Abstract
BAF chromatin remodeling complexes containing the BRG1 protein have been shown to be not only essential for early embryonic development, but also paramount in enhancing the efficiency of reprogramming somatic cells to pluripotency mediated by four transcription factors. To investigate the role of BRG1 in regulating pluripotency, we found that Oct4 and Nanog levels were increased immediately after BRG1 knockdown. While Nanog levels remained elevated over the investigated time period, Oct4 levels decreased at later time points. Additionally, OCT4 target genes were also found to be upregulated upon Brg1 knockdown. SiRNA-mediated BRG1 knockdown in embryonic stem (ES) cells led to Oct4 and Nanog upregulation, whereas F9 cells showed primarily Oct4 upregulation. BRG1 knockdown upregulated the expression of differentiation markers in mouse ES cells as well as differentiated morphology under reduced leukemia inhibitory factor conditions. Our results show that BRG1 plays an important role in maintaining pluripotency by fine-tuning the expression level of Oct4 and other pluripotency-associated genes.
Introduction
Embryonic stem (ES) cells are derived from the inner cell mass of blastocysts and can give rise to all three germ layers of the developing embryo. ES cells are not only capable of self-renewing, but also of exhibiting pluripotency, a feature maintained by the core network of transcription factors comprising OCT4, SOX2, and NANOG. 1 The expression levels of Oct4, Sox2, and Nanog are subject to an autoregulatory feedback loop. The pluripotency network has been the subject of investigations for many years. MicroRNA-145, the most abundant miRNA in normal vascular walls and freshly isolated vascular smooth muscle cells, 2 was recently shown to have a negative effect on Oct4, Sox2, and Klf4 expression in human ES cells. 2 In addition to transcription factors and small RNA, chromatin remodeling factors influence gene expression patterns by modifying the chromatin state of the underlying DNA either by posttranslational modifications of histone tails, also referred to as the histone code, 3 or by hydrolysis of ATP to noncovalently restructure, mobilize, or eject nucleosomes for modulating the access of transcription factors to chromosomal DNA.4–6 Of the second class of enzymes, five chromatin remodeling complexes have been described to date: SWI/SNF, ISWI, CHD (Mi-2), INO80, and SWR1. The individual components of these remodeling complexes play an important role in pluripotency. 7 Mouse and human cells possess two distinct SWI2/SNF2-like ATPase subunits: BRM and BRG1. Brahma-related gene 1, or simply BRG1, is a conserved subunit of the SWI/SNF family of ATP-dependent chromatin remodeling complexes. A knockout of Brg1 was shown to cause embryo lethality at the peri-implantation stage of mice, and attempts to derive Brg1-knockout ES cells have not been successful. 8 Furthermore, attempts to define the role of the BRG1-associated complex in cell survival via deletion of BAF155 9 and BAF47 10 have led to embryonic lethality. Additionally, BRG1 and BAF155, components of the BAF complex, were shown to increase the efficiency of somatic cell reprogramming when used together with the four factors (OCT4, SOX2, KLF4, and c-MYC). 11 Similarly, BRG1 has been shown to be an essential nuclear factor for reprogramming somatic cells using the approach of somatic cell nuclear fusion. 12
ES cell potency was recently shown to fluctuate in cultures. A rare transient cell population of ES cells was found to lack Oct4, Sox2, and Nanog but to express those transcripts present in two-cell stage mouse embryos. Nearly all ES cells were shown to cycle in and out of this privileged state, which is partially controlled by histone-modifying enzymes. 13 Interestingly, BRG1 has been shown to be important for zygotic genome activation at the two-cell stage of the embryo. These observations point to the important role of the BRG1-containing BAF chromatin remodeling complex in regulating pluripotency of ES cells.
