Abstract
Abstract
Background:
DNA methylation is one of the crucial mechanisms by which a cell maintains its epigenetic landscape. Regulation of many genes takes place via DNA methylation. The silencing of tumor suppressor genes in cancer through hypermethylation of their promoters is one such example. To study the expression patterns of several genes in various disease conditions, there are many ways employed. One such mechanism is to reverse the methylation in these gene promoters and observe their upregulation and associated phenotypes.
Method:
To reverse DNA methylation, there are DNA methyltransferases inhibitors in use such as 5-azacytidine and 5-aza-2′-deoxycytidine. These drugs are clinically used for treating Myelodysplastic disorders. However, their use as an epigenetic reprogrammer for various cancer cells has been a commonly explored aspect these days.
Results:
There are many studies involving the usage of these nucleoside analogs for reversing the methylation of genes in context of various cancers, such as cancers associated with breast, lungs, digestive system, immune system, reproductive system, head and neck, thyroid gland, and nervous system.
Conclusion:
The usage of 5-Azacytidine and 5-Aza-2′-deoxycytidine as epigenetic reprogrammers is a very widely employed mechanism to understand the role of several genes in various disease contexts specifically cancer.
Introduction
Genes act like the scripts for the synthesis of messenger RNAs (mRNAs) and proteins. The genetic information stored in genes is transcribed and translated to functional proteins, which then regulate cellular functions. Such a vast flow of information needs strict checkpoints to ensure smooth functioning. Hence, other than the various regulatory mechanisms at transcriptional and translational levels, damage repair pathways, and post-translational modifications, another way of gene regulation called epigenetic regulation exists. Epigenetic regulation encompasses various mechanisms that control gene expression without altering the underlying DNA sequence. Several types of epigenetic regulations have been identified, each playing a crucial role in cellular development, differentiation, and disease progression.1 One such example for epigenetic regulation is histone modification. Histones, theproteins around which DNA is wrapped, undergo modifications such as acetylation, methylation, phosphorylation, and ubiquitination. These modifications alter the chromatin structure, either promoting or inhibiting gene transcription.
Similarly, noncoding RNAs, such as microRNAs (miRNAs/miRs), and long noncoding RNAs also contribute to epigenetic regulation. miRs bind to target mRNAs, mostly leading to their degradation or inhibition of translation, sometimes leading to the activation of gene targets depending on the binding site and extent of complementarity.2 Epigenetic regulation is also done through chromatin remodeling. It is a process that modifies the accessibility of DNA to transcriptional machinery. Adenosine triphosphate-dependent chromatin remodeling complexes utilize energy to reposition nucleosomes and expose or hide specific DNA regions, thereby influencing gene expression.1 Genomic imprinting is another epigenetic phenomenon where specific genes are expressed in a parent-of-origin-dependent manner due to differential DNA methylation or histone modifications. DNA methylation is one of the most prominent epigenetic modifications involving adding a methyl group to residues in DNA. Methylation influences gene expression by inhibiting the binding of transcription factors or by recruiting proteins that silence gene activity, whereas acetylation of histones leads to the activation of gene expression. Overall, these diverse epigenetic regulations work in concert to establish and maintain gene expression patterns in different cell types, and dysregulation of these processes contributes to various diseases, including cancer, neurological disorders, and developmental abnormalities.1
