Abstract
The study of extremophilic microorganisms has sparked interest in understanding extraterrestrial microbial life. Such organisms are fundamental for investigating life forms on Saturn’s icy moons, such as Enceladus, which is characterized by potentially habitable saline and alkaline niches. Our study focused on the salt-alkaline soil of the Al Wahbah crater in Saudi Arabia, where we identified microorganisms that could be used as biological models to understand potential life on Enceladus. The search involved isolating 48 bacterial strains, sequencing the genomes of two thermo-haloalkaliphilic strains, and characterizing them for astrobiological application. A deeper understanding of the genetic composition and functional capabilities of the two novel strains of Halalkalibacterium halodurans provided valuable insights into their survival strategies and the presence of coding genes and pathways related to adaptations to environmental stressors. We also used mass spectrometry with a molecular network approach, highlighting various classes of molecules, such as phospholipids and nonproteinogenic amino acids, as potential biosignatures. These are essential features for understanding life’s adaptability under extreme conditions and could be used as targets for biosignatures in upcoming missions exploring Enceladus’ orbit. Furthermore, our study reinforces the need to look at new extreme environments on Earth that might contribute to the astrobiology field.
Introduction
Astrobiology is an emerging interdisciplinary field that investigates the origin, evolution, and distribution of life in the universe, on Earth and beyond, and includes the possibility that life exists in other planetary bodies in our solar system (https://www.astrobiologyprimer.org; Schultz et al., 2023a). The discovery of extraterrestrial environments that are analogous to some extreme environments on Earth and the study of the organisms that inhabit them have produced results that further support the plausibility of life beyond Earth (Wagner et al., 2022). Extremophilic microorganisms isolated from these terrestrial environments have become the main models for possible extraterrestrial life in harsh environments (Lopez et al., 2019; Merino et al., 2019; Schultz et al., 2023a).
Haloalkaliphilic organisms are particularly attractive models in astrobiology because they are evolutionarily ancient, physiologically versatile, and capable of surviving in diverse extreme planetary environments. Of these organisms, polyextremophilic characteristics, such as tolerance to high salinity, anaerobic conditions, high levels of ionizing ultraviolet radiation, and temperatures ranging from subzero to >120°C, make these organisms an ideal model for studying the possibility of life on Enceladus, the icy moon of Saturn (Wagner et al., 2022). Observations of the Cassini–Huygens and Galileo spacecraft have revealed that Enceladus has a significant ocean of liquid water beneath its icy crust (Kivelson et al., 2000; Waite et al., 2009). The use of many scientific instruments coupled with the Cassini spacecraft has shown that the ocean of Enceladus is highly alkaline, has a pH around 10–12, and is rich in phosphate salts. Furthermore, Enceladus has been shown to be 100 times richer in phosphate than Earth’s oceans (Glein et al., 2020; Postberg et al., 2023). In addition, it is hypothesized that the bottom of this ocean is potentially dominated by hydrothermal sources and governed by volcanic activity (Deamer and Damer, 2017). Recent studies have shown that the ocean of Enceladus contains macromolecules of up to 200 Da, in addition to small molecules (Mora et al., 2022; Postberg et al., 2018). These observations have sparked discussions within the scientific community about the possibility of these molecules being associated with microbial activity. Consequently, several space agencies are now focused on finding life in the ocean of Enceladus (Brockwell et al., 2016; Deamer and Damer, 2017; Neveu et al., 2020). To better understand icy moons, astrobiologists and planetary scientists are highly interested in finding remote locations on Earth with physicochemical conditions similar to those present in Enceladus’ subglacial ocean and aim to conduct fieldwork to exploit their microbiomes in search for novel biological models (Kanik and Paul de Vera, 2021; Steinle et al., 2018).
Saudi Arabia has many volcanoes and extensive basaltic lava fields, known as harrats, that are located along the western part of the Arabian Shield subparallel to the Red Sea (Wahab et al., 2014). Notably, the Al Wahbah crater within the Harrat Kishb volcanic area emerges as a significant site for extraterrestrial analog studies and a potential new source of astrobiological resources such as extremophiles. Despite its Earth-bound location, the Al Wahbah crater is an intriguing analog for studying microbial life akin to that possibly present on Enceladus. The similarities between Enceladus’s physical and chemical environments—particularly its ocean and hydrothermal vents—and the phosphate-rich, highly alkaline soil of the Al Wahbah crater are striking. The crater comprises soil with a high concentration of sodium phosphates and chlorates, being highly alkaline. These chemical characteristics arise due to leaching processes and the formation of a temporary lake in the middle of the crater during rainy periods. This seasonal lake in Al Wahbah exhibits pH and salinity levels that parallel those of Enceladus’s ocean. Soil analysis at the crater’s center reveals high phosphate and chloride concentrations, with a pH between 9 and 10 (Wahab et al., 2014). Similarly, Enceladus’s ocean is considered alkaline, with a pH between 9 and 12, and replete with various ions, which suggests that potential microbial inhabitants would need to be halophilic and alkali-tolerant. The moon’s potential for hosting microbial life is especially high near its hydrothermal vents, where volcanic activities could foster favorable life-sustaining conditions (Reh et al., 2016). The volcanic nature of Al Wahbah, characterized by basaltic minerals, phosphate crystals, and olivines, coupled with the desert’s elevated temperatures, renders it an exemplary terrestrial counterpart for Enceladus’s potentially habitable environments. Furthermore, a molecular study revealed a diverse microbial community capable of thriving in these saline and alkaline conditions (Albokari et al., 2018). Given the physicochemical similarities between Enceladus’s ocean and the alkaline clay in Al Wahbah, this crater represents a prime location for exploring thermo-haloalkaliphilic (heat, salt, and alkali-loving) microorganisms that could serve as biological proxies for life on Enceladus.
Although the microbial metabolism of potential extraterrestrial living organisms may differ from that of terrestrial microorganisms, the search for molecular biosignatures in other worlds could be initiated by characterizing micromolecules and their respective metabolic pathways in organisms that inhabit extreme conditions on Earth. With the current advancement in instrumental techniques, particularly mass spectrometry (MS), and the popularity of platforms for annotating organic molecules, such as Global Natural Products Social (GNPS), it has become easier to propose classes or families of compounds produced by microorganisms cultivated under various conditions (Wang et al. 2015; Seyler et al., 2020).
Currently, we are aware that, in the coming years, new mass spectrometer generations will be sent to the gas giants of our solar system with the intent to investigate the possibility of life on their icy moons (Arevalo et al., 2020). One of these instruments, the Mass Spectrometer for Planetary Exploration/Europa (MASPEX), is scheduled to be sent in 2024 on the Europa Clipper mission. MASPEX is a time-of-flight mass spectrometer known for its high resolution and sensitivity, which allows for the detection of traces (ppb) of organic compounds. Selected by NASA for the Europa Clipper mission, MASPEX will analyze samples of the sprayed atmosphere on the moon’s surface to evaluate any evidence of habitability and life (Brockwell et al., 2016; Sephton et al., 2018). This same technology and approach are being considered for use in missions investigating organic molecules in the oceans of Enceladus in the coming decades (Reh et al., 2016). In addition, other mass spectrometers are being developed exclusively for use in astrobiology research, such as the CosmoOrbitrap, which features an Orbitrap analyzer, the same analyzer used in the present study. This equipment will have the capability to produce results similar to the equipment used in Earth’s laboratories (Arevalo et al., 2018; Arevalo et al., 2023).
Given the imminent deployment of mass spectrometers to Enceladus, understanding the molecules potentially present on the moon through laboratory experiments that integrate extremophilic microorganisms with the physicochemical conditions of these extraterrestrial oceans can provide valuable insights into the possible molecular signatures that could be discovered. This approach is crucial for advancing our understanding of life’s limits and detecting environments as extreme as those found on Enceladus.
