Abstract
Cellular plasticity obtained by natural adaptation or by forced expression of developmental stage and lineage-specific transcription factors converts differentiated cells into three different levels of potency: pluripotent, multipotent (dedifferentiation), and oligo- or unipotent (transdifferentiation). These processes hold promise for regenerative medicine to replace damaged cells and organs through the identification of transcription factors that determine the fate of cellular plasticity. In this study, we report that ectopic expression of Oct4 and Klf4 induces transdifferentiation of human preadipocytes into three different lineages: hematopoietic, neuronal, and pancreatic endocrine cells. Furthermore, ectopic expression of Oct4 is sufficient to induce transdifferentiation of preadipocytes into glucagon-expressing pancreatic α cells. This will provide a simple and efficient method to produce functionally competent cells for therapeutic regenerative medicine.
Introduction
Overexpression of defined transcription factors can convert one cell type into another and has important implications for regenerative medicine. Ectopic expression of key transcription factors in somatic donor cells has been used to generate many different cell types, including cells resembling blood cells,1,2 brown fat cells, 3 hepatocytes, 4 sertoli cells, 5 and various cell types of the neural lineage.6–9
Expression of Oct4, partially in conjunction with Klf4, is critical to maintain pluripotency at early developmental stages, including in the blastomere, inner cell mass (ICM), and epiblast. Oct4 and Klf4 have been reported as components of four defined transcription factors (Oct4, Sox2, Klf4, and cMyc) that trigger reprogramming of somatic cells into pluripotent stem cells (induced pluripotent stem [iPS] cell) that are functionally identical to embryonic stem cells. 10 Ectopic expression of Oct4 alone, or ectopic expression of Oct4 and Klf4, in combination with small molecule treatment, is sufficient to generate iPS cells from neural stem cells, mouse embryonic fibroblasts (MEFs), and dermal papilla cells.11–14 Oct4 activates early epithelial-to-mesenchymal transition (EMT), which is critical to somatic cell reprogramming. 15 Interestingly, Szabo et al. demonstrate direct conversion of human dermal fibroblasts into multipotent blood progenitors by ectopic expression of Oct4. 16 This suggests that Oct4 functions as a hematopoietic lineage differentiation marker as well. In this study, we report that Oct4 and Klf4 induce transdifferentiation of human preadipocytes into three different lineages, including hematopoietic, neuronal, and pancreatic endocrine cells. Furthermore, ectopic expression of Oct4 alone is sufficient to induce transdifferentiation of preadipocytes into glucagon-expressing pancreatic α cells.
Results and Discussion
Ectopic expression of Oct4 and Klf4 induces transdifferentiation of human preadipocytes
Primary cultures of preadipocytes from stromal vascular fraction of adipose tissues are known as a rich source of CD34 and alkaline phosphatase-positive adult mesenchymal stem cells. 17 Preadipocytes are capable of differentiating toward mesodermal lineages, including osteocyte, adipocyte, and chondrocyte (Supplementary Fig. S1a). We generated two-factor (OK) and four-factor (OKSM) transfected cells using highly proliferative, low passage preadipocytes (<3 passages) that had been demonstrated to have ∼50% transfection yields using a lentiviral vector (Supplementary Fig. S1b). Our general strategy is outlined in Figure 1a. One day before lentiviral infection, cells were seeded on a six-well plate at the concentration of 105 cells per well with Dulbecco's Modified Eagle's Medium (DMEM) containing 10% fetal bovine serum (FBS). For the lentiviral infection, total 25 multiplicity of infection (MOI) of lentivirus that expresses Oct4, Klf4 under control of cytomegalovirus promoter (SBI and Cellomics), or polycistronic lentivirus STEMCCA that expresses four iPS factors (Oct4, Sox2, Klf4, and cMyc) were added to the medium with 5 μg/mL polybrene (Sigma-Aldrich) (Fig. 1a). The lentiviral infection was repeated the next day, and the culture medium was changed with fresh DMEM/5% FBS medium every other day. When cells became confluent, they were trypsinized, counted, and seeded on MEF feeder layer at a density of 5 × 104 cells/mL with the mTeSR1 medium (StemCell Technology). Emerging of round cells with colony formation was observed as early as 5 days post feeder layer cultures. The two-factor (OK)- transfected cells showed similar morphological changes when compared to the four iPS factor (OSKM)-transfected cells that were undergoing early stages of somatic cell reprogramming (Fig. 1a), with the exception that more dense colony formation and immortalization from the OSKM-overexpressed iPS colonies (adipose-derived iPS cell [AdiPS cells]) were observed. Interestingly, similar morphological changes have been shown from multilineage blood progenitors through overexpression of Oct4 in human adult dermal and neonatal foreskin fibroblasts. 16 We established nine OK cell lines and multiple iPS cell lines from OSKM-overexpressed preadipocytes by colony picking.

