Abstract
Bacillus anthracis, the etiological agent of anthrax, a zoonosis relatively common throughout the world, can be used as an agent of bioterrorism. In naturally occurring outbreaks and in criminal release of this pathogen, a fast and accurate diagnosis is crucial to an effective response. Microbiological forensics and epidemiologic investigations increasingly rely on molecular markers, such as polymorphisms in DNA sequence, to obtain reliable information regarding the identification or source of a suspicious strain. Over the past decade, significant research efforts have been undertaken to develop genotyping methods with increased power to differentiate B. anthracis strains. A growing number of DNA signatures have been identified and used to survey B. anthracis diversity in nature, leading to rapid advances in our understanding of the global population of this pathogen. This article provides an overview of the different phylogenetic subgroups distributed across the world, with a particular focus on Europe. Updated information on the anthrax situation in Europe is reported. A brief description of some of the work in progress in the work package 5.1 of the AniBioThreat project is also presented, including (1) the development of a robust typing tool based on a suspension array technology and multiplexed single nucleotide polymorphisms scoring and (2) the typing of a collection of DNA from European isolates exchanged between the partners of the project. The know-how acquired will contribute to improving the EU's ability to react rapidly when the identity and real origin of a strain need to be established.
Cutaneous anthrax is the most common and mild manifestation of the disease. However, since the 2000s, a novel form of cutaneous anthrax (termed injectional anthrax) that is characterized by severe soft tissue infection has been described in heroin users. 3 The pathology of this is especially devastating. 4 Pulmonary and gastrointestinal anthrax are less common, but both forms can be deadly if left untreated. Fatality rates can reach up to 90% for inhalational anthrax. 5 Prompt diagnosis has a major impact on the effectiveness of treatment.
Although many microbiological pathogens could be used as biowarfare agents, Bacillus anthracis is considered the single greatest bioterrorism threat. 6 The attractiveness of anthrax as a bioweapon is due to its high rate of infectivity and lethality, the extreme stability of its spores, and the ease of use. Because of its global dispersal pattern, the agent can be obtained from a variety of sources; anthrax is distributed throughout the world and is still endemic in many countries, particularly in the developing world. Once obtained, it is relatively easy to cultivate and process into a spore-containing powder or suspension. And because it is one of the most feared microorganisms, the anthrax agent has also a high potential to produce public panic and social disruption, as seen in the 2001 anthrax letter attacks in the United States.
Molecular Genotyping Methods
Forensic methods for investigating a biocrime and for studying infectious diseases or pathogens have evolved in similar fashion: Both have shifted away from reliance on biological phenotypes of the suspected perpetrator, such as fingerprints, toward more reliable and robust molecular markers, such as variations in DNA sequence. DNA polymorphisms have now largely replaced morphological comparisons. 7
B. anthracis is an evolutionarily young pathogen that has recently emerged from the B. cereus group. Numerous studies have demonstrated the lack of molecular variation in this species. Isolates collected from across the world show an extremely high genetic homology.8,9 The absence of horizontal transfer of genetic material, a slow rate of accumulation of genetic variations, and the paucity of large genome rearrangements within the genome of the bacterium contribute to its highly clonal nature. 10 Because of this lack of diversity, only modern molecular characterization techniques with high discrimination power—such as multiple-locus variable-number tandem repeat analysis (MLVA), single nucleotide repeat (SNR), genome-wide single nucleotide polymorphism (SNP) analyses, or even whole-genome sequencing—have proved effective in differentiating strains within this highly clonal species. A typing strategy called “progressive hierarchical resolving assay using nucleic acids” (PHRANA), which relies on a combination of genetic markers that are progressively less stable but have increasing resolving power (SNP, MLVA, and SNR), has been proposed for B. anthracis. 10 This approach, which had been used to subtype a collection of 1,044 isolates from 42 countries, has contributed to rapid advances in our understanding of the global diversity of anthrax.11,12
During the past decade, MLVA became one of the major assays for molecular typing of B. anthracis. Variable-number tandem repeat (VNTR) loci are regions in the genome that vary greatly from strain to strain and may exhibit a number of different allelic states. The basis for this diversity is a high mutational rate driven primarily by slip strand mismatch repair. 10 In 2000, the first MLVA system targeting 8 loci (MLVA-8) was developed. At that time, the method was able to identify 89 B. anthracis genotypes (GTs) among a collection of 426 isolates from 30 countries. 9 Since then, its use has been pivotal to the identification of many different subpopulations throughout the world.9,13-24 In search of higher discriminatory power, additional VNTR loci have been added by different research groups to upgrade the system into new schemes (MLVA-15, MLVA-25, or MLVA-31).12,25-27 Further progress in providing fine-scale resolution was also obtained by analyzing SNR sequences. SNR is a special category of VNTR in which the repeat length is only a single nucleotide. The evolution of these sequences is very rapid. It represents the most mutable and variable markers that could exist. SNR analysis is used to distinguish even very closely related isolates from an outbreak.10,28-30 Although VNTR and SNR loci are useful for resolving B. anthracis population, they are not appropriate for discerning deeper phylogenetic relationships. Being more prone to evolutionary reversals or convergent evolution (homoplasy), these highly variable markers can provide misleading phylogenetic information and consequently obscure population structures.
