Abstract
Objectives:
Personalized medicine emphasizes prevention and early diagnosis by developing genetic screening and biomarker assessment tools. Biobanks, including University of Piemonte Orientale (UPO) Biobank, support this effort by providing high-quality biological samples collected, processed, and stored using optimized standardized protocols. To determine the optimal long-term storage conditions for biospecimens used in biomedical research, we evaluated plasma and serum samples cryopreserved using two storage methods, cryovials and straws, across various analytical methodologies with differing sensitivity and robustness.
Design and Methods:
Plasma and serum samples cryopreserved in liquid nitrogen in vials and straw at the UPO Biobank were subjected to multiple analyses including standard biochemical laboratory analysis, targeted lipidomics, untargeted proteomics, and targeted metabolites quantification through mass spectrometry-based analytical techniques.
Results:
Our data demonstrate the robustness and applicability of both storage methods for standard laboratory analyses in evaluating clinically relevant markers in plasma and serum. Lipidomic analysis revealed slight disparities in lipid abundance, though these differences were mostly confined to specific lipid species, particularly fatty acids. Conversely, proteomic and metabolomic analyses uncovered variations in abundance in a significant, albeit limited, fraction of analytes between vials and straw-derived samples.
Conclusions:
By highlighting similarities and differences in samples stored in these conditions, this study provides significant insights into optimizing biobanking practices and understanding the factors that influence the integrity of cryopreserved biospecimens and the reliability of the data derived from them. Both straws and vials are convenient and efficient cryopreservation methods, essentially equivalent for samples dedicated to robust and relatively low-sensitive standardized analyses. However, our findings emphasize the need for caution when interpreting omics data from samples subjected to different cryopreservation methods, as subtle variations can arise even with different types of containers.
Introduction
Personalized medicine focuses on developing tools for individualized prevention, early diagnosis, and targeted therapy. 1 This approach is crucial given global demographic shifts, including an aging population and the rising prevalence of age-associated, high-impact chronic diseases. 2 These conditions diminish the quality of life and impose economic strain on health care systems, highlighting the need for proactive health care strategies with early intervention to improve outcomes and reduce the overall disease burden. 3
Biomarker discovery is crucial in precision medicine for identifying therapeutic targets in genetic, infectious, noncommunicable diseases and cancer. 4 Advances in genetic screening and biochemical biomarker analysis are improving early diagnosis 5 ; genomics and proteomics enhance our understanding of disease mechanisms and facilitate the discovery of new drug targets. 6 Metabolomics and lipidomics present promising opportunities for identifying biomarkers in complex diseases, offering insights beyond traditional technologies. 7 Also, hematochemical indicators, in combination with epigenetic markers, can predict individual organ/system functional decline and frailty and estimate lifespan. 8
Success in personalized medicine depends on large-scale data collection from numerous samples, 9 with high-impact advancements relying on research quality, reproducibility, and reliability. High-quality sample preservation is essential to meet these standards. 10
Cryopreservation at low and ultra-low temperatures is essential for the long-term storage of various biospecimens and ensures data quality from these samples. 10 Biobanks are crucial in this effort, particularly in the omics era, providing infrastructure for high-quality biospecimen collection for large-scale research. 11 However, high-throughput analytical methods present challenges in maintaining sample quality during preanalytic and storage phases. 12
In 2020, the University of Piemonte Orientale (UPO) Biobank was established as the Institutional Research Biobank of the UPO to support both epidemiological and translational research. 13 The UPO Biobank developed standard operating procedures (SOPs) for the collection, processing, and storage of biological samples by reviewing current knowledge and validated practices in the field.13–17 Several types of biological samples are preserved in UPO Biobank (i.e., whole blood, buffy coat, plasma, serum, peripheral blood mononuclear cells, saliva, urine, and stool) at different storage temperatures (i.e., +4°C, −80°C, and liquid nitrogen), to ensure optimal preservation.13,16,17
In addition to temperature, storage conditions can be affected by the choice of containers. 18 Cryovials are commonly used for blood samples but have limitations such as handling difficulties and space requirements. 19 Plastic straws, though fragile, offer a space-saving alternative and easy transportation in liquid nitrogen and dry ice, are widely used for cryopreserving livestock sperm, 20 and have also been adopted for biobanking human biospecimens. 21
The objective of this research was to evaluate the impact of storage containers on data obtained from various analytical methods. Plasma and serum samples preserved in liquid nitrogen and stored in either vials or straws were investigated to assess lipid, protein, and metabolite composition using both standard laboratory methods and advanced omics techniques. By comparing samples stored under these conditions, this study offers key insights for optimizing biobanking practices and improving the integrity of cryopreserved biospecimens and the reliability of the resulting data.