Ectopic expression of Oct4 and Klf4 in human preadipocytes.
To further characterize OK-transfected cells, we examined global gene expression patterns using microarray analysis (Illumina). Sample relations based on the expression of 20,909 genes showed that OK cells (single clone, mixed colonies) are distinct from preadipocyte and four-factor (OSKM)-induced iPS cells (AdiPS) (Fig. 1b). A majority of the pluripotency genes that were highly expressed in the fully reprogrammed AdiPS cells were expressed at lower levels in two-factor-induced OK cells, but at levels similar to or less than levels observed in preadipocytes (Fig. 1c). Expression of fibroblast and preadipocyte marker genes was expressed at lower levels in OK cells with the exception of EMT activators (Slug, N-Cad), which maintained higher expression levels (Fig. 1d). In contrast to a previous report, 15 this suggested that OK cells underwent an EMT rather than reprogramming processes. Epigenetic modifiers, including inhibitors for histone deacetylase (HDAC), histone methyltransferase (HMT), and DNA methyltransferase (DNMT), are known to facilitate the reprogramming process by targeting epigenetic barriers.18–20 Small molecule combination treatments (e.g., BIX-01294, BayK8644, RG-108) have been reported to enable two-factor (Oct4 and Klf4)-induced reprogramming of MEFs. 18 In our hands, exposure to the DNMT1 inhibitor zebularine did not affect the reprogramming or global gene expression patterns (data not shown).
Transdifferentiation of hematopoietic and neuronal lineages by Oct4 and Klf4 overexpression
Using a BioGPS gene annotation portal (www.biogps.org), we analyzed the lineage and tissue-specific expression profiles of the top 292 genes characterized by a minimum fourfold change induced by Oct4 and Klf4 overexpression. As summarized in Figure 2a, we observed the enrichments of specific genes from three germ layers (mesoderm, ectoderm, and endoderm) and ICM. Overall, 41% of selected genes are annotated as tissue-specific expression while the remaining is ubiquitously expressed. Specifically, a total of 45 genes are designated hematopoietic cell specific, 59 genes are brain and peripheral nerve specific, and 27 genes are exclusively expressed from pancreas and pancreatic islets. Oct4 is also known as a hematopoietic transcription factor that enhances the expression of pan-leukocyte marker CD45 in human fibroblasts. 16 In the article, Oct4-derived fibroblasts (CD45+ FibOct4) differentiate into mature blood cells by additional cytokine treatments. Unlike the Oct4-derived fibroblasts (FibOct4, CD45+ FibOct4), OK cells do not alter the expression of hematopoietic transcription factors and cytokines (Supplementary Fig. S2). However, OK cells showed upregulation of CD71+, early erythroid marker genes (CHST2, AMPD3, CD58, SREAP3, MINPP1, IDH2), lymphoblast marker genes (CD70, HLA-DRA, MX1, CD83, CD74, IFI44L, BST2, ISG15, CLIC6, PBX3, TOX2, HCP5, RAC2, RGS1), lymphocyte marker genes (STAT4, HLA-DMB, NT5E, HCP5, RAC2, BTG1), and CD14+ monocyte/dendritic cell marker genes (F13A1, FGL2, FOLR3, MX1, PSENEN, RGS1) (Fig. 2b).

Transdifferentiation of hematopoietic, neuronal, and pancreatic lineages by Oct4 and Klf4 overexpression.