SNPs represent ideal signatures to determine phylogenetic groups. SNPs are evolutionarily stable markers. Those nucleotide substitutions, with mutation rates of approximately 10−10 changes per nucleotide per generation, are probably generated by DNA replication errors that are not subsequently repaired. 31 Despite their paucity in B. anthracis, whole-genome sequencing of multiple strains allows for the efficient identification of thousands of SNPs scattered throughout the genome.11,32 By querying a large number of them against collections of diverse strains, representative SNPs that define major clades can be selected. The selection of strains used for genome-wide SNP discovery greatly affects the accuracy of the resulting SNP tree. The original choices of strains should maximize the amount of diversity that can be subsequently discovered to avoid biased phylogenetic reconstructions by incomplete taxon sampling.11,33
In 2007, a set of 13 diagnostic SNPs, termed “canonical SNPs” (canSNPs), representing key phylogenetic positions along the B. anthracis phylogenetic SNP tree were identified. 12 Analysis of these 13 markers enables the assigning of any B. anthracis strains into 3 major lineages (A, B, and C), with further subdivisions into 12 clonal groups: 7 distinct sublineages (C.Br.A1055, B.Br.KrugerB, B.Br.CNEVA, A.Br.Ames, A.Br.Australia94, A.Br.Vollum, A.Br.WNA) and 5 subgroups (B.Br.001/002, A.Br.001/002, A.Br.003/004, A.Br.005/006, A.Br.008/009). An additional 14th canSNP was recently reported that further divides the A.Br.008/009 group into 2 subgroups (A.Br.008/011 and A.Br.011/009). 34 Other SNPs that define the A.Br.Ames and A.Br.WNA lineages35,36 or that lie on various branches of the B. anthracis SNP tree 37 have also been published. SNP genotypes are now the gold standard in B. anthracis subtyping.
Next-generation sequencing technologies are powerful tools for the unbiased typing of any pathogen. By interrogating nearly every base of a genome, individualizing genetic markers can be reliably discovered, thus facilitating high-resolution strain tracking. Therefore, a whole-genome sequencing approach that focuses on identifying novel, strain-specific informative SNPs is clearly the future. As the cost and time required for sequencing and bioinformatic analyses is reduced, availability of whole genomes will increase. Currently, whole-genome sequences of several isolates can be obtained with high accuracy within a matter of days. However, although genomics enables characterization of microbes at an unprecedented level of resolution, whole-genome sequencing and comparison are still limited to a few “priority” strains, as exemplified by the Amerithrax investigation 38 and investigations undertaken following the 2009-10 anthrax outbreak among heroin users in Europe. 3
The application of modern molecular markers in forensic studies has led to some interesting findings, as illustrated by the following few examples. First, microbial forensic evidence gathered during the Amerithrax investigation assisted in identifying the Ames strain as the source of the material used in the 2001 postal anthrax attacks in the United States. The fact that this particular genotype is apparently rare in nature 39 but commonly used in a number of laboratories in the US made it unlikely that the source material was acquired directly from nature, pinpointing putative perpetrators of this act.38,40 Second, the B. anthracis bioterror strain dispersed by the Aum Shinrikyo religious cult in Japan was identified as the veterinary vaccine strain, Sterne 34F2, a low-virulent derivative, explaining why this malevolent release had been unnoticed in 1993 and had no victims. 41 And third, genetic evidence has also provided insight into the strains involved in the environmental release of weaponized spores from a Soviet biological warfare production facility in Sverdlovsk in 1979. The accident killed at least 68 people. 42
Current genotyping methods are also very useful for present or retrospective epidemiologic and epizootiologic studies, such as the monitoring of the spread and distribution of anthrax outbreaks in the Etosha National Park in Namibia 27 and the investigations to link clinical and environmental isolates associated with animal products used by anthrax patients in the US. 34
B. anthracis Molecular Diversity
CanSNP typing and whole-genome comparison of multiple strains from various origins have made the reconstruction of the B. anthracis phylogeny extremely accurate.11,12 As noted above, the substructure of the population is divided into 3 lineages (A, B, and C), with further subdivisions into 12 major sublineages or groups (Figure 1A).
