Abstract
An ideal life detection instrument would have high sensitivity but be insensitive to abiotic processes and would be capable of detecting life with alternate molecular structures. In this study, we propose that catalytic activity can be the basis of a nearly ideal life detection instrument. There are several advantages to catalysis as an agnostic life detection method. Demonstrating catalysis does not necessarily require culturing/growing the alien life and in fact may persist even in dead biomass for some time, and the amplification by catalysis is large even by minute amounts of catalysts and, hence, can be readily detected against abiotic background rates. In specific, we propose a hydrolytic catalysis detection instrument that could detect activity in samples of extraterrestrial organic material from unknown life. The instrument uses chromogenic assay-based detection of various hydrolytic catalytic activities, which are matched to corresponding artificial substrates having the same, chromogenic (preferably fluorescent) upon release, group; D- and L-enantiomers of these substrates can be used to also answer the question whether unknown life is chiral. Since catalysis is a time-proportional product-concentration amplification process, hydrolytic catalytic activity can be measured on a sample of even a minute size, and with instruments based on, for example, optofluidic chip technology.
Introduction
The three biology experiments on the Viking missions in 1976 were the first, and to date the only, attempt at life detection on another world. The pyrolytic release experiment (Horowitz et al., 1977) exposed martian soil samples to 14C-labeled CO2 and light to determine whether organic material was produced via photosynthesis. The labeled release (LR) experiment (Levin and Straat, 1977) measured the radioactivity in the headspace above a soil sample to which a solution containing 14C-labeled organics had been added. The gas exchange (GEx) experiment (Oyama and Berdahl, 1977) measured the gases in the headspace over a sample of martian soil as water vapor and a nutrient solution were added in succession. The results of the GEx and LR experiments were of interest.
In the GEx experiment when the samples were exposed to water vapor, the soil released O2 in amounts ranging from 70 to 770 nmol/cm3. Heating the sample to 145°C for 3.5 h reduced the amount of O2 released by about 50%. There was a slow evolution of CO2 when nutrient solution was added to the soil. The LR experiment showed a rapid release of CO2, followed by a prolonged slow release of CO2, when a nutrient solution was added to the soil sample. The effect was completely removed by heating to 160°C for 3 h, partially destroyed at 40–60°C, and relatively stable for short periods at 18°C, but lost after long-term storage at 18°C. Although not considered as one of the life detection instruments, a gas chromatograph mass spectrometer (GCMS) on Viking sought to detect organics in the samples following pyrolysis and reported no detection of organics other than contaminants.
The results of the Viking biology experiments and the lack of organics detection by the GCMS are now explained by the presence of high levels of perchlorate salts in the martian soil (Hecht et al., 2009; Glavin et al., 2013) and the reaction of this perchlorate with organics during pyrolysis in the GCMS (Navarro-González et al., 2010) as well as of reactive oxidants, such as superoxide and hydroxyl free radicals, and hydrogen peroxide in the soil, produced by the perchlorate when irradiated by ultraviolet radiation (Quinn et al., 2013) or cosmic radiation (Georgiou et al., 2017).
It is interesting to note that the only other approved missions with life detection as a stated science goal are the ExoMars Rover mission set to depart for Mars in a few years, and the Dragonfly mission set to arrive at Titan in 2034. ExoMars will search for biomolecules with Gas Chromatography-Mass Spectroscopy (GCMS) and laser desorption Mass Spectrometry (Vago et al., 2017). Dragonfly will detect organic molecules with a GCMS and search for H2 variations in time and location as a biosignature of CH4-based life (Lorenz et al., 2019; Barnes et al., 2021).
In contrast to Viking, the approaches for life detection being used on approved missions are based not only on detection of metabolism or growth but also on detection of the structural or functional molecules used by Earth biology, both their relative concentration and their chiral patterns (e.g., McKay, 2011; Vago et al., 2017). This was first suggested by Lovelock (1965) and further considered by Pace (2001), who argued that life anywhere would use the same biomolecules as life on Earth. These suggestions focused on the detection of molecules and patterns as found in life on Earth in lipids, amino acids, and informational molecules such as nucleic acids.
McKay (2004) generalized this approach to life different from Earth life but based on the same types of biomolecules. The same approach would hold for the panspermia hypothesis, which suggests that life may have originated on a particular planet through an interplanetary transfer of either life's fundamental components (molecular panspermia) or life itself (organism-based panspermia) (Mitton, 2022; Sithamparam et al., 2022). Similar lines follow the use for alien life detection of the so-called picogram-scale interstellar probes via Earth life-bioinspired engineering (Church, 2022).
Other approaches to life detection are also being considered. Abrahamsson and Kanik (2022) and Enya et al. (2022) provided a systematic listing of methods for life detection grouped in the following major categories: growth in culture, microscopic imaging, biomolecules, and metabolism. Metabolism is split into two subcategories: nutrient uptake and waste release, and catalysis. The three Viking biology experiments were examples of methods to detect nutrient uptake and waste release. Current mission concepts to search for life on Mars and the outer solar system that have been proposed or are at a high level of development—such as Mars Icebreaker (McKay et al., 2013), Europa Lander (Hand et al., 2022), and Enceladus (MacKenzie et al., 2021)—are examples of biomolecule detection. The ExoMars mission is likely to be the next example of this with instruments that can search for biomarkers and characterize their concentration and chirality (Vago et al., 2017). The GCMS on Dragonfly is an example of searching for biomolecules without specification as to the type used.