Direct conversion of mouse fibroblasts into neurons by the combinatorial expression of neural lineage-specific transcription factors (Ascl1, Brn2, Myt1l) has been demonstrated previously. 6 In addition, four reprogramming factor (Oct4, Sox2, Klf4, cMyc)-induced intermediate cells have been identified as neural progenitor cells. 21 Interestingly, the tissue-specific gene expression pattern analysis revealed that OK cells expressed increased levels of brain-specific markers (IFI6, HS3ST2, TAGLN3, SV2B, NCALD, ENC1, COBL, EEF1A2, KBTBD11, RGL1, CA11). Some genes are known to be exclusively expressed from fetal brain (NETO2, DOK5, TUBB2B, PCDH19, PTPRO, SATB2, FGF13, CXADR, LPPR4, MLLT11) or regions specific to the amygdala (TMEFF2, BRSK1, NRGN, TAC1PCDH19), pineal glands (CDH10, TPD52L1, HIST2H2BE, NCALD, CXADR), prefrontal and cortex (SLC16A9, NRGN, MEGF10, FMNL2, EEF1A2, KIAA1598, KBTBD11, RGL1, CA11, SATB2), hypothalamus and thalamus (DNER, CALB2, FMNL2, KIAA1598), and spinal cord (KAL1, DNER, MEGF10). Most of the brain-specific genes that were highly induced in OK cells were undetectable or at very low levels when compared to untransfected human preadipocytes (Fig. 2c). Neuronal differentiation, however, did not proceed any further: OK cells did not show neuronal morphological changes such as neuronal rosettes or nerve fibers (axon) (data not shown). In addition, the expression of neural lineage-specific transcription factors (Ascl1, Brn2, Myt1l) were not altered in OK cells. These results suggest that Oct4- and Klf4-induced neural transdifferentiation is independent of neural development, and that Oct4 and Klf4 overexpression directly converts mesoderm to ectoderm lineages.
Transdifferentiation of pancreatic α cells by Oct4/Klf4 or Oct4 alone
As a part of an endocrine organ, pancreatic α and β cells play vital roles maintaining blood glucose homeostasis by secreting glucagon and insulin, respectively. Three pancreatic lineage-specific transdifferentiation factors (Pdx1, Ngn3, Mafa) have been identified, which efficiently convert adult hepatocytes and pancreatic exocrine cells into insulin secreting pancreatic β cells.22–25 However, pancreatic α cell-specific transdifferentiation has not been reported. Global gene expression analysis indicated that 27 pancreatic islet-specific genes were enriched in OK cells, including pancreatic progenitor markers (RBPJ, Sox9, NeuroD, and ISL1) (Fig. 2d). The enriched genes are functionally associated with G protein-related cell signaling and are responsible for glucose sensing (RGS2, IL8, KCTD12, AKAP7), transcription regulation (ISL1, BEX1, CEPD, HIST2HAB, NEUROD, SOX9, RBPJ), and enzymes for pancreatic hormone processing (PCSK1, LOXL4, CPE, OAS1, PLTP, RNF128). Among the four pancreatic hormones that secret from distinct cells in the pancreatic islets, we observed upregulation of glucagon (GCG) gene (Fig. 2d), but not insulin, somatostatin, or pancreatic polypeptide (data not shown). These results suggest that human preadipocytes can be directly converted to pancreatic α cells by Oct4 and Klf4 overexpression. Pancreatic cell-specific gene expression has been confirmed with real-time RT-PCR using SYBR green-conjugated gene-specific primers that show distinct expression patterns across three types of pancreatic cells (α, β, and exocrine cells). 26 As shown in Figure 3a, the expression of pancreatic α cell markers (GCG, CNTN1, PCSK1, PDK4, RGS4, IRX2, LPPR4, LOXL2, KCTD12, KL) and β cell markers (ISL1, HDAC9, KCNJ2) were upregulated dramatically in OK cells, while the preadipocyte marker gene (CD36, FABP4) expression decreased (Fig. 3a). The expression of pancreatic exocrine cell-specific markers (AMY1C, AMT1B, AMY1A) was highly expressed in all samples (data not shown). To assess glucagon and NeuroD protein expression (a pancreatic α cell product and a pancreatic progenitor marker, respectively), OK cells were seeded onto Matrigel-coated cell culture plates and cells were examined by immunocytochemistry. Our results show that all of the colonies on matrigel stained with specific antibodies against glucagon or NeuroD except for the feeder cells (Fig. 3b). In an attempt to evaluate the secretion of glucagon from the transdifferentiated pancreatic α cells (OK cells, Oct4 alone), the level of glucagon was measured using an ELISA from the cell culture supernatants. Although we have detected the expression of glucagon from the cell extracts, we could hardly detect the glucagon secretion. This suggests that pancreatic α cells are missing a component for the glucagon processing and secretion, while the cells highly express pancreatic exocrine cell markers such as AMY1C, AMT1B, and AMY1A.