BRG1 knockdown was shown to lead to the differentiation of ES cells.14,15 Both studies found that expression of Oct4, a pluripotency-associated marker, was downregulated in mouse ES (mES) cells at a later time point after BRG1 knockdown, whereas no pluripotency-related gene expression changes were seen immediately after BRG1 knockdown. Although pluripotency-associated genes, such as Oct4 and Nanog, were found to be upregulated immediately after siRNA-mediated BRG1 knockdown, 16 similar observations were made in mouse blastocysts by using siRNA-mediated knockdown of BRG1. 15
To further clarify the role of the BRG1-containing BAF complex in regulating pluripotency, we utilized both the shRNA- and siRNA-based knockdown approaches in mES cells as well as F9 cells, mouse embryonal carcinoma cells, which react relatively robustly to nonphysiological stress. We found that immediately after BRG1 knockdown, Oct4 and Nanog levels were increased, whereas Sox2 levels were decreased. Although Nanog levels remained elevated over the investigated time period, Oct4 levels decreased at later time points. Furthermore, BRG1 knockdown was shown to upregulate OCT4 target genes. Additionally, BRG1 siRNA-mediated knockdown led to Oct4 and Nanog upregulation in ES cells, whereas F9 cells showed primarily Oct4 upregulation, indicating the cooperativity of BRG1 and leukemia inhibitory factor (LIF)/Stat3 pathway. This was confirmed by observations showing that mES cells undergo morphological differentiation immediately after BRG1 knockdown under reduced LIF conditions. Our results show that BRG1 maintains the pluripotency of mES cells by acting both as an activator and a repressor of the expression of Oct4 and other pluripotency-associated genes, thus regulating the levels of these key pluripotency genes.
Material and Methods
Cell culture, transfections, and alkaline phosphatase staining
F9 cells were cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% heat-inactivated fetal bovine serum (FBS). Mouse ES cells (mES OG2 and ESD3) were cultured under feeder-free conditions with knockout DMEM containing 4 mM L-glutamine, 1.5 g/L sodium bicarbonate, and 4.5 g/L glucose supplemented with 0.1 mM β-mercaptoethanol and 10% heat-inactivated FBS. The ES cell medium was completed with 2000 U/mL of LIF unless stated otherwise. F9 cells were transfected using Nucleofection (Lonza, Basel, Switzerland), whereas mES cells were transfected using Lipofectamine 2000 (Invitrogen, Carlsbad, CA). Alkaline phosphatase (AP) staining was performed with the ES Cell Characterization Kit (Millipore, Billerica, MA) according to the manufacturer's protocol.
ShRNA-mediated knockdown of BRG1 and time-course analysis
For shRNA-mediated knockdown of BRG1, SureSilencing shRNA constructs against BRG1 were purchased from SABiosciences (Frederick, MD). The best knockdown efficiency was achieved using ShRNA-Brg1 with the following sequence: 5′-GAC CAC CTA TGA ATA TAT CAT-3′. ShRNA with the following sequence served as a control: 5′-GGA ATC TCA TTC GAT GCA TAC-3′. Green fluorescent protein (GFP)-positive cells sorted by fluorescence-activated cell sorting (FACS) were collected at the indicated time points and total RNA was isolated.
SiRNA-mediated knockdown of BRG1
siRNA SMARTpool oligos against mouse Brg1 were purchased from Dharmacon (Schwerte, Germany; On-TARGET plus SMARTpool L-041135-00-0005) with the following sequences: GAG CGA AUG CGG AGG CUU AUU; CAA ACU GGG CGU AUG AAU UUU; GAG ACU AUC CUC AUU AUU CUU; and GAU CCU CAC UGG CAC AGA UUU.
Total RNA was isolated at 48 and 72 h posttransfection from ES and F9 cells, respectively.
BrdU and Annexin V staining
BrdU (10 μM; BD Biosciences, Franklin Lakes, NJ) was added to the cell culture medium for 45 min. BrdU-specific antibody was applied after membrane permeabilization according to the manufacturer's manual, followed by flow analysis. Annexin V staining was performed according to the manufacturer's instructions (BD Biosciences). All samples were analyzed on a FACSAria cell sorter (BD Biosciences). Data analysis was performed using FlowJo software (Tree Star, Ashland, OR).
