Abstract
The dengue virus (DENV) genome encodes 10 different genes including the NS3 gene, which has a protease and helicase domain used in virus replication. This domain is a potential target for antiviral agents against dengue. Due to a high mutation rate, DENV is classified into four major serotypes (DENV1–DENV4). This study was designed to perform conservancy analysis of all four serotypes by drawing a consensus sequence for each serotype and then drawing a global consensus sequence to study conserved residues in all four serotypes. A total of 127 NS3 sequences belonging to all four serotypes were retrieved and aligned using multiple alignment feature of CLC Workbench and were subjected to phylogenetic tree construction. Conservancy analysis of NS3 revealed conserved peptides with active site residues that can be important in developing antiviral agents against dengue virus. Among conserved residues, residues G142, Ser144, and G145 (catalytic pocket residues), A219, D220, and D221 (divalent cations binding residues), and His56, Asp79, Ser144, 146 were highly conserved among all the serotypes. Residues from L138 to L149 and from L226 to L245 were also considerably conserved in all serotypes, while lysine141 mutated to serine in serotype 3. A total of 14 peptides from the conserved regions of DENV NS3 protein were identified, which may be helpful to develop peptide inhibitors. The DENV NS3 phylogenetic tree showed the evolutionary relationship among all four serotypes, and all serotypes of dengue were found to have evolved from the dengue 4 serotype. Because of its high variability, DENV has become a global health concern. It is important to study residues that are present in protease, helicase, the catalytic pocket Mg2+ binding site, and the AAA domain. This study revealed peptides with active site residues that are highly conserved among all four serotypes. These regions of the NS3 sequence may be helpful in developing antiviral agents.
Introduction
Dengue infection has become a major health problem in more than 100 countries in Africa, Asia, America, the Western Pacific, and the Eastern Mediterranean. In the last decade, dengue infection has caused many endemics in Pakistan and thus has become a major health issue in Pakistan. Dengue infection is caused by a member of the Flaviviradae family known as dengue virus (DENV), which has four different serotypes (DENV1–DENV4). From a prevalence analysis of dengue infections in Pakistan during the last decade, it was found that DENV2 serotype is most prevalent serotype circulating in Pakistan. Dengue infection results in two type of infections ranging from a dengue fever to a more severe infection that can cause dengue hemorrhagic fever and dengue shock syndrome.
1
Dengue fever is an infectious disease transmitted by mosquitoes (
The mode of DENV transmission makes an attractive target to study DENV propagation. 6 Dengue virus is an enveloped, 11-kilobase-long RNA positive-strand virus 7 with three structural proteins, C (capsid), M (membrane), and E (envelop), 8 and seven nonstructural proteins, NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5, that are important in viral pathology.2,9,10 Among the nonstructural proteins, NS3 has enzymatic reactions that are important for viral replication. NS3 has two domains, the N-terminal protease domain, which is important for the proteolysis of polyprotein, and the C-terminus helicase domain. 11 The NS3 N terminal domain is about 180 residues long with a triad of active residues, histidine, aspartate, and serine, that form a catalytic triad that actively participates in the catalysis process of the enzyme. The C-terminal domain consists of conserved motifs that are found in several nucleoside triphosphatases (NTPases) and the DEXH family of RNA helicase12,13 and help to increase the overall length of NS3, which results in enhanced enzymatic activity. It has also been reported that NS3 has DNA unwinding activity. Despite the high mutation rate of DENV, some residues that actively participate in viral replication remain conserved. These conserved residues can be important in developing specific antiviral agents and inhibitors against DENVs. In this study, a novel approach is used to produce a consensus sequence of NS3, which will be useful in designing peptides that may help in finding a possible cure for dengue infection. The present study was designed to draw a global consensus sequence of the NS3 protein of DENV, study the DENV NS3 conserved domain function, and draw a phylogenetic tree.
Methods
Drawing a consensus sequence of DENV NS3
A total of 127 sequences of dengue NS3 protein were retrieved from the National Center for Biotechnology Information (NCBI). 14 Consensus sequences of dengue NS3 proteins for each serotype were drawn using multiple sequence analysis feature of CLC Workbench, a software that is use to analyze and visualize sequence comparisons. 15 Thirty sequences of serotype 1 (DENV1) reported from Cuba, Indonesia, Japan, United States, Thailand, and China were used to construct the serotype 1 consensus sequence. Thirty-three sequences of serotype 2 (DENV2) from China, United States, Taiwan, and Singapore were used to construct the serotype 2 consensus sequence. Thirty-one sequences of serotype 3 from the United States, Brazil, Singapore, and French West Indies, were used to construct the serotype 3 consensus sequence. Thirty-one sequences of serotype 4 (DENV4) retrieved from NCBI and originating in Taiwan, Indonesia, United States, China, Thailand, and Brazil were used to produce the consensus sequence of DENV4.