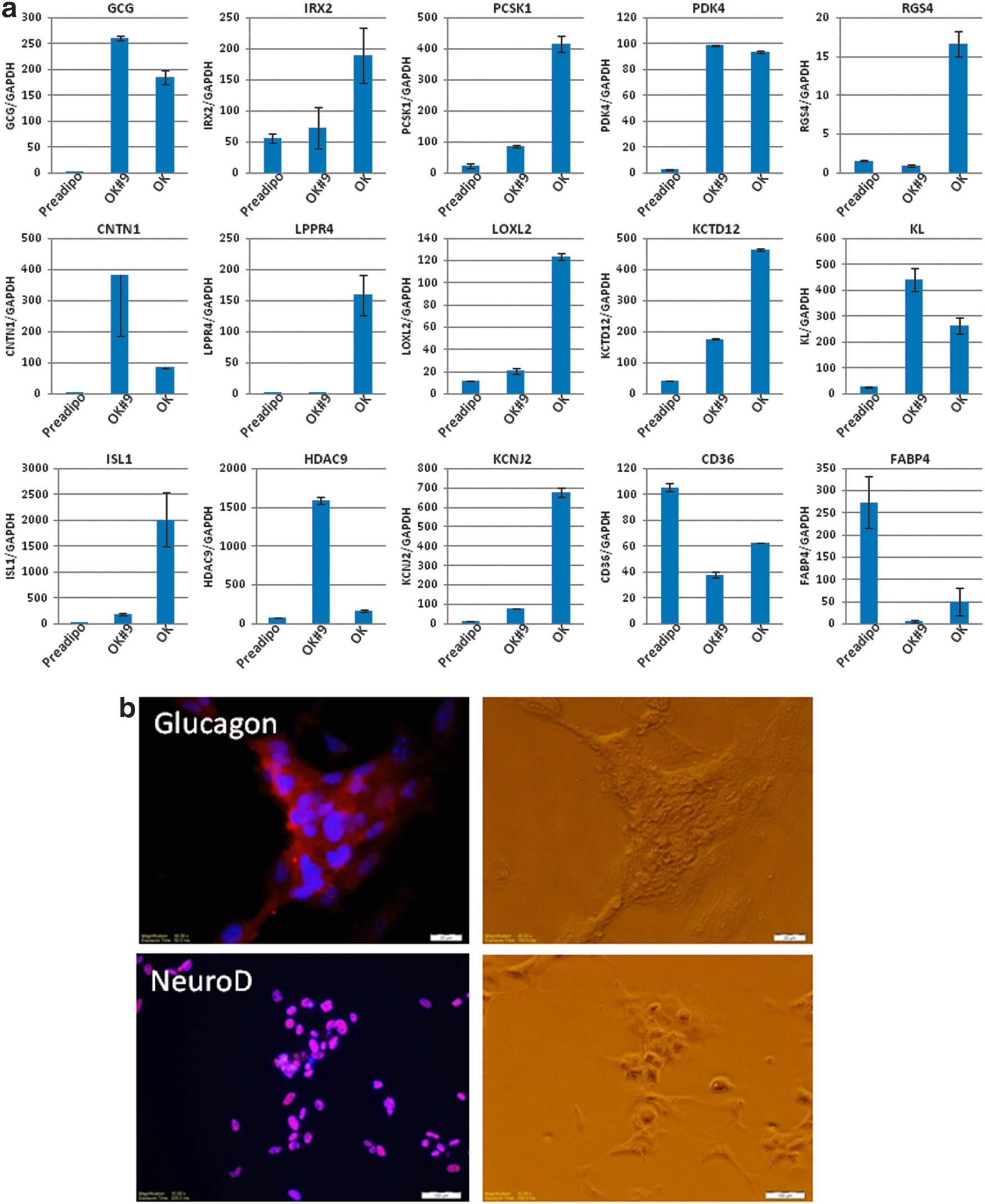
Transdifferentiation of pancreatic α cell by Oct4 and Klf4 overexpression.