Nuclear extract preparation
OG2 ES cells were cultured under feeder-free conditions and washed in phosphate-buffered saline (PBS) before trypsin digestion. Cells were centrifuged at 2500 g for 6 min, and the pellet was washed twice with PBS before it was resuspended in 5 packed cell volumes of prechilled Buffer A (10 mM KCl, 10 mM HEPES, pH 7.9, 1.5 mM MgCl2, 2 mM dithiothreitol, 0.5 mM phenylmethanesulfonyl fluoride, 0.2% IGEPAL CA 630). Cells were incubated on ice for 30 min before being centrifuged at 2000 g for 8 min at 4°C. All further work was performed at 4°C unless stated otherwise. The pellet was resuspended in 2 packed cell volumes of Buffer A plus 0.2% IGEPAL, and cell lysis was monitored. After lysis, the sample was centrifuged at 1000 g for 10 min and the supernatant was carefully removed. Nuclei were centrifuged at 4000 g for 10 min, the supernatant was removed, and the cell pellet was resuspended in 1 packed cell volume of RIPA buffer (50 mM Tris, pH 8.0, 300 mM NaCl, 1% IGEPAL CA 630, 0.5% sodium deoxycholate, 1 mM EDTA, 1× protease inhibitors [Roche, Mannheim, Germany], and 1× phosphatase inhibitors [Sigma, St. Louis, MO]). Nuclei were sonicated three times (10 sec, 1-msec pulses, 30% power) using a Bandelin Sonoplus ultrasonic homogenizer. The nuclear extract was centrifuged at 16,000 g for 20 min and the sample was frozen in liquid nitrogen.
Immunoprecipitation
All work was performed at 4°C, except for sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and chemiluminescence detection, which were performed at room temperature. Dynabeads (Invitrogen) were washed in 0.1 M citric acid (pH 5.0) and equilibrated in RIPA buffer (150 mM NaCl) before the beads were coated with bovine serum albumin and the appropriate antibody. Nuclear extract (2500 μg) was precleared by incubation with 20 μL of anti-goat IgG antibody for 2 h. Immunoprecipitation was performed with 25 μL of anti-BRG1 (N-15) antibody (Santa Cruz sc-8749; Santa Cruz, CA) for 30 min at 4°C. Twenty-five microliters of goat IgG antibody was used as a negative control. Captured immunocomplexes were washed three times in RIPA buffer (150 mM NaCl) using the MACS system (Miltenyi, Bergisch Gladbach, Germany). Specifically bound proteins were eluted at 95°C in Laemli buffer and analyzed on immunoblots using 8% SDS-PAGE gels. For Western blotting, anti-OCT4 (Santa Cruz) and anti–β-CATENIN (BD Biosciences) were used, each at a dilution of 1:1000.
Quantitative reverse-transcription polymerase chain reaction analysis
TaqMan Assays-on-Demand was used for the detection of the following genes (TaqMan primer ID): Brg1 (Mm01151948_m1), Oct4 (Mm00658129_gH), Sox2 (Mm00488369_s1), Nanog (Mm02019550_s1), Oct11 (Mm00478284_m1), Fgf4 (Mm00438917_m1), Fgf5 (Mm00438919_m1), Gata4 (Mm00484689_m1), Meox1 (Mm00440285_m1), and T (Mm00436877_m1). Quantification was normalized to endogenous Hprt1 (Mm00446968_m1). The gene within the log-linear phase of the amplification curve obtained for each probe–primers set was analyzed using the ΔΔCt method (ABI PRISM 7700 Sequence Detection System, user bulletin number 2). Total RNA was extracted using the RNeasy Kit (Qiagen, Hilden, Germany), and cDNA was generated using Superscript II (Invitrogen). Polymerase chain reaction was performed for 35 cycles for all marker genes.
Results
Oct4 and Nanog expression is upregulated upon BRG1 knockdown
ShRNA-mediated BRG1 knockdown was carried out to investigate the BRG1-mediated regulation of ES cell pluripotency. BRG1 knockdown was carried out by transfecting ES cells with a plasmid expressing GFP along with shRNA against Brg1. To study early phenotypes, GFP-positive cells were sorted by FACS at the indicated time points and further analyzed for the expression of pluripotency genes. ShRNA against Brg1 led to an 80% reduction in RNA levels at 24 h after transfection and up to a 90% reduction at 48 h posttransfection. RNA levels were reduced for at least 72 h posttransfection (Fig. 1A).