Motivated by the extreme characteristics of the Al Wahbah crater in Saudi Arabia and their resemblance to those of Enceladus, along with the growing emphasis on genomic characterization in bacterial taxonomy and functionalities for astrobiological purposes, we aimed to isolate and characterize extremophilic bacteria from the crater’s samples. Our isolates spanned 11 distinct bacterial genera and belonged to three phyla as follows: Actinomycetota, Bacillota, and Pseudomonadota. Moreover, the genomes of two thermo-haloalkaliphilic strains were sequenced and compared with those of closely related species available in public databases. Herein, we present an in-depth analysis of genomic characterization, metabolic potential, and chemical profiles of the bacterial strains cultured from the Al Wahbah crater and focus on the search for novel biosignatures that could be useful in the search for life on Enceladus. For this study, we hypothesized that microorganisms isolated from the Al Wahbah crater could provide insights into the existence of life on Enceladus. The unique features of the thermo-haloalkaliphilic strains indicate their potential compatibility with the extreme conditions of Enceladus’ subglacial ocean. Genomic and chemical analyses of these microorganisms can broaden our understanding of extremophilic life on Earth, serve as a biological model for hypothesizing about the nature of extraterrestrial life forms, and lay the groundwork for future astrobiological explorations to identify molecular biosignatures on celestial bodies and potentially enhance our understanding of the types of potential life forms that could one day be discovered in Enceladus’ ocean.
Materials and Methods
Study site and sampling strategy
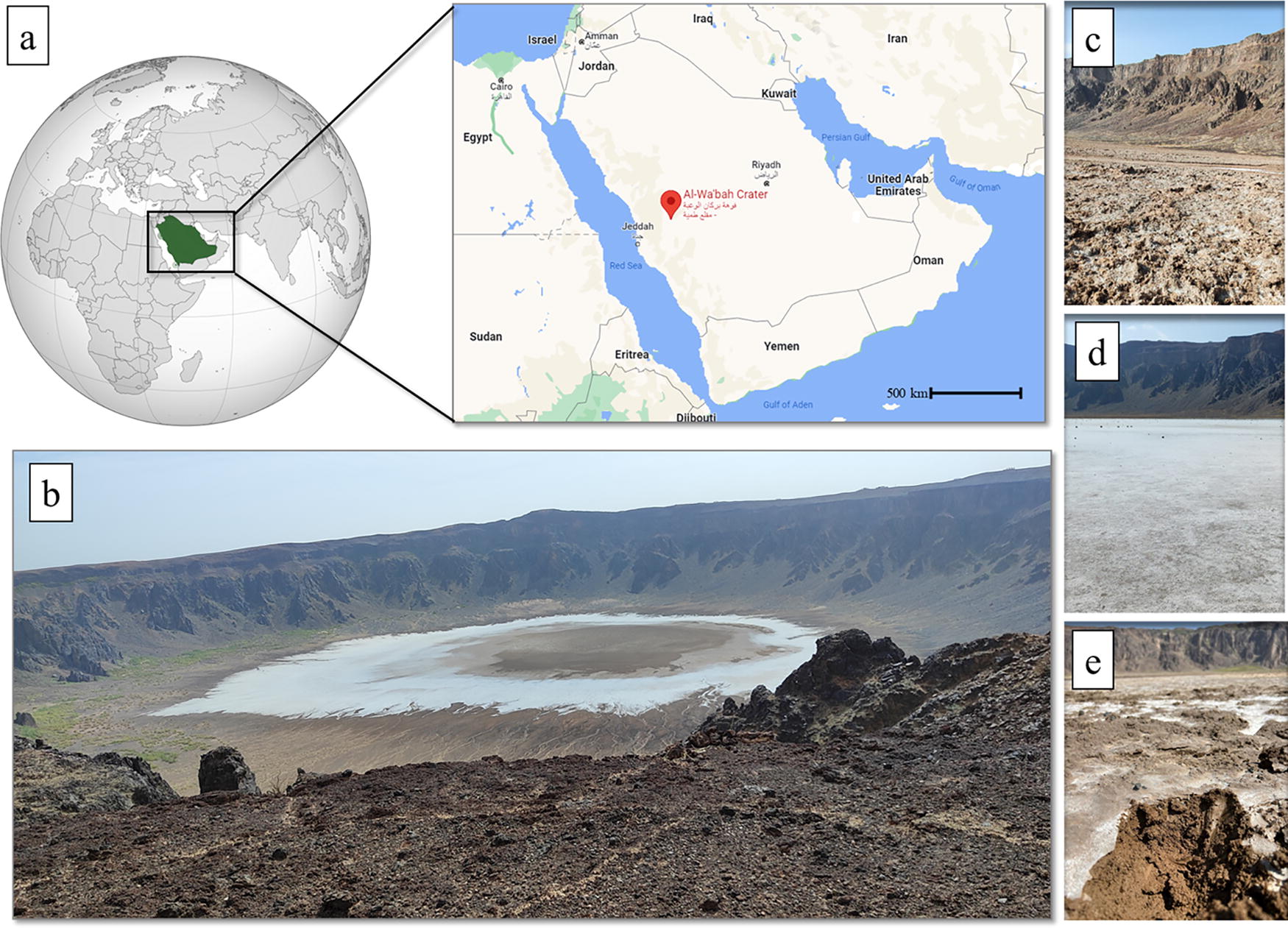
Sampling was performed in February 2023 at different locations in the Al Wahbah crater (lat. 22.90632, lon. 41.13849), which is located on the Harrat Kishb basalt plateau in the Hejazi region of Saudi Arabia. A total of nine samples were collected from the following three sites: (1) a site with a high concentration of sodium phosphate salt, (2) a dry portion at the edge of the site with salt, and (3) at clay-rich area in the middle of the crater (Fig. 1). Approximately 500 g of soil was collected from each sampling point (in triplicate) at 0–10 cm depth, placed into sterile plastic bags, and stored at 4°C until processing in the laboratory.
Physicochemical characterization
To characterize the collected soil samples, we evaluated the physicochemical parameters of the soils, which included pH, electrical conductivity, and elemental analysis. The analyses were performed in the Analytical Core Laboratory of the
Isolation and growth conditions
To retrieve the culturable fraction of bacteria from the soils of the Al Wahbah Crater, a soil-sprinkling strategy in agar Petri dishes was applied. Four different culture media were used as follows: Horikoshi alkaline medium (final pH, 10.0), modified marine medium (final pH, 10.3), halophilic medium (final pH, 9.0), and Bushnell–Haas medium (final pH, 7.4) (the media compositions are presented in Supplementary Table S1). For the sprinkling methodology, soil samples were homogenized, and 100 mg of each soil sample was sprinkled onto four culture media in triplicate and incubated at 35°C and 55°C as growth temperatures for up to 72 h. After incubation, morphologically distinct colonies were isolated by streaking on their respective media.
Pure bacterial colonies from both strategies were transferred to their respective liquid medium (15 mL) and incubated at the isolation temperature with shaking (150 rpm) for up to 48 h. An aliquot of the microbial culture was placed in a cryogenic vial that contained glycerol (20% v/v) in triplicate and stored at −80°C. All isolated microbial strains from Al Wahbah crater were deposited in the Saudi Extreme Culture Collection of the Laboratory of Microbial Ecogenomics and Biotechnology, KAUST. A summary of the experimental design is shown in Supplementary Figure S1.