The detection of H2 on Dragonfly is an example of searching for nutrient uptake, although in the environment and not in a growth chamber. Uptake of H2 has been suggested as an indicator of methane-based life on Titan (McKay and Smith, 2005; McKay, 2016). Systemic disequilibrium caused by biological activity alters its environmental surroundings and leaves behind traces (physical, chemical, and isotopic) that would not otherwise be present in nature without life, which has been proposed as a potential biosignature (Cabrol, 2016); the prime example is the chemical disequilibrium in planetary atmospheres (Hitchcock and Lovelock, 1967; Krissansen-Totton et al., 2018). Α Ladder of Life detection scheme, not meant to endorse particular biosignatures or instruments for life detection measurements, has been suggested for establishing a flexible set of criteria for distinguishing putative biosignatures from an abiotic background, such as “the response to substrate addition” as evidence for ongoing metabolism (Neveu et al., 2018), where extant life catalysis is implied—the approach of the Viking biological experiments (Klein, 1978).
More recently discussions of the so-called agnostic life detection have considered molecular patterns that might arise in an unknown biology on the basis of complexity without specifying the type of molecule (Marshall et al., 2017; Johnson et al., 2018, 2020; Chou et al., 2021). These approaches could define agnostic biosignatures as those that discern unidentified, molecular characteristics and chemistries that might indicate as-yet-unrecognized living processes.
Our approach is to detect and measure by instrumentation an organic function in extraterrestrial organic matter (EOM) that is common in all life types. In the present study, we extend catalytic activity from a universal functional marker of life, as previously presented in Georgiou (2018), to a feasible method for detecting agnostic life. Catalysis is a critical feature of life as it is a basic requirement for speeding up all biochemical organic reactions to a degree life requires to be sustained. Therefore, highly efficient and specific organic catalysts of different types must be a universal feature of biology everywhere, and among life's most fundamental, even defining characteristics. Supportive to our proposition are the following roles of catalysis suggested for the emergence of life on Earth and elsewhere in the Universe.
Catalysis feeds the evolutionarily stable disequilibrium that develops in life when primitive cells are selected to enhance internal molecule catalytic activity, while molecules tend to evolve to minimize it to optimize their relative metabolic fitness (Takeuchi et al., 2016). Moreover, catalysis can be the outcome of urability, a property of planetary bodies distinct from habitability, which allows the chemical reactions and molecular assembly processes required for life to begin (Deamer et al., 2022). Catalysis is an inherent part of the postulated evolutionary processes leading to the emergence of life (Calvin, 1956; Nowak and Ohtsuki, 2008). Life may emerge by an evolutionary process starting from abiotic materials, which involves the initial random formation of complex molecules from simpler ones.
Gradually, through processes of random variation, selection, and autocatalysis (the origin of catalytic activity), more complex systems emerge that carry the continuity and order of living systems (Calvin, 1956). In a more updated scenario, life may emerge with the ability of replication in a pre-life environment of active monomers that form random polymers, which can act as information carriers and catalysts, and as templates for their own reproduction. Yet, there must be a phase transition, where the effective replication rate by exceeding a critical value allows life to outcompete pre-life (Nowak and Ohtsuki, 2008). Once past this threshold, life gathers resources and energy and disperses quickly to all connected environments by acquiring a faithful reproduction, meaning that life's start place is likely local and very transient. Supportive to this view is the scenario pointing that before the emergence of life, very short pulses of energy cause the occurrence of extremely fast chemical reactions in the chemical–prebiotic organic microsystem.
They are the system's response to high frequency and multilevel fluctuations in temperature, pressure, and other physicochemical parameters. These induced impulses initiate a shift of the system to a new state, which is intermediate in the process of its transition to life. This shift is defined as a thermodynamic inversion, which is a key link in the transition of a nonliving prebiotic microsystem into a primary living state characterized by catalytic activity, when the contributions of free energy and information become dominant over the contribution of entropy (Kompanichenko, 2019). In that context, catalytic function besides speeding up organic chemical reactions is thought to be one of the decisive prerequisites for self-organization of Earth life's matter, by causing certain self-enhancing growth properties of the living system in combination with various feedback mechanisms (Eigen, 1971).
Evolutionary development (e.g., Darwinian)—arguably the definition of living systems (Benner, 2010)—involving both chance and competitive optimization, will result in the development of catalytic systems in any life-form that are specific and highly efficient and likely to be unique in molecular structure. Thus, organic catalysis must be a universal feature of life (Georgiou, 2018) as long as its functional manifestation on highly accelerated reactions takes place within enclosed spaces [most likely surrounded by polar lipids (Georgiou and Deamer, 2014)] to keep the concentrations of catalysts, substrates, and products from diluting out.
Catalysis for Agnostic Life Detection
Catalysis is the chemical activity by a given regenerated molecule (organic or inorganic, termed the catalyst) to accelerate reactions toward formation of products or substrates, or reversibly toward both. That is, catalyst has no effect on the equilibrium position of a reaction nor of its free energy (the net energy output produced or usable energy) but allows its equilibrium to be reached more quickly or established at a lower temperature (by lowering the activation energy required for the reaction to happen), thereby making reactions more metabolically suitable. Life cannot be envisioned without acceleration of the rates of the reactions taking place among its component organic molecules. If not catalyzed, organic reactions often require such a long time that is impossible to sustain life's metabolism.
For instance, the rate of the uncatalyzed breakage of an amide bond in a protein peptide by water (termed hydrolysis) at neutral pH and room temperature (RT) has a half-life (t 1/2) = ∼7 years (Kahne and Still, 1988); orotic acid is non-catalytically decarboxylated (in water at neutral pH and RT) with t 1/2 = 78 million years; spontaneous hydrolysis of the dimethyl phosphate bond of nucleic acids (e.g., DNA) has t 1/2 = 130,000 years (in water at neutral pH and RT). On the contrary, catalysis performed by certain proteins (enzymes) of Earth life can accelerate such very low reaction rates by >1014-fold. For instance, the hydrolytic decarboxylation rate of orotidine 5′-phosphate by the enzymatic catalyst decarboxylase is accelerated by 1017-fold (Radzicka and Wolfenden, 1995).
