Abstract
Introduction:
The Minimum Information About BIobank Data Sharing (MIABIS) is a biobank-specific terminology enabling the sharing of biobank-related data for different purposes across a wide range of database implementations. After 4 years in use and with the first version of the individual-level MIABIS component Sample, Sample donor, and Event, it was necessary to revise the terminology, especially to include biobanks that work more in the data domain than with samples.
Materials & Methods:
Nine use-cases representing different types of biobanks, studies, and networks participated in the development work. They represent types of data, specific sample types, or levels of organization that were not included earlier in MIABIS. To support our revision of the Biobank entity, we conducted a survey of European biobanks to chart the services they provide. An important stakeholder group for biobanks include researchers as the main users of biobanks. To be able to render MIABIS more researcher-friendly, we collected different sample/data requests to analyze the terminology adjustment needs in detail. During the update process, the Core terminology was iteratively reviewed by a large group of experts until a consensus was reached.
Results:
With this update, MIABIS was adjusted to encompass data-driven biobanks and to include data collections, while also describing the services and capabilities biobanks offer to their users, besides the retrospective samples. The terminology was also extended to accommodate sample and data collections of nonhuman origin. Additionally, a set of organizational attributes was compiled to describe networks.
Discussion:
The usability of MIABIS Core v3 was increased by extending it to cover more topics of the biobanking domain. Additionally, the focus was on a more general terminology and harmonization of attributes with the individual-level entities Sample, Sample donor, and Event to keep the overall terminology minimal. With this work, the internal semantics of the MIABIS terminology was improved.
Introduction
The Minimum Information About BIobank Data Sharing (MIABIS) is a biobank-specific terminology enabling the sharing of minimal biobank-related data for different purposes across a wide range of database implementations. The development of MIABIS was initiated in 2012 by the Biobanking and BioMolecular Resources Research Infrastructure of Sweden (BBMRI.se). 1 Since 2017, the Common Service IT (CS IT) 2 within the Biobanking and Biomolecular Resources Research Infrastructure—European Research Infrastructure Consortium (BBMRI-ERIC) has coordinated the further development of the MIABIS terminology. Originally, the main aim of MIABIS Core was to establish a means to describe administrative information about biobanks, which at the time was not systematically represented in any terminology. At present, MIABIS Core version 2 3 is used in several biobank registers and catalogs,4,5 it is implemented as part of data models for biobank and research information management systems, 6 and it is integrated in other data models 7 and biobanking ontologies. 8 The MIABIS terminology has recently been complemented with the extension of an individual-level component to describe samples, sample donors, and related events. 9
MIABIS Core v2, together with the MIABIS individual-level reference data model for Sample, Sample donor, and Event, render the terminology heavily sample-centric. Biobanks, however, can also be built around data generated from samples and/or obtained from donors and may not include biological samples at all. Imaging biobanks7,10 are an example of such biobanks. In addition, many biobanks today offer mostly data to researchers, as their infrastructure already contains extensive omics-data derived from biosamples. 11 The biobanking community also includes banks collecting nonhuman samples relevant to public health, 12 such as microbes from the environment. Owing to the increasing demand for collaborative work and large datasets, the need for research networks combining the expertise of individual network members13–15 has arisen.
Thus, there was a need to update MIABIS Core, which we addressed with the efforts and results described here.
Materials and Methods
We identified nine different use-cases representing different types of biobanks, studies, and networks operating with big data or specific sample/data types, not previously defined in MIABIS (see Supplementary Table S1). In addition, in 2020-2021, we collected 38 sample/data requests from BBMRI-ERIC national nodes (from Austria, Belgium, Finland, Germany, The Netherlands, and Poland) to make informed decisions on how the terminology should be adapted (summarized in Supplementary Table S2).
In addition, to describe the biobanks at a more practical level, we conducted a survey in September 2021 targeting over 500 biobanks in the BBMRI-ERIC network about the services that the biobanks offer to their clients or what they have included in their infrastructure, such as availability of sample storing services and facilities, possibility to provide access to additional data, possibility to provide analytical services to biobank samples, and possibility to provide cell culture services. Seven biobanks responded to the survey, and their answers were included in the Biobank entity revision.
The names and descriptions of the entities were evaluated to ensure their compatibility with the new domain extensions. Each attribute was then thoroughly reviewed, and suggested modifications were discussed by the working group until consensus was reached. The MIABIS working group consisted of experts in biobanking, data (such as antimicrobial data resistance data, omics, epidemiological and population-based lifestyle data, clinical information, imaging data), interoperability, and IT. The assembly of experts in the MIABIS working group included the authors of the article and individuals mentioned in the acknowledgments.