DNA extraction, 16S rRNA gene sequencing, and phylogenetic analysis
A Wizard® Genomic DNA Purification Kit (Promega, USA) was used to extract total genomic DNA from the bacterial strains according to the manufacturer’s instructions. Isolates were identified based on Sanger sequencing of the 16S rRNA gene for molecular identification. A Master Mix AmpliTaq 360 (Promega, USA) was used to amplify an approximately 1450-bp fragment of the 16S rRNA gene with universal primers 27F (5′AGA-GTT-TGA-TCM-TGG-CTC-AG-3′) and 1492R (5′-TAC-GGY-TAC-CTT-GTT-ACG-ACT-T-3′) (Weisburg et al., 1991) using polymerase chain reaction (PCR), according to the manufacturer’s instructions. The amplification was performed in a thermocycler as follows: an initial denaturation step was performed at 95°C for 10 min followed by four consecutive cycles of 98°C for 30 s, 98°C for 1 min, and 72°C for 1 min. This step was followed by 30 cycles of 54°C for 30 s, 72°C for 1 min, and a final extension step at 72°C for 15 min. PCR products were quantified using Qubit 4.0 with the Qubit® dsDNA HS Assay Kit (Life Technologies, USA); stained with SYBR Safe stain (Invitrogen, USA) in 1% agarose for examination; run at 90 V and 500 A for 45 min; and visualized using Gel Doc™ EZ System. The PCR products were then sent to Macrogen, Inc., Seoul, South Korea, for Sanger sequencing of the 16S rRNA gene. Subsequently, BioEdit (v.7.2.5) was used to check and trim the raw sequences and align the sequences from both primers. BLAST was then used to compare the resulting sequences with homologous sequences in the National Center for Biotechnology Information (NCBI)-GenBank database. Aiming to check the evolutionary distance between our isolates, the neighbor-joining statistical method with 1,000 bootstrap replications was used to construct the phylogenetic tree in IQTREE (v.2.2.0.3; Minh et al., 2020), and the interactive Tree Of Life (iTOL; v.6.7) was used for annotation and editing (Letunic and Bork, 2021).
Whole-genome sequencing, assembly, and characterization
To better understand the taxonomy and functionalities of polyextremophilic bacteria for astrobiological purposes, two bacterial strains capable of growing under thermo-haloalkaliphilic conditions were selected for whole-genome sequencing and in-depth characterization of their genomes. Hence, an amount of 5 μg/μL of gDNA was used to construct paired-end sequencing libraries (2 × 150 bp) of 350-bp inserts following the manufacturer’s protocol for the Illumina TruSeq Nano DNA Library Preparation Kit. Quality analysis of the final libraries was performed on an Agilent 2100 bioanalyzer (Agilent Technologies). The bacterial sample was sequenced on the Illumina NovaSeq 6000 platform, as recommended by the manufacturer, with a minimum data output of 10 Gb per sample.
For genome assembly, FastQC (Andrews, 2010) was used to check the quality of the raw reads, and SPAdes (v.3.13.0) (Bankevich et al., 2018) and Unicycler (v.0.4.8) (Wick et al., 2017) were used to perform de novo assembly. Metrics to evaluate the quality and choose the best assembly were generated by QUAST (Gurevich et al., 2013). The Bacillus halodurans strain C-125 genome sequence from NCBI (accession BA000004.3) was used as the reference to generate a scaffold in CONTIGuator (Galardini et al., 2011). MOB-suite (v3.0.3) (Robertson and Nash, 2018) was used to screen plasmid sequences, and BLASTn was used to check plasmid contigs against the nonredundant database of NCBI. GFinisher (v.1.4) (Guizelini et al., 2016) and GapBlaster (v.1.1.2) (De Sá et al., 2012) were used to fill gaps within genome and plasmid sequences. CheckM (Parks et al., 2015) was also used to check each assembly for completeness and contamination. The default parameters were used in all of the software.
To perform nucleotide-level comparisons of these two genomes within their respective genera, we used the NCBI command line tool datasets (v.15.23.0) and obtained all validly described representative genomes of this genus (https://github.com/ncbi/datasets). We then used JSpeciesWS to compute the pairwise Average Nucleotide Identity (ANI) computations with the novel strains as a query with representative genomes (Richter et al., 2016). Furthermore, to estimate digital DNA–DNA hybridization (dDDH), we used the Genome-to-Genome Distance Calculator (GGDC; v3.0) online tool with the recommended Formula 2 and the BLAST+ alignment tool (Meier-Kolthoff et al., 2013).
Whole-genome sequencing (WGS)-based phylogeny
To validate the taxonomy and phylogeny placement of the two polyextremophilic strains within their respective genera, we generated two phylogenetic trees based on the pangenome analysis that capture the complete genetic diversity within species. Hence, first, we used the OrthoFinder program that accepts .faa amino acid sequence fasta files for each genome to perform all-vs-all BLASTp to identify orthologous genes. OrthoFinder uses a Markov clustering algorithm to group proteins to determine orthologous genes (Emms and Kelly, 2015). We used 29 species (27 Bacillus species and 2 newly sequenced genomes, 3Ho3b and 4Ho3b) for the pangenome, core genome, and singletons calculation (Supplementary Table S2). The newly sequenced genome strain 3Ho3b was used as the reference genome for the analysis with the default OrthoFinder parameter. Furthermore, we used our own in-house Perl script for the pangenome, core genome, and singleton set calculations. The extrapolations of the pangenes were calculated according to Heap’s Law for both the open and closed pangenome (Guimarães et al., 2015; Jaiswal et al., 2020; Soares et al., 2013) and the least-squares fit of the exponential regression decay for the core and singletons calculations. For the pangenome, core genome, and singleton, the methodology of Jaiswal et al. (2020) was used. Subsequently, the phylogenetic trees were constructed at the genus level (27 Bacillus species and 2 newly sequenced genomes 3Ho3b and 4Ho3b) and also based on the core genome proteins (shared by all of the species) obtained from the pangenome analysis. The species Streptococcus agalactiae ATCC13813 and S. gordonii ATCC10558 were used as outgroups for tree construction. All species and their protein sequences used for the analysis were downloaded from the NCBI genome database. Using GToTree (v.1.8.2), which is a command line tool that uses hidden Markov models, we aligned and identified single-copy genes from the entire genome and generated concatenated protein alignments (Lee, 2019). Because the strains were affiliated with Bacillota, 119 single-copy genes associated with Firmicutes were applied to position both strains. Next, we used IQTREE v.2.2.0.3 (Minh et al., 2020) with ModelFinder-Plus to construct the phylogenetic tree from the protein alignment generated by GToTree with 1,000 ultrafast bootstrap replicates. Then, iTOL v.6.7 (Letunic and Bork, 2021) was used to visualize the WGS-based trees.
Genome annotation and in silico genome mining to search for potential biosignatures
Prokka (v.1.11) (Seemann, 2014) was used to perform annotation, and Subsystems Technology (RAST-tk) (Aziz et al., 2008) and rapid annotation of microbial genomes were used to perform functional annotations. The results were used to infer functions at the subsystem level. In addition, the NCBI clusters of orthologous genes (COG), PFAM, and KEGG databases in Anvi’o (v.7.1; Eren et al., 2015) were used to analyze the predicted contigs. Manual filtration for data mining was performed in the functional tables generated by NCBI COG. To predict biosynthetic gene clusters, the web-based platforms antiSMASH 6.0 (Blin et al., 2021), BAGEL4 (van Heel et al., 2018), and PRISM 4.0 (Skinnider et al., 2015) were used. To identity prophage sequences, Phage Search Tool Enhanced Release (PHASTER) was used (Arndt et al., 2016).
Cultivation and extraction of metabolites from polyextremophilic bacteria
Initially, a sterile loop was used to plate the two bacterial strains isolated at a temperature of 55°C onto R2A culture medium plates in quintuplicate and supplemented with 15 g/L of sodium bicarbonate and 15 g/L of sodium chloride to simulate the chemical conditions of Enceladus’ ocean. The bacteria were incubated at 55°C for 7 days, and after incubation, the bacteria were completely scraped from the culture medium. In addition, five 10-mm agar plugs were placed in a solution of ethanol and methanol acetate (1:1 v/v). The solution was then placed in an ultrasonic bath for 45 min. The obtained extract was dried in a rotary evaporator in vacuum. Subsequently, the dried samples were suspended in high-pressure liquid chromatography (HPLC)-grade methanol, centrifuged, and transferred to vial flasks for ultra-HPLC-Q-Orbitrap high-resolution MS.