Analysis of the regulatory transcription factor binding sites indicated that the promoter region of most genes that were upregulated in OK cells contain putative POU domain binding sequences (5′-ATTTGCAT-3′), but not Klf4 binding sequences, suggesting Klf4 might be dispensable for the transdifferentiation processes. To test this hypothesis, we transfected preadipocytes with Oct4 only using a lentiviral vector and repeated the experiment using the same conditions used previously. The general outline is shown in Figure 4a. Similar to the previous two-factor overexpression, we observed colony formation at 5 days after culturing on feeder cell layer. The cells had virtually identical morphology as the two-factor (Oct4, Klf4)-induced OK cells (Fig. 4a). Next, we analyzed the pancreatic cell-specific gene and protein expression from established (Oct4#1, Oct4#5) and mixed colonies (Oct4). As shown in Figure 4b, the expression of pancreatic cell markers from Oct4 single factor-overexpressed cells is upregulated at levels similar to two-factor (Oct4, Klf4)-induced OK cells, except for LOXL2, RGS4, and PCSK1. Glucagon and NeuroD protein expression was confirmed by immunohistochemistry (Fig. 4c). These results suggest that ectopic expression of Oct4 alone is sufficient to induce transdifferentiation of human preadipocytes into glucagon expressing pancreatic α cells. These results are consistent with previous findings that Oct4 and Nanog play roles in maintaining pluripotency, while Klf4 and cMyc regulate cellular division. 27 This report also indicates that Klf4 is dispensable in the transdifferentiation processes. We have expanded the Oct4-induced pancreatic α cells into more than 10 passages without any loss of cellular proliferation properties or pancreatic marker gene expression (data not shown). Glucagon producing pancreatic α cells comprise 0.2–0.5% of pancreatic islets and show great plasticity to become insulin-secreting β cells after severe β cell loss in mouse model. 28 Furthermore, Bramswig et al. have recently demonstrated that HMT inhibitor treatment facilitates the conversion of pancreatic α cell to insulin-secreting β cells. 26 These findings regarding the plasticity of adipose stem cells and transdifferentiation into pancreatic endocrine cells and the development of a simple and efficient method to produce pancreatic α cells will provide a model to advance therapeutic strategies for metabolic disease.
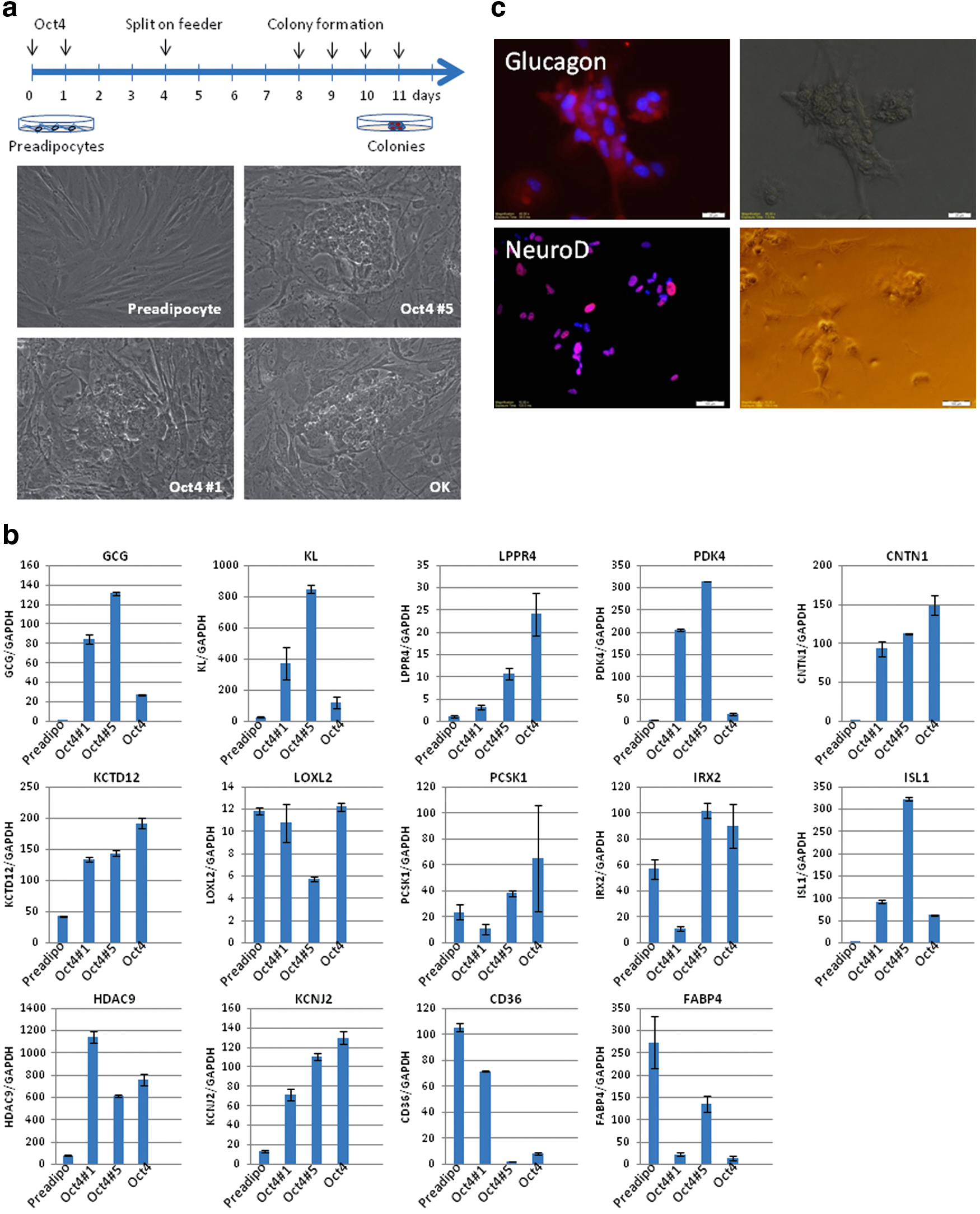
Transdifferentiation of pancreatic α cells by Oct4 overexpression.
Materials and Methods
Cell culture and lentiviral transfection
