Abstract
Abstract
Background:
The emergence of multicellularity required the evolution of sophisticated intercellular communication, coordination, and adaptive mechanisms. In this context, the formation of syncytia—multinucleated cells resulting from cell fusion—along with polyploidy (the condition of having multiple sets of chromosomes) and epigenetic alterations (heritable changes in gene expression without DNA sequence modification), plays a pivotal role in enhancing cellular plasticity. These processes are not only critical during normal development, such as embryogenesis and organogenesis, but also in adaptive responses to environmental stress and in the progression of diseases like cancer.
Method:
This comprehensive review integrates current literature on the molecular and cellular interplay within syncytia. An extensive analysis was conducted of studies addressing both germline and somatic cell syncytia, focusing on the regulatory impact of epigenetic mechanisms—including DNA methylation, histone modifications, and non-coding RNA-mediated control—on ploidy variation and cell fate determination. Comparative insights from experimental models and clinical observations were synthesized to elucidate the evolutionary and pathological implications of these processes.
Results:
The analysis revealed that epigenetic regulation is critical in mediating ploidy changes and orchestrating the functional dynamics of syncytia. Polyploidy, often arising from mechanisms such as cell fusion and endoreplication, contributes to increased genomic and transcriptomic diversity, thereby enhancing cellular resilience in dynamic environments. Furthermore, the review highlights striking similarities between polyploid giant cancer cells and syncytia, suggesting a shared evolutionary mechanism that drives both adaptive responses and tumor progression. These findings underscore the role of syncytia as key facilitators of intercellular cooperation, resource sharing, and adaptive plasticity.
Conclusion:
The interplay between syncytial organization, polyploidy, and epigenetic control is fundamental to cellular evolution, adaptation, and the development of pathological states such as cancer. This review provides novel insights into how these integrated processes shape cell fate decisions and underscores their potential as targets for therapeutic intervention. Future studies aimed at unraveling the specific molecular pathways involved will be essential to advance our understanding of cellular plasticity and to develop innovative strategies for disease treatment.
Introduction
The emergence of multicellularity was a pivotal evolutionary step, enabling the compartmentalization of cells into specialized units with distinct functions that underpin the complexity of life. This specialization fostered intercellular communication, coordination, and adaptability, crucial for maintaining equilibrium in a dynamic and highly entropic environment.1,2 Among the evolutionary innovations facilitating cellular plasticity, syncytia—structures formed by the fusion of multiple cells into a multinucleated entity—stand out for their critical roles in both normal physiological processes and pathological conditions.3–5
Syncytia contribute to biological adaptability by enabling intercellular cooperation and sharing of resources, such as DNA and RNA, which bolster resilience against environmental stressors and lethal mutations. This mechanism provides an efficient response to external chaos and ensures the resilience of cellular systems.6,7 Syncytial dynamics establish a complex network of molecular, cellular, and physiological processes that are significantly influenced by the environment.8–10
For instance, in skeletal muscle development, myoblasts fuse to form multinucleated myofibers, which are essential for muscle contraction and function. Similarly, in the placenta, the syncytiotrophoblast (STB) layer facilitates nutrient and gas exchange between the mother and fetus, playing a crucial role in fetal development. In the immune system, osteoclasts are multinucleated cells formed by the fusion of monocytes and are responsible for bone resorption, a process vital for bone remodeling and calcium homeostasis. However, syncytia formation also occurs in pathological contexts. Certain viruses, such as human immunodeficiency virus and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), induce the formation of multinucleated giant cells by fusing infected cells with neighboring uninfected cells, contributing to viral dissemination and tissue damage. In cancer, particularly in aggressive tumors, the formation of multinucleated giant cells has been observed, which may contribute to tumor progression and resistance to therapy.8–12
Similarly, polyploidy—the presence of multiple sets of chromosomes within a cell—emerges as another evolutionary strategy to enhance cellular diversity and adaptability. Polyploidy contributes to developmental processes, tissue repair, and stress responses in normal contexts, while in diseases like cancer, it supports genomic instability and tumor heterogeneity.8–10,12,13 Both syncytia and polyploidy represent cellular adaptations that rely heavily on epigenetic regulation, where environmental cues drive heritable changes in gene expression without altering the underlying DNA sequence, thereby enabling phenotypic plasticity and functional versatility.