In the context of cancer, an interplay of various epigenetic dysregulations of cancer-causing or preventing genes navigates the disease progression. If the genes are associated with regulating cellular proliferation, extra-cellular matrix maintenance, cell growth, essential signaling pathways, or other migration-invasion-related processes, the slightest change in their epigenetic pattern would cause dysregulation of the cellular processes they are involved with. Since epigenetic modifications are reversible, there have been attempts to restore the expression levels of cancer-associated genes (e.g., tumor suppressors) to normal levels to rescue cells from cancer and find therapeutic interventions. To epigenetically target a gene, there are multiple ways. It could be through the alteration of epigenetic modifying machinery, by changing the accessibility of genes to the machinery or by removing the modification that has already been made. Expression of specific genes is activated by histone acetylation at promoter or enhancer regions of the gene. At the same time, histone deacetylases remove the acetyl groups and reverse the gene activation. Similarly, depending on the methylation or demethylation, a gene will be expressed or silenced. Methylation at the promoter region silences genes, whereas the methylation on the gene body leads to the expression of specific genes. Out of all the epigenetic modifications, DNA methylation is studied the most in the context of cancers.1 DNA methylation occurs through the assistance of DNA methyltransferases (DNMTs), which facilitate the transfer of a methyl group from S-Adenosyl methionine to substrates. Mammalian cells contain four types of DNMTs: DNMT1, DNMT2, DNMT3a, and DNMT3b. DNMT1 predominates among them and is primarily tasked with preserving DNA methylation patterns. Conversely, DNMT3a and DNMT3b are responsible for initiating de novo DNA methylation in newly synthesized DNA strands. DNMT2, however, is recognized for its role in methylating RNA molecules. The presence of these enzymes is indispensable for regulating gene expression appropriately, and their inhibition can lead to modifications in gene methylation profiles.3
DNMT inhibitors are of two categories: nucleoside analogs and non-nucleoside analogs. Nucleoside analogs consist of modified bases that are incorporated into DNA during replication. For example, 5-azacytidine and 5-aza-2′-deoxycytidine are two of the most studied DNMT inhibitors with modified cytidine residues (Fig.1).4 DNMTs covalently bind with the adducts of DNA formed by the incorporation of modified nucleoside analogs and inhibit DNA methylation. Zebularine is another modified nucleoside that gets incorporated in RNA. Both 5-azacytidine and 5-aza-2′-deoxycytidine are Food and Drug Administration (FDA)-approved drugs for myelodysplastic syndromes (MDS) and chronic myelomonocytic leukemia (CML), as these drugs efficiently bind with the new DNA content in dividing cancer cells. They have also shown therapeutic potential in various other cancers.4 While they share similarities in their mechanisms of action, there are some notable differences between the two drugs. Both 5-azacytidine and 5-aza-2′-deoxycytidine work by incorporating themselves into DNA during replication, leading to the inhibition of DNA methyl transferase enzymes. By inhibiting these enzymes, along with promoting the passive demethylation during replication, they also prevent adding any more methyl groups to DNA, resulting in DNA hypomethylation and subsequent reactivation of silenced genes. One key difference between the two drugs lies in their chemical structures. 5-Azacytidine is a pyrimidine analog, while 5-aza-2′-deoxycytidine is a cytidine analog (Fig.1). This structural difference may contribute to variations in their pharmacokinetic properties and bioavailability.4
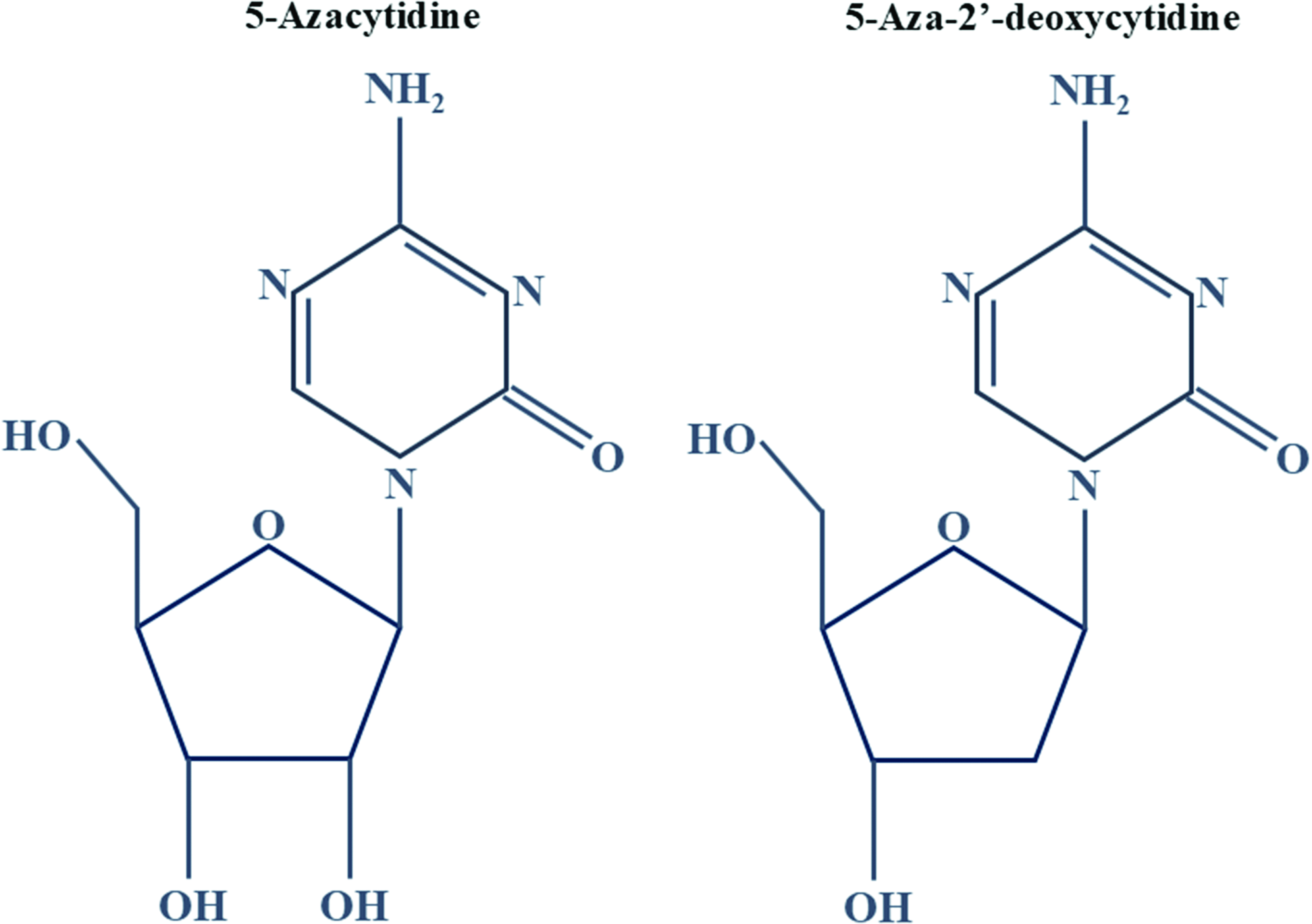
Chemical structure of 5-azacytidine and 5-aza-2′-deoxycytidine.