Results
Moving from MIABIS Core v2 to v3
In this work, we propose an update for the MIABIS Core v2 entities Biobank, Sample collection, and Study, as well as an update of the structure of the MIABIS Core by including the new entity Network, renaming the former entity Study to Research resource, and defining it as an optional entity in the Core. We will first show our main aims and describe the important aspects of the update process before presenting the new MIABIS Core version 3.0.
Aims and conduction of the MIABIS update
We pursued the following aims with the MIABIS update:
Incorporate the concept of data-driven biobanks and data collections to support biobanks providing mostly data: As accompanying data and sample-derived data are becoming ever more relevant in biobanking, the descriptions of Collection and Research resource entities were rephrased to widen their applicability to data derived from samples or donors. We also revised attributes, e.g., the “Data categories” attribute to better reflect data-driven biobanks. Synchronize MIABIS Core attributes and values with the MIABIS individual-level component Sample, Sample donor, and Event: To achieve synchronization with the individual-level entities Sample, Sample donor, and Event, it was necessary to update some MIABIS Core attributes, e.g., “Storage temperature” and “Material type,” which were subsequently renamed as “Sample type.” Allow the description of nonhuman sample and data collections related to biomedical research: To be able to include samples of nonhuman origin, new sample types were added to the “Sample type” attribute value list, such as “Specimen from environment or food,” and in general, attribute descriptions were changed to be more generic and include nonhuman samples. Restructure of the MIABIS Core: We included an entity on biomedical networks, which has become more and more important for biobanks and in biomedical research. Additionally, we renamed the entity Study to Research resource and made this entity an optional one.
Thus, the MIABIS Core 3.0 consists of the following three entities: Network, Biobank, and Collection. Together, they form a more natural combination of biobank data sharing, and the information provided by these entities needs to be already used in several catalog solutions, such as the BBMRI-ERIC Directory. 4 We made the Research resource optional, as this entity is not relevant in current biobank catalog solutions but may receive more attention in the future to reflect active and successfully conducted projects of biobanks and network infrastructures. The updated MIABIS structure is shown in Figure 1, which also includes perspectives of an additional component on Dataset entities.
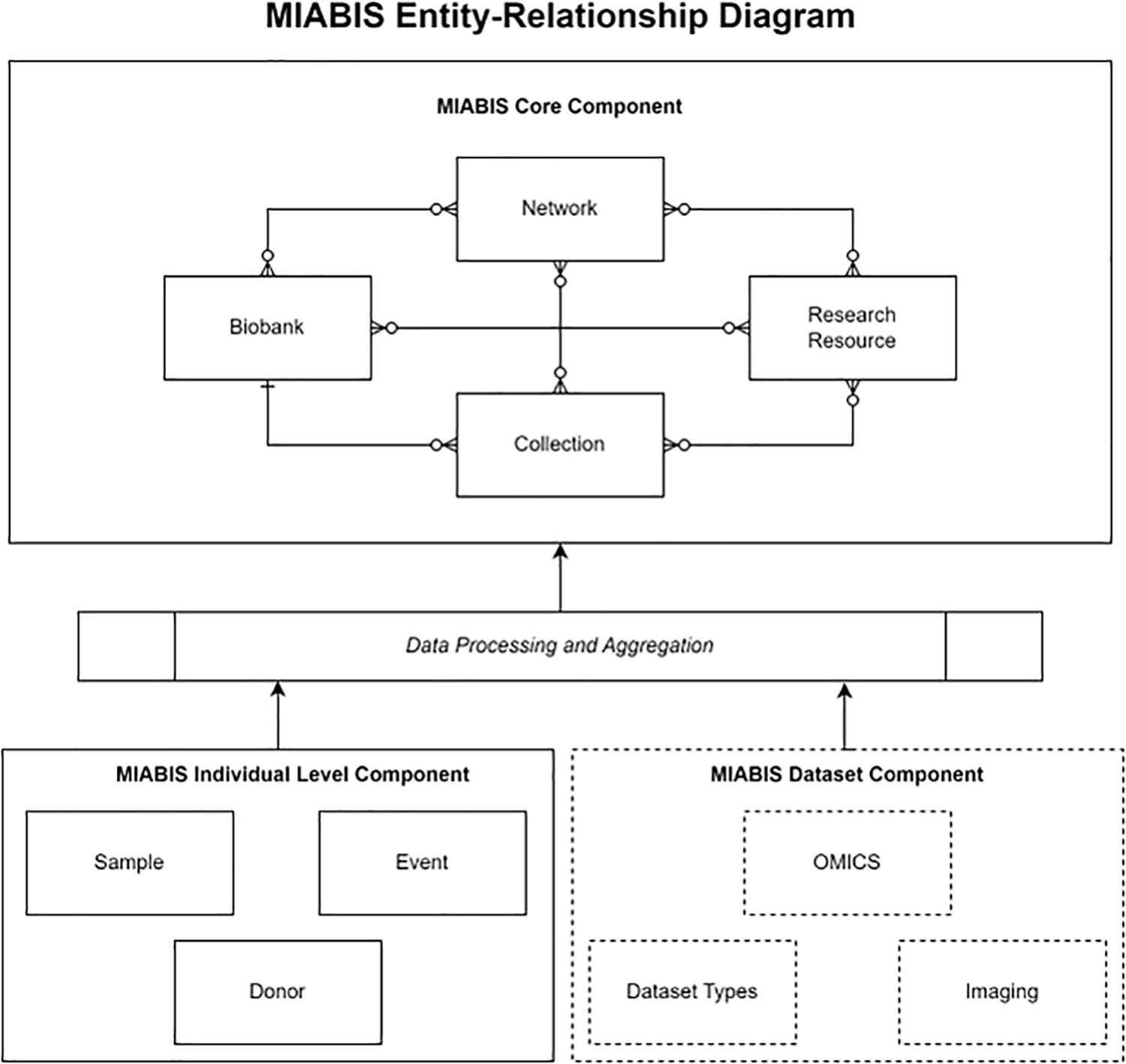
The MIABIS terminology structure consists of aggregate-level and individual-level components. Together the Biobank, Collection, and Network form the Core, with the optional Research resource being maintained along the Core component. The individual-level component Sample, Sample donor, and Event are a basis to provide information from which some data elements of the Collection can be aggregated. Here, we already anticipate the next component on datasets, omics, and imaging, which will be on both aggregate and individual levels. This diagram uses crow’s foot notation to depict the relationship between the Core entities. MIABIS, Minimum Information About BIobank Data Sharing.