This article aims to explore the molecular and cellular interplay within syncytia, emphasizing the critical role of epigenetics in mediating ploidy changes and influencing cellular fates. By elucidating the evolutionary significance of polyploidy and its epigenetic regulation, this work highlights how syncytial complexity contributes to biological processes, particularly in adaptive responses and disease contexts such as cancer. A focus on polyploidy serves as the foundation for understanding the functional mechanisms driving cellular plasticity and the potential role of syncytia in shaping evolutionary outcomes. This interdisciplinary approach seeks to provide a clearer understanding of the intricate molecular and epigenetic connections that define syncytial organization and its broader implications in cellular evolution and pathology.
A Brief History of Epigenetics
Epigenetics represents a highly complex and dynamic field, distinguished by its capacity to regulate cellular processes without altering the underlying DNA sequence.8 Central to its function is the regulation of gene expression and the generation of diverse phenotypes from a single genome, contributing to tissue differentiation, polyphenism, and phenotypic plasticity.9,10 This dynamic regulation is particularly evident in the distinction between germline and somatic DNA in eukaryotes, a process that relies heavily on epigenetic mechanisms such as histone modification, DNA methylation, and the activity of noncoding RNAs.14
The evolutionary history of epigenetics underscores its profound role in genomic organization and regulation, especially in the context of mobile genetic elements. During eukaryogenesis, the segregation of germline and somatic lineages emphasized the dual roles of epigenetic modulation: in the germline, it facilitates genome reprogramming and transgenerational inheritance of epigenetic information, while in somatic cells, it is associated with processes like polyploidy, genomic rearrangements, and the regulation of extrachromosomal DNA.14 These mechanisms highlight the adaptability and plasticity conferred by epigenetic regulation across biological systems.
Heritable “epimutations” further exemplify epigenetics’ evolutionary significance, providing a basis for phenotypic variation and evolution.15 Moreover, parental genome merging and global epigenetic reprogramming establish epigenome asymmetry, which directs cell differentiation and fate.16
The mechanisms underlying epigenetic regulation include DNA methylation, histone modifications, and noncoding RNAs. DNA methylation, predominantly occurring at cytosine bases to form 5-methylcytosine, is a key means of transmitting epigenetic information during cell replication.17,18 Histone modifications, including methylation and acetylation, regulate chromatin structure and gene expression. These modifications are mediated by enzymes like histone methyltransferases and acetyltransferases and can influence or be influenced by DNA methylation.17,18
The discovery of regulatory noncoding RNAs (ncRNAs) has further expanded our understanding of epigenetic regulation. While early research focused onthe maintenance of RNAs such as ribosomal RNAs and transfer RNA, recent studies highlight the role of regulatory RNAs, including short interfering RNAs, microRNAs, PIWI-interacting RNAs and long non-coding RNAs, in processes like messenger RNA degradation, splicing, translation, and post-translational modifications.19
Additionally, protein complexes like Polycomb and Trithorax groups play critical roles in maintaining gene expression patterns. These complexes, with opposing actions, regulate developmental processes, cell differentiation, and environmental responses by activating or repressing transcription. Polycomb group proteins, particularly Protein Regulator of cytokinesis 1 and Polycomb repressive complex 2, and Trithorax group proteins are recruited to specific genomic regions by DNA-binding proteins or ncRNAs, underscoring the intricate interplay between chromatin structure and epigenetic regulation.20
Epigenetic mechanisms are also closely linked to pathological processes, particularly cancer. Dysregulation of DNA methylation, histone modifications, and noncoding RNAs contributes to tumor progression and therapeutic resistance.21 The convergence of these mechanisms highlights the centrality of epigenetics in shaping cellular behavior, with profound implications for development, evolution, and disease.