Materials and Methods
Biological samples
This research used plasma and serum samples from eight participants (Supplementary Table S1) in the Novara Cohort Study (NCS), stored in the UPO Biobank (https://biobank.uniupo.it/). NCS is a multidisciplinary, longitudinal study on aging, collecting biological samples and data from a minimum of 10,000 participants aged 18 or older in Novara Province, Italy. All participants gave informed consent for biobanking and research use of their samples/data as per the NCS protocol (approved by the Local Ethical Committee, Comitato Etico Interaziendale A.O.U. Maggiore della Carità di Novara, EC number 703/CE).
Blood samples were collected by trained nurses who collaborated in the NCS project in the morning from fasted subjects using a 21-gauge needle in ethylenediaminetetraacetic acid (EDTA) plastic tubes (BD Vacutainer), lithium-heparin plastic tubes (PST™ II Plus, BD Vacutainer), and SST II™ Advance plastic tubes (BD Vacutainer) gently inverted per instructions. UPO Biobank operators processed the samples. EDTA blood samples were centrifuged at 500 ×g for 15 minutes at room temperature to separate the cellular components. The plasma-EDTA (PLE) and the lithium–heparin (PLH) blood samples were centrifuged at 1500 ×g for 10 minutes at +4°C to remove platelets. Serum samples (SE) were centrifuged at 1250 ×g for 10 minutes at +4°C. Both plasma and serum samples were split in aliquots; 500 μL were manually aliquoted into 1.2 mL sterile vials (Biosigma SpA) and 250 μL into 300 μL mL CBS™ high-security straw (Cryo Bio System) through an automated aliquoting system, stored at −80°C for 24 hours, and then transferred to liquid nitrogen for long-term preservation.
Biochemical measurements
For standard biochemical analyses, fresh PLH, PLH, and SE samples stored in both straw (PLH_S and SE_S) and vials (PLH_V and SE_V) were employed. Aspartate transaminase, alanine aminotransferase, gamma-glutamyl transferase, high-sensitivity C-reactive protein, transferrin, and alkaline phosphatase quantifications were performed using the ADVIA® 1800 Clinical Chemistry System (Siemens Healthcare Diagnostics); ferritin and thyrotropin were analyzed with the ADVIA Centaur® XP Immunoassay System.
Lipidomics and untargeted proteomics
Lipidomic and proteomic analyses were performed on PLE and SE samples stored in both straw (PLE_S and SE_S) and vials (PLE_V and SE_V) (detailed procedures in Supplementary Data). Data were analyzed with Metaboanalyst.ca (accessed from July to September 2024). Protein–protein interaction network and gene ontology biological process analyses were performed with string-db.org (accessed in September 2024).
Metabolomics
Targeted analysis of selected metabolites (N-acetyl cysteine, cysteine, cystine, glutathione, glutathione disulfide, S-adenosyl-methionine, allantoin, methylglyoxal, methionine sulfoxide) was carried out using Liquid Chromatography-Tandem Mass Spectrometry (LC-MS/MS) technique through Multiple Reaction Monitoring (MRM) identification (quantifier and qualifier), followed by quantification with a 6-point calibration curve following UNI EN ISO 15189:2022. The analytical protocol for this determination is the exclusive property of Crescendo Care Srl (Milano, Italy), which analyzed the samples.
Results
Effectiveness of plastic straws versus conventional vials in preserving hematochemical biomarkers
Standard hematochemical biomarkers are fundamental in clinical practice and research for their affordability, reproducibility, and accessibility. Consistent biomarker assessment in both fresh and cryopreserved samples is key to maintaining data accuracy in large cross-sectional and longitudinal studies, ensuring reliable trend identification.