Time-course analysis was performed by using shRNA directed against Brg1.
ShRNA-mediated BRG1 knockdown led to Oct4 upregulation at 24 h posttransfection. Oct4 was downregulated at 48 and 72 h posttransfection (Fig. 1B). Interestingly, Nanog mRNA levels were also upregulated at 24 h posttransfection, but remained elevated continuously over the investigated time period of 72 h (Fig. 1C).
To further confirm our initial results from the shRNA study, we carried out BRG1 knockdown using a commercially available siRNA SMART pool (i.e., a pool of four different siRNAs). We observed about 85% knockdown in Brg1 RNA levels at 48 h posttransfection (Fig. 1D). Western blot analysis at 48 h posttransfection confirmed BRG1 knockdown at the protein level as well (Fig. 1E).
SiRNA-mediated BRG1 knockdown led to Oct4 upregulation by about 50% and Nanog by about 70%, whereas levels of the HMG-box transcription factor Sox2 were slightly reduced at 48 h posttransfection (Fig. 1F). These observations show that BRG1 regulates the pluripotency genes Oct4, Nanog, and Sox2 in a gene-specific manner. Whereas Oct4 and Nanog mRNA levels are upregulated, Sox2 mRNA expression levels are downregulated immediately after BRG1 knockdown. At later time points, Oct4 mRNA expression is downregulated, whereas Nanog mRNA levels remain elevated.
Brg1 knockdown leads to Oct4, but not Nanog, upregulation in F9 cells
ES cells exhibit sensitivity to stressful environments—induced by changes in culture or various experimental conditions—and tend to differentiate under those conditions. We therefore repeated the analysis using the more environmentally robust model of embryonal carcinoma F9 cells. SiRNA SMART pool against Brg1 was transfected into F9 cells using nucleofection, leading to 90% knockdown in Brg1 RNA levels at 72 h posttransfection (Fig. 2A).
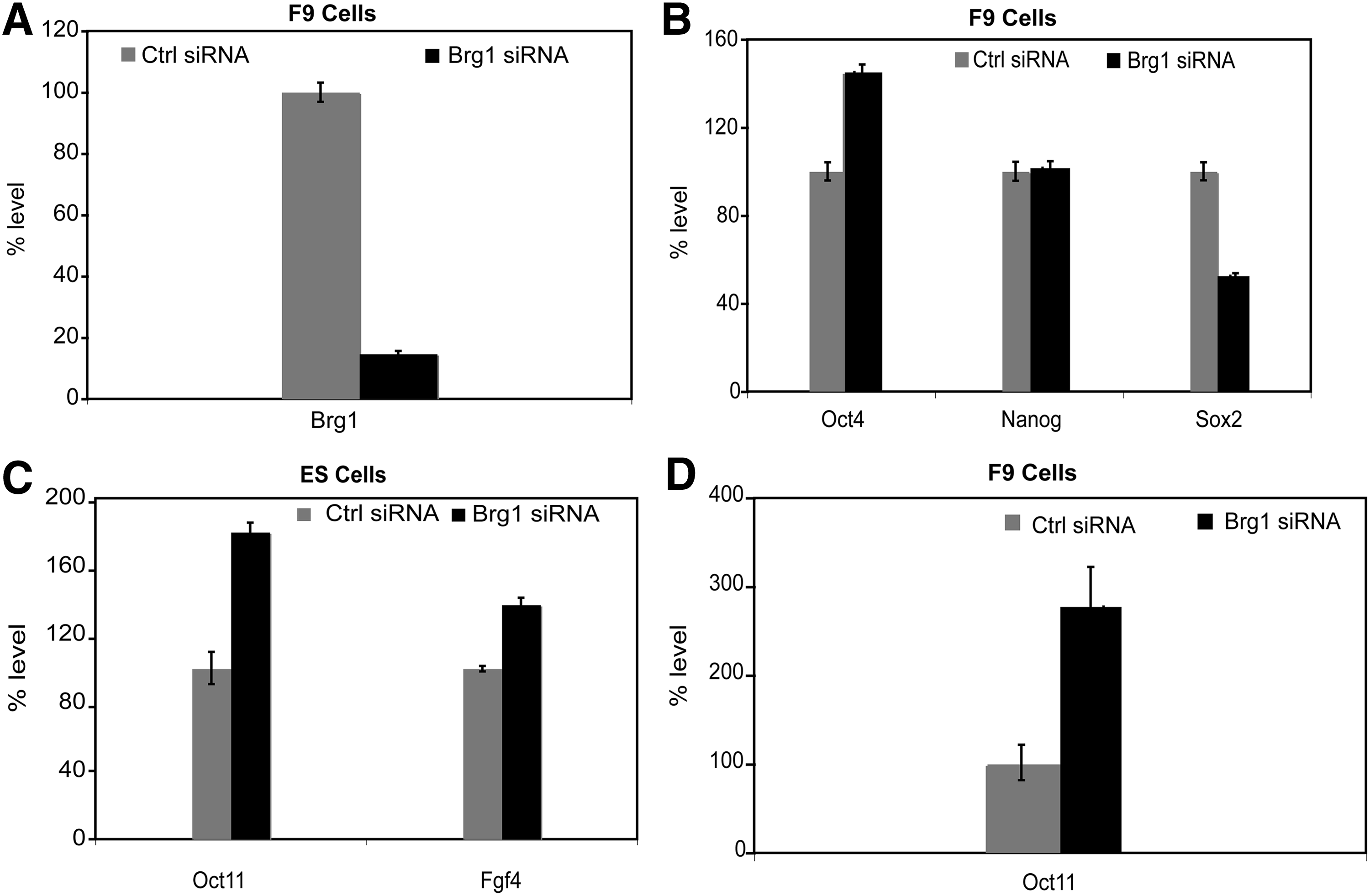
BRG1 knockdown in F9 cells using SMARTpool siRNA and OCT4 target genes.