Beginning of the Path to Syncytia
Polyploidy and syncytia represent significant cellular phenomena with intricate interconnections, playing crucial roles in physiological and evolutionary processes. Polyploidy, defined as whole genome duplication (WGD), is observed across various taxa, including plants, fungi, invertebrates, and vertebrates.22 This phenomenon originates through mechanisms like endoreplication, cell fusion, and cellular cannibalism, providing adaptive advantages such as enhanced biomass production, increased tolerance to genomic stress, and a pivotal role in tissue regeneration.3–5 However, unregulated polyploidy often results in genomic instability.23,24 To integrate a general compilation of polyploidy in different organisms, we highlight advantages and disadvantages of polyploidization (and associated outcomes) in Fig.1.

Advantages and disadvantages of the polyploidization process. Syncytia offer several biological advantages. These include heterosis, where genetic diversity leads to enhanced biological functions, redundancy of genes that provide a buffer against mutations, and self-fertilization mechanisms in plants that ensure reproduction under challenging conditions. Syncytia also allow greater plasticity of the genome, enabling organisms to adapt to various environments. Homologous recombination in syncytia ensures efficient DNA repair, while their ability to synthesize proteins at high rates supports survival in restrictive environments like fungi and plants. The restricted phenotypic expression of deleterious recessive mutations in syncytia prevents the manifestation of harmful traits and aids in tissue repair and regeneration, highlighting their role in developmental and adaptive processes. However, there are significant disadvantages tied to epigenetic instability and syncytial structures. Changes in gene regulation and impaired meiosis can disrupt normal cellular processes, leading to a broader range of mutational targets and increased genetic degradation. The accumulation of deleterious mutations over time poses long-term risks, exacerbating genetic instability. Syncytia also come with a high metabolic cost, as maintaining a multinucleated state requires significant energy expenditure. Epigenetic instability further complicates the scenario by altering gene expression patterns, potentially leading to diseases or reduced adaptability. Thus, while syncytia and epigenetic processes play vital roles in cellular function and adaptation, they present a complex balance of benefits and challenges that depend on the context and environment in which they occur. Note: figure created by Biorender.
Polyploidy is categorized into euploidy, involving alterations in entire chromosome sets, and aneuploidy, which affects parts of the genome. These categories canfurther be divided into autopolyploidy and allopolyploidy, reflecting whether the chromosomes derive from the same or different species.3 The physiological outcomes of polyploidy include increased DNA content, augmented RNA and protein production, enhanced substrate availability for DNA repair, and heightened cellular complexity.25–27
Evolutionarily, polyploidy offers resilience against double-strand breaks and deleterious mutations in oxygen-rich environments. Notably, all angiosperms have undergone WGD, and 30–80% of plants are polyploid. Although less common in animals due to developmental constraints, polyploidy has played a role in vertebrate evolution.28,29 Despite its advantages, polyploidy carries risks, such as epigenetic instability, aneuploidy, and disruptions in cellular architecture, which may lead to chaotic gene regulation.25,30,31 Furthermore, one of the practical examples of the influence of ploidy variation is that of polyploid giant cancer cells (PGCCs),13,32 critical for tumor survival, resistance, and immortalization in different tumors, such as breast33 and lung34 cancers.
Epigenetic mechanisms, including chromatin modifications and derepression of bivalent genes, significantly influence polyploidy. Transcriptional remodeling arises from interactions between divergent parental genomes in allopolyploids and from genome duplication itself.35,36 While polyploidy initially induces genomic instability, subsequent epigenomic modifications can stabilize the genome, suggesting a transient yet critical adaptive phase.37–39
The interplay between polyploidy and syncytia is evident in their shared generative mechanisms, such as endoreplication, cell-in-cell (CIC) structures, mitotic slippage, and cell fusion. These processes exemplify the dynamic relationship between genomic and epigenomic modifications, contributing to both normal and pathological states, as highlighted in Fig.2.40–42 Syncytia formation and polyploidization provide a cellular strategy to navigate evolutionary and physiological challenges, emphasizing their critical roles in maintaining cellular and organismal complexity. By integrating these concepts, this discussion highlights their interconnected roles in shaping cellular plasticity and adaptability.