While 5-azacytidine and 5-aza-2′-deoxycytidine share similarities as DNA demethylating agents, they differ in their chemical structures, approved indications, dosing schedules, and potentially their side effect profiles. Their distinct characteristics provide clinicians with alternative options for tailored treatment approaches for different malignancies. These drugs are also being tested as therapeutic modalities in solid cancers. Nucleoside analogs tend to cause genomic instability and mutagenicity as side effects.4 It is interesting to investigate the roles of these drugs in various clinical and research contexts along with their history, current practices, and future implications.
Discovery of 5-Azacytidine and 5-Aza-2′-Deoxycytidine
The pyrimidine analogs 5-azacytidine and 5-aza-2′-deoxycytidine were discovered in the mid-1960s by Pliml and Sorm.5 Shortly thereafter, 5-azacytidine was isolated microbiologically from the fermentation beer of Streptoverticillium ladakanus.6 5-Azacytidine distinguishes itself from cytosine primarily through the nitrogen present at position 5 (Fig.1). The hypomethylating effect seems to hinge primarily on the existence of this modified C5 position. Initially, these drugs were thought to be just antimetabolites, but eventually, various roles of these two drugs came into picture. Between the two, 5-azacytidine was synthesized as a therapeutic for acute myeloid leukemia (AML), but there were early reports where this drug was considered as a mutagen and inducer of chromosomal breakage.7 Recognized early on for their potential as anticancer agents, these compounds were first clinically studied in the 1980s.6 But progress was achieved when lower dosages were explored, which proved noncytotoxic yet capable of reactivating epigenetically silenced genes. Nearly 40 years later, 5-azacytidine finally received FDA approval in 2004/2005 for the treatment of certain subtypes of MDS. In 2006, the FDA further approved 5-aza-2′-deoxycytidine for MDS treatment. The endorsement of low-dose regimens by the FDA, along with supportive data, has not only refined their application in hematological malignancies but also holds promise for potential use in solid tumors.6
Mechanism of Action of 5-Azacytidine and 5-Aza-2′-Deoxycytidine
It has been well established that DNA methyl transferases act by interacting with the 6th Carbon residue of cytosine residue in the DNA through a cysteine residue in their enzyme catalytic site. This process enhances the flow of electrons to 5th Carbon, leading to the attack on the methyl group of S-Adenosyl methionine, the methyl group donor. Extracting a proton from the 5th Carbon and subsequent beta-elimination facilitates the reestablishment of the double bond connecting Carbons 5 and 6, causing the release of the enzyme.8 Consequently, the DNA retains a methylated cytosine. Given that the covalent bond formed between the nucleotide analogs and DNMT is irreversible, the release of the DNMT is impeded, resulting in enzyme inhibition. This mechanism, identified in 1984, was proposed as a potential mode of action for both 5-azacytidine and 5-aza-2′-deoxycytidine. Studies revealed that at high doses, these drugs are cytotoxic, while at low doses, they induce differentiation and demethylation (Fig.2).8

Action of 5-azacytidine and 5-aza-2′-deoxycytdine at high and low doses leading to various cellular outcomes.