Consistent with our findings in mES cells, BRG1 knockdown in F9 cells caused an upregulation in Oct4 mRNA expression by about 40%, but a downregulation in Sox2 mRNA expression by 50%. However, Nanog levels were not altered by BRG1 knockdown (Fig. 2B). Taken together, our results confirm that BRG1 knockdown leads to an immediate increase in Oct4 transcript levels, but an immediate decrease in Sox2 levels in pluripotent cells such as mES cells and F9 cells.
Oct4 target genes are upregulated in Brg1-knockdown cells
To assess whether upregulation of Oct4 transcripts is reflected at the OCT4 protein level, we examined the expression of the downstream OCT4 target genes Oct11 and Fgf4.17–19 Oct11 RNA levels were increased by more than twofold in F9 cells; RNA levels were also increased in ES cells (Fig. 2C,D). Brg1 knockdown also led to a significant increase in Fgf4 levels in ES cells (Fig. 2C). The activation of these OCT4 target genes demonstrates that OCT4 upregulation mediated by BRG1 knockdown is not limited to the RNA level, but is reflected at the functional level of the OCT4 protein.
OCT4 interacts with the BRG1-containing BAF complex
To uncover the molecular mechanism underlying BRG1 regulation of mES cell pluripotency, we investigated whether BRG1 interacts directly with OCT4. BRG1 was immunoprecipitated from the nuclear extract of mES cells using a polyclonal antibody against BRG1. We assayed for the presence of OCT4 in the pull-down and found that OCT4 co-immunoprecipitates with Brg1. β-CATENIN has been demonstrated to interact with Brg1 20 and was used here as a positive control (Fig. 3A). These data reveal that OCT4 is a direct binding partner of the BRG1-containing complex in mES cells. This interaction is likely responsible for BRG1 recruitment onto the promoters of Sox2, Nanog, and Oct4.

Interaction of BRG1 with OCT4, and the role of BRG1 in cell cycle and apoptosis.