Mass spectrometry analysis and data processing
An untargeted screening study of the crude extracts of thermo-haloalkaliphilic bacteria was conducted that used ultra-HPLC coupled to an Orbitrap ID-X MS (UHPLC-Orbitrap ID-X MS). Electrospray ionization in positive (ESI+) and negative (ESI−) modes was applied to the studied compounds, and the following parameters were used: vaporized temperature = 100°C, voltage = 3500V, sheath gas = 30L/min, auxiliary gas = 15L/min, ion-source fragmentation = 35V, and capillary temperature = 300°C. The equipment was set to perform automated operations. A 10-µL aliquot of each sample was injected through a loop injector into a C18 column (Acquity CSH 100 × 2.1 mm, 1.7 µm). The 5-µL samples were pumped through the UHPLC system along with the mobile phase and separated on a C18 column. The flow rate was set to 0.5 mL/min, and a 15-min gradient was used to achieve the required separation.
A molecular network was created using the online workflow (https://ccms-ucsd.github.io/GNPSDocumentation/) on the GNPS website (http://gnps.ucsd.edu). The data were filtered by removing all MS/MS fragment ions within ±17 Da of the precursor m/z. MS/MS spectra were window-filtered by choosing only the top six fragment ions in the ±50-Da window throughout the spectrum. The precursor ion mass tolerance was set to 0.02 Da, and the MS/MS fragment ion tolerance was set to 0.02 Da. A network was then created in which edges were filtered to have a cosine score >0.65 and more than four matched peaks. Furthermore, edges between two nodes were kept in the network if, and only if, each of the nodes appeared in each other’s respective top-10 most similar nodes. Finally, the maximum size of a molecular family was set to 50, and the lowest-scoring edges were removed from the molecular families until the molecular family size was below this threshold. The spectra in the network were then searched against the GNPS spectral libraries. The library spectra were filtered in the same manner as used for the input data. All matches between network and library spectra were required to have a score >0.65 and at least four matched peaks.
Data availability
All Sanger sequences were deposited in NCBI GenBank under accession numbers OR888764–OR888811 in the BioProject PRJNA1047907 (Table 1). The BioSample accession numbers for the assembled genomes described here are SAMN38599447 (strain 3Ho3b) and SAMN38599448 (strain 4Ho3b). The genomes used for comparison are listed in Supplementary Table S2. The data obtained through MS using a molecular networking approach are publicly available for reference and use by the scientific community. For the ESI+ MS-technique-acquired data, interested parties can use GNPS with the identifier ID = b609721100ce491e9cde0cced1909f84 to access them. The ESI-MS data are available in the same repository, and the identifier ID = f36714ac5bd54798982cab5afda5cb97 can be used for access.
Results
Physicochemical parameters of the Al Wahbah crater
The pH and electrical conductivity values were 8.9 and 66.22 dS/m in the salt-crust, 9.1 and 28.6076 dS/m in the clay soil, and 9.4 and 42.8028 dS/m in the dry edge of the salt-crust soil sample, respectively. The clay-soil sample had the highest number of elements with the highest concentrations, including Fe, C, Ca, Mg, Ti, and P. The highest concentrations of Al, K, Na, and N were in the soil with salt crust, whereas the highest concentration of Si was in the soil-only sample. The salt-crust soil sample had the highest concentration of sodium (59.92 g/kg) compared with 28.14 g/kg in the soil-only sample and 23.21 g/kg in the clay-soil sample. In contrast, the Mg concentration was higher in the clay-soil sample (32.88 g/kg) than in the samples with only soil (15.38 g/kg) and salt crust (12.72 g/kg). The total nitrogen content was 0.21 g/kg in the soil-only and clay-soil samples and 0.3 g/kg in the samples with salt crust. The total carbon concentration was highest in the clay soil (11.37 g/kg), followed by salt-crust (9.75 g/kg) and soil-only (7.17 g/kg) samples (Table 1).
Physicochemical Properties and Elemental Composition of Al Wahbah Crater Soil Samples
Data are presented as the mean (±95% confidence level) of a triplicate.
Culturing haloalkaliphilic bacteria from the Al Wahbah volcanic crater
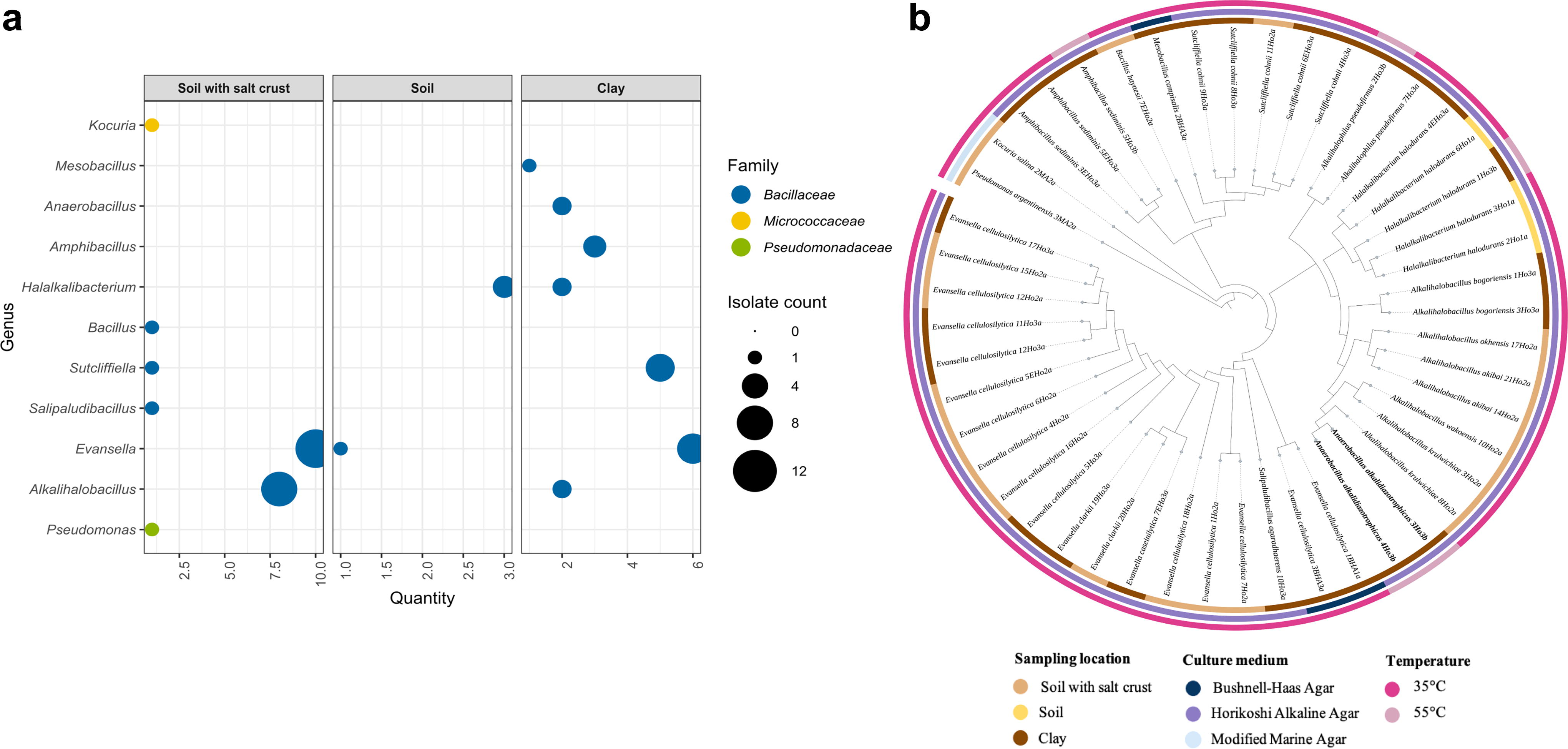
A total of 48 bacterial isolates, spanned into three families, were obtained from the soil samples collected from three sites at the Al Wahbah crater (Fig. 2a, Supplementary Table S3). The phylogenetic tree showed four clearly distinct clusters (Fig. 2b), with three large clades comprising the Bacillota phylum. The clay soil yielded the highest number of strains and generic diversity (25 isolates), followed by the salt crust soil sample (20 isolates). Most of the cultured strains belonged to family Bacillaceae, with more representatives of the genera Evansella (n = 18) and Alkalihalobacillus (n = 10). Moreover, strains of Kocuria and Pseudomonas genera were isolated only from soil samples containing salt crust. The majority of bacteria (n = 43) were isolated at 35°C, whereas five strains were recovered at 55°C, including members of Halalkalibacterium, Alkalihalophilus, and Amphibacillus genera. Two of them were exclusively cultured in thermophilic conditions, identified as Anaerobacillus alkalidiazotrophicus strains 3Ho3b and 4Ho3b based on 16SrDNA sequencing (Fig. 2b), and were further selected for WGS and taxonomic and functional analyses.
