Abstract
Avian influenza virus (AIV) and Newcastle disease virus (NDV) infect various avian species including domestic poultry. Clinical manifestations vary from subclinical or mild to severe multiorgan systemic disease with a near 100% mortality rate. Severe disease is caused by highly virulent specific virus strains, termed highly pathogenic AIV and velogenic NDV. Recent controversial influenza H5 adaptation studies in ferrets have highlighted the importance of preparedness against AIV as a bioterrorism agent. Furthermore, NDV also has zoonotic potential, although symptoms in humans are mild and self-limiting for naturally occurring viruses. Thus, both of these viruses pose a direct biothreat to domestic poultry but also indirectly to humans via zoonotic transmission. For diagnosis and rapid containment of disease, it is crucial to differentiate highly pathogenic AIVs and NDVs from frequently occurring low pathogenic variants. Recently, we developed a novel strategy for pathotyping of AIV and NDV that we review here. The method should be ideal for rapid testing and surveillance in food safety, for wild bird monitoring, and for combating acts of bioterrorism.
Flu pandemics usually occur by recombination of avian flu virus genes (or flu from other species such as swine) with those of a native human influenza virus. The global cost for a human flu pandemic can be enormous, depending on the severity of the virus. In a recent simulation study applied to the United Kingdom, it was concluded that, depending on the fatality rate of the disease (0.06%-0.35%), the GDP would decrease by 0.3% to 0.6%. Taking into account the cost for implementation of disease mitigation policies, the decrease could become almost 4% (£47.5 billion in 2011). NDV lacks the strong connection to human health of AIV, but the disease is very common in all regions of the world and causes severe problems mainly in developing countries. 6 Of the 15 List A diseases reported to OIE during 1997, only the number of foot-and-mouth disease (FMD) outbreaks exceeded the number of ND outbreaks. 1 In 1992 it was estimated that, in Nepal, 90% of the chickens each year died from Newcastle disease. 6 Thus, it is clear these 2 viruses have rightfully been considered prime candidates for agroterrorism actions. 1
Highly Pathogenic Avian Influenza Virus
Avian influenza viruses are pathogens that constantly evolve and cause seasonal epidemics and epizootics with large socioeconomic impacts and severe consequences for human and animal health. 7 The influenza subtypes, defined based on the antigenicity of the hemagglutinin (HA) and neuraminidase (NA) glycoproteins, currently consist of 17 (H1-H17) and 10 (N1-N10) variants, respectively, which occur in many combinations.8,9 They infect various host species including humans, but wild aquatic birds constitute the natural reservoir. 10 Infections by all AIV subtypes are usually of low pathogenicity (LP), but introduction of H5 or H7 LP virus subtypes to poultry may result in mutation to the highly pathogenic (HP) variants. Poultry infections by H5 and H7 subtypes, irrespective of pathotype, are classified as notifiable avian influenza (NAI), and the OIE Terrestrial Code recommends that poultry commodities should be traded only between countries, zones, or compartments that are free of notifiable AIV. 11 Thus, HPAI poultry infections can have a devastating effect on meat and egg production and trade in affected areas. 12
High pathogenicity is related to the cleavage of the AIV HA glycoprotein during the infection cycle. An RXXR↓G motif situated at the cleavage site (CS; denoted by↓) in combination with elongation compared to the LP-variants via insertions leads almost invariably to high pathogenicity.13-15 Very high diversity of the CS is observed in H5 and H7 AIVs, and current standard procedures for molecular pathotyping of NAI specimens involve nucleotide sequencing across the CS.16,17
The circulation of H5N1 viruses among wild and domestic birds provides opportunities for continuing human infections and potential adaptation of the virus for more efficient poultry-human and human-human transmission. 18 This scenario has become even more likely in view of recent results from controversial influenza H5 adaptation studies in ferrets.19,20 In these studies it was shown that by relatively simple methods, such as serial passaging, highly pathogenic H5N1 viruses could be adapted to the airways receptors in ferrets and become lethal to this host. Since it is well known that humans and ferrets can be infected by the same influenza viruses, the results implied that these viruses potentially also could infect humans. Furthermore, this virus also acquired capacity for airborne transmission. Before publication, it was intensely debated if the results should be made public since they essentially provide a blueprint for the manufacture of a powerful biological weapon. However, it was decided the papers should be published.19,20 On July 6, 2012, 607 human cases of influenza A(H5N1) had been recorded; 358 of these (59%) were fatal. 21 It is easy to see that if a human adapted HPAIV capable of airborne transmission were released, it could have devastating consequences.