Syncytium formation mechanism. Based on current syncytium research, the main origin mechanism is the cell fusion, which is based on numerical highlights from [1] to [5] in “A”, “B,” and “C”. “A” highlights an initial process of contact between cells [1] and [2], mainly because of chemoattraction, and, thus, there are contacts between transmembrane fusogenic proteins that allow cell surface contact and union between uninuclear cells. In “B,” the union of uninuclear cells product makes two different paths: (i) nuclear fusion, resulting in the ability to generate hybrids with reciprocal genetic exchange (horizontal transfer of genes) responsive to the microenvironment; (ii) no nuclear fusion results in syncytia generation, that is, an enlarged multinucleated cell with a predominance of internuclear communication due to gap junctions and chemical compounds in the medium, mainly. “C” points to a third moment in which, after syncytial formation, countless other cells may be subjected to the numerous mechanisms of “A,” whereas with the expression of fusogenic proteins and under the influence of the biochemical microenvironment [3], [4], and [5] manage to merge and add more nuclei. Note: figure created by Biorender.
Epigenetics Behind the Origin of Syncytia
The relationship between genomic quantity and phenotypic outcomes becomes particularly significant when considering the role of polyploidy and its parallels with the formation of syncytial structures. Syncytia, defined as multinucleated cells sharing a common cytoplasm and functioning as a single unit, are observed across diverse taxa. Examples include myxogastric amoebas, protosporangiids, dictyostelids, vampirelids, and schizoplasmodids in protists.43 Fungi, renowned for their multinucleate states, rarely exhibit mononuclear forms.44 In plants, syncytial structures are exemplified by the placental endosperm, while in animals, muscle cells, osteoclasts, and the placenta serve as prominent examples.45,46
One key process in syncytia formation is cell fusion, a mechanism capable of inducing “genomic catastrophes.” This process involves the merging of two cells mediated by fusogenic proteins, resulting in hybrids with novel phenotypic, genotypic, and functional characteristics. Fusion can occur between homologous (homotypic) cells or heterologous (heterotypic) cells, with the latter potentially forming a heterokaryon. Upon nuclear fusion (karyogamy), the heterokaryon transitions into a synkaryon. Syncytial formation thus introduces significant complexity, especially in cases of both homologous and heterologous fusions, as shown in Fig.3.47,48
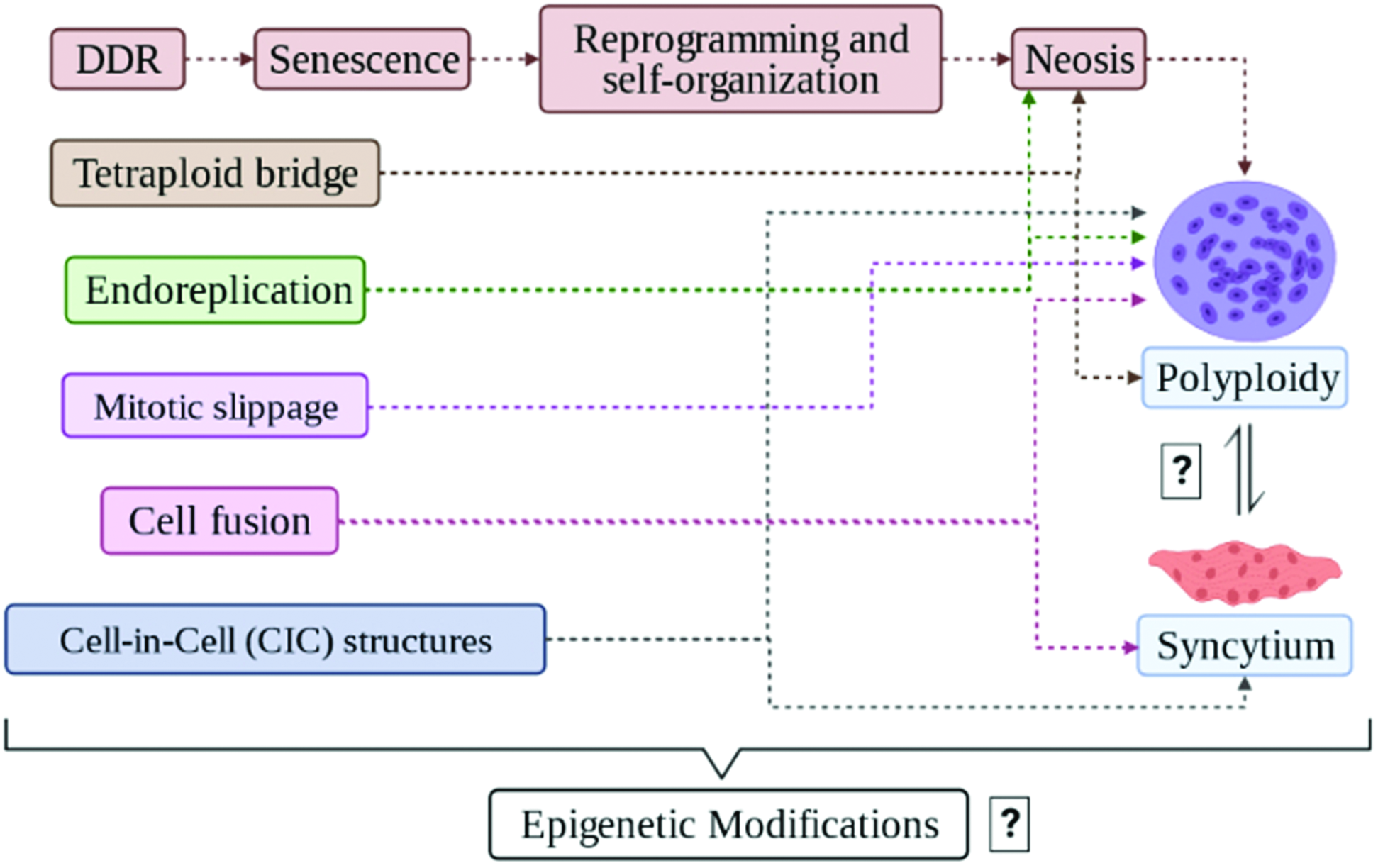
