Various international efforts are underway to catalog the genomic similarities and variations
in the human population. Some key discoveries such as inversions and transpositions within
the members of the species have also been made over the years. The task of constructing
a phylogeny tree of the members of the same species, given this knowledge and data, is an
important problem. In this context, a key observation is that the "distance" between two
members, or member and ancestor, within the species is small. In this paper, we pose the
tree reconstruction problem exploiting some of these peculiarities. The central idea of the
paper is based on the notion of minimal consensus PQ tree T of sequences. We use a modified
PQ structure (termed oPQ) and show that both the number and size of each T is
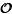 (1). We
further show that the tree reconstruction problem is statistically well-defined (Theorem 7)
and give a simple scheme to construct the phylogeny tree and the common ancestors. Our
preliminary experiments with simulated data look very promising.
(1). We
further show that the tree reconstruction problem is statistically well-defined (Theorem 7)
and give a simple scheme to construct the phylogeny tree and the common ancestors. Our
preliminary experiments with simulated data look very promising.