Peptides designing for potential peptide vaccine
The consensus sequences of all four serotypes (DENV1–DENV4) were drawn in CLC Workbench and aligned to get the global consensus sequence. The consensus sequence was used to study variations in different motifs and domains of the DENV NS3 region. Short peptides from the highly conserved regions of the DENV NS3 protein were selected from the consensus sequence analysis; these peptides would be the best targets to be tested as a potential peptide vaccine.
Phylogenetic tree analysis
To draw a phylogenetic tree of the DENV NS3 gene belonging to different serotypes, we used 125 sequences; 30 sequences from DENV1, 33 from DENV2, 31 from DENV3, and 31 from DENV4. All 125 sequences were first aligned in CLC Workbench, and the aligned file was then subjected to the unweighted pair group method with arithmetic mean (UPGMA) method to draw a phylogenetic tree with boot strap value 100. The UPGMA method is based on pairwise similarity/dissimilarity distance matrix. 16
Results
NS3 protein has two domains (serine protease and NTPase helicase) and is considered the second largest protein of DENV genome. NS3 has become an important drug target because both of its domains are involved in viral replication. The NS3 consensus sequence of each DENV serotype was drawn in CLC Workbench, and the sequences were aligned to study the residues that were highly conserved among all the serotypes. Figure 1 shows the alignment of the consensus sequence of all four DENV serotypes. The alignment of all the consensus sequences will help us to study the strongly conserved residues in the DENV NS3 protein. Short peptides of 9 to 18 amino acids were designed from the highly conserved regions of the DENV NS3 consensus protein sequences; the sequence and position of these peptides are shown in Table 1. These positions are highly conserved and may serve as the targets to design peptide vaccines or site-specific inhibitors. A phylogenetic tree of 127 DENV NS3 sequences belonging to serotypes DENV1–DENV4, reported from all over the world, was constructed using the UPGMA method in CLC Workbench as shown in Figure 2. A default value of 100 was used in bootstrap analysis, and the values are present at each branch. Sequences from different serotypes are clustered together. The tree shows that the different DENV serotypes have evolved from the serotype DENV4.

Multiple sequence alignment of consensus sequences of dengue NS3 belonging to all serotypes (DENV1–DENV4). The global consensus sequence is shown at the base. Conserved residues are shown with their corresponding symbols while the highly variable amino acids are denoted by an “x” symbol. DENV, dengue virus.
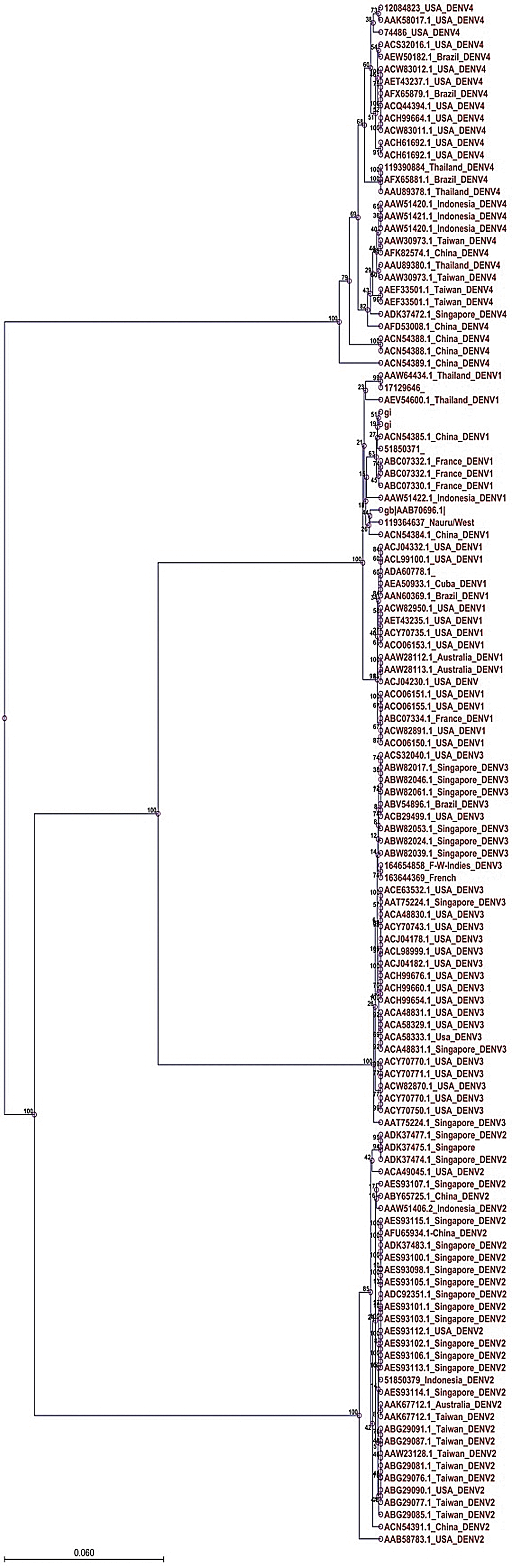
Phylogenetic tree showing evolutionary relationship among the four serotypes of dengue reported from different regions of the world.