Comparative Efficacy and Clinical Considerations
The culmination of three consecutive Cancer and Leukaemia Group B studies, including a phase III randomized controlled trial, comparing 5-azacytidine with best supportive care, has highlighted its efficacy. Throughout these studies, the dosage of 5-azacytidine remained consistent at 75 mg/m2. In phase III clinical trials involving patients with MDS, both 5-azacytidine and 5-aza-2′-deoxycytidine showed similar response rates, but their impact on overall survival varied. 5-Azacytidine resulted in a notably longer median overall survival among higher-risk MDS patients compared to standard care. Conversely, 5-aza-2′-deoxycytidine did not show a significant survival improvement in a comparable trial.6 5-Azacytidine got its FDA approval in 2004, whereas 5-aza-2′-deoxycytidine was approved in 2006 as an intravenous drug for MDS with a recommended dose of 20 mg/m2 daily for 5 days in a 28-day cycle. The storage for both drugs is similar. 5-Azacytidine exhibits chemical instability due to nitrogen substitution at position 5, necessitating its manufacture as a lyophilized powder. Refrigerated storage maintains stability for up to 8 h; however, it must be administered within 1 h of reconstitution. Both intravenous and subcutaneous administration of azacytidine offer comparable bioavailability. Subcutaneous administration leads to rapid absorption, reaching peak concentration within 30 min.6 5-Azacytidine has a half-life of 1.5–2.3 h and becomes undetectable 8 h post-administration. Common side effects of both drugs include myelosuppression, gastrointestinal disturbances, and fatigue, but the specific incidence and severity of adverse events may differ. While in the case of invitro experiments, the treatment dosage and duration are decided based on the cytotoxicity exerted by different doses of these drugs.6
Absorption and Intracellular Processing
After administration, 5-azacytidine and 5-aza-2′-deoxycytidine can be absorbed through various routes depending on the formulation and administration method, such as oral ingestion or intravenous injection. Once absorbed, they may undergo cellular uptake facilitated by specific transporters present on the cell membrane. The transporters involved in the uptake of 5-aza-2′-deoxycytidine and 5-azacytidine have not been fully elucidated, but research suggests that equilibrative nucleoside transporters and concentrative nucleoside transporters are implicated in facilitating the passage of nucleoside analogs across cell membranes.9 Within the cell, 5-azacytidine can undergo phosphorylation by enzymes such as uridine-cytidine kinase, leading to the formation of its active metabolites. These metabolites can then be incorporated into newly synthesized DNA, where they exert their pharmacological effects by inhibiting DNMTs and promoting DNA demethylation. Additionally, 5-azacytidine may undergo metabolic transformation by enzymes such as cytidine deaminase, leading to the formation of various metabolites that can be further processed and eliminated from the body.
Controversies, Challenges, and Future Implications
While the FDA approval for both 5-azacytidine and 5-aza-2′-deoxycytidine stands for the treatment of MDS and AML, there are several clinical trials going onto decide more suitable treatment duration and regimens. But, there are several controversies associated with the usage of these drugs. While both drugs have shown efficacy in treating certain hematological malignancies such as MDS and AML, there may be variability in response rates among different patient populations. Some patients may experience significant improvements in overall survival and disease progression, while others may have limited benefit. The optimal dosing regimen for these drugs remains a topic of debate even today. While standard dosing schedules have been established, there is ongoing research to assess alternative dosing strategies that may enhance efficacy or minimize toxicity. Additionally, questions persist regarding the duration of treatment and the potential benefits of maintenance therapy. The 5-azacytidine treatment is associated with various adverse effects, including myelosuppression, gastrointestinal symptoms, fatigue, and injection site reactions. Managing these adverse effects while maintaining treatment efficacy can be challenging, and balancing the risks and benefits of therapy is crucial. Resistance to 5-aza-2′-deoxycytidine treatment can develop over time, leading to disease relapse or progression. The mechanisms underlying 5-aza-2′-deoxycytidine resistance are not fully understood, but they may involve genetic mutations, epigenetic alterations, or changes in drug metabolism pathways. Managing and overcoming resistance to 5-aza-2′-deoxycytidine therapy represent significant challenges in clinical practice. The 5-azacytidine therapy can be expensive, particularly for patients without adequate insurance coverage or access to financial assistance programs. Cost considerations may limit access to treatment for some patients, leading to disparities in care. There is ongoing interest in exploring combination therapies involving azacytidine with other agents, such as targeted therapies or immunotherapies, to improve treatment outcomes. However, identifying optimal combinations and understanding potential interactions and synergies remain an area of active research. Beyond MDS and AML, these drugs are being investigated in clinical trials for the treatment of other hematological malignancies, such as chronic myelomonocytic leukemia (CMML) and CML, as well as solid tumors like lung cancer and ovarian cancer.10–11 These trials aim to explore its efficacy, safety, and potential in different cancer types and patient populations. There is ongoing research into the role of these drugs as maintenance therapy following induction chemotherapy or stem cell transplantation in patients with AML or MDS. Maintenance therapy aims to prolong remission duration and prevent disease recurrence. The role of 5-azacytidine and 5-aza-2′-deoxycytidine in genomic imprinting involves its ability to influence epigenetic modifications, particularly DNA methylation patterns, which play a crucial role in regulating gene expression in a parent-of-origin-specific manner. Genomic imprinting is an epigenetic phenomenon where specific genes are expressed in a parent-of-origin-dependent manner, meaning that only the maternal or paternal allele of the gene is expressed while the other allele is silenced through DNA methylation. This process is essential for normal development and growth, and disruptions in genomic imprinting can lead to various genetic disorders and diseases, including cancer. Studies have shown that treatment with 5-azacytidine can lead to changes in DNA methylation patterns, including alterations at imprinted gene loci. By reversing aberrant DNA methylation associated with imprinting disorders or abnormal gene expression, 5-azacytidine has the potential to modulate genomic imprinting and restore normal gene expression patterns. 5-Aza-2′-deoxycytidine has been studied for its impact on genomic imprinting in various contexts, particularly in diseases where aberrant DNA methylation patterns contribute to pathogenesis, such as cancer and imprinting disorders like Beckwith–Wiedemann syndrome and Prader–Willi syndrome.12 By inducing DNA hypomethylation, 5-aza-2′-deoxycytidine may potentially restore normal imprinting patterns and gene expression profiles in these disorders. Overall, the role of these drugs in genomic imprinting involves its ability to modulate DNA methylation patterns, thereby influencing the expression of imprinted genes and potentially contributing to the treatment of imprinting disorders and associated diseases.