Catalysis as a life detection approach (Georgiou, 2018) was presented in the NASA's 2022 Future of the Search for Life Workshop, aiming to identify measuring/sampling technologies, 20 years or more in the future, for in situ searches for life elsewhere in our solar system. Among the new approaches to life detection discussed, catalysis was one of the three possible indicators of life selected as highly promising avenues for further investigation; the others were “dynamic disequilibrium” and “informational polymers” (Neveu et al., 2023). Catalysis was also recognized as biosignature in a recently proposed taxonomy of potential biosignatures, and has been placed in the “activity” category among the three proposed; the others were “chemistry” and “structure” (Davila et al., 2023).
There are several advantages to catalysis as an agnostic life detection method. Catalysis may be demonstrated without culturing or growing the alien life, and it can even survive in dead biomass for extended periods of time. An example is the detection of phosphatase activity across the hyperarid regions of the Atacama Desert (Kobayashi et al., 2022) and in other extreme environments (Takano et al., 2005, 2006), as it is correlated with other biomarkers and proposed as a life detection method. The amplification by catalysis is large and, hence, can be readily detected against abiotic background rates.
Catalysis could also answer to the question whether unknown life is chiral by testing catalytic activities on the D- and L-enantiomers of chiral substrates. Benner (2017) has argued for the necessity for homochirality for the construction of polymers by a genetic code—the production of complex polymeric systems from simple chiral molecules (such as Earth life's polysaccharides that are made of right-handed sugars, proteins/peptides of left-handed amino acids, and nucleic acids of the compositionally asymmetric sugars D-deoxyribose and D-ribose forming the respective DNA and RNA backbones) that possess varied forms of molecular recognition and selectivity—both of which are crucial in life.
Agnostic life may not possess polymeric systems such as those found in Earth life (and hydrolyzed by, e.g., peptidases, nucleosidases, glycoside hydrolases), and one task to be addressed for the detection of catalytic chirality is to synthesize D- or L-artificial substrates for more than one chiral hydrolytic catalytic activities (e.g., peptidase, nucleosidase, glycoside hydrolase, phosphatase, lipase, epoxy hydrolase). More than the hydrolytic activity detected in an EOM sample can serve as a criterion for excluding the abiotic nature of the detected organic catalysts. The prevalence of the D-substrate over the L-substrate and vice versa may imply a preferred chirality by alien life.
The most important forms of catalysis in Earth life are dehydration synthesis and its opposite hydrolysis (Georgiou, 2018). Reactions involved in catalytic dehydration synthesis are mostly endergonic, that is, they require input of energy (via losing one water molecule). This energy input requirement makes this form of catalysis an unreliable marker of unknown life because its energy sources cannot be known. In contrast, hydrolytic catalysis is a potential marker of unknown life because hydrolytic reactions (where a molecule is broken down by water to two or more smaller molecules) release energy in general; usually, they do not need an energy input, except, for example, P-O bonds are broken as in ATP hydrolysis.
This form of catalysis is a suitable marker for the additional reason that Earth life's class of hydrolytic enzymes (hydrolases) is the largest in numbers and most diverse class of enzymes (among the six main classes); it is composed of more than 200 enzyme catalytic types/subtypes, with some common ones being esterases, proteases, glycosidases, lipases, and peptidases (Shukla et al., 2022). Therefore, hydrolytic catalysis offers many energy-independent modes for detection of catalysis.
However, nonliving materials can also act as catalysts, as this is a common characteristic of industrial processes, while a variety of processes in planetary settings are catalyzed by mineral surfaces. Therefore, it is essential to consider that catalysis may also have an abiotic background, depending on the specifics of the catalytic activity measurements. Our approach for the detection of hydrolytic catalytic activity in the EOM sample minimizes the possibilities of being due to abiotic background by searching for many different catalytic activity types, using more than 200 different matched artificial substrates.
Another criterion is the attained catalytic timescales, where those suitable for life are 10−3–105 s (over the biological temperature range from −20°C to +120°C) in contrast to the timescales 10−3–1016 s of uncatalyzed reactions (Arcus and Mulholland, 2020). In addition, the chemical structures of the catalysts in the EOM samples [identified by, e.g., NASA's Mars Organic Molecule Analyser (MOMA) instrument (Goesmann et al., 2017)] would be of extinct/extant life origin if they also comply with the suggested molecular pattern complexity criteria described elsewhere (Marshall et al., 2017; Johnson et al., 2018, 2020; Chou et al., 2021).
Operational Principles of a Hydrolytic Catalysis Instrument
In this study, we present a hypothetical conceptual model for the operation of a hydrolytic catalysis detection (HCD) instrument on EOM samples. In principle, EOM samples are tested for certain hydrolytic catalytic activity types after incubation with catalytically matched artificial organic substrates resistance to cosmic radiation, which are designed to be broken down into known and easily measured products. These substrates are synthesized to carry a chromogenic group bound to them, which, upon its catalytic release, either fluoresces or absorbs at known wavelengths (Georgiou, 2018). Thus, hydrolytic catalytic activity and its specific type exhibited in an EOM sample can be identified and quantified either by the fluorescence or absorbance exhibited by the released chromogenic group, when a matching of the artificial substrate and the corresponding catalyst type in the EOM sample is met (Georgiou, 2018).
Artificial substrates have been already developed to match various types of hydrolytic catalytic activity (e.g., esterase, lipase, amidase) in studies for new enzyme discovery, differentiation between similar enzymes, quality control purposes, and so on (Wahler et al., 2001; Reymond, 2004, 2008). The main hydrolytic catalysis identification/quantification methodologies available are: (1) Fluorogenic, based on a periodate (NaIO4)-coupled β-elimination of umbelliferone (Badalassi et al., 2000) and p-nitrophenol (Beisson et al., 2000) (the latter also absorbing in the UV-Visible spectrum); (2) chromogenic such as the back titration method with adrenaline (Fluxá et al., 2008).
