Abstract
An RNA secondary structure is saturated if no base pairs can be added without violating the
definition of secondary structure. Herewe describe a new algorithm, 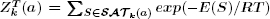 ,
where the sum is over all saturated secondary structures of a which have exactly k base pairs,
R is the universal gas constant and E(S) denotes the free energy with respect to the Turner
nearest neighbor energy model. By dynamic programming, we compute
,
where the sum is over all saturated secondary structures of a which have exactly k base pairs,
R is the universal gas constant and E(S) denotes the free energy with respect to the Turner
nearest neighbor energy model. By dynamic programming, we compute 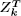 simultaneously for all values of k in time O(n
5) and space O(n
3).Additionally,
simultaneously for all values of k in time O(n
5) and space O(n
3).Additionally, 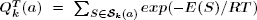 , where the sum is over all secondary
structures of a which have k base pairs; the latter computation is performed simultaneously
for all values of k in O(n
4) time and O(n
3) space. Lastly, using the partition function
, where the sum is over all secondary
structures of a which have k base pairs; the latter computation is performed simultaneously
for all values of k in O(n
4) time and O(n
3) space. Lastly, using the partition function 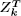 [resp.
[resp. 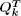 ] with stochastic backtracking,
] with stochastic backtracking, 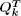 this provides a parametrized form of
this provides a parametrized form of
Get full access to this article
View all access options for this article.