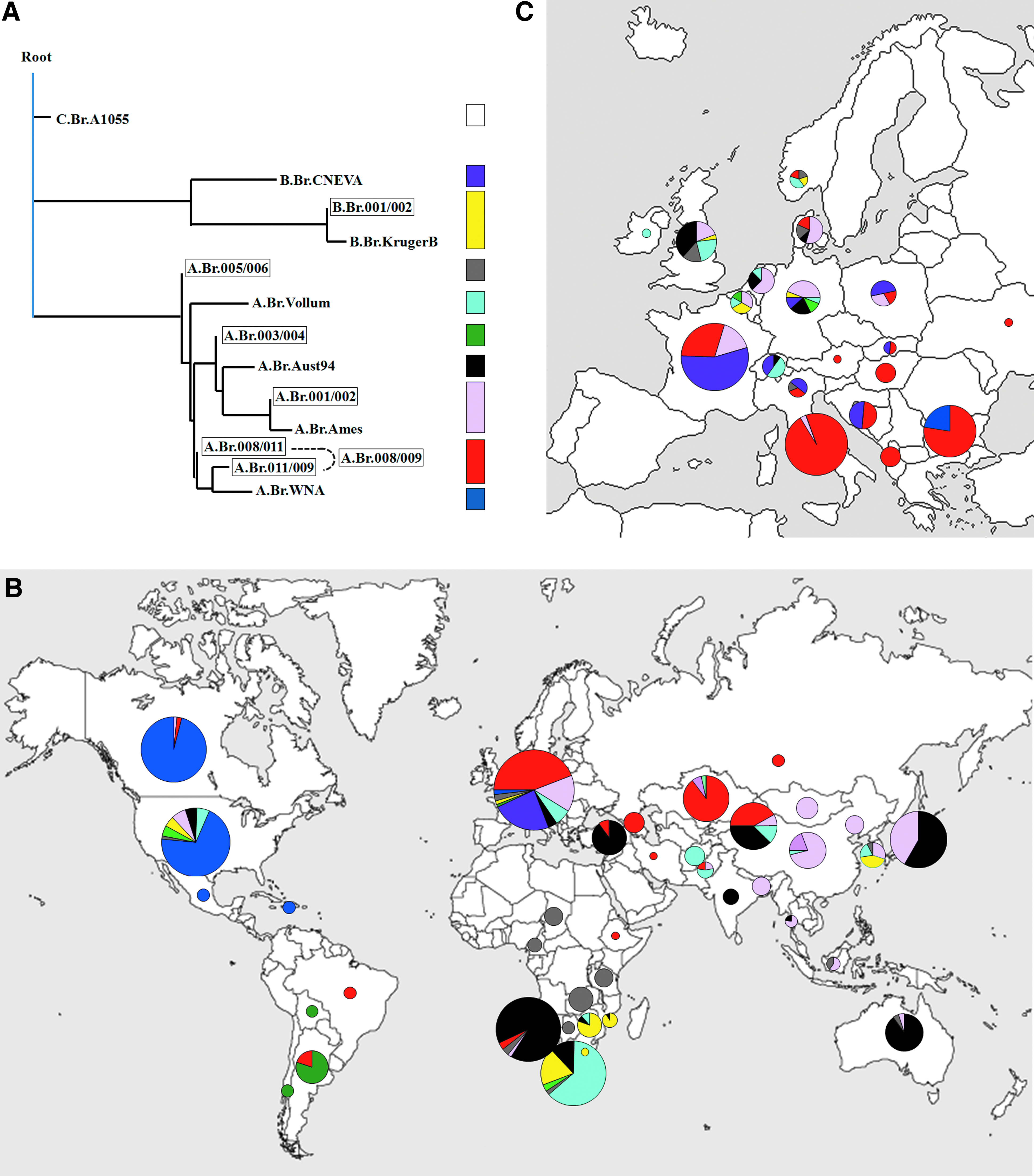
Strains belonging to the A lineage account for the majority of anthrax cases reported around the world.8,9,43 The 8 subgroups that comprise this clade (ie, A.Br.001/002, A.Br.Ames, A.Br.003/004, A.Br.005/006, A.Br.Aust94, A.Br.Vollum, A.Br.008/009, A.Br.WNA) are globally distributed but exhibit distinct geographical patterns across the 5 continents (Figure 1B, Table 1). Co-occurrence of several subgroups in a single region is generally observed, although only 1 dominant sublineage often exists. The A.Br.008/009 subgroup, also known as the Trans-Eurasian group (TEA), is one of the most common groups of B. anthracis in the world. TEA's isolates are widespread throughout Europe, Russia, Kazakhstan, the Caucasus, and the western province of the People's Republic of China.12,17,18,36,44
B. anthracis Genotyping Studies
Note: WGS: whole genome sequencing; KNG: Kruger National Park; Sv: Sverdlovsk outbreak; Bosnia-H.: Bosnia-Herzegovina; GT: genotype; pagA: sequencing of the pagA gene.