Genomic features and relatedness indices
The genomes of strains 3Ho3b and 4Ho3b, both cultured from clay-soil samples in Horikoshi alkaline agar at 55°C, were assembled de novo. The genome size was approximately 4.4 Mb, with 43.4% GC content and ∼4,530 protein-coding sequences in 48 and 49 contigs, respectively. The general genomic features of the two strains 3Ho3b and 4Ho3b are summarized in Table 2. Their circularized genomes with additional information of the contigs, GC%, CDS on both strands and gene content are shown in Supplementary Figure S3.
General Features of the Genome of Strains 3Ho3b and 4Ho3b
WGS, Whole-genome sequencing.
Aiming to confirm the identity of the bacterial strains, we performed ANI and dDDH analyses with the closest species from the public database (accession number GCF_000011145.1), and both strains, 3Ho3b and 4Ho3b, were closely related to Halalkalibacterium halodurans, with 99% similarity for ANIb and 94.5% for dDDH (Supplementary Table S4). Moreover, ANI indices (<95%) and dDDH values (<70%) were above the threshold levels of bacterial species identity, confirming that the examined Al Wahbah crater strains belonged to Halalkalibacterium halodurans species.
Phylogenomic analysis of Halalkalibacterium halodurans strains
We generated whole-genome based phylogenies to identify closely related species in the isolated genomes. For that, first, pangenome analysis was performed with our 2 newly sequenced genomes—3Ho3b and 4Ho3b—and 27 genomes from family Bacillaceae (selected on the basis of their core proteins analyses), aiming to capture the complete genetic diversity within species. Based on the resulting pangenome, core genome, and singleton analysis (Supplementary Fig. S4, Supplementary Table S5), we identified a total of 15,752 genes as a pangenome and 879 genes were identified as the core genome, keeping genome 3Ho3b as a reference. We also identified approximately 537 genes as part of the core genome and 29 genes as singletons of our novel strains of Halalkalibacterium halodurans (Supplementary Fig. S4, Supplementary Table S5).
Subsequently, we created two phylogenetic trees that were based on the whole genome of 27 species and 2 newly sequenced genomes of family Bacillaceae (Fig. 3) and based on core proteins, keeping strain 3Ho3b as the reference genome for the core protein (Supplementary Fig. S2). We found that newly sequenced genomes 3Ho3b and 4Ho3b were in the same clade as the species Halalkalibacterium ligniniphilum and Halalkalibacterium halodurans. Similarly, the phylogenetic tree based on the core protein exhibited the same pattern, confirming the phylogeny placement of our strains into Halalkalibacterium halodurans species.
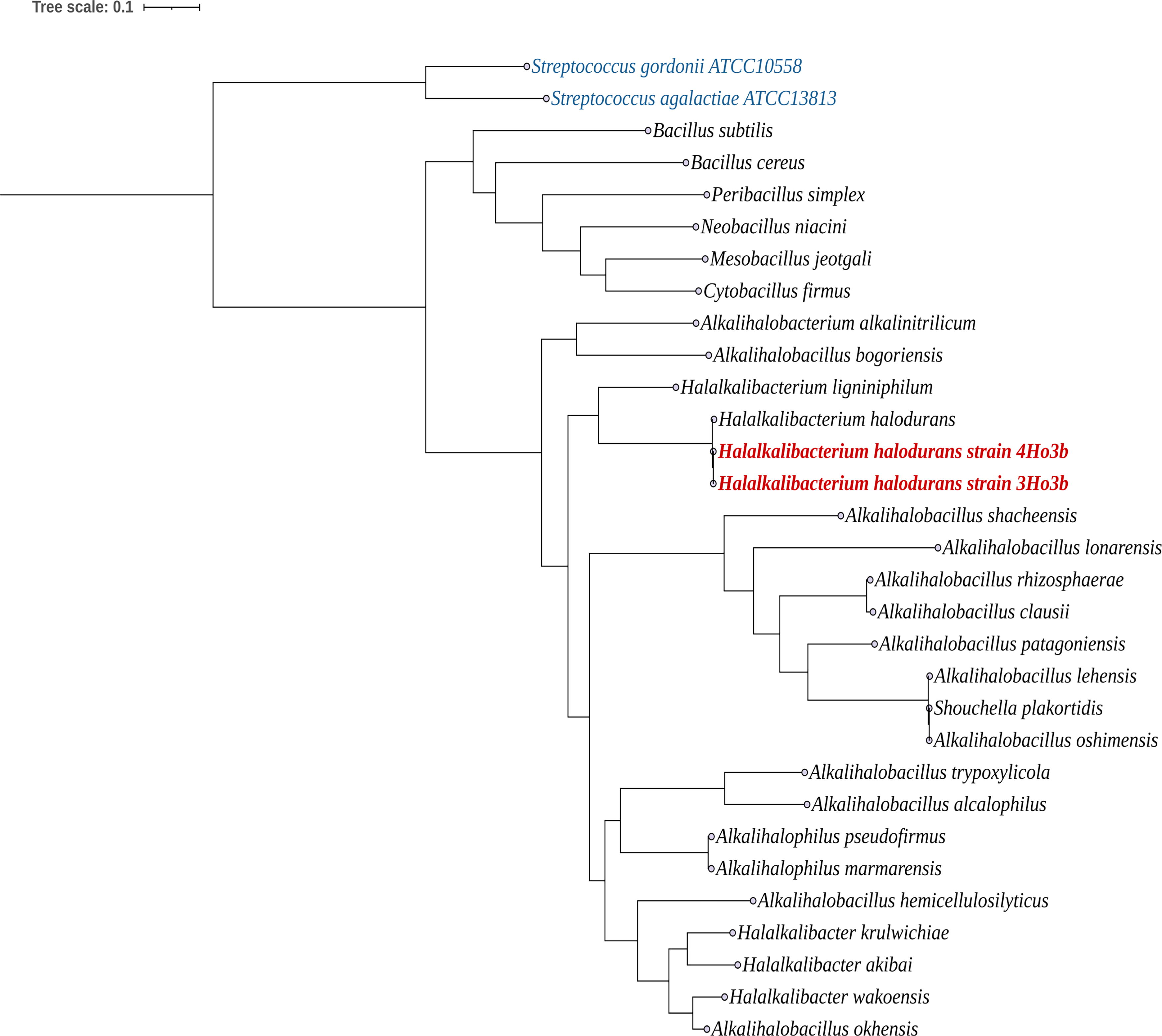

Whole genome sequencing phylogenetic tree of our newly sequenced genomes from 3Ho3b and 4Ho3b strains. The phylogenetic tree was constructed using the protein sequences of halophilic bacteria (27 Bacillus species) that were downloaded from the NCBI genome server and our two genomes from 3Ho3b and 4Ho3b strains. Species from Streptococcus were used as outgroups. A total of 119 single-copy genes associated with Bacillota were applied to position both strains. ModelFinder-Plus was used to construct the phylogenetic tree from the protein alignment with 1,000 ultrafast bootstrap replicates. NCBI, National Center for Biotechnology Information.
Functional characterization and data mining for genes with potential applications in astrobiology
To investigate the genetic characteristics of the two polyextremophilic strains, we used RAST-tk and NCBI COG to perform a comprehensive genome annotation. The subsystem mapping resulted in 455 subsystems for both Halalkalibacterium halodurans strains (3Ho3b and 4Ho3b). Among the annotated subsystems, the top categories based on average gene counts were related to amino acids and their derivatives, carbohydrates, cofactors, vitamins, prosthetic groups, pigments, and protein metabolism, which showed the highest numbers of assigned genes in descending order. With COG, 4,317 genes were predicted for the 4Ho3b strain and 4,315 for the 3Ho3b strain.