It is clear that surveillance of HPAI is a very complex task, with low and highly pathogenic viruses co-circulating, with asymptomatic birds or vaccinated birds able to shed virus, and with wild birds able to transmit the virus over large distances. 7 It is also important to be prepared if AIV or NDV is deliberately released. Thus, there is a need for fast and relatively inexpensive diagnostic methods for implementation under field conditions and for preparedness against bioterrorism.
Velogenic Newcastle Disease Virus
Newcastle disease virus, or as it is also denoted avian paramyxovirus type 1 (APMV-1), belongs to the family Paramyxoviridae. ND is a very serious problem for the poultry industry worldwide. 22 Signs of NDV infection can range from very mild respiratory problems and mild enteric disease caused by lentogenic strains to mesogenic viruses and, most seriously, the extremely high mortality rates in flocks caused by velogenic strains. 23 NDV strains exhibit wide-ranging heterogeneity, and the respiratory signs presented by birds infected with NDV may be similar to other severe avian respiratory infections, including those caused by AIV. Reliable laboratory diagnostics are therefore essential for the rapid and accurate differential diagnosis of ND. The diversity within NDV variants makes the prompt identification and the accurate pathotype discrimination of NDV variants very difficult. At present pathogenicity can be determined by either experimental study in vivo or sequencing of the F-gene cleavage site analogously to the AIV hemagglutinin pro-protein cleavage site sequencing. But in this case the cleavage site variants signifying high pathogenicity is RQ/RK/RR↓F. However, these methods are not suitable for rapid diagnosis and field settings, which would be desirable to efficiently counter a bioterror act. It is crucial that new methods be developed for rapid and accurate pathotyping of NDV strains.
Humans are among many species that can be infected by NDV (except the more than 241 avian species). (See reference 2 for a review.) In humans the most common sign of infection is conjunctivitis. General symptoms include headache, discomfort, and slight chills that occur within 48 hours of the initial infection. The disease in humans is typically self-limited, and the risk of severe illness is low. 2
A New Method for Pathotyping
Recently, we presented a novel strategy for highly multiplexed interrogation of a sample, and we employed it both for pathotyping of H5 and H7 AIVs 14 and for the discrimination of lentogenic from velo- and mesogenic strains of NDV. 24 The method provides a fast and simple alternative to DNA sequencing, which can be implemented for large-scale testing in both disease diagnosis and surveillance settings. In a typical advanced laboratory setting, sequencing is a 24-hour procedure, while the PCR method presented here can be completed within 2 hours. Thus, the presently described method should be ideal for rapid testing and surveillance to efficiently contain outbreaks caused by naturally occurring infections or by deliberate acts of bioterrorism. Furthermore, since the PCR-based method is easily adapted to portable PCR instruments, complete diagnosis can be obtained a few hours after a case is discovered in the field. Sequencing is not currently suitable for field settings, and transportation to a well-equipped laboratory is necessary, which further underscores the advantage of the method described here. Although the sequence interrogation is carried out by means of an extensive primer mixture, the primer cost is actually smaller than with conventional PCR since the concentration used is 1000-fold less. In the present assay construct, plexor chemistry was used, 25 but in principle other fluorescent primer variant such as LUX primers 26 could also be used. This method is unique in the sense that, to our knowledge, no other PCR-based method has been published that is able to in a single assay distinguish the pathotype of all H5 and H7 viruses from both the eastern and western hemispheres.