Brg1 knockdown leads to cell cycle arrest and apoptosis
We attempted to generate a stable Brg1-shRNA cell clone but were not successful. BrdU and Annexin V–APC analysis showed that BRG1 is required for cell cycle completion and cell survival, since shRNA-mediated BRG1 knockdown led to arrest of cells at the G1 phase of cell cycle (Fig. 3B) and increased apoptosis within 72 h of transfection (Fig. 3C), which is consistent with the role of the BRG1-containing chromatin remodeling complex in cell cycle progression, apoptosis, and cell differentiation.
BRG1 knockdown causes the differentiation of ES cells
To test whether reduced levels of BRG1 lead to differentiation of ES cells, we assessed the expression of differentiation markers for ectoderm, mesoderm, and endoderm in ES cells at 48 h after siRNA-mediated BRG1 knockdown. Levels of the endodermal marker Gata4 and the mesodermal marker Brachyury (T) were not changed, whereas those of the mesodermal marker Meox1 were reduced by about 50%. Only levels of the primitive ectodermal marker Fgf5 were elevated by more than twofold, indicating that BRG1 knockdown leads to the differentiation of ES cells, possibly toward the ectodermal lineage (Fig. 4A).

BRG1 knockdown leads to the differentiation of ES cells.
However, siRNA-mediated knockdown of BRG1 cells did not lead to morphological changes in ES cells (data not shown). We previously observed that Nanog levels were upregulated in ES cells, which may prevent early changes in ES cell colony morphology. To test this hypothesis, we carried out Brg1 knockdown in medium with a lower LIF concentration (1000 U/mL). As expected, knockdown of BRG1 under reduced LIF conditions prevented the formation of ES cell colonies, whereas mock-knockdown cells maintained their ES cell morphology under the same culture conditions. Colonies from both samples were further analyzed by AP staining. BRG1-knockdown cells showed reduced AP expression compared with mock-knockdown cells, indicating that BRG1 knockdown leads to the differentiation of ES cells under reduced LIF conditions (Fig. 4B).
Taken together, our results show that BRG1-mediated chromatin remodeling complexes play an important role in regulating the pluripotency of mES cells by maintaining the expression levels of key pluripotency genes.
Discussion
Interestingly, two studies showed that BRG1 knockdown leads to differentiation of the ES cells.14,15 Both studies showed that Oct4, a pluripotency-associated marker, was downregulated upon BRG1 knockdown. In contrast, another study from one of these groups 16 showed that pluripotency-associated genes, such as Oct4 and Nanog, were upregulated upon BRG1 knockdown. Similarly, other reports have produced mixed results on the role of BAF155 in ES cells. BAF155 shows high expression in ES cells,14,21 and reduction in BAF155 expression leads to aberrant ES cell colony morphology 22 and decreased OCT4 expression in ES cells. 14 However, in differentiating ES cells, loss of BAF155 results in perturbed chromatin condensation and increased OCT4 expression. 21 Based on these results, speculation has swirled around the stoichiometry of the different BAF subunits, and not their actual levels, as determining subunit function, perhaps reconciling the contradictory results in the literature. 7 However, the reported contradictory effects of BRG1 knockdown on the expression of pluripotency-associated genes require further investigation, since BRG1 knockdown may provide another layer at which BRG1-containing BAF complexes regulate ES cell pluripotency.