To explore the potential for producing secondary metabolites in the newly discovered strains of the Al Wahbah crater, we used antiSMASH, BAGEL, and PRISM tools to predict secondary metabolites. This analysis revealed a total of eight gene clusters in both analyzed genomes, two of which exhibited 100% similarity with lantibiotic haloduracin and bacillopaline, and the other six clusters were similar to ectoine, schizokinen (NI siderophore), fengycin (betalactone), 7-deoxypactamycin (T3PKS), and RiPP-like and cyclic lactone autoinducer. In addition, PRISM 4 detected three clusters, including the same types observed with antiSMASH-class II lanthipeptide, ectoine, and NI siderophore, whereas with BAGEL, haloduracin was identified in one area of interest (Fig. 4a). Analysis with the PHASTER program detected only one intact prophage sequence in the genomes, which was related to Staphylococcus with a GC content of 40.41%, a 38.8-kb region length, and four incomplete prophages with similarity to Bacillus and Peanebacillus phages.
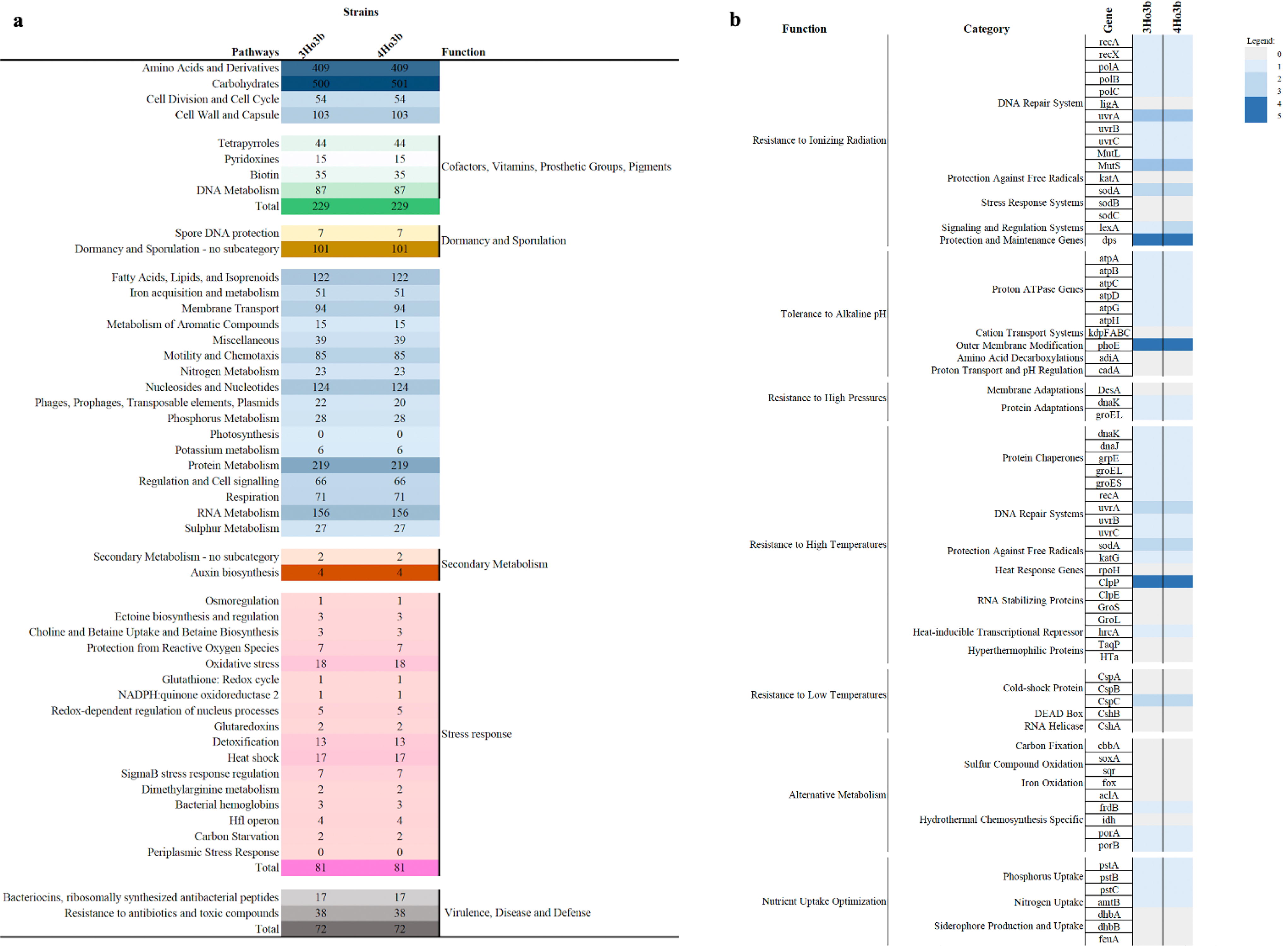
Annotated genes of metabolisms from Halalkalibacterium halodurans strains (3Ho3b and 4Ho3b). Panel
To understand more deeply the applicability of the analyzed strains from the Al Wahbah crater to the field of astrobiology, we conducted further investigations into the metabolic potential of these strains by manually mining genes of interest for different functionalities in the strain genomes, such as resistance to ionizing radiation, tolerance to alkalinity and extreme temperatures, resistance to high pressure, the presence of alternative metabolisms, and optimization of nutrient uptake. We then made some noteworthy observations (Fig. 4b). Overall, we used COG to predict genes related to genome stabilization, DNA repair systems, and adaptation to high temperature and alkaline conditions. A complete list of detected genes is presented in Supplementary Table S6.
Regarding the adaptation of the strains to thermal stress, both genomes exhibited chaperones involved in repairing heat-induced protein damage, DnaK and DnaJ, as well as small Hsps (hsp20), htpX, and ClpP genes, and GroEL and GroES. Genes related to DNA repair systems, such as recA, uvrA, uvrB, and uvrC, and those that protect against free radicals, sodA and katG, were also detected. Genes associated with cold-shock proteins were not commonly detected, and only the ClpC gene was observed in the analyzed genomes. Genes involved in adaptation to alkaline pH were identified, including the antiporter NhaC, proton ATPase genes atpA to atpH, and phoE, which is a gene involved in outer-membrane modification. To protect against ionizing radiation, the genes recA, recX, polA-C, uvrA-C, MutL, and MutS, which are involved in DNA repair, and sodA, lexA, and dps, which are involved in the stress-response system, were identified. Interestingly, many genes involved in carbon, sulfur, iron, and nitrogen metabolism were absent. We predicted hydrothermal chemosynthesis-specific genes, such as frdB, porA, and porB, and genes related to phosphorus (pstA, pstB, and pstC) and amtB for nitrogen uptake were observed.
Molecular network-based dereplication of microbial extracts
To analyze the chemical inventory of Halalkalibacterium halodurans strains, we used tandem MS (MS–MS) data processed on the GNPS platform. This approach enabled the execution of detailed chemical annotations and creation of a global molecular network. The integration of GNPS with dereplication, assisted by a natural products library, facilitated the identification of various chemical structural classes and their analogs. As indicated in Table 3, the molecules were annotated in accordance with level-2 identification, consistent with the Metabolomics Standards Initiative guidelines by Sumner et al. (2007).
Compounds of Halalkalibacterium Halodurans Strains Annotated Using the GNPS Database: Molecular Families, Ionization Mode, Retention Time, and Detected Masses
GNPS, Global Natural Products Social; RT, retention times.