General processes linked to syncytization and polyploidization. Polyploidy comes from multiple processes: endoreplication, cytokinesis failure, cell fusion, CIC structures (cell cannibalism, entosis, emperipolesis, and others), mitotic slippage, tetraploid bridge, and neosis. Syncytia are formed from processes in common with polyploidy, such as endoreplication, cell fusion, CIC structures, and others that are poorly understood. The possible relationship of the common origin of both processes is the transience that is subjected to epigenetic modifications still very elusive. Note 1: polyploidization can result in amoeboid supergiant nurse cells, P1 oncogenerative cells, aneuploid cells, polyploid giant cancer cells (PGCCs), polyploid cyst-like structures (aCLSs), and numerous other cell types. Note 2: figure created by Biorender.
The influence of epigenetics on cell fusion has gained increasing attention. Symmetrical fusion (between homologous cells) and asymmetrical fusion (between heterologous cells or a mix of homologous and heterologous cells) provide ideal conditions for epigenetic reprogramming and the emergence of new phenotypes. This phenomenon is notably observed in pleomorphic giant cells (PGCs), which are multinuclear structures derived from tumor cells. While the origins of PGCs—whether through cell fusion or division failure—remain unclear, their multinucleated state suggests a syncytial nature.48–50
The evolutionary and functional significance of syncytia raises several questions. For instance, are multinucleated forms across eukaryotic supergroups evolutionary convergences, or do they reflect an ancestral state? The multinucleated condition may have been advantageous in ancestral cellular environments, allowing nuclei to complement one another. This complementation would mitigate the effects of mutations that might otherwise be lethal in mononuclear cells, thus enhancing the survival of multinucleated organisms.7,43
From an evolutionary perspective, syncytia in ancestral environments functioned more as populations of nuclei and mitochondria than as individual cells. In such populations, genetic diversity accumulated within the shared cytoplasm, with advantageous alleles in one nucleus compensating for deficiencies in others. This collective functioning underscores the adaptive benefits of the syncytial state, particularly in the context of environmental pressures and cellular survival.43
Despite these insights, the molecular mechanisms underlying syncytia formation remain poorly understood. Questions persist regarding the evolutionary origins, molecular content provisioning, and precise functions of syncytia. Epigenetics emerges as a critical factor in addressing these gaps, highlighting its central role in shaping syncytial structures and their broader implications across biological systems. Further exploration of this interplay promises to unravel the complexities of syncytial biology and its evolutionary significance.