In addition to their involvement in diverse therapeutic protocols and potential usage in reversing genomic imprinting, researchers are investigating the potential of both 5-azacytidine and 5-aza-2′-deoxycytidine as hypomethylating agents to elucidate hypermethylated genes associated with various diseases, notably cancer. This perspective positions these drugs as invaluable research tools as epigenetic reprogrammers in scientific inquiry.
5-Azacytidine and 5-aza-2′-deoxycytidine as epigenetic reprogrammers in different cancers
Breast cancers
Breast cancer ranks as the second most prevalent cancer worldwide, according to data from the GLOBOCAN 2022 database (https://gco.iarc.fr/en) with the highest occurrence observed among females. Exploring genes linked to breast cancer is imperative for refining drug treatments. In investigations concerning breast cancer, particularly triple-negative breast cancer, researchers have utilized both 5-azacytidine and 5-aza-2′-deoxycytidine to investigate gene methylation patterns. In one of the studies, breast cancer cell lines (e.g., MCF7 and MDA-MB-231) treated with 5-azacytidine exhibited increased expression of PTPRO, a gene from protein tyrosine phosphatase receptor family, previously shown to be methylated and silenced in breast cancer.13 Other than PTPRO, the functional significance of demethylation in breast cancer was also supported by the induction of another proto-oncogene BRF2 following 5-azacytidine treatment.14 Furthermore, treatment with 5-azacytidine resulted in an upsurge in E-cadherin expression, confirming its involvement in the silencing of this gene through methylation. Moreover, the 5-azacytidine treatment effectively sensitized MCF7-TAM breast cancer cells to tamoxifen, bolstering the potential of these drugs as sensitizers.15 The other examples are summarized in
Cancers associated with digestive system
Digestive system cancers encompass malignancies affecting organs like the esophagus, stomach, liver, pancreas, colon, and rectum. Risk factors vary but often include diet, smoking, alcohol, and genetics. Symptoms may notmanifest until later stages, highlighting the importance of screenings. Treatment involves surgery, chemotherapy, radiation, and immunotherapy tailored to the specific cancer type and stage. A significant number of studies in the context of these cancers utilized both 5-azacytidine and 5-aza-2′-deoxycytidine for checking the methylation status of genes involved in the pathogenesis. Few of the initial examples are of RUNX3, CDH1, and SOX17. Under hypoxic conditions, the esophageal cancer cell line Eca109 displayed methylated RUNX3, and the degree of methylation at its promoter decreased following 5-azacytidine treatment, resulting in increased gene expression.16 Like PTPRO in breast cancer cells mentioned earlier, the 5-azacytidine treatment of hepatocellular carcinoma cells was used to restore the expression of PTPRD due to its hypermethylated status detected by Combined Bisulphite Restriction Analysis.17 While CDH1 and SOX17 showed upregulation post-demethylation by 5-azacytidine,18 the same treatment increased the expression of KLF4 in human esophageal squamous cell carcinoma cell lines.19 After this, in 2006, using microarrays comprising probes for 22,284 genes, the contrasting RNA expression patterns after the treatment with 5-aza-2′-deoxycytidine between diffuse gastric cancer and intestinal-type gastric cancer were examined. Through this analysis, 16 methylated genes were identified, among which several were previously undiscovered.20 Other examples are tabulated in
Cancers associated with respiratory system
Lung-associated cancers encompass a range of malignancies affecting the lungs and nearby structures. Primarily caused by smoking, they include non-small cell lung cancer, small cell lung cancer, and mesothelioma. Symptoms often emerge late, leading to a challenging prognosis. Treatments vary but may involve surgery, chemotherapy, radiation, and immunotherapy. In A549 lung cancer cells, the administration of 5-azacytidine led to the hypomethylation of the ABCB1, subsequently resulting in decreased expression of the ABCB1 protein.21 In approximately one-third of the lung cancer cell lines examined (e.g., SPCA1 and A549), hypermethylation was identified in the promoter region of KEAP1. The impact of this hypermethylation on the modulation of KEAP1 mRNA levels was further validated through invitro treatment with 5-azacytidine in lung carcinoid, small lung cancer, and adenocarcinoma cell lines.22 The methylation of the DAPK promoter might have contributed to the downregulation of the DAPK gene and protein, consequently affecting the sensitivity of gefitinib in lung adenocarcinoma lines and inducing gefitinib resistance. However, demethylation of the DAPK promoter by 5-aza-2′-deoxycytidine could reverse gefitinib resistance.23 Other examples are mentioned in
Cancers associated with nervous system