Figure 1 outlines the operational and methodological principles of an HCD instrument using umbelliferone as a fluorescence-based chromogenic detector of hydrolytic catalytic activity, using as example the hydrolases acylase, phosphatase, lipase, esterase, and epoxide hydrolase, using nonfluorescent artificial substrate matches that they all contain the same chromogenic (fluorescent) group of umbelliferone. Detailed lists of the numerous artificial organic substrates (each corresponding to a certain type of hydrolytic catalytic activity) and the involved methodologies are presented extensively elsewhere (Badalassi et al., 2000; Reymond, 2001, 2004, 2008; Wahler et al., 2001; Goddard and Reymond, 2004a, 2004b) and have been developed in co-author Reymond's laboratory (
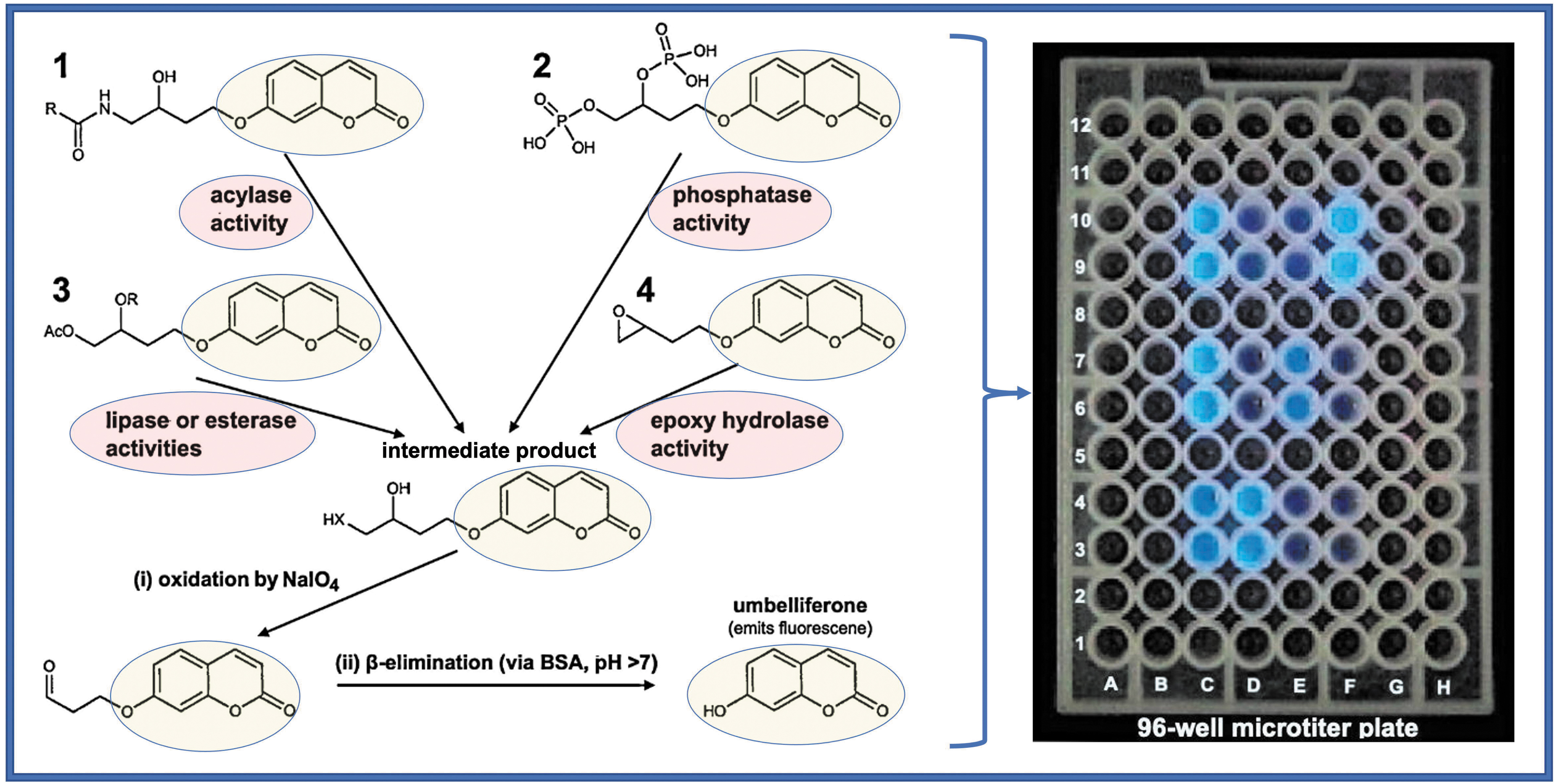
The principle of the periodate (NaIO4)-coupled fluorogenic assay operation-based HCD instrument. The indicative catalytic activities it measures are of the hydrolytic types, acylase (1), phosphatase (2), lipase or esterase (3), and epoxide hydrolase (4), all encircled in purple, using their correspondingly matching nonfluorescent artificial substrates 1, 2, 3, and 4. Each of these four different substrates contains the same chromogenic (fluorescent) group, that of umbelliferone (encircled in yellow). Upon mixing the sample of each hydrolytic catalysis type with its artificial substrate match, each of the four different artificial substrates is converted to the same nonfluorescent intermediate product, which upon a two-step processing (i/ii via NaIO4/BSA) releases the same blue fluorescent product, umbelliferone, by a process completed in only <15 min. The four different catalytic type fluorogenic reactions take place in a 100 μL sample/substrate mixture, placed inside separate round bottom microwells positioned as arrays (e.g., in a microtiter polypropylene plate). The reaction rate of each catalytic hydrolytic type is determined by recording, over time, the fluorescence emission intensity of umbelliferone at ex/em 360/460 nm by a microtiter plate reader (Reymond, 2001; Wahler et al., 2001), shown as intense light blue emission at the endpoint of the measurement. Lane C contains reference amino alcohol or diol products from the primary hydrolysis of #1 and #4 or #3, respectively. Enzyme reactions: D3-F4: amide #1, D6-F7: ester #3, D9-F10: epoxide #4. Enzymes: lane D: penicillin G acylase, lane E: Pseudomonas species lipase, lane F: Aspergillus niger epoxide hydrolase. The figure is a modification/combination of figures from previous studies with methodological details (Badalassi et al., 2000; Reymond, 2001; Wahler et al., 2001). A similar procedural principle can be applied to an EOM sample, the types of catalytic activities present can be identified by mixing it with a matched set of selected artificial substrates (e.g., as in Badalassi et al., 2000; Wahler et al., 2001; Goddard and Reymond, 2004a). In an EMO sample, up to 48 different hydrolytic catalytic types (including the corresponding reagent controls) can be tested on a 96-well microtiter polypropylene plate. EOM, extraterrestrial organic matter; HCD, hydrolytic catalysis detection.
