Abstract
The Life Detection Knowledge Base (LDKB) is a community webtool developed to test and evaluate strategies to search for evidence of life beyond Earth, with an emphasis on recognizing potential false-positive and false-negative results. As part of the LDKB framework, we developed a taxonomy of potential biosignatures. The taxonomy brings together a broad array of life-detection strategies into a common and systematic structure that allows for equitable evaluations based on a specific set of criteria, chosen to assess the likelihood of false-positive and false-negative interpretations. The taxonomy is also a tool to organize life-detection strategies in a way that streamlines their infusion into robotic spaceflight missions. This article describes the structure of the taxonomy and its functional qualities. Two accompanying articles detail the overall LDKB framework and the set of criteria used to evaluate potential biosignatures.
Introduction
The last few decades of space exploration have revealed many potentially habitable worlds within and beyond our solar system (cf. Hand et al., 2009; Borucki et al., 2013; Grotzinger et al., 2014; Cable et al., 2020; Carrier et al., 2020). In the coming decades, robotic spaceflight missions that are currently in development will seek evidence of life on such worlds (e.g., McKay et al., 2013; Hand et al., 2017; Bolcar et al., 2018; MacKenzie et al., 2021; Quanz et al., 2022). To pursue this bold endeavor, the 2019 National Academies of Sciences, Engineering, and Medicine consensus study report (NASEM, 2019) recommended that NASA “should support the community in developing a comprehensive framework for assessment—including the potential for abiosignatures, false positives, and false negatives—to guide testing and evaluation of in situ and remote biosignatures.” Hence, there is a desire to evaluate and organize existing life-detection strategies to maximize the likelihood of detecting and recognizing evidence of life beyond Earth, if it exists.
The problem of organizing life-detection strategies into a coherent, functional framework has multiple solutions (e.g., Neveu et al., 2018; Enya et al., 2022). The task is not trivial, because astrobiology knowledge is diverse, often taking forms that do not map clearly to planetary exploration; diffuse, in that it crosses many disciplines and a wide-ranging literature; and, in some key areas, it is incomplete. For these reasons, an ideal framework is unlikely to exist. But following the NASEM (2019) recommendation, any functional framework ought to include considerations for false positives—test results that incorrectly indicate the existence of life; false negatives—test results that incorrectly indicate the absence of life; and abiosignatures—a test result that indicates abiotic processes and phenomena.
Environmental context must also be considered, as it bears on biological potential—conditions that affect the abundance, diversity, and productivity of life in an environment; and biosignature potential—conditions that affect the survivability and detectability of signatures of life in an environment. The framework should also be flexible, so that it can accommodate new knowledge, and unbiased, so that claims of potential signatures of life ultimately meet rigorous standards of proof that increase confidence and consensus in interpretations. Finally, a functional framework for life detection/evaluation that is applicable to planetary missions will inherently intersect science and technology fields; hence, it must also consider the principles and practices that apply to the design, development, and implementation of robotic spaceflight projects.
A community webtool, the Life Detection Knowledge Base (LDKB), was developed to address these challenges and the NASEM charge (Pohorille et al., 2025 in this issue). The LDKB helps test and evaluate life-detection strategies, with an emphasis on recognizing potential false-positive and false-negative results. The LDKB framework is built on two critical elements: (1) a taxonomy of potential biosignatures; and (2) a set of criteria to evaluate each potential biosignature. A potential biosignature is a measurable attribute of an object or a substance that, by itself or in combination with others, could reveal the presence of life in an environment. Examples of potential biosignatures include “enantiomeric ratios of amino acids,” the “size distribution of laminations,” or the “motion of cell-like objects.” Measurements that seek to detect potential biosignatures have four possible outcomes. A result consistent with biology is a potential biosignature, or a false positive. A result not consistent with biology is a potential abiosignature (a signature of abiotic processes) or a false negative (Section 2). The criteria used to evaluate potential biosignatures were chosen to assess the likelihood of each potential outcome in a scientifically rigorous manner based on existing knowledge.