Nervous system-associated cancers, such as gliomas, meningiomas, and neuroblastomas, originate from abnormal growth within the brain, spinal cord, or peripheral nerves. They can arise in any age group and present diverse symptoms, including headaches, seizures, or neurological deficits. Diagnosis typically involves imaging studies like magnetic resonance imaging scans and biopsy confirmation. Treatment approaches depend on factors like tumor type, location, and patient health, often comprising surgery, radiation therapy, chemotherapy, and targeted therapies. Despite advancements, challenges persist due to the intricate nature of the nervous system and potential for tumor recurrence, necessitating ongoing research for improved therapies and outcomes. While investigating the genes associated with the pathogenesis of nervous system-associated cancers, the promoter-silencing mechanism of cognitive function-associated gene EHD3 in gliomas was studied using sodium bisulfite sequencing, and it was found that there was removal of hypermethylation in response to 5-azacytidine treatment.24 Over a period of 28 days, glioblastoma cells upon the exposure to 5-azacytidine underwent astrocyte-induced differentiation, and the demethylation of exon 2 of POLGA was identified.25 A set of demethylation assays using 5-azacytidine was conducted in both macro-neuroblastoma cells, and the expression of PHOX2B was found to be upregulated in the former.26 On the other hand, neuroblastoma cells treated with 5-azacytidine exhibited selective upregulation of Human Endogeneous Retro Viral gene-like sequences.27 There are several other genes studied in the context of neuro oncology as mentioned in
Cancers associated with reproductive system
Reproductive cancers encompass a spectrum of malignancies affecting the reproductive organs, including the ovaries, uterus, cervix, vagina, testes, and prostate. These cancers can significantly impact fertility, sexual health, and overall well-being. Risk factors vary, including genetics, hormonal imbalances, infections, and lifestyle factors. Screening methods such as Pap smears, mammograms, and prostate-specific antigen tests aid in early detection. Treatment options range from surgery and radiation to hormone therapy and chemotherapy, tailored to the specific cancer type and stage. Despite advances, challenges remain in preserving reproductive function and improving outcomes for affected individuals. In ovarian cancer cell lines, the expression of both mRNA and protein levels of the cell adhesion molecule opioid binding protein was restored after 5-azacytidine treatment, indicating the hypermethylation associated silencing of this gene.28 In cervical cancer, both RNA and protein levels of a known tumor suppressor KLF4 exhibited a significant increase upon treatment with 5-azacytidine, indicating promoter methylation. KLF4 belongs to the Kruppel family of transcription factors.29 Gene expression restoration assay using 5-azacytidine was also employed in the context of endometrial carcinoma to check the methylation levels of LNPP5K and MYO1C. Both these genes were showing downregulation in the endometrial carcinoma cells.30 In HeLa cells, the methylation level of the SALL3 promoter showed a decrease after treating with 5 μM and 10 μM of 5-azacytidine. Additionally, an increase in the degree of promoter demethylation was observed, and this increase was dependent on the dosage of 5-azacytidine. The usage of 5-azacytidine acted like an epigenetic reprogrammer in this context as well.31 In prostate cancer, aggressive tumor cells have epigenetically silenced SPARC, which was demethylated and reactivated after 5-azacytidine treatment indicating the preexisted hypermethylation.32 Other examples are mentioned in
Cancers associated with immune system
Immune system cancers, also known as hematologic malignancies or lymphomas, originate from abnormal growth within the body's immune cells, including lymphocytes, plasma cells, and myeloid cells. These cancers encompass a diverse group of diseases, such as leukemia, lymphoma, and multiple myeloma, each with unique characteristics and treatment approaches. Symptoms may vary widely and can include fatigue, unexplained weight loss, and recurrent infections. Diagnosis often involves blood tests, bone marrow biopsies, and imaging studies. Treatment strategies typically include chemotherapy, immunotherapy, targeted therapy, and stem cell transplantation, tailored to the specific cancer type and patient's condition. Ongoing research aims to uncover novel therapies and improve outcomes for patients withimmune system cancers. The administration of 5-azacytidine to Burkitt’s lymphoma cells led to the induction of a zinc finger gene PRDM1, which was accompanied by the demethylation of PRDM1 promoter, indicating that DNA methylation plays a role in the transcriptional silencing of PRDM1.33 The CD20 expression, which was lost upon rituximab treatment in follicular lymphoma cells, was also shown to be restored post-5-azacytidine treatment.34 In primary leukemic cells obtained from seven relapsed AML patients, 5-azacytidine exhibited the ability to restore ATPsyn-β mRNA expression and enhance the sensitivity to adriamycin.35 In addition, the administration of 5-azacytidine also resulted in a reduction in the expression of DNMT1 and the restoration of KLF4 expression in mantle cell lymphoma cell lines too.36 The increase in PCDH17 mRNA expression mediated by 5-azacytidine was notably lower compared to the increase induced by trichostatin A (an histone deacetylase (HDAC) inhibitor) in leukemia cells.37 More examples are given in
