover, we were also able to differentiate between expression of pluripotent markers in pluripotent stem cells and GMSCs (Table 5).
mRNA expression of pluripotent genes in GMSCs that have not been reported before
Comparison of pluripotent genetic markers expressed in pluripotent stem cells and gingival mesenchymal stem cells
However, in our research, we observed many unexpressed pluripotent genes as well. The pluripotent genes that did not show expression in gingival mesenchymal stem cells were 60 in number. These findings denote that pluripotency genes, such as DNMT3B, GABRB3, and GDF3, are only expressed in pluripotent stem cells (Fig. 3).
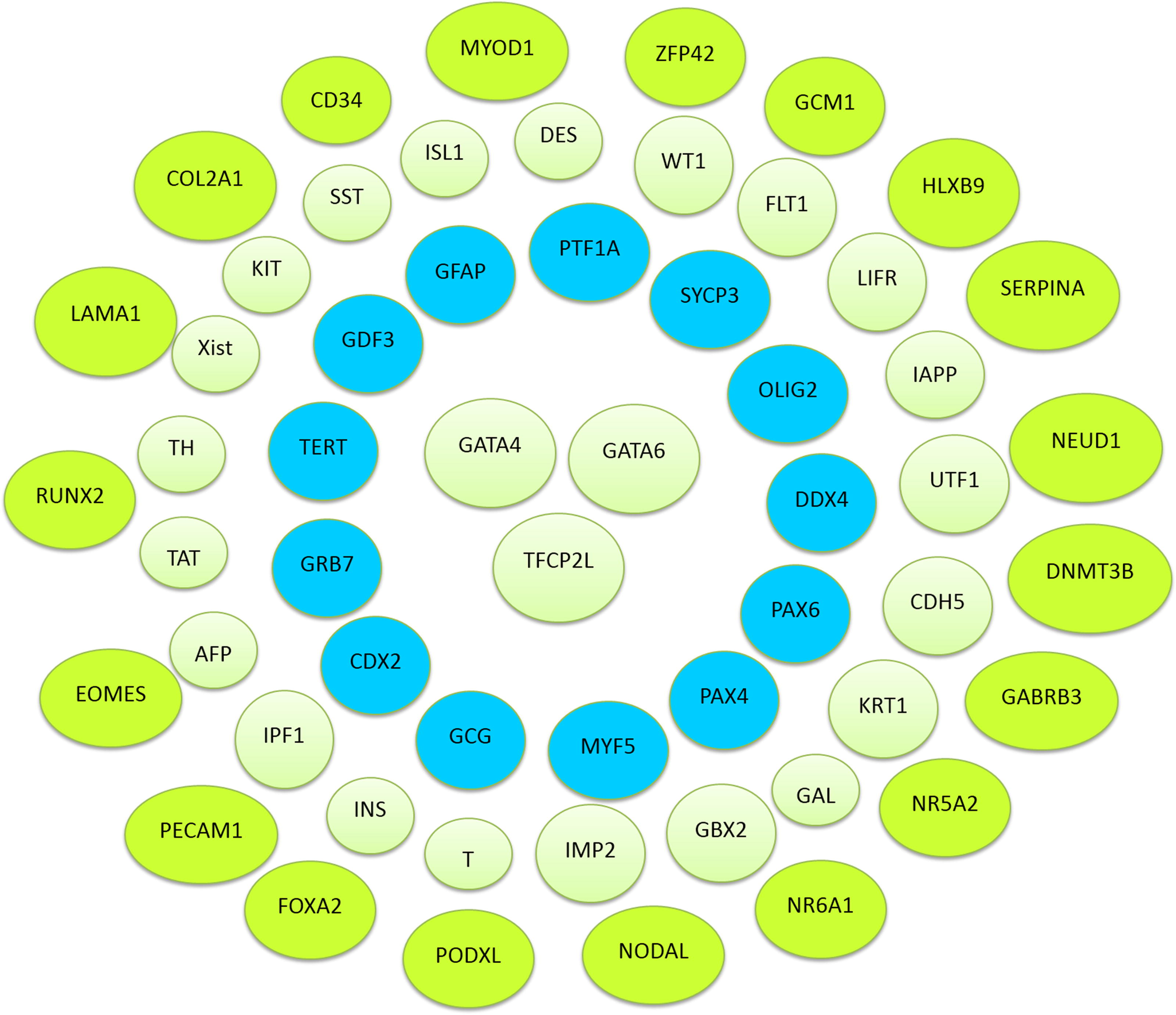
Genes unexpressed in real-time PCR. The above diagram shows the pluripotent genes that were totally unexpressed in gingival mesenchymal stem cells.