Such instrument can also determine whether unknown life is chiral or not. This is carried out by measuring the hydrolytic catalytic activities in the EOM sample by incubating it, ideally separately, with the L- and D-chiral form of a given artificial substrate. Chirality, then, can be deduced by determining whether there is a statistically significant catalytic activity difference between the L- and D-chiral artificial substrate. In other words, the chirality activity demonstrated by the enantioselective catalyst in the EOM sample can be assumed ideally when there is very high catalysis rate by the enantiomeric catalyst for the artificial L-substrate and near zero for the D-substrate, and vice versa.
However, chirality can be deduced even if the artificial substrate exists as a 50%/50% mixture of the L- and D-form. In theory, an enantioselective catalyst should hydrolyze only half of a chiral substrate existing as a 50%/50% L-/D-mixture, which would be immediately observable at the end of the reaction by the chromogenic intensity of the released product. 7-[(3-cyano-3-benzoyloxyprop-1-yl)oxy]-2H-1-benzopyran-2-one is an example of such artificial substrate, where only 50% of it was used up by Candida cylindracea lipase, as determined by the fluorescence of liberated umbelliferone and validated by chiral-phase high-performance liquid chromatography analysis of the unreacted ester (Leroy et al., 2003). In any case, L- or D-chirality catalytic activity will then be assumed as also correspondingly reflecting the L- or D-chiral structures of the unknown natural substrates used by the EOM catalyst.
Hydrolytic catalytic activity of an EOM sample is measured over a short period time (e.g., ∼10 min) and at different concentrations of the given artificial organic substrate (e.g., in the μM range), where the reaction rate is measured (as a linearly proportional ΔμΜ increase over a set time interval), by quantifying the increasing concentration of the released from the substrate fluorogenic/chromogenic product. The fluorogenic product can be detected within a 5–20 pM range (the detection limit of fluorescein) and the chromogenic within the limit (±0.005) of a typical absorbance instrument. HCD instruments for chromogenic measurements suitable for space mission applications could be based on, for example, optofluidic chip technology (Yin et al., 2006).
As this proposed hydrolytically catalyzed reaction rate is set for single-substrate reactions catalyzed by the EOM sample, its hydrolytic catalysis kinetics may follow an enzyme-like Michaelis–Menten single-substrate kinetics (Kargi, 2009), a general feature of Earth life's enzymatic catalysis (Albe et al., 1990). If such kinetics is also displayed by the hydrolytic catalytic activities of the EOM sample, this sets an additional criterion to further strengthen the reliability of hydrolytic catalysis as marker of unknown life.
The minimum quantity of the EOM sample needed for testing its hydrolytic catalytic activity types by a hydrolytic catalysis instrument can be predicted from the methodological requirements in measuring Earth life's catalysis. Single cells are known to internally transform streams of relatively simple molecules, such as sugars, into a complex biomass mixture (proteins, nucleic acids, lipids, etc.). The catalytic enzymes performing this metabolic task comprise the largest fraction of the cell proteome, with Escherichia coli being a typical example (Valgepea et al., 2013). Thus, enzymatic catalysts (hydrolytic ones included) in Earth life comprise the largest fraction of total cell dry mass, and this may hold true for alien life as well. The total proteins contained in a single cell can be as low as 4 fg (10−15 g), and in E. coli, they comprise ∼55% of its dry mass (Milo, 2013; Zotter et al., 2017).
Assuming the worst-case scenario where the EOM sample has been extracted and concentrated from a single cell of an extant/extinct unknown life-form, the question is whether it is practically feasible to measure one of its possible types of hydrolytic catalytic activities by current methodologies? This may be possible when considering that catalysis is a time-proportional product-concentration amplification process and that there are kinetics measurement methodologies even for single cells; the latter can measure various classes/types of enzymatic catalytic activities (Kovarik and Allbritton, 2011) for enzyme concentrations from 0.01 to 10 μM (Zotter et al., 2017), and even do enzyme kinetics (Di Carlo et al., 2006).
Moreover, the concentrations of the artificial organic substrates—we propose to be used in the μM range for EOM hydrolytic catalysis measurements—fall in the concentration range (1–100 μM) of the metabolic substrates in Earth life's cells (Albe et al., 1990; Zotter et al., 2017). Nonetheless, catalytic activity detection sensitivity largely depends on the background activity (blank and/or activity by inorganic materials), which is due to the activation energy of the hydrolysis. For example, background of (carboxylate) esterase is usually high, while that of phosphatase is low. Moreover, background activity can originate by unidentified planetary chemistry, which may create circumstances that are favorable for the breakdown of the artificial organic substrates. In any case, we assume that such breakdown rates are much lower than those attained by catalysis.