Results
Assay Design
The method is based on a 3-level semi-nested PCR system, which achieves highly multiplexed interrogation of the CS sequences corresponding to 2 pathotypes (Figure 1). Initially, a larger genomic region surrounding the CS region is preamplified by conventional PCR. Then the pathotypes of the CSs are recognized by a subsequent recognition step using “selection primers” directly binding to the CSs. The large increase of the amount of template by the PCR preamplification allows selection primer concentrations in the subnanomolar range. This permits a high degree of multiplexing of these primers.
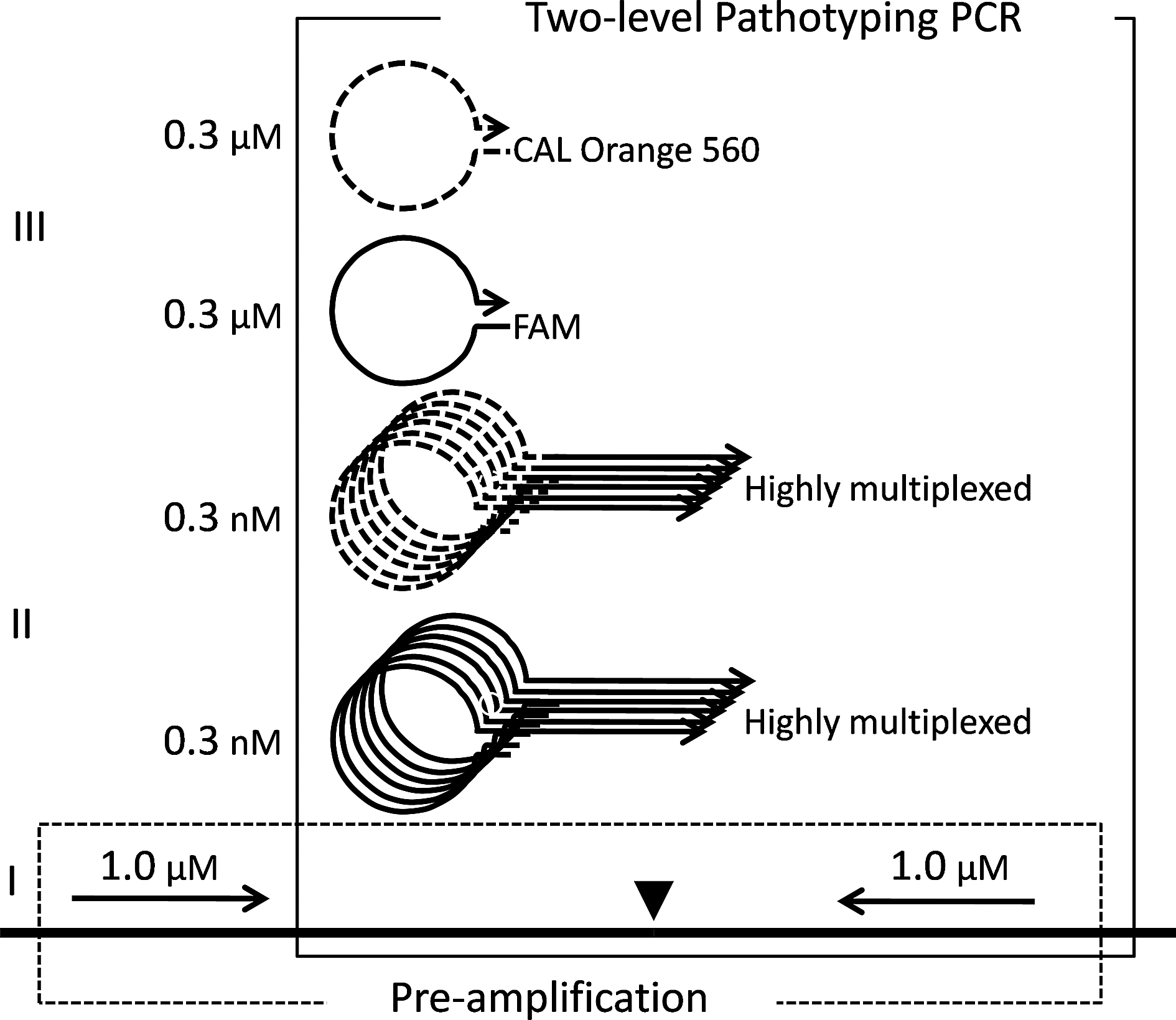
Outline of the scheme for the semi-nested 3-level real-time PCR method designed for AIV and NDV pathotyping. A triangle marks the CS. Typical primer concentrations are shown. The preamplification PCR, denoted by “I,” and the 2-level pathotyping PCR are indicated and boxed with dashed and full lines, respectively. The bipartite selection primers are denoted by “II,” with the tag sequence signifying high and low pathogenic viruses shown with full and dashed lines, respectively. The detection primers are denoted by “III” and have sequences identical to the corresponding tag sequence. These primers initiate the amplification of the PCR products originating from selection primers to detectable levels. The dyes tethered to the detection primers that label products originating from highly (FAM) and low (CAL Orange 560) pathogenic viruses are indicated.
In total, 484 and 320 selection primers were included in the assays for AIV and NDV, respectively. These numbers give a rough estimate of the variability and high mutation rate of the cleavage sites of the 2 viruses. The procedure to design these primers is very simple and generic and basically only includes adjustment for getting a similar melting temperature over the whole set.14,24 The selection primers are bipartite with a 5′-tag region that is unrelated to the target but of 2 different kinds, depending on the HP/LP status of the targeted CS sequences. Each of the 2 tagging sequences corresponds to a fluorescently labeled plexor “detection primer” 25 with the same sequence, but the 2 labels are different and thus indicate the pathogenicity of the CS; HP and LP cleavage sites are labeled with FAM and CAL orange-560 (CO-560), respectively.
