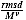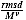A theory for assessing the statistical significance of structure alignment is developed using
a random or Gaussian chain model. In this model, we consider the statistical distribution
of the root mean square distance (rmsd) of the alignment between two random chains of
equal length and common center of mass (referred to as Case 1). We demonstrate that
the rmsd
2 is distributed as a sum of independent Gamma variables. Analytic results on
the mean and variance of the rmsd
2 are presented. Since rmsd is strongly dependent on
the length, we define the dimensionless quantity, reduced rmsd, as the rmsd divided by the
radius of gyration. We find that the reduced rmsd can be accurately approximated by an
extreme value distribution (EVD) that is independent of chain length and of bond length.
The parameters of the EVD can be calculated from the mean and the variance of the
rmsd
2. We also consider the case of two chains with a common center of mass that are then
rotated to minimize the rmsd (Case 2). In this case, the distribution of reduced rmsd can
again be accurately approximated by an EVD, which is independent of the chain length
and expected bond length. This distribution is used to calculate the p-value for a given
reduced rmsd. Performing an analogous comparison for proteins, we find that <rmsd> ∼M
ν
and ν = 0.28 and 0.32 for Case 1 and Case 2, respectively, where M is the chain length.
This result for Case 2 exactly matches previous scaling results and suggests that
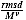 is an appropriate metric for protein structure alignment and will be independent of chain length.
We also find that the new score roughly follows the EVD.
is an appropriate metric for protein structure alignment and will be independent of chain length.
We also find that the new score roughly follows the EVD.