The A.Br.001/002 subgroup is a major part of the B. anthracis population in China. The rare A.Br.Ames sublineage, which has been introduced into Texas (US), has emerged from this group in Inner Mongolia (China). 44 North and South America are dominated by the A.Br.WNA and A.Br.003/004 subgroups, respectively, while A.Br.005/006 is primarily found in Africa. The southern part of Africa and parts of Asia (including Afghanistan and Pakistan) are dominated by the A.Br.Vollum sublineage, whereas collections from Turkey, India, Namibia, and Australia are dominated by A.Br.Aust94.12,20,27,36,44
The B lineage is divided into 2 genetically distinct subclades, geographically restricted to specific regions. The B1 subclade (including the B.Br.001/002 and B.Br.Kruger groups) is ecologically established in southern Africa, in particular in the Kruger National Park, where it coexists with strains from the main A.Br.Vollum sublineage.12,16 The B2 sublineage (B.Br.CNEVA) is exclusively found in continental Europe,12,19,45-49 except for a single report from California. 12 Finally, the C lineage (C.Br.A1055) is an uncommon cluster consisting of only 3 isolates of unknown origin.12,13
In Europe, the population structure of B. anthracis is largely diversified, with the co-occurrence of up to 6 phylogenetic subgroups in a single country (Figure 1C, Table 1). Three sublineages appear to be more largely distributed: the A. Br.008/009, B.Br.CNEVA, and A.Br.001/002 canSNP groups. They may reflect ancient introductions of B. anthracis into Europe followed by dispersal and regional differentiation. The B.Br.CNEVA sublineage is ecologically established in the middle of Europe, spreading from the Pyrenees (southwest France) to northeast Poland, along the Italian Alps, Switzerland, Germany, Croatia, and Slovakia.12,45-49 In these countries, B.Br.CNEVA usually co-dominates with A.Br.008/009, although both phylogenetic groups exhibit distinct geographic localization in many regions.45,46,49 The TEA group has been found in most European countries. It is well established in southern and eastern Europe and represents the dominant subgroup in Italy, Bulgaria, Hungary, and Albania.12,22,50 The last major subgroup, A.Br.001/002, is distributed at higher latitudes in western Europe. It accounts for a significant part of the B. anthracis distribution in Denmark, Belgium, the Netherlands, and the United Kingdom (this study). Strains belonging to this subgroup have also been found in northeastern France, where they caused an outbreak associated with the death of 39 animals in 2008. 51
Minor subgroups found in Europe include A.Br.005/006, A.Br.Aust94, A.Br.Vollum, B.Br001/002, A.Br.Ames, and A.Br.003/004. 12 This assortment of genetic types is likely a consequence of human activities and international trade exchanges. In the United Kingdom, for instance, the presence of genotypes that are dominant in portions of Asia is consistent with reports tracing anthrax infection to imported animal products (eg, hair and wool, hides and skins, bone meal, shaving brushes) from these regions during the 19th and early 20th centuries. 12 The worldwide occurrence of A.Br.Vollum, in particular, including in European countries such as Belgium and Switzerland, is thought to be due to the extensive trade of cashmere wool from the Indian subcontinent.9,14,47 Colonial-era importations of contaminated animal products from Africa and southeast Asia could also have affected the population structure in Europe, with the introduction of strains related to A.Br.005/006, B.Br.001/002, or A.Br.Aust94 subgroups in specific countries.