Syncytia and the Intimate Relationship with Epigenetics
The relationship between syncytial formation and epigenetics reveals intricate mechanisms that integrate genetic and epigenetic modifications with cellular differentiation and adaptability. Syncytia, such as the STB in the placenta, represent examples where epigenetic processes play a fundamental role. Genetic alterations in signaling pathways, including mitogen-activated protein kinase, Transforming growth factor, and Neurogenic locus notch, have been observed during the formation of STB, reflecting the interplay between signaling and epigenetic regulation.51 Histone modifications, such as deacetylation and acetylation of histone H3 residues, appear to regulate critical genes involved in cytotrophoblast differentiation, such as TEA domain transcription factor 4, tumor protein p63, ovo like transcriptional repressor 1, Chorionic gonadotropin, LIM Homeobox 4, and synapse defective Rho GTPase homolog 1.52 These findings highlight the dynamic relationship between histone modifications and syncytial development.
Epigenetic changes in DNA and histone methylation have also been documented in trophoblastic cell line models such as forskolin-stimulated BeWo cells. Histone markers associated with active transcription, such as H3K4me3, H3K9ac, and H3K27ac, were upregulated during syncytial formation, suggesting that epigenetic and transcriptional changes are deeply interwoven with the syncytial state.51
The relationship between syncytial formation, epigenetics, and viral mechanisms further underscores the complexity of these processes. Viruses, including endogenous retroviruses and infectious agents such as human papillomavirus, have been shown to induce genomic and epigenomic destabilization through cell–cell fusion, often mediated by viral proteins such as syncytin-1. Viral mechanisms, such as the methylation of CpG islands genomic regions that contain a high frequency of cytosine and guanine nucleotides, connected with a phosphodiester bond, facilitate processes like endocytosis and syncytial formation, demonstrating how viral epigenetic modifications can drive cell fusion and structural reorganization.53,54
Syncytia also play a critical role in cancer biology, where their presence has been linked to genetic and epigenetic instability. Multinucleated giant cells, often seen in tumors, exhibit characteristics of syncytia and may arise through mechanisms such as mitotic catastrophe or cell fusion, leading to polyploid cells capable of overcoming genomic instability.40,48,55 These giant cells may serve as reservoirs of genetic diversity, facilitating tumor evolution and adaptation. The connection between syncytia and ancestral unicellular survival strategies offers a compelling framework for understanding tumor resilience and heterogeneity.
Oncogenic-enveloped viruses further demonstrate the role of syncytia in cancer. That phosphonic proteins, including syncytin-1, are utilized by these viruses to mediate cell fusion, resulting in the reprogramming of epigenetic landscapes.50 Syncytin-1, which is essential for STB formation, exhibits dynamic regulation in placental tissue through methylation levels.50 Intriguingly, similar processes are observed in tumors, where hypomethylation of retrotransposon-derived genes, including syncytin, contributes to tumor invasiveness.56
Studies provide additional evidence for the parallels between placental and cancer biology. Doello57 proposed that the hypoxia-mediated overexpression of methyltransferases could induce methylation of CpG islands in tumor cells, conferring trophoblastic characteristics to malignant cells. Similarly, Macaulay etal.58 observed that both placental and cancer cells undergo global hypomethylation, resulting in the activation of retrotransposons and facilitating cellular invasion. This epigenetic reprogramming is a key factor in the syncytial and invasive behaviors shared by cancer and placental cells.
Finally, the potential for syncytial transdifferentiation highlights the versatility of these structures. Schichor etal.59 demonstrated that human mesenchymal stem cells (hMSCs) can fuse with embryonic stem cells, adopting the recipient cell's phenotype. This capacity for transdifferentiation, while promising for cell therapy, also poses risks in the context of tumors, as hMSCs may integrate into glioma cells, emphasizing the dual-edged nature of syncytial mechanisms.