In contrast to most other real-time PCR techniques, the fluorescence will decrease in the course of the amplification reaction. 25 The preamplification system and the 2-level (patho)typing system use the same reverse primers, and the whole assay is consequently a 3-level semi-nested PCR system. The detection primers and thus the 5′-regions of the selection primers have “panhandle” self-structures to minimize cross-interactions (Figure 1). The preferential binding of selection primers corresponding to HP or LP cleavage sites are then measured by reading one fluorescence channel (CO-560) normalized against the other (FAM) in real-time. The normalization procedure will produce a decreasing fluorescence for low pathogenic isolates (decreasing CO-560 signal; no or late decrease of FAM signal) and an apparent fluorescence increase for highly pathogenic isolates (decreasing FAM signal; no or late decrease of CO-560 signal). Thus, increasing normalized signals are produced by highly pathogenic viruses and decreasing by low pathogenic viruses.
Pathotyping of AIV
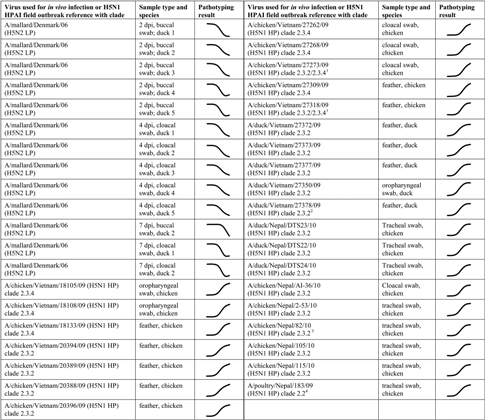
The assay construct was run on 39 H5 clinical specimens from LP- and HP-infected birds as shown in Table 1. The samples originated both from field specimens and experimentally infected birds and included 26 specimens from H5N1 HPAI “Asian lineage” outbreaks. Swabs, tissue samples, and feathers from infected birds were included. Matrix gene TaqMan PCR experiments 27 indicated a wide range of viral loads. Despite these variations in the sample origin and viral titer, all 39 samples were correctly pathotyped in accord with CS sequencing results. 14
Pathotyping results for 39 H5 clinical specimens from experimentally infected ducks and field-infected birds. The actual experimental result curves are inserted. Copyright © American Society for Microbiology, Journal of Clinical Microbiology 2011; 49: 3860-3873. doi:10.1128/JCM.01179-11
Specimen not sequenced but obtained from a large industrial holding with mixed clade 2.3.2 and clade 2.3.4 infection.