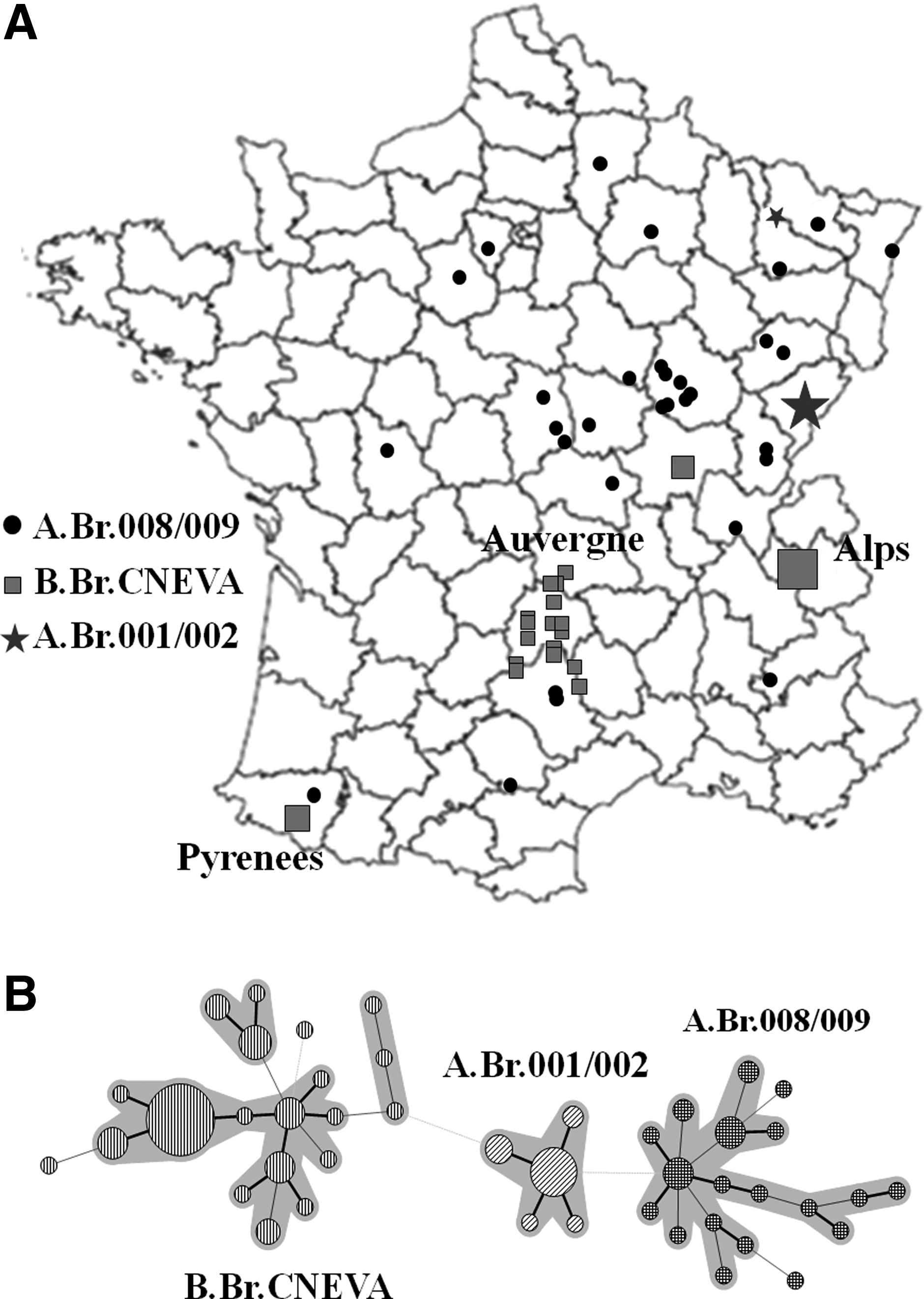
In France, for example, we have detected the co-occurrence of 3 canSNP subgroups (Figure 2A). The dominant B.Br.CNEVA sublineage was found to be prevalent in the Alps, the Pyrenees, and the Auvergne region. Strains affiliated with the TEA subgroup were observed throughout most of the country but only associated with single, sporadic cases. The minor A.Br.001/002 subgroup was restricted to northeastern France. 45 Using MLVA-30, the 3 canSNP subgroups found in France were further resolved into 55 different genotypes, with particular geographical clustering (unpublished data, Figure 2B).