Our primary objective was to assess the impact of cryopreservation methods using vials and straws on clinically relevant hematochemical parameters routinely assessed in NCS samples. The results indicated that the integrity of these diagnostic indicators was comparable to fresh samples, with no significant differences between PLH stored in vials or straws (t-test, p > 0.05 for all comparisons: fresh PLH vs. PLH_S, fresh PLH vs. PLH_V, and PLH_S vs. PLH_V) (Table 1). In population studies or specific patient groups, 22 the volume of collected blood is often limited, underscoring the need for alternative matrices for biomarker analysis. Thus, we aimed to evaluate the use of cryopreserved serum for assessing blood biomarkers typically measured in plasma by our healthcare laboratory facilities. Our results demonstrated that serum is equally suitable for measuring the selected parameters, providing fully comparable values regardless of the cryopreservation method used (t-test, p > 0.05 for all comparisons: fresh PLH vs. SE_S, fresh PLH vs. SE_V, and SE_S vs. SE_V) (Table 1).
Hematochemical Biomarkers Assessment in Plasma and Serum (Data Represent Mean ± SD)
ALP, alkaline phosphatase; CRP, C-reactive protein; GGT, gamma-glutamyl transferase; GOT/AST, aspartate transaminase; GPT/ALT, alanine aminotransferase; SD, standard deviation; TSH, thyrotropin.
This finding highlights the robustness of both preservation techniques, affirming their reliability for maintaining essential biomarkers and supporting their incorporation into biobanking SOPs.
The choice of storage method impacts the lipidomic profile of plasma and serum samples
Despite various technical challenges, including the diversity of lipid species and the required sensitivity in analytical procedures, lipidomics has rapidly evolved as a crucial tool for understanding lipids’ roles in metabolism and disease mechanisms.23–25 Thus, we aimed to evaluate how storage methods affect the lipid profiles of PLE and SE samples from eight subjects’ samples stored in the UPO Biobank (Supplementary Table S1). To this end, a basic standard protocol was applied, targeting 171 species across major lipid classes, including fatty acids (FA), glycerolipids, glycerophospholipids, and sphingolipids (refer to Materials and Methods).
A total of 120 lipid species were identified in at least one sample, with 90 detected in both matrices (Supplementary Table S2). In PLE, 107 lipid species were identified, with 11 lipids exclusive to PLE_S and 8 to PLE_V. In SE, 103 lipid species were identified, including six unique to SE_S and seven to SE_V.