Specimen not sequenced but obtained from a duck backyard experiencing clade 2.3.2 infection.
Specimen not sequenced but other specimens collected and sequenced from the same location were clade 2.3.2.
Specimen not sequenced but other specimens collected and sequenced from the same location were clade 2.2.
Pathotyping of NDV
Nine genotypes have been defined for NDV, and pathotyping results for genotype I and II are shown in Figure 2. The lentogenic genotype I is represented by 5 isolates. Four of these were from 3 different continents: Asia (HK99/75; HK138/80), Europe (SE/94), and Australia (Queensland V4/66), while Ulster/67 is a vaccine strain from the US. Genotype II is represented by 3 lentogenic strains, the vaccine strains LaSota and B1, and a Canadian strain from 1963 (CA12/63). The 3 genotype II velogenic isolates were isolated from Europe and US. As described above lentogenic and meso/velogenic strains should yield negative and positive normalized signals, respectively. It is seen from Figure 2 that the 5 lentogenic genotype I strains (dashed lines) as well as the 3 lentogenic genotype II strains (dotted lines) indeed produce negative signals, while the 3 highly pathogenic genotype II strains (full lines), including the mesogenic Baudette C strain, give positive signals. Strains from the other 7 genotypes gave similar pathotyping results with the novel method in agreement with sequencing results. 24
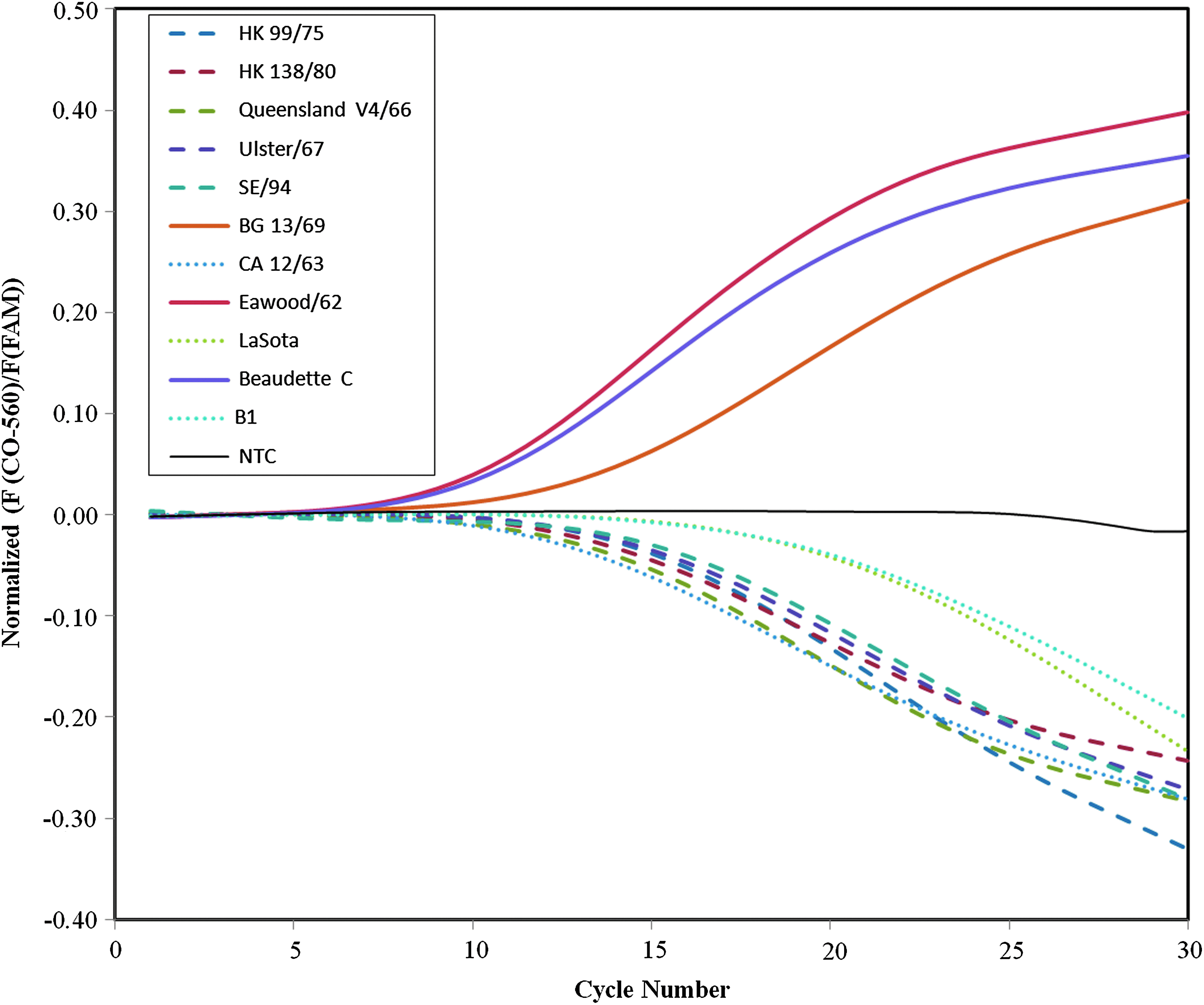
Amplification curves for genotype I lentogenic strains (dashed lines), genotype II meso/velogenic strains (full lines), and lentogenic strains (dotted lines) displayed as CAL Orange-560 fluorescence normalized against the FAM fluorescence with corrected baseline. This data processing means that CSs of lentogenic and meso/velogenic strains will yield negative and positive signals, respectively. Adapted from Yacoub et al., reference 24. Color images available online at www.liebertpub.com/bsp
Conclusions