Another critical factor affecting catalytic activity detection sensitivity is the source of the EOM sample and the concentration of its organic content. In that respect, the strategy for testing catalytic activity may vary depending on whether the target EOM samples are soils (e.g., Mars) or water (e.g., Europa). The catalytic activity detection method outlined in the present study can be easily applied in the latter situation assuming that the water EOM sample contains sufficient concentration of organics. However, additional soil EOM sample extraction procedures are required in the former case, which can also apply to water EOM samples in case fractionation, and concentration maximization of their organics content is required. In principle, water-dissolved organic matter (WDOM) can be easily obtained from dry soil samples (preferably from sedimentary locations, which presumably have the highest probability for accumulating biological organic matter) via their agitation with water, and from ice samples by their melting to water.
Subsequently, WDOM is filtered (to remove dust particles) and is subsequently fractionated into various organic molecular classes. Among the many cartridge-packed resin-based solid-phase fractionation methodologies used in water chemistry research (Green et al., 2014; Sandron et al., 2015), we choose as example the resin-based solid-phase fractionation methodology (Rho et al., 2019), because its main advantages are fast processing speed (∼4.5 h), regeneration (reuse) of the involved resins, and its potential for microfluidic configuration. It consists of passing the WDOM through two cartridges positioned in line, which are packed with macroreticular cross-linked aromatic polymers (the first cartridge with the Amberlite XAD 8 resin and the second with the Amberlite XAD 4 resin) to selectively separate three main organic molecular fractions, the hydrophobic (HPO), the transphilic (TPI), and the hydrophilic (HPI) fraction, which are isolated as illustrated in Fig. 2.
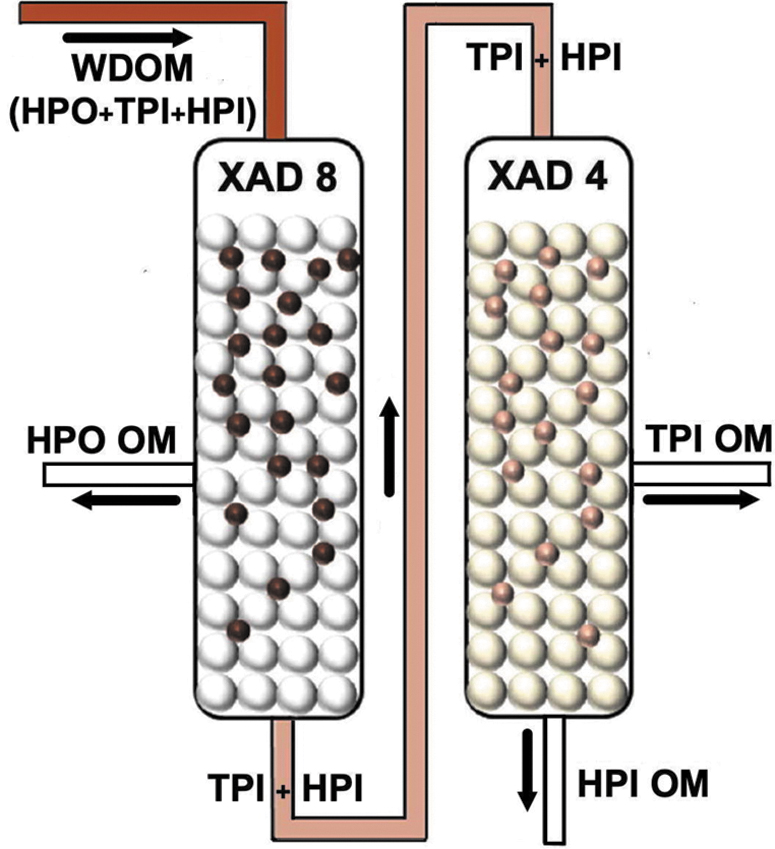
Schematic concept for the fractionation steps of WDOM into HPO, TPI, and HPI OM by sequentially passing it through the XAD 8 and XAD 4 resins (modified from Rho et al., 2019). HPI, hydrophilic; HPO, hydrophobic; OM, organic matter; TPI, transphilic; WDOM, water-dissolved organic matter.
Briefly, the HPO fraction is separated by selective adsorption onto the XAD 8 resin cartridge (and can be selectively eluted with alkaline H2O, e.g., 0.1 N NaOH). The TPI fraction [mostly containing the least HPI organic acids (Hood et al., 2005)] absorbs selectively onto XAD 4 resin cartridge (and can be selectively eluted also with alkaline H2O). Finally, the HPI fraction goes through both cartridges as it does not adsorb onto either XAD 8 or XAD 4 resins. The molecular characterization of organics in the HPO, HPI, and TPI fractions can be performed by instrumentation already developed [such as NASA's MOMA instrument (Goesmann et al., 2017)].
Catalysis has been proposed as a driving force of the evolution of primitive life. Protocells, for example, evolve by maximizing their internal molecules' catalytic activity, whereas individual molecules tend to do the opposite as they strive to boost their own relative fitness within a protocell. In other words, genetic drift and competing evolutionary pressures at various evolutionary scales cause protocell lineages to alternate endlessly between high and low intracellular catalytic activity (Takeuchi et al., 2016).
Hydrolysis-based catalysis is not only a reliable method for detecting unknown life but can also answer to the question whether extinct/extant life's biochemistry is homochiral (based on enantiomers of either L- or D-chirality) or heterochiral (based on enantiomers of L- and D-chirality). The catalytic hydrolytic chirality detection approach of the present study can identify unlimited types of organic molecules of alien life, which are unknown and unpredictable. In contrast, the current approaches search mainly for the prevailing homochirality of L- and D-amino acids, the reliability of which as biomarkers has not been established (Avnir, 2021; Georgiou, 2018; Glavin et al., 2020).