Principal component analysis (PCA) separated SE and PLE samples but distinguished them also based on the freezing method used (Fig. 1A). A volcano plot analysis (Fig. 1B, C) identified lipid species significantly different between vials and straws for both matrices. Sixteen lipids differed significantly between PLE_S and PLE_V, while 17 lipids differed between SE_S and SE_V. In both biological matrices, FA and FA_B comprised over 80% of the affected lipids. Stearic acid_B (Log2FC 4.52) and linoleic acid_B (Log2FC 2.69) were the most significantly overrepresented lipids in SE and PLE, respectively, while eicosatrienoic acid_B (Log2FC 10.31) was the most underrepresented in SE, and butyric acid_B (Log2FC 7.96) in PLE (Supplementary Table S3). These results underscore the variability introduced by storage conditions, highlighting the need to standardize preanalytical variables in lipidomics protocols for data consistency. Although not the main objective, we also compared the lipidomic profiles between SE and PLE. The comparison of PLE_V and SE_V revealed 19 significantly different lipids (log2FC> ±2 and p < 0.05), including diacylglycerols (e.g., dag18:0_20:4, dag18:0_22:6) (Supplementary Fig. S1A, B), highlighting the clotting process’s impact on serum lipid levels. 26
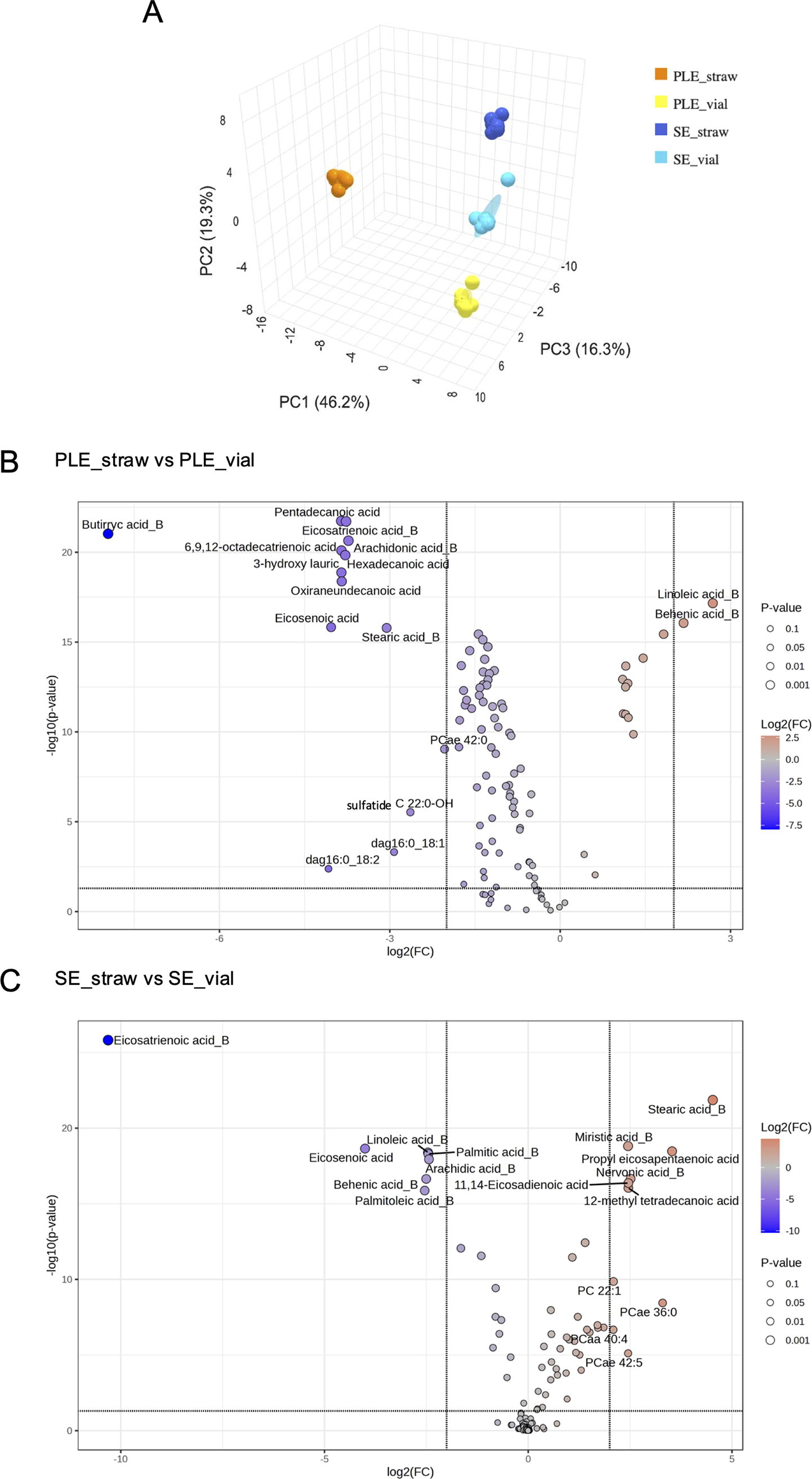
Targeted lipidomic analysis of PLE and SE samples stored in vials and straws. Principal component analysis (PCA) plot illustrating the multivariate distribution of pooled plasma and serum samples stored in vials or straws
Protein abundance is influenced by the choice of the storage method
Proteins are a valuable source of information about human health and disease in omics research and analyzing the blood proteome can offer insights into the coordination of different organ systems, enhancing the understanding of human health. 27
To evaluate the impact of the two storage methods on the proteomic profiles of PLE and SE samples, we conducted an untargeted proteomic analysis on the selected NCS subjects (Supplementary Table S1). Eighty-six proteins were identified, with 50 detected across all samples, regardless of biological matrix or storage method. Detailed analysis revealed 64 proteins in PLE_S and 61 in PLE_V, while 69 proteins were identified in SE_S and 63 in SE_V (Supplementary Table S4). PCA confirmed the expected differences between PLE and SE and highlighted distinctions between vials and straws (Fig. 2A).