Cancers associated with head and neck
Head and neck cancers (HNCs) encompass a group of malignancies that originate in the oral cavity, throat, larynx, nose, sinuses, and salivary glands. These cancers can significantly impact vital functions such as breathing, swallowing, and speaking. Major risk factors include tobacco use, alcohol consumption, and infection with certain strains of human papillomavirus. Symptoms may include persistent sore throat, hoarseness, difficulty swallowing, and a lump in the neck. Diagnosis involves physical examination, imaging tests, and biopsy. Treatment options include surgery, radiation therapy, chemotherapy, targeted therapy, and immunotherapy, often combined for optimal outcomes. Early detection and multidisciplinary care are crucial for managing HNCs effectively. In paired nasopharyngeal carcinoma clinical samples, a high percentage of methylation at the CLDN11 promoter correlated with a low mRNA expression level, as confirmed using 5-azacytidine.38 In oral squamous cell carcinoma, a subtype of HNCs, 5-azacytidine was used to identify tumor suppressor miRs.39 Few more genes have been analyzed using the DNMT inhibitors in HNCs, and they are given in
Cancers associated with thyroid cancer
In 2013, it was shown that the thyroid cancer cells exhibited upregulation of melanoma-associated gene MAGE4 following 5-azacytidine treatment, indicating reactivation of previously methylated genomic regions.40 The expression of HOXD10 was found to be decreased in both papillary thyroid cancer cell lines and tissues compared to adjacent normal thyroid tissues. However, the treatment with 5-azacytidine led to a reversal of this downregulation as well.41 Following the administration of 5-azacytidine, thyroid cancer cells exhibited an upregulation of HORMAD2 expression, which, when overexpressed, demonstrated a remarkable suppression of thyroid cancer cell viability, mobility, and invasiveness.42
Other cancer-associated genes that have been studied in various cancer contexts are mentioned in
Usage of DNMT inhibitors in epigenetically reprogramming of noncoding genes
Along with reprogramming the protein coding genes epigenetically, 5-azacytidine and its derivatives were also used to study several noncoding genes. In chronic lymphocytic leukemia (CLL) cells, 5-azacytidine was used along with trichostatin A to induce the expression of miR-34b/c synergistically.43 Methylation sensitive polymerase chain reaction showed decreased methylation of the MIR137 promoter upon 5-azacytidine treatment, leading to increased expression of miR-137.44 In prostate cancer cells, miR-34a mimic transfection and demethylation of the MIR34A promoter by 5-azacytidine led to its increased expression.45 The expression of miR-124 increases post-5-azacytidine treatment in gastric cancer cells, leading to decreased expression of SMOX.46 A novel microRNA called miR-2 located in the TRkC gene shows differential expression in colorectal cancer specimens post-5-azacytidine treatment. This microRNA was shown to regulate Wnt signaling pathway.47 Using the treatment of 5-azacytidine, miR-495 was shown to be a tumor suppressor in breast cancer, and this microRNA was found to silence the STAT3 oncogene to promote apoptosis and reduce cell proliferation and invasion.48 It was shown that the miR-335 expression increases upon the 5-azacytidine treatment of multiple myeloma cells, and it leads to the downregulation of insulin-like growth factor receptor l along with decrease in cell migration.49 Restoration of miR-34a expression by 5-azacytidine treatment in non-small cell lung cancer cells leads todownregulation of HMGA1, HMGB2, and BCL2 expression.50 There have been more studies where 5-azacytidine was shown to be the epigenetic reprogramming tool for miRs.51
Being the other derivative of 5-azacytidine, which we discussed earlier in detail, 5-aza-2′-deoxycytidine was also used as an epigenetic reprogrammer for miRs. In 2014, it was shown that the castration-dependent prostate cancers show a hypermethylated state of MIR146a. Demethylation upon the 5-aza-2′-deoxycytidine treatment resulted upregulation of miR-146α, leading to the delayed progression of cancer.52 In gastric cancer cells, miR-495 levels increased post-5-aza-2′-deoxycytidine treatment, which led to decreased levels of its gene target PRL3. The overexpression of this miR led to reduction in the ability of invasion and metastasis.53 Treatment with 5-aza-2′-deoxycytidine resulted in the reexpression of miR-34b/c in MEC1 cells of CLL.42 The involvement of miR-34a-5p in colorectal cancer as a tumor suppressor was also identified using the treatment of 5-aza-2′-deoxycytidine.54 Methylation in the CpG islands upstream of the MIR10b gene was identified after the treatment