The purpose of this article is to describe the taxonomy of potential biosignatures and its functional qualities. Two accompanying articles detail the overall LDKB framework (Pohorille et al., 2025 in this issue) and the set of criteria used to evaluate potential biosignatures (Parenteau et al., 2025 in this issue).
Taxonomy of Potential Biosignatures
A taxonomy is a system of classification that allows for organizing things (taxa) into groups or types for equitable comparisons. A taxonomy of potential biosignatures supports the primary goal of the LDKB framework to help evaluate the likelihood that a search for a potential biosignature, or group of potential biosignatures, may yield a false-positive or a false-negative result based on existing knowledge as applied to a defined set of criteria (Parenteau et al., 2025 in this issue). The taxonomy also allows identification and organization of knowledge relevant to a potential biosignature, or group of potential biosignatures, that may help explain a particular experimental outcome. For example, a result that incorrectly indicates the absence of life (false negative) could be due to an absolute lack of signal (e.g., no cell-like movement under a microscope because cells are sessile, dead, or dormant) or to a signal that falls below possible detection limits (e.g., no cell-like movement under a microscope because the size of the cells falls below the image resolution). In the first case, the false negative stems from the biological prevalence of the potential biosignature (cell-like movement); in the second case, it stems from its biological signal strength. Knowledge captured in the LDKB regarding the “motion of cell-like objects” can help address this ambiguity.
Generic structure of the taxonomy
The taxonomy consists of three main categories: (1) Chemistry; (2) Structure; and (3) Activity (Section 2.2), which are considered to be universal hallmarks of life. It is anticipated that measurable attributes of life on other worlds will fall within one of these three categories.
Each category comprises two tiers or taxonomic levels (Fig. 1). The first tier are the measurable attributes of that category. In the Chemistry category, these include chemical attributes of substances (Section 2.2.1); in the Structure category, they include physical attributes of objects (Section 2.2.2); in the Activity category, they include time-dependent processes that affect substances or objects (Section 2.2.3). The second tier are the specific substances and objects of interest that are measured. Hence, potential biosignatures in each taxonomic category are all defined consistently, and with the same level of detail, by combining one element from the first tier (the measurement of a physical/chemical feature or time-dependent process) and one element from the second tier (a specific substance or object of interest) (Fig. 1). Tier 1 and Tier 2 taxonomy entries are not necessarily all inclusive, and it is expected that their contents will expand over time through community input.