Define a set of attributes for services and capabilities the biobanks offer to their customers apart from providing retrospective samples.
Adjust the MIABIS terminology to provide better search capabilities for available data and samples for the main stakeholder group of biobanks: researchers.
From the 38 researchers’ queries, we were able to identify similar features and group them by their type (Supplementary Table S2). We ensured that the researchers’ needs are included in the Collection entity update, especially in the “Dataset type” attribute. We included most of the needs, but some requirements were too specific and would be better represented in their own extension, such as clinical data.
Review entity names and descriptions and MIABIS metadata for overall consistency.
One goal of this revision was to achieve a level of generalization, to make the entities and attributes as reusable and self-explanatory as possible. Therefore, entity names and descriptions were revised and updated as shown in the next chapter. We aimed to keep the entity descriptions short and informative but still generic to avoid the need for constant adaptation. This way, we will ensure the broad usability of the terminology in different settings.
We reviewed the descriptive structure and metadata of the MIABIS terminology. All the attribute descriptions were synchronized across the component, and the original description field from MIABIS v2 was split into two columns to accommodate descriptions and constraints. In addition, all the attribute codes were updated throughout the terminology so that each attribute has a unique and informative entity-specific code using the coding scheme described earlier. 9 As part of the metadata revision, the data types for allowed values were also revised to be more accurate by changing the “Text list” data type to “Enumerated values.”
MIABIS Core 3.0 entities
The entities included in the MIABIS Core v3 are defined as follows:
In addition, we adapted the definition of a sample to include samples of nonhuman origin.
We aimed to keep the entity descriptions short and informative, but still generic to avoid the need for constant adaptation. This way we will ensure the broad usability of the terminology in different settings.
Attribute lists for Core entities
When revising the attributes, we carefully considered backwards compatibility with previous MIABIS versions. However, some carefully considered breaks were accepted to ensure a more accurate representation of attributes.
Biobank entity
The existing attributes in the given Biobank entity, presented in Table 1, were found to be mostly adequate to describe organizational and administrative information about biobanks. Only “Contact information” was updated by merging it with the structured attribute “Principal investigator” from MIABIS Core v2 Study (structured attributes are presented in Supplementary Table S3A), as both attributes were very similar. In addition, based on the survey of biobanks, we added three new attributes for biobank capabilities and one for quality. The “Quality Management standard” attribute is the first quality attribute included in the MIABIS dataset and gives the opportunity to describe the quality accreditation or certification status of biobanks. The proposed list of values is minimal; however, we recommend that it be expanded to include local requirements when implemented.