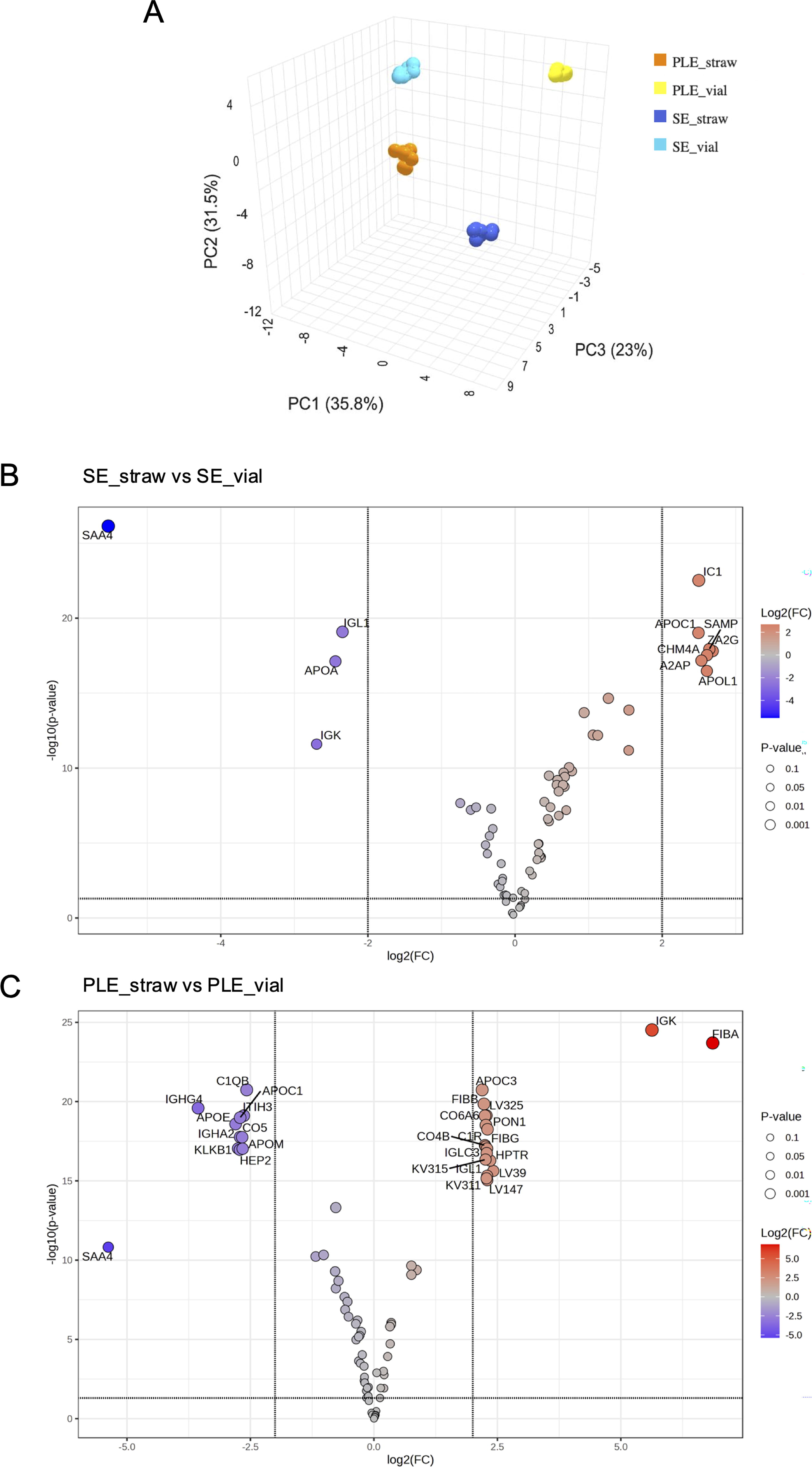
Untargeted proteomic analysis of PLE_straw versus PLE_vial and SE_straw versus SE_vial. PCA plot illustrating the multivariate distribution of plasma and serum samples stored in vials or straws
A volcano plot was generated to further explore proteins significantly affected by storage methods, as illustrated in Figure 2B, C, and Supplementary Table S5. In both matrices, we observed a notable enrichment of specific proteins in straw-stored samples compared to vials, with 18 proteins in SE and 7 in PLE showing this pattern. Fibrinogen alpha chain and immunoglobulin kappa light chain (IGK) were significantly overrepresented in SE_S [log2(FC) >5], while zinc-alpha-2-glycoprotein and serum amyloid P-component were overrepresented in PLE_S [log2(FC) >2.5]. Conversely, 11 proteins were overrepresented in SE_V and 4 in PLE_V, with serum amyloid A-4 protein (SAA4) being the only protein with a log2(FC) >5, consistently more abundant across all vials-cryopreserved samples.