Taxonomy Definitions
Taxonomy Definitions
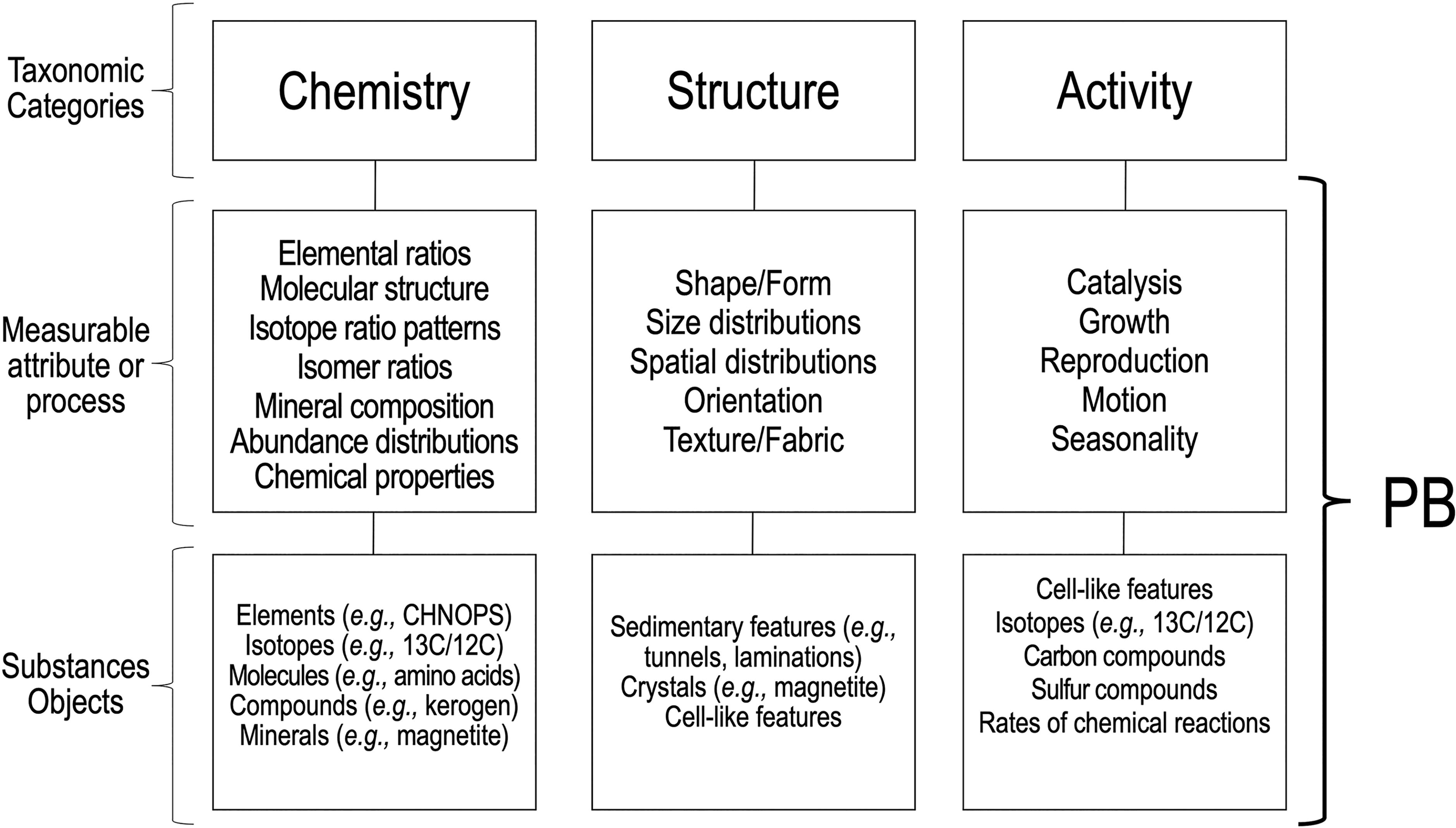
Generic structure of the taxonomy of potential biosignatures (PB). The taxonomy consists of three generic categories (chemistry, structure, activity) thought to represent universal attributes of life. Each category comprises two tiers or taxonomic levels. The first tier are measurable attributes of substances and objects or time-dependent processes characteristic of that category. The second tier are the specific substances and objects whose attributes are measured. Potential biosignatures in each category are defined by combining one element from the first tier and one element from the second tier.
While this generic taxonomic structure enables the grouping of many potential biosignatures into discrete and comparable bins, some potential biosignatures do not conform to the two-tier taxonomic structure. For completeness, specific solutions were implemented that make it possible to capture those “non-conformable” potential biosignatures (<5% of the potential biosignatures captured in the taxonomy) within the appropriate category, as explained in the next section.
Chemistry category
The Chemistry category consists of measurable attributes of substances (e.g., elements, molecules, compounds, minerals) known to be affected or modified by life (Fig. 2). Chemical synthesis in biological systems seeks to maximize fitness and adaptive utility and evolves continuously through a process of natural selection. In contrast, chemical synthesis in abiotic environments is primarily dictated by reaction kinetics and thermodynamics. This difference can result in diagnostic chemical properties of substances that serve as proxies for biological or abiotic sources, including the structure of organic molecules, the relative abundance or prevalence of certain enantiomers, or the isotopic makeup of organic and inorganic compounds. In addition, biological processes can modify the chemical composition of an environment from the cellular to the planetary scale (e.g., atmospheric gasses, chemical composition of crustal minerals and rocks) in ways that are distinguishable from abiotic processes. There are seven measurable chemical attributes in the Chemistry category.