To understand how BRG1 contributes to the maintenance of ES cell pluripotency, we focused on the immediate effect of BRG1 knockdown on Oct4, Sox2, and Nanog expression. We showed that Oct4 mRNA levels increase immediately after BRG1 knockdown, followed by a decrease in Oct4 transcripts. These observations were confirmed by siRNA-mediated knockdown of BRG1 in two different pluripotent cell lines. We further demonstrated that downstream OCT4 target gene expression is also upregulated at the RNA level. BRG1 has been recently shown to be recruited onto the Oct4 promoter.15,16 This suggests that Brg1 is directly involved in the transcriptional regulation of Oct4. Our immunoprecipitation experiment demonstrates that BRG1 interacts with the OCT4 protein. Thus, it is likely that OCT4 plays an important role in recruiting BRG1 onto the promoter of Oct4 and downstream target genes, such as Sox2 and Nanog. Since Nanog levels remain elevated for at least 72 h post–BRG1 knockdown, downregulation of Oct4 expression reflects a state of cellular differentiation rather than a general impairment in the transcriptional capability of BRG1-knockdown cells. Although our data demonstrate that BRG1 represses Oct4, we can only speculate about the molecular function of BRG1. BRG1 may provide an interaction surface for the recruitment of repressors onto the Oct4 promoter. An interaction between BRG1 and DNMT3a 23 as well as DNMT3b 14 has already been described in mES cells. In addition, Nanog and Oct4 have been shown to physically interact with a transcriptional repressor complex containing BRG1 together with HDAC1/2 and MBD3. 24 Although we cannot exclude the possibility that BRG1 is crucial for the expression of a repressor that keeps OCT4 expression under a critical threshold, the direct interaction of BRG1 and OCT4 and the short time span between BRG1 knockdown and Oct4 upregulation both suggest that BRG1 is directly involved in the repression of OCT4 expression. Interestingly, BRG1 knockdown triggers the differentiation of ES cells toward an ectodermal cell fate, as indicated by high Fgf5 expression, rather than expected endodermal cell fate. 25 Since NANOG overexpression has been shown to rescue ES cells from differentiation into endodermal cells, 26 it is possible that elevated levels of NANOG override the endodermal cell fate triggered by transient OCT4 upregulation, thus accounting for ES cell differentiation into the ectodermal cell fate. Increased NANOG expression upon BRG1 knockdown may therefore explain why the LIF concentration had to be lowered to induce morphological changes in ES cells. Since LIF has been described to induce NANOG expression via STAT3 signaling in mES cells, 26 it follows that ES cells expressing high levels of NANOG bypass the need for LIF and exhibit clonal expansion. 27 Interestingly, BRG1 was recently shown to potentiate LIF signaling by facilitating STAT3 binding thus validating our observations that increased LIF levels can alleviate the effect of Brg1 depletion. 28
In this study, we show that key pluripotency factors respond to loss of BRG1 by either the up- or downregulation of gene expression in a gene-specific manner. It is very likely that BRG1 plays an important role in stabilizing the autoregulatory feedback loop of the core pluripotency network comprising OCT4, SOX2, and NANOG by acting as a platform for activators and repressors. We show that BRG1 knockdown leads to elevated levels of OCT4 and speculate that this in turn triggers ES cell differentiation. The numerous functions of BRG1 on various promoters, however, suggest a multilayer scenario for the role of BRG1 in maintaining ES cell pluripotency. The mechanism underlying the differential recruitment of activators and repressors onto the target promoters of Oct4, Sox2, and Nanog remains unclear. The function of BRG1 in balancing expression levels of the key pluripotency factors OCT4, SOX2, and NANOG is essential for maintaining mES cells in a pluripotent state. We have recently shown that the Brg1-containing BAF complex enhances the efficiency of reprogramming by opening up the chromatin structure through DNA demethylation and increased H3K4me3 formation in the promoter region of important transcription factors. 11 In addition, the results of this study also lend credence to the possibility that the BRG1-containing BAF complex regulates OCT4 expression, maintaining appropriate OCT4 levels during the reprogramming process, and thereby enhancing reprogramming efficiency. Therefore, much interest lies in uncovering the precise molecular role of BRG1 in recruiting activators and repressors onto the promoters of pluripotency factors during reprogramming.
Footnotes
Acknowledgments
We thank Sandra Heising (Department of Cell and Developmental Biology, Max Planck Institute) for assisting with cell culture; Claudia Ortmeier, Martina Sinn, and Gaby Verberk (Department of Cell and Developmental Biology, Max Planck Institute) for performing quantitative reverse-transcription polymerase chain reaction; and Areti Malapestas and Shelley Herron (Department of Neurosciences, University of California, San Diego) for proofreading the manuscript. This project was funded by the Max Planck Society and DFG grants DFG SI 1695/1-2 (SPP1356).
Author Disclosure Statement
No competing financial interests exist.