Human preadipocytes (LaCell, LA) were maintained with DMEM supplemented with 10% FBS, 50 U/mL penicillin and 50 μg/mL streptomycin at 37°C, and 5% CO2 in a humidified incubator. High-titer lentivirus, overexpressing human Oct4, Klf4, or polycistronic lentivirus, which expresses four iPS factors (STEMCCA), was purchased from System Biosciences, Cellomics, and EMD Millipore. The day before lentiviral transfection, human preadipocytes were trypsinized, counted, and seeded in six-well plates at a density of 105 cells/well. The next day, the culture medium was replaced with a prewarmed medium containing 5 μg/mL polybrene (Sigma-Aldrich) and 25 MOI of lentivirus. Lentiviral infection was repeated the next day, and the culture medium was changed with a fresh preadipocyte growth medium. When the cells reached confluence, cells were trypsinized, counted, and seeded on a prepared MEF feeder layer at a density of 5 × 104 cells with the mTeSR1 medium (StemCell Technology).
Microarray and gene expression analysis
Total RNA was prepared from cultures using Trizol Reagent (Life Technology) and the RNeasy Mini RNA isolation kit (Qiagen) with DNase I digestion. The RNA quality was verified using an Agilent Bioanalyzer 2100 (Agilent Technologies) using the RNA 6000 Pico Assay. Generation of double-stranded cDNA, preparation and labeling of cRNA, hybridization to HumanHT-12 v4 Expression BeadChip (Illumina), washing, and scanning were all performed according to the standard Illumina protocol. Quantitative PCR to measure mRNA expression levels was performed with PrimeTime qPCR assays (Integrated DNA Technologies) using a 7700 real-time PCR system in the genomic core facility at Pennington Biomedical Research Center. Expression levels were compared to known standard samples and were normalized to GAPDH.
Immunocytochemistry
Cells were fixed with 4% paraformaldehyde in phosphate-buffered saline (PBS) for 10 min and incubated for 1 h with antibodies specific for glucagon and NeuroD1 (Abcam). After washing thrice with PBS, cells were incubated for 1 h with fluorescent conjugated secondary antibody (Invitrogen). Nuclei were detected by DAPI staining (Vector shield).
Footnotes
Acknowledgments
The authors thank Ms. Susan Newman for technical support for microarray. We also acknowledge the use of Genomics, Cell Biology, and Imaging Core facilities that are supported, in part, by COBRE (NIH P20-RR021945) and NORC (NIH 1P30-DK072476) center grants from the NIH.
Author Disclosure Statement
All authors, with the exception of P.L., A.D., J.R., K.S., C.A., and J.K. acknowledge financial interests in the form of stock and/or stock options in NuPotential, Inc.
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