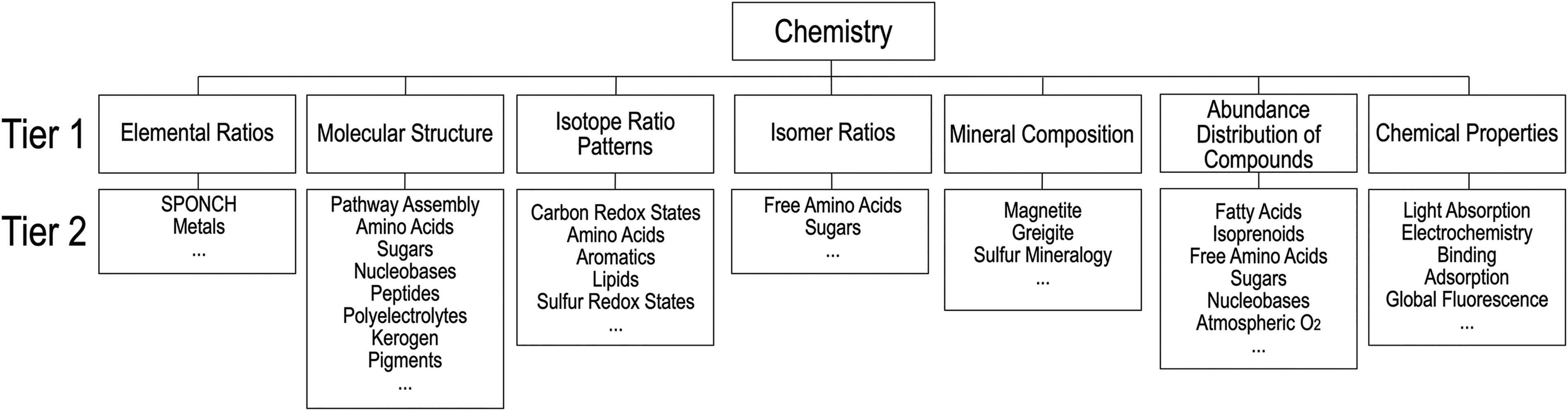
Taxonomic elements of the Chemistry category. Tier 1 and Tier 2 contents can be added without changing the overall structure of the taxonomy.
(1) Elemental ratios: The main elements composing life as we know it are carbon, hydrogen, nitrogen, oxygen, phosphorus, and sulfur (CHNOPS). Enrichments of these compounds in a sample could be indicative of biological activity, and some of these elements can display remarkably stable ratios in both living systems and the environments they occupy, such as the well-known Redfield ratio of C:N:P = 106:16:1 in ocean water (Redfield, 1958). In addition, biological activity can affect the concentrations of trace metals in the environment, and organisms can generate patterns of trace metal abundances that reflect specific aspects of their biochemistry, such as the metallome requirements of methanogenesis (Cameron et al., 2012).
(2) Molecular structure: Molecular structure refers to the three-dimensional arrangement of atoms in a substance. Living systems can produce complex molecules that are not observed in abiotic systems (Marshall et al., 2017). For example, the presence in a sample of a complex polymer with repeating units (e.g., peptide) or with a repeating charge (i.e., polyelectrolyte) could be indicative of biological synthesis (Benner, 2017). Some “building blocks” of polymers are also sufficiently complex that their presence alone could point to a biological source, such as the proteinogenic amino acids tryptophan and phenylalanine, or sugar monomers with five or more carbons in their structure. Structural richness in complex mixtures of organic compounds can also point to a diversity of molecular functions that is indicative of biological processes.
(3) Isotope ratio patterns: Isotopes of a light element typically differ noticeably in their chemical properties. Accordingly, reactions between chemical species can create patterns in the relative abundances of their stable isotopes that reflect the reaction mechanisms and environmental conditions involved. Enzymes and metabolic reaction networks create distinct isotopic patterns (Hayes, 2001). For example, the biosynthesis of straight-chain lipids involves isotopic discrimination during the formation of 2-carbon subunits that are then linked together, creating alternating different 13C/12C values along the lipid's carbon backbone (e.g., Monson and Hayes, 1982). Isotopic differences between groups of biogenic amino acids reflect each group's biosynthetic pathway, and these pathways are highly prevalent in life (e.g., Scott et al., 2006). But isotopic patterns should be viewed as additional pieces of evidence to aid interpretations of other potential biosignatures. Interpretations of isotopic patterns among organic compounds, minerals, and other features require observations of associated environments, processes, and geochemical reservoirs (e.g., Des Marais, 2001; Lloyd et al., 2020). Also see Shkolyar et al. (2025 in this issue) for discussion of how environmental context of the biosignature is crucial to Structure category biosignatures as well. Investigating isotope ratio patterns promotes an interdisciplinary approach toward determining environmental contexts that help interpret all types of potential biosignatures (e.g., Chan et al., 2019).