We compared the proteomic profiles of PLE_V and SE_V samples, revealing significant enrichment of coagulation proteins among differentially expressed proteins (log2FC > ±2 and p < 0.05) as shown in the volcano plot (Supplementary Fig. S2A, B). This finding was further validated through STRING software analysis (Supplementary Fig. S2C).
Overall, our results show differences in protein abundance between SE and PLE samples based on storage methods, underscoring the impact of storage conditions on protein preservation and biomarker discovery.
Metabolite identification through targeted MS reveals variability between the two storage methods
To assess the impact of storage methods on blood metabolites related to aging and chronic degenerative diseases, we conducted a targeted metabolomic analysis of metabolites involved in redox metabolism regulation and oxidative stress protection (Table 2).
List of Metabolites Quantified Through Targeted Mass Spectrometry Analysis
LOQ, limit of quantification; nd, not detected; SE, serum sample; PLE, plasma-ethylenediaminetetraacetic acid sample.
All metabolites, except one, were detected and quantified in both plasma and serum. Methionine sulfoxide was undetectable in all samples, indicating minimal oxidative damage and effective sample storage management. 28 Cysteine, GSH, and GSSG levels were low across all samples, likely due to their inherent instability in cryopreserved blood derivatives stored under standard conditions without stabilizers. 29 We compared metabolite concentrations between the two matrices stored in vials versus straws (Table 2, Fig. 3). No significant differences were found for NAC and ALL, but MGO and cystine concentrations were significantly higher in SE_V than SE_S. SAM levels were also significantly higher in straws than in vials for both PLE and SE.
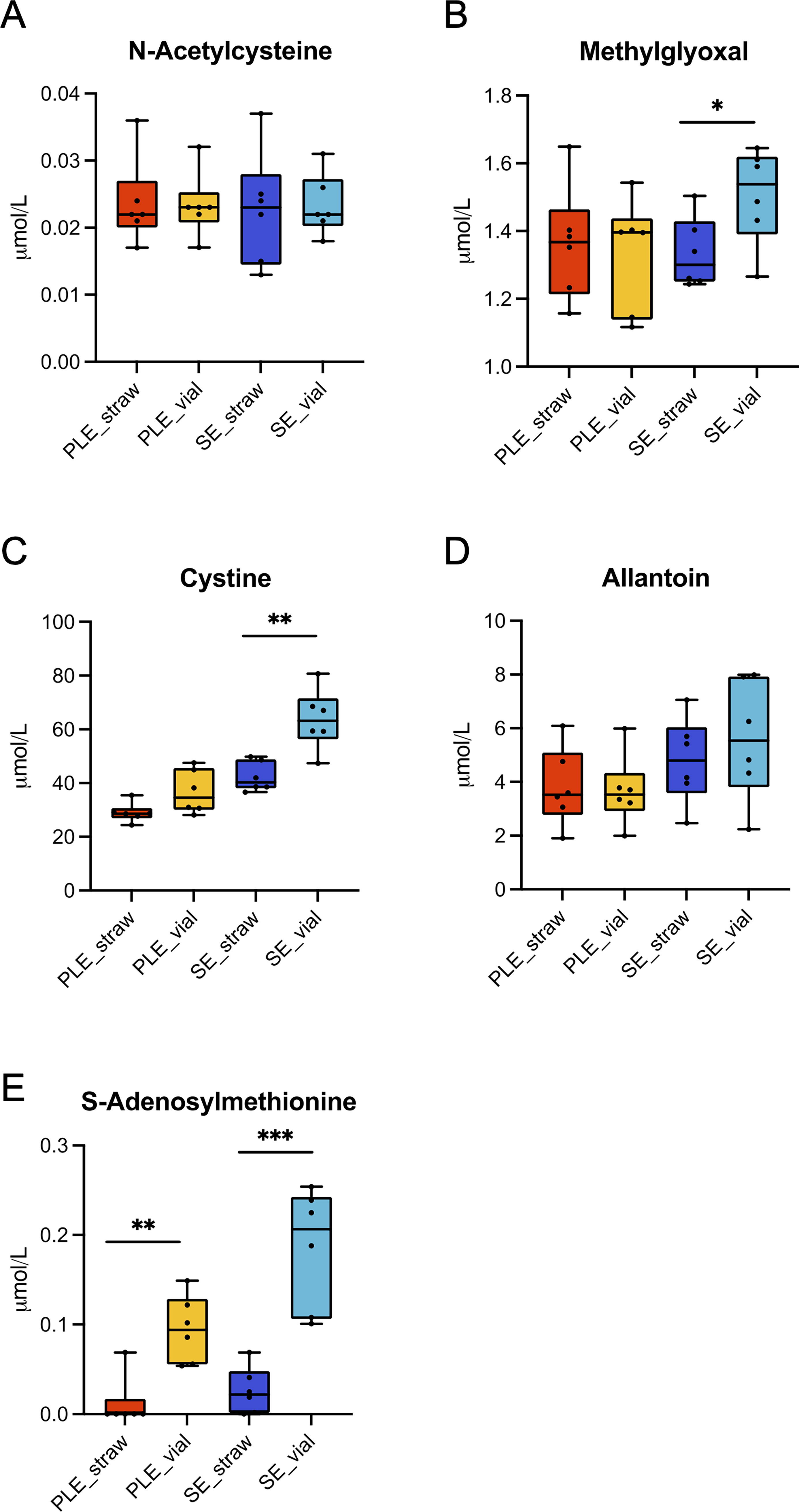
Metabolite quantification in PLE and SE samples through targeted mass spectrometry analysis. Graph representing metabolites quantification in PLE and SE samples. Each dot represents an independent subject. Statistical significance of SE_vials versus SE_straw and PLE_vials versus PLE_straw was determined through Student’s t-test (p < 0.05; *p < 0.05; **p < 0.01; ***p < 0.001).
These findings indicate that while storage procedures were effective, certain metabolite concentrations varied by storage method, highlighting the need for prestorage treatments to stabilize metabolites and careful cryopreservation practices for accurate analysis of oxidative biomarkers.
Discussion