with 5-aza-2′-deoxycytidine of gastric cancer cells.55 Even in lung cancer cells, the aberrant hypermethylation and silencing of the MIR137 promoter were reversed after 5-aza-2′-deoxycytidine treatment.56 In triple negative breast cancer cells, it was shown that PTPN2 is epigenetically regulated by miR-124 using 5-aza-2′-deoxycytidine.57 The expression of miR-378 is reactivated after 5-aza-2′-deoxycytidine treatment in AML.58 Tumor suppressor miRs (e.g., miR-34a and miR-203) were reactivated in Hodgkin’s lymphoma post-5-aza-2′-deoxycytidine treatment.59 Different cell lines of esophageal squamous cell carcinoma showed reactivation of miR-218, followed by the treatment with 5-aza-2′-deoxycytidine to promote the aggressiveness of the cancer.60 The DNA methylation status, target gene levels, and the expression of miR-149 were checked in glioblastoma cells after 5-aza-2′-deoxycytidine treatment, and it was identified that the miR-149 levels increase post-treatment.61 CARF was identified as a new target for miR-451 in various cancer cells post-5-aza-2′-deoxycytidine treatment.62 The involvement of miR-19a/b in multiple drug resistance of gastric cancer cells was identified using 5-aza-2′-deoxycytidine treatment.63 In the same model system, it was also shown that demethylation by 5-aza-2′-deoxycytidine led to the demethylation and induction of both HOXA10 and miR-196b-5p.64 Ecotropic viral integration site 1 (EVI1)-induced hypermethylation of miR-9 in leukemia was reversed by treatment with 5-aza-2′-deoxycytidine to overcome the EVI1-associated resistance in leukemia cells.65 The impact of miR-342-3p on autophagy in B-cell lymphoma cells was investigated by demethylating its promoter using 5-aza-2′-deoxycytidine.66 In astrocytoma, miR-204-5p acts like a tumor suppressor to downregulate ezrin expression. This was identified using treatment by 5-aza-2′-deoxycytidine.67
Conclusion
DNMT inhibitors like 5-azacytidine and 5-aza-2′-deoxycytidine were employed to induce epigenetic alterations in a variety of cancer cells. This approach proved effective in reactivating genes that had been silenced due to hypermethylation of their promoter regions. Notably, most tumor suppressor genes exhibited similar expression patterns following treatment with DNMT inhibitors. This method found application across numerous cancer types, facilitating the reprogramming of various genes. It allowed researchers to investigate the roles of individual genes across different cancer types and the interplay of multiple genes within the same cancer. For instance, researchers employed 5-azacytidine to explore the tumor suppressor activity and promoter methylation status of KLF4 in cervical cancer, breast cancer, and mantle cell lymphoma. Furthermore, researchers identified methylation of certain members of the thyroid phosphatase family in multiple cancer types using this method. Detailed investigations into RASSF1 family members across various cancers were also conducted using these agents. 5-Aza-2′-deoxycytidine was more frequently utilized than 5-azacytidine in these studies. Epigenetic reprogramming with 5-azacytidine was primarily carried out in common cell lines associated with breast cancer, thyroid cancer, lung cancer, and colorectal cancer. In some cases, these treatments were administered in isolation, while in others, they were combined with compounds like trichostatin A, UNC1999, sodium valproate, 4-phenyl butyric acid, and CMP5 to enhance their effectiveness.68–71 Notably, 5-aza-2′-deoxycytidine could also assess the methylation levels of MLH1 in both breast cancer and colon cancer.72,73 The restoration of gene expression following 5-azacytidine treatment was evaluated either directly through RNA/protein levels or indirectly by examining changes in other cellular processes influenced by altered gene expression.
Another derivative of 5-azacytidine, known as 5,6-dihydro-5’-azacytidine, was initially employed for the purpose of demethylating the androgen receptor gene.74 Some genes, such as SPARC and PTPN6, were investigated in different cancer types using both 5-azacytidine and 5-aza-2′-deoxycytidine.32,75,76 This technique of epigenetic reprogramming, using either 5-azacytidine or 5-aza-2′-deoxycytidine, was not limited to protein-coding genes; it was also applied to numerous miR genes. Through this method, alterations in methylation patterns were identified, leading to differential expression of miRs. Many tumor suppressor miRs, including miR-34b/c, were investigated using this approach.39,77 The capacity of these DNMT inhibitors to reshape the epigenetic landscape of cancer cells positions them as suitable therapeutic agents and valuable tools for conducting research related to gene expression studies in the field of molecular biology.
Declaration
During the preparation of this work, the author(s) used ChatGPT to rephrase and edit the grammar of the review. After using this tool/service, the author(s) reviewed and edited the content as needed and take(s) full responsibility for the content of the published article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