Hydrolytic catalysis may also provide an answer to the questions on the origin of homochirality in Earth life (Green and Jain, 2010) and possibly by extension to putative life beyond Earth in association with catalysis. It may shed light on catalysis/chirality interdevelopment from the early stages of life as we know it, where the proteinic enzyme-based catalysis may start and develop in its imperfect state (of low reaction rates/poor substrate selection) on a heterochiral organic molecular background (present in primitive forms of life, e.g., protocells), upon which perfect catalysis is established on a trial-and-error-based selection of a homochiral organic background (present in advanced forms of life, e.g., todays cells) (Green and Jain, 2010).
For an organic catalyst of unknown life to support accelerated reaction kinetics, it should possess some structural and/or molecular functional characteristics that would produce geometric, electronic, and confinement effects, which can influence how the reacting “substrates” approach and interact with the catalyst, as it also happens with enzymes as catalysts. That is, an EOM organic catalyst is expected to differ structurally/functionally from the so-called nanomaterial “catalysts” (e.g., CeO2, Fe3O4, MnO2, carbon nitride, Fe-doped carbon, gold nanoparticles, metal–organic frameworks), which have been named, wrongly, “nanozymes” due to their ability to disproportionate H2O2 (convert it to H2O and O2) (Scott et al., 2020). In that sense, hydrolytic catalysis of unknown life is quite possible to share certain catalytic groups that have been incorporated quite successfully in the enzymes of Earth life in its 3.7 billion years of evolution.
Most enzymes use only a small part of their protein structure for manifesting their catalytic activity, namely the active site, which is composed of one to three amino acids usually scattered within the peptide chains. When the protein folds, these amino acids have the terminal groups of their side chains converging into that site, where these groups coordinate with each other to perform the catalytic activity of the enzyme. The amino acids most frequently participating in enzyme active sites are characterized by their high catalytic propensity (Fig. 3), which is a measure of how frequently an amino acid is involved in catalysis, compared with random chance (Georgiou, 2018; Ribeiro et al., 2018).
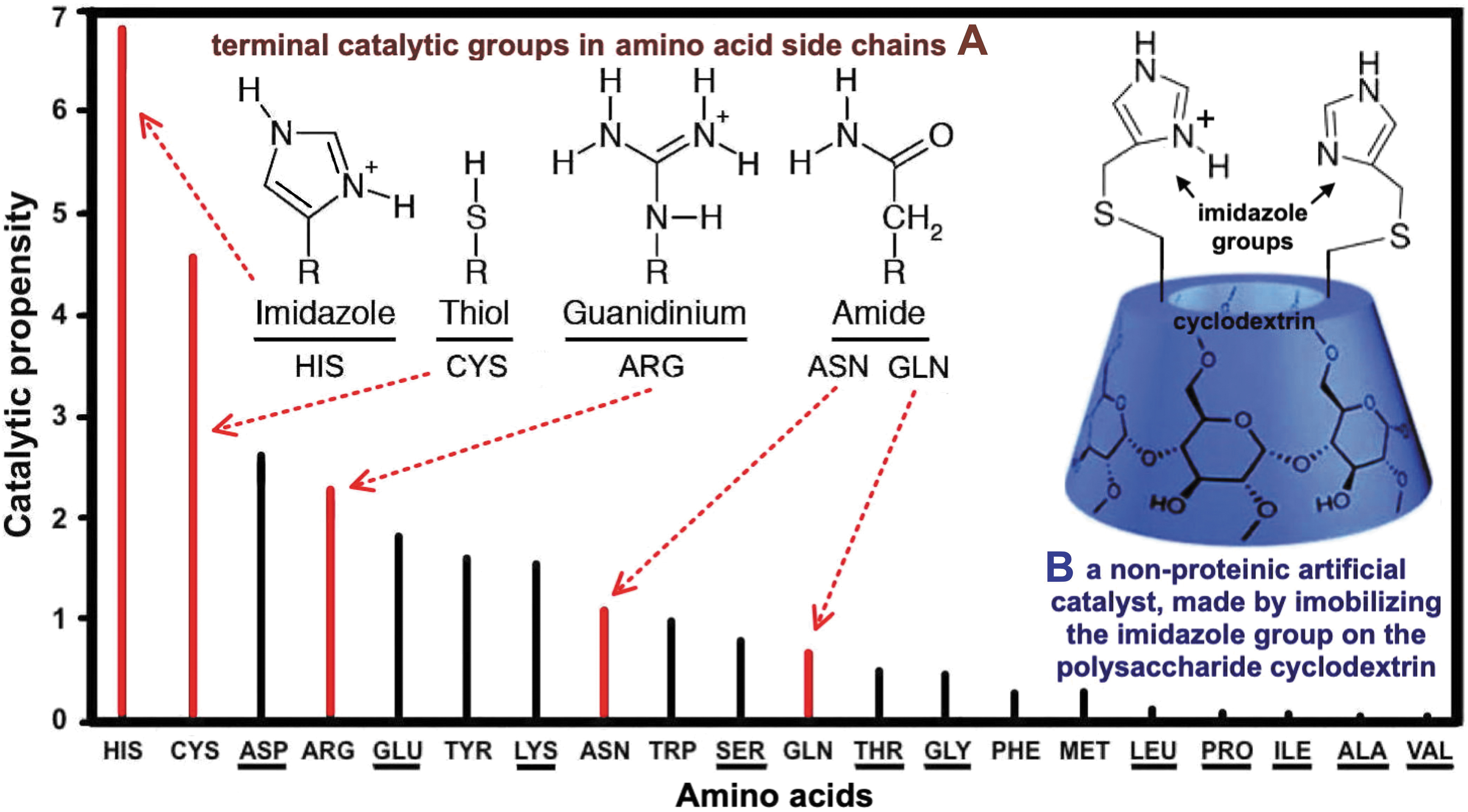
Unknown life catalytic organic groups possibly based on amino acid side chain catalytic groups. The main figure [a modification adapted from elsewhere (Bartlett et al., 2002; Georgiou, 2018)] is graphical illustration of the catalytic propensity of Earth life amino acids, including those (underlined) also found in meteorites (produced by hydrothermal prebiotic chemistry).