Position and Sequence of the Peptides and the Potential Domain to Be Used for Antiviral Drug Development
Discussion
Dengue infection has become a global threat to human health. Dengue virus has four serotypes, which hinders development of a successful vaccine. 17 NS3 domains, because of their conservancy, are considered as promising targets for drug development. NS3 has three conserved domains: peptidase S7 family domain, helicase C-terminus conserved domain, and flavivirus DEAD domain. 18 Residues (1–180) of the NS3 protease domain forms two β-barrels, and between these β-barrels there are three catalytic sites (His51, Asp75, and Ser135). All the remaining residues (180–618) of NS3 forms three subdomains (I, II, III) of the helicase domain. 19 The helicase domain belongs to the P-loop NTPase family. The helicase domain family is found in other helicase-related proteins and is restricted to DEAD/H helicase.
The consensus sequence alignment shows that His56, Asp79, Ser144, and Ser146 are highly conserved among all serotypes. Site-directed mutagenesis has revealed the catalytic role of these triad residues (His, Asp, Ser). 20 Thus, it can be inferred that this conserved triad is involved in a catalysis mechanism among serine proteases. The consensus sequence alignment shows that L138 to L149 are highly conserved among all serotypes, while L141 has mutated to a serine in serotype 3.
This peptide fragment encodes the peptidase S7 superfamily, which helps in processing of the NS3 poly-protein precursor into mature proteins; this region also encodes trypsin-like serine protease. Consensus sequence analysis shows that G142, Ser144, and G145 are highly conserved and form the oxyanion hole.
20
Mutation in residues (Y159 and G160) to alanine can completely abrogate the enzyme activity.
21
The residues GXGKT are found to be conserved in all four serotypes, except serine is replaced by alanine in DENV2 and DENV4. The results obtained from consensus sequence analysis show that the peptide fragment from residue L226 to L245 encodes the P-loop NTPase superfamily. This region has two motifs [Walker A, GK(S/T), and Walker B, Dex (D/H)] which are present in nucleotide binding proteins and participate in many cellular functions. Thus, designing antipeptides against the Walker A/Walker B region can significantly reduce RNA processing efficiency.22,23 Any impairment in helicase activity halts DENV replication.
24
The consensus sequence analysis shows that the string of amino acids from I349 to R357 encodes a part of the helicase DEAD-box domain involved in nuclear transcription, nucleic acid unwinding, pre-mRNA splicing, nucleocytoplasmic transport, ribosome biogenesis, translation, RNA decay, and organelle gene expression. In addition, it also contains a putative Mg2+ binding and ATP binding site.
25
The tripeptide string of A219, A220, and A221 is involved in DNA transportation.
26
The consensus sequence alignment shows that the peptide string from 465 to 473 (QRR267GR469XGRN) is the part of the helicase domain mutation in R467 and R469 that will reduce the activity of helicase.
27
In this study, we have drawn a phylogenetic tree of 127 DENV NS3 sequences reported from different countries. The tree was constructed by the UPGMA method as shown in Figure 2. The tree shows that the serotype DENV4 occupies the root of the tree, and the first serotype to evolve from DENV4 was DENV2. DENV2 bifurcates into two wings; from one wing, DENV3 evolved, and from the second, DENV2. Regato
Footnotes
Disclosure Statement
No competing financial interests exist.