History of Anthrax in Europe
Anthrax has been a widespread disease in livestock and humans for millennia and has been known in Europe since antiquity. There are descriptions of anthrax involving animals and humans in the early literature of Greeks and Romans. In the Georgics book III, the Latin poet Virgil provided in 25
The first anthrax pandemic in Europe, known as “Black Bane,” was recorded in 1613. It caused at least 60,000 human deaths and many more in domestic and wild animals.1,6 Epidemics of anthrax were common in Europe during the 18th and 19th centuries. In the 1850s, outbreaks of occupational anthrax were frequently reported in the industrial European countries. Human cases occurred not only in people who had contact with ill or dead animals, but also among workers in industries that process animal products including those in woolen industries, abattoirs, and tanneries and those who process hides, hair, bone, and bone products.53,54 The victims of “woolsorters' disease” (also termed “industrial anthrax”) also included people and cattle that had been infected through the environmental release of contaminated waste. 55
The identification of the anthrax bacillus by Robert Koch (in 1876), followed soon after by the development of live vaccines for animals (by William Greenfield and Louis Pasteur in 1880-81), led to a steady decrease in animal and, indirectly, human cases. Improved industrial and animal husbandry practices—among which were the development of wool disinfection measures, the immunization of livestock, and proper carcass disposal with the implementation of knackeries to avoid the burying of carcasses with further replenishments of spores into soils—also succeeded in reducing the incidence of the disease in the developed countries.53,54,56 Active control measures and large animal vaccination programs aimed at breaking the cycle of infection were undertaken during the first part of the 20th century. 6 Over time, soil contamination was sharply reduced in many European countries. In 1950, only a few sporadic cases in livestock were still reported in France, for instance. 56
Current Situation in Europe
Currently, although an increasing number of European countries are probably free of the disease (including Cyprus, Austria, the Czech Republic, Denmark, Finland, Latvia, Luxembourg, and Malta), anthrax is not yet a disease of the past. 57 B. anthracis spores are still widely distributed in the soils of Europe, though in very low concentrations. Anthrax spores are extremely resistant and can persist for very long periods of time in the environment (over 100 years). 1 Anthrax is relatively common in countries adjoining the Mediterranean Sea and in eastern Europe (eg, Albania, Romania, Greece, and Spain), but Turkey continues to be the main afflicted region. 57 In the rest of Europe, anthrax is generally considered to be a sporadic disease that almost exclusively affects animals at pasture, 54 with a majority of cases reported during the summer months. New animal cases are generally linked to climate changes (ie, with incidents of warmer and extreme weather) or human activities that interfere with soil surface (eg, works of drainage or excavation, digging, or cleaning out of dams and ditches) and are found in areas where outbreaks have occurred in the past (eg, old knackery sites, former tanneries, or buried sites). In France, for instance, sporadic cases in cattle with one to several dead animals may be recorded each year, but larger outbreaks can still occur. 58 In southern Italy, where anthrax is hypo-enzootic, 2 severe epidemic-like anthrax outbreaks, probably spread by the actions of haemophagic flies, have been recently recorded.59,60 In northern Europe, foci of animal disease are either absent or found only in very sporadic cases, as illustrated by the situation in Sweden where a few cows died from anthrax in 2008 and 2011 after an apparent absence of the disease for several decades. 61
Human anthrax is currently a very rarely reported disease in the European Union (Table 2). It involves people such as farmers, workers in textile plants, or drummers who handle infected animals or animal products. Most recent cases have been reported from Spain, Bulgaria, Greece, Romania, and the UK. Only 1 significant outbreak has been reported during the past decade. 62
Recent Cases of Human Anthrax Reported in Europe
Confirmed case.