The use of a molecular network was crucial for differentiating chemical species within the same molecular family, organizing them into clusters based on fragmentation spectral-alignment algorithms, independent of retention times (RT). This methodology significantly expedited the annotation of known molecules. Figure 5 shows the resulting global molecular network, with data from both the negative-ionization mode (Fig. 5a) and positive-ionization mode (Fig. 5b). The molecular networks reveal various chemical entities, accounting for approximately 710 nodes in the negative mode and 737 nodes in the positive mode. Specific clusters are formed for both ionization modes (Fig. 5). By comparing the spectra obtained with those deposited in the GNPS database, it was possible to annotate some clusters according to molecular family categories. This is observed, for example, in the five clusters identified in the negative mode (phospholipids, nucleotide derivatives, nonproteinogenic amino acids, pyridoxine derivatives, and tryptophan derivatives) and three clusters in the positive mode (phospholipids, lipoamino acids, and cyclopeptides).
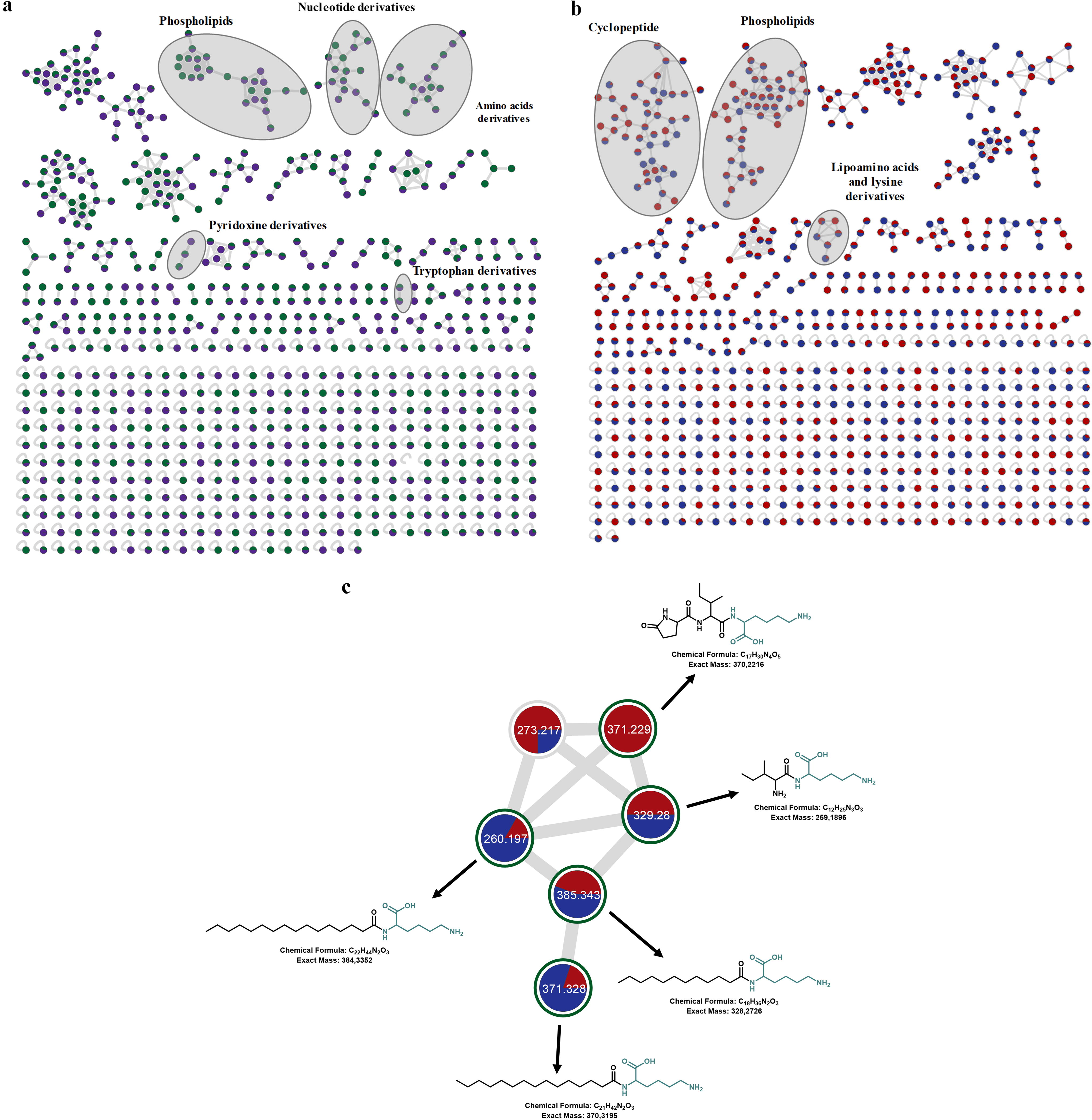
Global molecular network of the methanolic extracts obtained from the two strains of Halalkalibacterium halodurans. Each node represents the MS/MS spectrum of a feature (high-resolution mass and retention times).
In the analysis of the molecular network, some entities are specific to a single strain. The Venn diagram (Supplementary Fig. S5) shows that, for the negative-ionization mode, 16% of the nodes belong exclusively to strain 3Ho3b, 15% belong only to strain 3Ho4b, and 69% represent chemical entities that appear in both bacteria. Similarly, for the positive-ionization mode, 19% of the nodes belong exclusively to strain 3Ho3b, 18% belong only to strain 3Ho4b, and 63% are chemical entities present in both strains.
Discussion
Saudi Arabia harbors vast extreme environments with the potential to be exploited as analog habitats of extraterrestrial environments, such as Enceladus. The extremophilic microbiome thriving in these environments could be used for astrobiological applications, but no research of this type has been conducted at this location so far. With the aim of characterizing potential novel biological models, this study is the first to assess the culturable bacterial fraction of the Al Wahbah crater with a focus on its habitability and relevance to the habitability of Enceladus.
Due to the tidal forces exerted by Saturn, Enceladus’s core is active, which allows for volcanic activity on the ocean floor. In addition, recent discoveries indicate that Enceladus’ ocean possesses high concentrations of phosphate ions, which is highly unusual compared to Earth’s oceans, with about 100x to 500x the average abundance of phosphate (Postberg et al., 2023). This alkaline and saline environment, presenting high temperatures, offers a mystery regarding the potential life and types of polyextremophilic microorganisms that could be thriving there. On our planet, environments with this combination of physicochemical factors are quite rare. Recently, the shallow lakes of Last Chance and Goodenough were considered as a potential environmental analog to understand the origin of life and possible forms of life on Enceladus, which have similar chemical characteristics and present high phosphate concentrations and elevated pH (Haas et al., 2024).
In this sense, the search for other environments that exhibit these characteristics becomes of great importance to the astrobiology community. Therefore, the Al Wahbah crater, which is visually different from the icy moon, emerges as a very interesting environment to seek microbial models for Enceladus’s ocean and possible hydrothermal sources. Our physicochemical results of the Al Wahbah soils (Table 1), along with the studies of Wahab et al. (2014), show that the crater presents a high concentration of phosphates and presence of crystals at the bottom of the crater, also high pH, and temperatures that can exceed 50°C in the summer. It is worth noting that, during the rainy season, the interior of the crater becomes flooded, which gives rise to a temporary lake. Moreover, Albokari et al. (2018) used independent-cultivation techniques and detected a great diversity of halophilic and alkaline-tolerant microorganisms in the crater’s soils, which exhibited an interesting and diverse extremophilic microbial community.
Using different culture media (Schultz et al., 2023b), we isolated >48 strains from three sites. Phylogenetic analysis classified most isolates as belonging to Bacillaceae. Members of this family present distinct morphological and physiological characteristics, such as the formation of heat-resistant endospores, which enables them to thrive under extreme conditions, such as extreme environments in Saudi Arabia and potentially on Enceladus. Upon examining Al Wahbah bacterial strains, two polyextremophilic strains capable of growing under extreme temperatures, high salinity, and alkaline pH (conditions similar to potential hydrothermal sources in the subterranean ocean of Enceladus, Waite et al., 2017) were selected for deeper genomic characterization and chemical profile analysis with a focus on astrobiology. Both were phylogenetically close to Halalkalibacterium halodurans, a bacterium that lives in alkaline saline conditions, which was first described by Boyer et al. (1973), isolated from dried sewage sludge, and later reclassified by Joshi et al. (2021). Our strains, 3Ho3b and 4Ho3b, of Halalkalibacterium halodurans matched haloduracin, a two-peptide lantibiotic described as having potent antimicrobial activity (Oman and van der Donk, 2009), and bacillopaline, a metallophore used to acquire trace metals from the environment, respectively (Reitz and Medema, 2022). Moreover, clusters similar to ectoine, schizokinen, fengycin, T3PKS, and RiPP-like and cyclic lactone autoinducer were observed in both strains. These natural products may have a role when encountering unfavorable environmental conditions.