Attribute Definitions in the Biobank Entity
The table shows the entity-specific attribute code, terminology mapping to the previous Core version, attribute name, value types, attribute descriptions and value lists, any constraints on implementation in databases, and cardinalities on implementation.
Collection and Research resource entities
The Collection and Research resource attribute lists (presented in Tables 2 and 3, respectively) needed more extensive revision. The attribute lists of the two entities Collection and Research resource were merged, because they contained many similar attributes.
Attribute Definitions in the Collection Entity
The table shows the entity-specific attribute code, terminology mapping to the previous Core version, attribute name, value types, attribute descriptions and value lists, any constraints it may have when implemented in databases, and cardinalities for implementation.
Attribute Definitions in the Research Resource Entity
The table shows the entity-specific attribute code, terminology mapping to the previous Core version, attribute name, value types, attribute descriptions and value lists, any constraints it may have when implemented in databases, and cardinalities for implementation
Regarding synchronization between attributes in the Core component and in the individual-level component, the “Detailed sample type” and “Sample storage temperature” attributes were the references for value list updates to the Collection’s “Storage temperature” and “Material type,” subsequently renamed as “Sample type.” In addition, when expanding the scope of MIABIS to include samples of nonhuman origin, we added new sample types to the list, such as “Specimen from environment or food,” and made the attribute description more generic. Because the list of sample types in the Sample entity’s “Detailed sample type” attribute is extensive, we aggregated the list in the mapping table (Table 4). For the “Storage temperature” attribute, we synchronized the value list with the same attribute in the Sample entity, where the “Sample storage temperature” values are based on Standard PREanalytical Code (SPREC) v3. 18 The value lists for the “Age low unit” and “Age high unit” attributes were also synchronized with the Sample entity.
Mapping Table of the Sample Entity’s “Detailed Sample Type” Values to More Aggregated “Sample Type” Attribute Values
All the new Aggregated values compared with MIABIS Core v2 are shown in bold.
The Collection entity has been extended from its previous version to include new attributes for “Dataset type,” “Use and access conditions,” “Publications,” “Sample source,” “Collection setting,” and “Collection status.” We replaced the original “Data categories” attribute—a mix of data types and sources—with the “Dataset type” attribute. “Dataset type” is a more refined attribute that is used to categorize available data. Its value list combines the MIABIS Core v2 “Data categories” value list, its revision in the Sample donor entity, and the requirements identified from use-cases collected sample/data requests. In addition, another attribute from the Sample entity, “Sample use restrictions,” is introduced in the Collection entity. To make it more generic, we renamed the attribute to “Use & Access conditions” and revised the value list. We recognize that it may not always be possible to specify usage restrictions at the collection level. However, if these access conditions apply to the whole collection, this information is very valuable to researchers.
To support the representation of nonhuman samples, we introduced a new “Sample source” attribute to describe the source from which the sample was collected or isolated, e.g., human, animal, or environment. In addition, the value “Specimen from environment or food” was added to the “Sample type” attribute. The possibility to include nonhuman sample collections is making certain attributes optional when implemented (e.g., age) and new null values to attributes, e.g., “Not applicable” value was added to the “Sex” attribute. However, using these new null values results in a loss of information because the reason why the value is null is not known; therefore, the textual collection descriptions in the “Description” attribute become more important. Finally, we renamed the “Total number of participants” attribute to “Total number of subjects” to also allow the description of nonhuman sample collections, and we rephrased the definition of “Inclusion criteria” for the same reason.
To make the entities more generic and to support a variety of different types of sample/data collections, we renamed the “Collection type” attribute to “Collection design” and added a complementary “Sample collection setting” attribute to describe the primary setting for the sample/data collection. The value lists for both attributes were revised and new values added. New attributes include “Publications,” as citations in publications are considered a way to convey confidence in the sample provider’s credentials and track record. 19 In addition, we added “Collection status” to describe the activity and current state of the collection; this in particular can be important when searching for active collaborators for specific topics. The Collection entity also uses the structured attribute “Disease”, now allowing for disease categories in addition to specific disease codes (see Supplementary Table S3B). Furthermore, we omitted the attribute “Total number of samples” from the merged Collection and Research resource attribute list, as it required more active updating by data providers than the other attributes.
Subsequently, after updating the Collection and Research resource attributes, we acknowledged the need to adjust the Sample and Sample donor attributes accordingly.