The presently described molecular pathotyping method provides new opportunities for surveillance and diagnostics of AI and ND. These highly infectious transboundary animal diseases are among the most feared maladies threatening domestic poultry. Considering their highly infectious nature and their ability to cause serious losses as well as their zoonotic capacity, AI and ND have been implicated as potential weapons in agroterrorism.1,2 Validation of the assay with a population of clinical specimens showed that it accurately obtains the same molecular pathotyping result that is determined by sequencing. Incorrect pathotyping was never obtained either on virus isolates or on clinical specimens.14,24
A distinctive strength of the technique is that it is expandable in the sense that any novel previously unknown cleavage sites can be added to the selection primer mixture. This is in contrast to conventional probe-based real-time PCR methods, where mutation in the probe may make the assay unusable. The method is suitable for adaptation to portable PCR instruments 28 and can complement sensitive type-specific detection assays with pathotyping, and thus provides complete diagnosis of AIV and NDV on a portable real-time PCR platform in the vicinity of outbreaks. It could also be useful for direct rapid onsite monitoring of production and market chains for poultry commodities.
Footnotes
Acknowledgments
This work was supported by the Award of Excellence of the Swedish University of Agricultural Sciences (2007) granted to SB, a grant from the Swedish Civil Contingencies Agency (2:4 Krisberedskap), and a grant in the framework of the EU project AniBioThreat (Grant Agreement: Home/2009/ISEC/AG/191) with financial support from the Prevention of and Fight against Crime Programme of the European Union, European Commission—Directorate General Home Affairs. This publication reflects the views only of the authors, and the European Commission cannot be held responsible for any use that may be made of the information contained therein.