The mRNA fold change was evaluated through relative quantification (2−ΔΔCT) strategy and then statistical analysis was performed using by one-way ANOVA. There were a total of 92 genes in the array card, from which 32 pluripotent genes were upregulated in GMSC. Some genes increased mRNA expression, although their fold change was statistically insignificant, while approximately 12 pluripotent genes depicted statistically significant upregulated results in GMSCs (Fig. 4).

mRNA expression of pluripotent genes.
Discussion
In the field of regenerative medicine, mesenchymal stem cells (MSCs) isolated from oral structures are considered superior than MSCs obtained from bone marrow and adipose tissue as these orally derived MSCs can be easily isolated with less invasive procedures and extracted teeth and its associated structures are usually discarded as a waste. Among dental tissue-derived MSCs, GMSCs have gained a lot of attention due to their higher growth rate, easy availability, stable structural features, immunomodulatory properties, and multidirectional differentiation capacity. 7
Previous studies have reported that GMSCs can be differentiated into multiple cell types such as fat cells, bone cells, neural cells, keratinocytes, smooth muscle like cells, endothelial-like cells, and cartilage cells. These studies demonstrate not only mesodermal differentiation potential of GMSCs but also differentiation into ectodermal and endodermal cell types. 20 Our study tries to figure out the expression of various pluripotent genes in GMSCs to understand the differentiation potential of these cells.
There are a few pluripotency genes such as Nanog (gene encodes for homeobox protein NANOG), Oct4 (octamer-binding transcription factor 4), and Sox2 (SRY-Box transcription factor 2), whose expression in GMSCs has been frequently reported in many prior studies. 21 These genes are related to the maintenance of the pluripotency family and their expression was significantly upregulated in this study, which is comparable with plenty of earlier research.10,22–24 It has been stated that the pluripotency effect of these genes is mediated by their interaction with other essential mRNA and proteins. 25 Forkhead box D3 (FOXD3) gene is a member of correlation of stemness and its upregulated expression displayed that GMSCs have the ability to produce neural cells for nerve regeneration.26,27 Similarly, the gene NES (Nestin) was statistically increased in GMSCs that displayed its role in neuronal cell differentiation. 28 Lin28 gene (Lin-28 homolog A) also belongs to stemness related family and is a RNA-binding protein that controls developmental timings and pluripotency. In our study, this gene was significantly amplified and these findings were consistent with our previously published work on GMSCs. 7 In earlier studies, higher mRNA expression of interleukin 6 cytokine family signal transducer reduced the inflammatory effect in GMSCs and its expression was also increased in the present study.29,30
The Choriogonadotropin-beta gene has identical subunits with the pituitary gonadotropin hormones and is a differentiation marker. Previously, CGB scaffolds were used with human umbilical cord MSCs and it promoted neuronal differentiation of cells. 31 Teratocarcinoma-derived growth factor 1 gene is expressed in undifferentiated cells and gives rise to mesodermal cells. This gene was not reported before in GMSCs but its expression was previously seen in induced pluripotent stem cells 32 and bone marrow stem cells. 33 IFITM1 and IFITM2 genes belong to correlation of stemness group and have antiproliferative and osteoblast differentiation function. Another novel gene was cellular retinoic acid binding protein 2 and its role has been identified in human dental pulp stem cells. 34 Presence of the above pluripotent genes in GMSCs has not been reported before and they are the novel as well as significantly upregulated genes of our study.
Conclusion
In conclusion, gingival tissue provide a better source of MSCs because of superior cellular properties. The repetitive scar less repair distinguishes them from other body tissues. The earlier studies demonstrated that GMSCs can form many cells types other than mesodermal lineage. Therefore, we tried to rule out the pluripotent gene expression in GMSCs, and to the best of our knowledge, this is the first study that has focused on studying comprehensive gene expression analysis for 92 pluripotency genes in GMSCs and has identified several novel genes that were not previously reported to be expressed in these stem cells. The data have been normalized with housekeeping genes and statistical analysis has been carried out in detail.
Footnotes
Authors’ Contributions
M.W. designed and supervised the study and all the experimental work including cell culture, trilineage differentiation, qPCR, low density array experiments and analysis, and reviewed the manuscript. R.J. performed the experiments, analyzed the data, and wrote the manuscript. R.H. provided technical support and prepared the manuscript. O.S. provided technical support in experiments and prepared the manuscript. All authors read and approved the final manuscript.
Ethical Approval
Institutional Review Board of Dow University of Health Sciences (IRB-2039/DUHS/Approval/2021). The project comes under IRB exemption.
Author Disclosure Statement
All authors declare no conflict of interest.
Funding Information
This study was not funded by any agency in public, commercial, or not-for-profit sectors. It is self-funded.