Network entity
Many of the use cases that influenced the MIABIS Core v3 update were research resources operating as networks rather than being stand-alone biobanks or sample/data collections (see Supplementary Table S1). This highlighted the importance of introducing a new dimension into the administrative information presented in MIABIS, and thus, the new entity, Network, was added into the Core component. We anticipate that a Network can contain not only biobanks but also Collections or other Networks directly. Thus, Networks can also form recursive structures. An example of such is the Belgian BBMRI network BBMRI.be, where the biobanks contain multiple collections, and which then can form networks. 20 Examples of Networks within Networks are BBMRI-ERIC Network consisting of National Node Networks, 21 and RD Connect, a global network for rare diseases, which includes an Italian Telethon Network of Genetic Biobanks in addition to biobanks hosting rare-disease collections.22,23
The Network attributes are based upon the attributes describing Biobanks, as outlined in Table 5. To describe Networks more specifically, network collaboration attributes were added to describe the members of a Network and the type of collaboration taking place within the Network. In addition, the “Network status” attribute allows distinguishing between active and dormant networks, which can be important when trying to find collaborators in specific domains.
Attribute Definitions in the Network Entity
The table shows the entity-specific attribute code, terminology mapping to the previous Core version, attribute name, value types, attribute descriptions and value lists, any constraints it may have when implemented in databases, and cardinalities for implementation.
Discussion
In the current work, we have succeeded in synchronizing the existing Core attributes with the MIABIS individual-level component Sample, Sample donor, and Event to make the aggregate-level entities compatible with the individual-level ones. We also adapted and extended the existing Core component to meet the requirements of typical use-cases not only for human samples but also for animal or environmental samples as far as biobanking is concerned. At the same time, we added the new Core entity Network to cover an additional organizational level where biobanks often operate.
One of the main goals of this update was to adjust the terminology to better account for data-driven biobanks and digital samples. The working group discussions in this area were complex, but in this revision, we made a first effort to include digital samples and to better represent the different types of data stored in biobanks. It is expected that the dataset types will be expanded over time as biobank samples are used in research resources and projects and transformed into data with better-defined structures. Therefore, further work is needed to describe specific datasets in more detail, e.g., for omics and imaging. We will follow international activities in these areas, e.g., the European Genome-Phenome Archive (EGA), 24 to adhere to their respective data models. In addition, a generic component for datasets to reflect the numerous possibilities of sample use for biomedical data generation will be among the next efforts of the MIABIS working group. As with the current update of the MIABIS Core to version 3, based on the MIABIS individual-level component for Sample, Sample donor, and Event, a new Core update to version 4 may be required to accommodate the work on the dataset component. Focusing our future efforts on addressing the need to describe datasets in more detail will also further facilitate the move toward individual-level data integration in various database implementations such as federated search platforms.
With this work, we updated the semantic interoperability of the MIABIS terminology and improved its actual use and suitability, despite implementation challenges and barriers.
Footnotes
Acknowledgments
The authors thank the members of the MIABIS Core v3 Working Group Andrea Gutierrez, Andres Metspalu, Blazej Marciniak, Christoph Doellinger, Eline Vinke, Enrico Glerean, Heiko Ritzmann, Jens Haberman, Joanna Vella, Jörg Geiger, Kristjan Metsalu, Lisa Scheefhals, Luciano Milanesi, Marije van der Geest, Mario Schattschneider, Mia Liljeström, Michael Hummel, Morris Swertz, Noemi Deppenwiese, Riad Gacem, Ruben Haeck, Sara Casati, Tomas Snäckerström, and Zdenka Dudová for their contributions and support of this work. The authors are also grateful to all the reviewers who provided their valuable expert opinions at various stages of the work
Authorship Contribution Statement
N.E.: Conceptualization, Resources, Investigation, Data Curation, Writing—Original Draft. C.E.: Resources, Investigation, Data Curation, Writing—Review & Editing. M.N.: Investigation, Validation, Writing—Review & Editing. A.S.: Investigation, Validation, Visualization, Writing—Review & Editing. E.V.E.: Resources. R.B.: Resources, Writing—Review & Editing. M.B.: Resources. A.D.: Resources, Writing—Review & Editing. A.V.D.L.: Resources. H.M.: Resources. L.P.: Resources. P.R.Q.: Investigation, Resources. E.U.: Investigation, Resources. P.H.: Resources, Funding Acquisition, Writing—Review & Editing, Supervision. K.S.: Conceptualization, Resources, Writing—Review & Editing, Supervision. G.A.: Writing—Original Draft, Supervision.
Author Disclosure Statement
The authors declare no conflicts of interest.
Funding Statement
This work was supported by BBMRI-ERIC Common Service IT, EU Horizon 2020 supported projects EOSC-Life (grant agreement no. 824087), European Joint Programming for Rare Diseases (EJP RD) (grant agreement no. 825575), and A European-wide foundation to accelerate Data-driven Cancer Research (EOSC4Cancer) (grant agreement no. 101058427).
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