Considering the possible physicochemical conditions, such as high pressure, excess salts, alkaline pH, and elevated temperatures of potential hydrothermal vents on the Enceladus ocean, the metabolism of any potential microbial community there must meet specific requirements to survive in that environment. With regard to specific genes and metabolisms of interest for astrobiology, genome mining has led to the identification of key functional genes related to stress response and survival strategy. Our strains of Halalkalibacterium halodurans can flourish under thermophilic, highly saline, and alkaline conditions, and as such they developed different approaches to survival in extreme environments. To thrive under harsh thermal conditions, strategies related to DNA-stabilizing mechanisms, efficient DNA repair systems, and adapted proteins are required (Wang et al., 2015), and our strains exhibit these modifications. Genes related to protein folding under high temperatures (dnaK, dnaJ, groEL, and groES) and genes encoding DNA repair and protection against free radicals (recA, uvr, sodA, and katG) were detected. They are crucial in maintaining the functioning and integrity of the microbial system (Coker, 2019; Paulino-Lima et al., 2013; Somayaji et al., 2022). Genes encoding cold-shock proteins were also analyzed because of their wider role in stress tolerance both in cold tolerance and in osmotic stress (Schmid et al., 2009); however, only the CspC protein was observed. When facing a temperature downshift, the bacterial cells might counteract some harmful effects (Phadtare, 2004). In addition, proton ATPase genes and phoE encoding a phosphate-limitation-inducible outer-membrane pore protein were observed in our strains. These genes are crucial for maintaining intracellular pH under alkaline conditions and help to pump protons into the cell (Padan et al., 2005). In addition, in response to adverse environmental conditions, microorganisms enduring numerous physicochemical stresses may adopt reversible metabolic strategies, such as dormancy and spore formation (Leung et al., 2020).
Screening of the chemical profile of Halalkalibacterium halodurans strains using the molecular network approach revealed that the bacteria also differ minimally in their metabolite composition, as expected. However, the analysis showed several molecular classes essential for bacterial life that could be used as potential biosignatures. The plan to send mass spectrometers to Enceladus again is becoming increasingly close to reality. We know that the next generation of these instruments, aimed at searching for biosignatures, is very similar to the mass spectrometers used in Earth’s laboratories (Arevalo et al., 2020). With this advancement in space exploration, it will soon be possible to integrate bioinformatics tools for annotating natural products to help identify molecules from other celestial bodies. The approach of generating a molecular network from MS/MS spectra, along with comparison with spectral libraries, is very efficient for obtaining a preliminary understanding of an unknown sample. Thus, having an idea of the possible complex molecules that we could detect there, as a result of the metabolism of extremophilic microorganisms cultivated under similar conditions, would greatly assist in future missions (Lopez et al., 2019).
Notably, a phospholipid cluster was detected in both ionization modes in our study. As fundamental components of bacterial cell membranes, these molecules form the base structure of the lipid bilayer, which significantly contributes to the maintenance of structural integrity and membrane fluidity and facilitates selective substance transport. Another detected class was nucleotides and their derivatives, which are omnipresent in all life domains and have key roles in DNA synthesis, enzyme regulation, and energy transport, as in ATP and GTP cases. In addition, the production of pyridoxine derivatives (vitamin B6) was observed. These molecules are critical cofactors in various enzymatic reactions, particularly in amino acid synthesis and degradation. Because of their chemical structure similar to that of some osmolytes, it is possible that pyridoxine may also be biosynthesized by bacteria for this purpose (Sleator and Hill, 2002).
The following two molecular classes have attracted considerable interest: cyclopeptides and lipoamino acids or lipopeptides. Cyclopeptides are notable for their variety of biological activities. In the literature, these molecules are often associated with biological control and the capture of certain metabolic cations, acting as chelators (Gisin et al., 1978; Kubik, 2022; Ribeiro et al., 2022). Lipoamino acids, illustrated in the molecular cluster in Figure 4c, are molecules that feature one or more amino acid units linked to a fatty acid, usually attached to the amino group. These molecules demonstrate the ability to confer stability to macromolecules under chemical and thermal stress conditions (Vicente-Garcia and Colomer, 2023). The presence of a hydrophobic chain differentiates them from peptides and amino acids, thereby endowing them with amphiphilic properties. Thus, these molecules can act in micelle formation to protect nutrients and other cellular molecules in alkaline and saline environments (Vicente-Garcia and Colomer, 2023).
Although the molecules presented are not novel, their identification in microorganisms isolated from the Al Wahbah crater is novel. Analyzing the chemical profile of these strains reveals the importance of investing in studies on metabolites produced by extremophilic microorganisms. Our study showed that, even when using specialized databases, it was possible to annotate only <10% of the entities detected in the extracts. This finding indicates that exploring the secondary metabolism of extremophiles is a promising field that, if further explored, could broaden our understanding of astrobiology and the molecular diversity produced by microorganisms in extreme environments on our planet. In the search for extraterrestrial life, identification of these molecules as biological markers in upcoming space missions could be significant, particularly if their patterns are not easily attributable to nonbiological processes. Detection of these molecules in the oceans of Enceladus could suggest the presence of microbial activity.
Conclusion
In summary, these study findings broaden the range of analogous extraterrestrial environments in astrobiology, including the Al Wahbah crater in Saudi Arabia. We investigated and characterized cultivable bacteria from the crater to uncover potential biological models and biosignatures pertinent to the search for extraterrestrial life, particularly on Enceladus, an icy moon of Saturn. Specific attention was devoted to exploring the distinctive attributes of Halalkalibacterium halodurans strains 3Ho3b and 4Ho3b, which led to demonstrating their ability to thrive under thermo-haloalkaliphilic conditions and provided valuable insights into the adaptations necessary for survival in extreme environments. The molecules reported by chemical profile analysis show that the MS analytical technique can be useful for identifying the components that characterize life in an astrobiological context. Furthermore, chemical profile MS analysis underscores the potential of this technique as a powerful tool for identifying life-associated components in an astrobiological context. The molecules identified through this analysis open avenues for further research, emphasizing the need for a more extensive exploration of metabolite production by extremophilic microorganisms. However, the quest to understand the production of metabolites by extremophilic microorganisms remains challenging and underinvestigated because of the scarcity of chemical annotations.
Overall, these initial insights from integrated omics data, encompassing both genomics and metabolomics, suggest that bacterial strains isolated from the Al Wahbah crater in Saudi Arabia may serve as terrestrial proxies for hypothetical microbes on Enceladus. This comparison provides a foundation for identifying potential biosignatures and potentially offering novel biosignatures for use in the search for life beyond Earth.
Footnotes
Acknowledgments
The authors thank Tahira Jamil and Isabela Dal’Rio for their assistance with the phylogenetic and phylogenomic trees. The authors remember Andrii Romanenko, who is sadly no longer with them and to whom they owe much more than words.
Authors’ Contributions
A.S., J.S., and A.S.R. conceived the study and wrote the article. A.S., A.R., and J.S. conducted the microbiology experiments. J.S., F.M., A.K.J., and L.G. performed the bioinformatics analysis and created the figures. A.S., M.A.T., and E.R.F. analyzed and interpreted the data from the chemical profiles and generated the figures. A.R. and J.S. conducted the physicochemical analyses. A.S.R. funded the project. All authors read, revised, and approved the final article.
Author Disclosure Statement
The authors declare that this research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Funding Information
This work was supported by Prof. Alexandre Soares Rosado’s KAUST Baseline Grant (
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
Supplementary Table S6
Associate Editor: Don A. Cowan
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