Figure 3 helps in setting the following criteria (a, b) for predicting molecular structures of catalysts possibly present in an EOM sample (Georgiou, 2018): (a) Catalytic organic groups, free or bound to non-amino acid-like organic molecules, which, however, are structurally same or near identical to the terminal catalytic groups present in the side chains of those biogenic amino acids that are classified in the highest range of catalytic propensities. (b) Organic molecules with trace elements bound/associated with them. Criteria a, b for identifying organic structures with a hydrolytic catalytic potential in an EOM sample are further specified as follows: (a) Unknown life catalytic groups could serve the imidazole group (in His), the thiol group (in Cys), the guanidinium group (in Arg), and the amide group (in Asn and Gln) (Fig. 3A). In order for these groups to be catalytically functional in an EOM sample, they do not need to be located in organic molecules such as the amino acids present in Earth life. For example, the catalytic potential of the imidazole group, alone, has been displayed in the designing of non-protein-based artificial enzymes, where this group has been chemically immobilized on organic compounds, such as polysaccharides (Bogliotti and Dalko, 2007; Breslow, 2006), tripeptides (Albada et al., 2008; Jarvo et al., 2002), and ribonucleoside-5′-phosphorimidazolides [in the nonenzymatic RNA copying (Walton et al., 2019)]. An important such example is the immobilization of imidazole on the polysaccharide cyclodextrin (Fig. 3B) for the design of an artificial hydrolytic catalyst with ribonuclease activity (Breslow, 2006). (b) Hydrolytic catalytic activity can be also exhibited by an EOM sample that contains trace elements such as Cr, Se, Cu, Fe, Mg, Mo, Ni, and Zn (Georgiou, 2018). In Earth life, such metals can serve as enzyme catalytic cofactors via binding to the terminal groups in the side chains of the amino acids Asp, Asn, Glu, Gln, Gly, Ser, and Cys (Mestek et al., 2012). Moreover, they have been shown to retain activity even complexed to synthetic oligopeptides. Example is the Ni-binding peptide of 13 amino acids that catalyze hydrogen production, mimicking a structure similar to that found in NiFe hydrogenases and the NiNi cluster in acetyl-CoA synthetase (Timm et al., 2023). Nonetheless, it is doubtful whether such catalysts would have survived biochemical evolution in an amino acid polymer-based alien life, unless the trace metal–organic catalysts are part of life based on polymers structurally different from Earth life. Therefore, a minimal requirement for any trace element to be considered as a potential catalysis biomarker is to be bound to organic ligands present in the EOM sample. Provision should be taken that the inorganic matter bound to organic molecules can be removed in order for the latter to be tested as control.
In any case, possible catalysts in an EOM sample can be searched in terms of molecular structures and for evaluating their synthetic complexity (Marshall et al., 2017; Johnson et al., 2018, 2020; Chou et al., 2021) by instrumentation already available, such as the NASA's MOMA instrument developed for EOM extraction/fractionation from martian subsurface (Goesmann et al., 2017).
The catalytic acceleration of basic organic reactions, such as the addition (hydrolysis) or removal (dehydration) of water is likely to be a universal feature of life regardless of the molecular structures used. Assuming that life's most fundamental, even defining, characteristic is evolutionary (e.g., Darwinian) refinement, then it is likely that highly efficient, specific catalysts are a universal feature of biology everywhere. Their synthetically complex organic structure, space constraining, metabolic loop coordination, and subjection to environmental evolutionary pressures are what distinguish them from nonlife catalysts. We have focused our search for alien life on the detection of hydrolytic catalysis because it is energetically favored and of many types, as displayed by their wide range in Earth life.
We have also presented a conceptual model for the design of an HCD instrument for use in a life detection mission. Such an instrument has clear application to worlds with habitats of past or present water, such as Mars, Europa, and Enceladus. However, life need not be only water based in terms of its hydrolytic catalyzed reactions to occur. That is, hydrolytic catalysis needs access to a water molecule only as to act as a nucleophile by attracting toward the lone pair of electrons of its oxygen a positive charge for bond cleavage to occur, regardless of water source of origin. Besides water, alkenes, sulfides, sulfoxides, amines, and phosphines can act as nucleophiles (Bach et al., 1995), thereby supporting hydrolytic catalysis in nonaqueous planetary environments.
Phosphines have been identified in the cloud decks of Venus (Greaves et al., 2020), sulfides on Mercury's surface (Renggli et al., 2022), and amines are believed to be present in aerosols on Titan (Mahjoub et al., 2012; Raulin et al., 2012). Therefore, an HCD instrument can be applicable even for worlds lacking suitable liquid water habitats such as Titan, Mercury, and Venus. Hydrolytic catalytic activity is a sufficient and useful identification criterion for agnostic life to be considered in planning or devising life detection missions.
Footnotes
Acknowledgments
We thank David Deamer for a careful review of this article, and we also thank the reviewers for their constructive and timely reviews.
Authors' Contributions
C.D.G. conceived of this study. C.M. and J.-L.R. contributed to the content and reviewed the final version.
Author Disclosure Statement
The authors declare no conflicting interests.
Funding Information
C.D.G was institutionally supported by the Department of Biology, University of Patras (Patras, Greece), and by the Greek Ministry of Education. C.M. was supported by the NASA Astrobiology Program.
Associate Editor: Norman Sleep