From December 2009 to November 2010, the United Kingdom and Germany reported cases of injectional anthrax among drug users. By the end of the outbreak, there were at least 55 confirmed cases of heroin-related anthrax in the United Kingdom, mainly in Scotland and the London area, and in Germany, including 21 deaths.63,64 Strains that caused infections in heroin users have recently been shown to cluster in the widespread TEA subgroup (A.Br.008/011). All share a single, indistinguishable, novel genotype that is closely related to a few strains from Turkey isolated from infected goats. 3 Epidemiologic and microbiologic evidence supports the conclusion that the spores responsible for the outbreak likely came from a single batch of heroin trafficked into Europe from Afghanistan. The heroin contamination could have arisen somewhere along the drug trafficking route, probably in Turkey, through contact with animal hides used to smuggle heroin into Europe. Although this outbreak was officially declared over at the end of December 2010,63,64 new cases among European heroin users that might be linked to the 2009-10 outbreak emerged in 2012 in Germany, France, the United Kingdom, and Denmark.65,66
Threat Recognition
Faced with a suspect case of anthrax, it is of utmost importance to identify and quickly confirm the causative infectious agent. Anthrax is a notifiable disease by law in most European countries. Disease regulatory actions and control procedures have to be enforced by government veterinary services: quarantine, immunization and treatment, proper disposal of carcasses, disinfection of the affected area, a ban on animal movements, and restrictions on human activities in the area.
In the event of a biological animal terrorism threat, early detection and warning are crucial to countering an attack, underscoring the need to increase animal health services' vigilance and preparedness. In an outbreak, keeping in mind a few questions could contribute to raising our recognition of a biological animal terrorism incident when it occurs. Could there be something behind this outbreak, other than a naturally occurring infection? Is there any clue that might suggest we are faced with a malevolent act (an inadvertent or deliberate release of spores, a bioterrorism attack)? To properly address this issue, it must be established whether the index strains are indeed representative of ecologically established strains for the region or have been imported from other areas. Molecular typing techniques are now available for anthrax national reference laboratories to genotype and discriminate B. anthracis strains. However, determining the genotype of a strain with no further knowledge on the diversity found within a putative geographical area of origin is not necessarily helpful in tracing an outbreak strain back to its possible origin. Without in-depth understanding of the anthrax “background,” we can draw false conclusions and make wrongful accusations. Extensive typing data about autochthonous but also worldwide isolates are required for comparison so that accurate hypotheses can be made about isolate origins.
Work in the AniBioThreat Project
The focus of the AniBioThreat project is to improve the EU's capacity to counter animal biothreats in terms of awareness, prevention, and contingency. In work package 5.1 of the AniBioThreat project, we aim to (1) improve and standardize real-time PCR detection assays for B. anthracis; (2) increase the use of polymorphic sequence markers (notably VNTR and canSNP) to characterize European collections of strains and create an extensive polymorphism database; and (3) develop a highly precise and robust subtyping tool based on suspension array technology and SNP scoring for strain traceability. The work undertaken will contribute to a better understanding of the B. anthracis population present in Europe and enhance the EU's ability to distinguish between intentional and naturally acquired infections caused by a bioterror microorganism that is widespread in the environment.
Routine analysis of canSNPs for molecular typing requires an assay that is robust and provides high capacity. A wide variety of techniques and platforms exist for screening SNPs,
67
including allele-specific hybridization, oligonucleotide ligation assays, mini-sequencing, and real-time fluorescent PCR methods, such as TaqMan-minor groove binding allelic discrimination assays (Taq-MAMA) or High-Resolution Melting assays (HRM).68,69 During the project, we have developed a suspension array method for the rapid and high-throughput canSNPs analysis. The luminex platform and its
The dramatic improvement of Next-Generation-Sequencing technologies allows for their increasing use in systematic characterization of B. anthracis. Large-scale comparative genome sequencing offers a powerful approach for identifying novel phylogenetically informative SNPs. The sequencing in progress of a few selected strains from around Europe will add resolving power to current genotyping tools. New DNA signatures that discriminate further within sublineages or specifically define a particular strain will be searched for and developed into new HRM and luminex assays. PCR-HRM assays are cost-effective and can be rapidly designed around SNPs to determine the extent to which each marker varies in the B. anthracis population. Available B. anthracis collections will be screened and validated markers added to the developed
Footnotes
Acknowledgments
This research was supported by and executed in the framework of the EU project AniBioThreat (Grant Agreement: Home/2009/ISEC/AG/191) with financial support from the Prevention of and Fight against Crime Programme of the European Union, European Commission—Directorate General Home Affairs. This publication reflects the views only of the authors, and the European Commission cannot be held responsible for any use that may be made of the information contained herein.
