Abstract
In this paper, we study the problem of computing the similarity of two protein structures
by measuring their contact-map overlap. Contact-map overlap abstracts the problem of
computing the similarity of two polygonal chains as a graph-theoretic problem. In 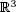 , we
present the first polynomial time algorithm with any guarantee on the approximation ratio for
the 3-dimensional problem. More precisely, we give an algorithm for the contact-map overlap
problem with an approximation ratio of σ where σ = min{σ(P
1), σ(P
2)} ≤ O(n
1/2)
is a decomposition parameter depending on the input polygonal chains P
1 and P
2. In
, we
present the first polynomial time algorithm with any guarantee on the approximation ratio for
the 3-dimensional problem. More precisely, we give an algorithm for the contact-map overlap
problem with an approximation ratio of σ where σ = min{σ(P
1), σ(P
2)} ≤ O(n
1/2)
is a decomposition parameter depending on the input polygonal chains P
1 and P
2. In 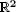 ,
we improve the running time of the previous best known approximation algorithm from
O(n
6) to O(n
3 log n) at the cost of decreasing the approximation ratio by half.We also give
hardness results for the problem in three dimensions, suggesting that approximating it better
than O(n
∊), for some ∊ > 0, is hard.
,
we improve the running time of the previous best known approximation algorithm from
O(n
6) to O(n
3 log n) at the cost of decreasing the approximation ratio by half.We also give
hardness results for the problem in three dimensions, suggesting that approximating it better
than O(n
∊), for some ∊ > 0, is hard.
Get full access to this article
View all access options for this article.