The storage and preservation of plasma and serum samples are crucial for translational and biomedical research, particularly in the context of personalized medicine and biomarker discovery.10,11 In biobanks, large quantities of blood and blood-derived samples are typically stored at low or ultralow temperatures, which slow down chemical processes and biological processes.7,9,11 Biobanks are currently facing an emerging challenge aiming at defining strategies to decrease sample composition variability between heterogeneous handling procedures, storing methods, SOP, and thus, among biobanks. 30
Plastic straws are emerging as a preferred alternative to standard tubes and vials for long-term cryopreservation due to their compact storage, lower contamination risk, and enhanced sample viability, making them a cost-effective option for biobanks.21,31
The preanalytical phase can affect the quality of blood samples. Factors such as the type and brand of blood tubes, processing time, and storage time can overall contribute to the preanalytical variability.32,33 In this light, herein, we evaluated the effect of the two methods used in UPO Biobank (vials and straws) on the biochemical and omics profiles of cryopreserved samples, as strongly recommended by the international quality standard ISO 23118:2021. In line with previous findings, 34 standard laboratory practice did not highlight significant differences, supporting the robustness of routine hematochemical analyses in clinical diagnostics and emerging research.
Lipidomics is acquiring an increasing interest in precision medicine, despite technical challenges (e.g., lipid degradation or artifacts from oxidized or hydrolyzed metabolites).35,36 Our results indicate that both cryopreservation methods similarly preserve major lipid classes in plasma and serum. However, serum stored in straws showed significantly lower levels of bound FA while preserving oxidation-sensitive lipids, including polyunsaturated FA (PUFAs) such as 11,14-eicosadienoic acid and propyl eicosapentaenoic acid, a derivative of eicosapentaenoic acid, as well as phosphatidylcholines (PCae 36:0, PCae 42:5, PC22:1). Conversely, plasma samples in straws displayed higher levels of the oxidation-sensitive omega-6 FA, linoleic acid_B, while vials contained more arachidonic acid and gamma-linolenic acid (6,9,12-octadecatrienoic acid). Notably, both plasma and serum samples in straws displayed low levels of free eicosenoic acid and bound eicosatrienoic acid, indicating that vials may be more suitable for their cryopreservation. To our knowledge, this is the first report of these findings, and limited data exist in the literature on the underlying mechanisms.