These findings collectively underscore the centrality of syncytia in the crosstalk between epigenetic regulation, viral processes, and cancer biology. The robust adaptability of syncytia, mediated by epigenetic and genomic reprogramming, highlights their role as evolutionary and pathological innovations. Understanding the molecular underpinnings of syncytial formation offers new perspectives on cellular plasticity, evolution, and disease progression.
Correlations Between Syncytia and Cell Fates
The relationship between changes in cell fate and epigenetic control reveals a complex and dynamic system that governs cellular development, plasticity, and specialization. This intricate process is guided by gene regulatory networks (GRNs) and transcriptional regulatory complexes (CoRCs), which ensure precise transitions in spacetime to achieve cellular specification and proliferation.60,61 Gene expression and epigenetic mechanisms play central roles in cell specification, as demonstrated in studies on transcriptional progressions and their regulation during cell state transitions.61,62
Conrad Waddington’s epigenetic landscape, developed in 1957, serves as a foundational framework for understanding these transitions, highlighting how cells navigate through developmental paths dictated by transcriptional changes and epigenetic modifications.63 These processes often involve points of divergence where cell states shift until reaching attractor states, which represent stable, yet not necessarily permanent, cellular outcomes.61,64 Recent findings challenge the idea of deterministic and unidirectional differentiation, suggesting that even differentiated cells retain plasticity, enabling reprogramming and adaptation to oscillatory stimuli via epigenetic mechanisms, nuclear organization, and cellular metabolism.65–68
This adaptability underscores the role of syncytia in epigenetic regulation and cell fate transitions. Syncytia, which are multinucleated cell structures formed by cell fusion, exemplify the capacity of epigenetic mechanisms to influence cellular outcomes in dynamic environments. Studies by Chaturvedi and Vijayraghavan69 reveal that syncytial reprogramming in the anterior alar muscle ofDrosophila involves de-differentiation and re-differentiation regulated by Hippo signaling and c-Jun N-terminal kinase, demonstrating how subtle epigenetic controls govern cellular plasticity.
Viral mechanisms also exploit epigenetic reprogramming to influence cell fate, as seen in the SARS-CoV-2 virus, which utilizes CpG methylation and syncytium formation to evade host immune responses, contributing to severe disease outcomes.70 Similarly, Physarum syncytia maintain nuclei with distinct transcriptome states regulated epigenetically, enabling rapid transcriptional responses to environmental demands.2
Human embryonic stem cells further illustrate the connection between syncytia, epigenetics, and cell fate. These cells exhibit remarkable plasticity and can transition into trophoblastic lineages under epigenetic modifications. Such transitions involve the expression of GATA binding protein 3 for syncytia formation, cell fusion with maternal immune cells, and Wnt signaling to determine trophoblastic lineages.71–73 Additionally, the TWIST helix-loop-helix (bHLH) transcription factor gene mediates myogenic plasticity during syncytial muscle cell transdifferentiation, with implications for metastasis through the regulatory axis Twist/Yorkie/fibroblast growth factor receptor.74
Cancer biology offers profound insights into the relationship between syncytia, epigenetics, and cell fate changes. PGCCs mirror evolutionary processes, using syncytialization and depolyploidization to recover from DNA damage and generate genetic diversity.48,75 Impaired cytokinesis and polyploidy in cancer cells promote genetic and epigenetic instability, fostering tumor adaptability.55 Moreover, multinucleated cancer syncytia facilitate immune evasion via heterotypic CIC structures, altering immune cell fates and enhancing malignancy.76 Wu etal.77 further describe therapeutic opportunities in targeting these syncytial bodies using oncolytic viruses, demonstrating the interplay between immune responses and epigenetic reprogramming.
Epigenome dynamics also regulates cell fate in embryonic and extraembryonic lineages. Bivalent enhancers in human trophoblastic stem cells mediate histone modifications that balance gene activation and repression, influencing differentiation and tumorigenesis.78 Furthermore, the biophysical and biochemical properties of connexin channels in tumor syncytia provide a platform for metabolic cooperation and rapid signaling, regulated by epigenetic modifications.79 These properties contribute to the pluripotency and cellular reprogramming observed in cancer.