The preservation of PUFA and highly unsaturated FA (≥20 carbons, ≥3 double bonds)—which undergo degradation during storage—relies on the freezing rate and temperature storage, with fast ultra-cold storage essential for preventing degradation and maintaining stability. 37
Our targeted metabolomics revealed that cryopreservation protocols significantly affected the representation of key metabolites involved in redox processes, showing that storage conditions and matrix type influenced several metabolites. While metabolomics is promising in biomedical research, its efficacy can be compromised by preanalytical conditions that lead to variability and degradation of biological samples. 38
Proteomics is emerging as another key area of personalized medicine research and several publications examined the effect of the processing methods on the proteomic profile of both fresh and stored samples. 39 Our analysis, minimizing sample manipulation, showed similar overall proteomic profiles between storage methods, with more differentially represented proteins between SE_straw and SE_vial than between PLE_straw and PLE_vial. Sera stored in straws showed a higher level of α-, β-, and γ-fibrinogen subunits, suggesting that this storage method may be preferable for studies investigating residual serum fibrinogen. 40 Conversely, the abundance of immunoglobulin (IG) light (e.g., IGK, IGL1, IGLC3) and heavy chains (IGHG4, IGHA2) differed notably between straw and vial samples across both matrices. These findings underscore how ex vivo biological processes, such as coagulation, when combined with cryopreservation methods, can significantly affect the representativeness of serum proteins compared to plasma proteins. 41 Furthermore, factors such as pH, temperature, and freeze/thaw cycles are well-documented as influencing protein stability, especially for IGs. 42 Additionally, protein loss due to adsorption onto the surfaces of vials or straws is another critical consideration, as it can further alter the protein profile and should be evaluated to ensure accurate data analysis. 43 An illustrative case is the protein SAA4, which was found to be underrepresented in both serum and plasma stored in straws. Also, given that SAA4 is known to be sensitive to low temperatures in other biological matrices in cattle, 44 it is likely that the rapid freezing associated with straw storage may adversely impact its cryopreservation. Further analysis is needed to investigate this issue.
In conclusion, the effect of containers on the omics representation of various analytes can be attributed to several chemical and physical factors. Literature indicates that the volumes and surface areas involved in the freezing process differ between vials and straws, along with the distinct chemical compositions of the two types of containers. 45 These differences can influence the freezing rate and the uniformity of temperature distribution, potentially affecting analyte preservation. Also, surface reactivity with specific analytes may modify or reduce their concentration, further impacting the accuracy of the results. 43
While our study has some limitations, such as the small sample size (n = 8), the absence of a fresh control for comparison, and the focus on a single time point, the findings provide a valuable starting point for further research in this field. The signatures derived from our samples, although estimated from a relatively small dataset, exhibit high consistency across different sample types. This indicates that even with this limited sample size, the generated signatures are likely to be comparable with those derived from significantly larger datasets. While increasing the sample size could enhance the robustness of the signatures, this study effectively consistently highlights critical preanalytical factors. We are actively aiming to address these issues in future studies, and the publicly accessible nature of the derived signatures allows for easy updates and expansion as additional data become available. Therefore, more extensive analyses are recommended to thoroughly characterize the multi-omics profiles in plasma and serum samples preserved in vials and straws. This will help ensure the reliability of data derived from samples subjected to these different cryopreservation methods.
Footnotes
Authors’ Contributions
A.An.: Formal analysis, investigation, visualization, writing—original draft, and writing—review and editing. V.B.: Formal analysis, investigation, writing—original draft, and writing—review and editing. J.V.: Investigation and writing—review and editing. S.V.C.: Investigation. E.M.: Writing—review and editing. G.G.: Investigation. M.V.: Investigation. C.F.: Investigation. M.S.: Investigation. A.Am.: Investigation and writing—review and editing. R.R.: Investigation and writing—review and editing. D.C.: Conceptualization, funding acquisition, supervision, and writing—review and editing.
Author Disclosure Statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Funding Information
This study was (partially) funded by the Italian Ministry of University and Research (MUR) program Departments of Excellence 2023–2027, AGING Project—Department of Translational Medicine, Università del Piemonte Orientale (D.C.). This publication is part of the project AGE-IT, which has received funding from the MUR—M4C2 1.3 of PNRR with grant agreement no. PE0000015 (D.C.).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