Syncytia and polyploid cells thus represent an evolutionary bridge between unicellularity and multicellularity, demonstrating antifragile epigenetic regulation that allows adaptation to diverse environments. Their study offers profound insights into cell identity, evolution, and potential therapeutic strategies for complex diseases like cancer. These findings highlight the transformative potential of understanding syncytia as key players in the regulation of cell fate, advancing thefields of developmental biology, epigenetics, and translational medicine. This is supported by some pioneering research and attempts that will be developed ahead with a central approach in cancer by highlighting the ability of syncytia under the influence of epigenetics to reach a threshold of change in cellular fates as shown in Fig.4.

Crosstalk between syncytia, polyploidy, epigenetics, and cell fate. Continuous stress and unstable entropy are responsible for biological responses to “Genome Chaos,” “Epigenome Chaos,” and “Quantum Chaos” (1). Such distinct responses provide a basic need for cell survival based on new CoRC (“core regulatory complex”) and GRN that provide their own identity (2). Thus, in cases of polyploidy and syncytia, the gathering of numerous uninuclear cells leads to an association of multiple identities and attractor states, providing a broad genomic, epigenomic, transcriptomic, proteomic, and metabolomic structures, as well as the benefits provided by polyploidization and syncytialization (3). Therefore, there is a high genetic and epigenetic heterogeneity, benefiting the development of a sustainable cellular transitory state by reprogramming and maintenance of genetic and epigenetic integrity together with numerous mechanisms (4). Through such a transient state, the cell becomes capable of responding to multiple stresses much faster and accessing different cellular destinations, ensuring survival and perpetuation in the face of “extinction,” maintaining a chemical, physical, and biological balance (5). Note: figure created by Biorender.
In light of such information, studies about the interplay between polyploidy, syncytia, and epigenetics reveal a promising field. Polyploidy and aneuploidy drive genome evolution in fungi, highlighting their relevance for cellular adaptation. The role of PGCCs intumor macroevolution underscores how ploidy alterations impact tumor plasticity, while dividing polyploid cells can reduce genetic instability, a factor linked to neoplasia development.80–82 Syncytia, on the other hand, represent a key mechanism of cellular interaction. The integration of these findings with advances in epigenetics provides critical insights into cellular regulation. Additionally, exploring these interactions from an integrative perspective, as proposed in evolutionary parallels between plant and animal systems, broadens the scope of research.83 Future studies could delve deeper into the molecular mechanisms that govern the interplay between polyploidy, syncytia, and epigenetic regulation, with a focus on identifying specific pathways that mediate cellular resilience and adaptability (provide unprecedented insights into how these processes dynamically shape cellular behavior).84–88
Conclusion
The transition from unicellular to multicellular life marked a pivotal point in evolution, bringing about profound changes in cellular organization and necessitating the development of sophisticated intercellular communication, coordination, and interaction mechanisms. This complexity is exemplified by the interplay between syncytia formation, polyploidy, and epigenetic regulation, which together drive cellular plasticity, adaptation, and survival in dynamic and entropic environments. A growing body of research underscores the critical role of epigenetic control in regulating gene expression and phenotypic plasticity within these processes, offering unprecedented potential for numerous fields of study. Despite this, the literature remains fragmented, with limited integration of polyploidy, syncytia, and epigenetics, creating a narrow perspective on their collective impact. By reconstructing the evolutionary trajectory of polyploidy and syncytia through a holistic approach grounded in recent and foundational studies, this work bridges these gaps, highlighting the intricate molecular and epigenetic mechanisms that shape cellular fate and biological outcomes. Moreover, this analysis sheds light on the syncytium-epigenetics-cell fate axis, emphasizing its significant implications in pathological states such as cancer and its potential for personalized therapeutic innovations. The convergence of syncytial complexity, epigenetic regulation, and polyploidy not only advances our understanding of cellular evolution and adaptation but also paves the way for groundbreaking applications in oncology and beyond.