(4) Isomer ratios: Isomers are compounds with the same formula but a different arrangement of atoms in the molecule, which can confer different properties. There are two general types of isomers: (1) Constitutional (or structural) isomers and (2) Stereoisomers (which include diastereomers and enantiomers). A hallmark of living systems is their selectivity toward specific isomers involved in biochemical roles. This selectivity can lead to some readily identifiable and measurable molecular qualities that can be used as potential biosignatures (Summons et al., 2008). For example, enantiomers are pairs of the same molecule that are mirror images of each other. Life on Earth often displays a preference toward specific enantiomers (e.g., amino acids; sugars) to build functional polymers (e.g., peptides; nucleic acids), which can result in high levels of enantiomeric excesses in a sample, which might serve as signatures of biological activity (Avnir, 2021). Abiotic organic synthesis tends to form equal mixtures of enantiomers in the absence of an asymmetric chiral influence (i.e., a physical or chemical process that favors a specific enantiomer), although abiotic organic mixtures, such as those found in carbonaceous chondrites, sometimes display enantiomeric imbalances, which point to abiotic mechanisms of enantioselectivity (e.g., Glavin et al., 2020).
(5) Mineral composition: Biology can affect the chemical composition of crystalline substances in ways that might be distinguishable from abiotic processes. For example, certain organisms precipitate minerals whose properties serve important biochemical/physiological functions, such as magnetosomes in magnetotactic bacteria (Faivre and Schüler, 2008). Evolutionary pressures can optimize those functions by affecting the chemical composition of the minerals involved (e.g., by increasing chemical purity) to extents that are not observed in their abiotic counterparts. In addition, biologically induced and biologically controlled mineralization can result in “out-of-equilibrium” minerals and mineral assemblages (e.g., coexisting reduced and oxidized phases) that may be difficult to explain by abiotic processes alone.
(6) Abundance distribution of compounds: Living systems utilize only a small subset of monomers, such as amino acids, sugars, and nucleobases, to synthesize more complex polymers such as peptides, saccharides, and nucleic acids. The relative abundance of each monomer within that subset depends on the structure and functionality of polymers, which are the result of evolutionary adaptations. In contrast, when the same monomers are produced abiotically their relative abundance is a function of their energy of formation, so smaller molecules tend to dominate. The different patterns in the relative abundance of molecules within a certain class of compounds can therefore be used to discriminate biotic sources from abiotic ones (Dorn et al., 2011). Measurements of abundance distributions can also be conducted on planetary scales. For example, the relative abundance of gases such as O2 and CH4 in an atmosphere can be indicative of biological processes driving atmospheric composition out of equilibrium.
(7) Chemical properties: This taxonomic category is an outlier in that it does not conform to the Tier 1 (chemical parameters) and Tier 2 (substances) structure. However, it was included in the taxonomic scheme to capture physical/chemical phenomena indicative of compounds that can serve important biochemical or physiological roles (e.g., light absorption, electrochemical properties). Measurements of those chemical properties can point to the presence of life even if the compounds are unknown (the physical/chemical phenomena can sometimes be measured remotely without the need for compositional analyses). For example, absorption of light within specific regions of the solar spectrum could be indicative of a substance that can capture and convert light energy into chemical energy, akin to biological pigments such as chlorophyll. Such measurements are often considered in the context of searching for spectral evidence of life on exoplanets by means of remote sensing methods.
The Structure category consists of physical attributes of objects known to be produced or influenced by life and includes life's morphological traits, such as the 2D shapes, 3D forms, and size ranges of organisms (or their specialized structures), and their spatial distribution (Fig. 3). Biological behavior and metabolic processes can modify or influence the physical appearance of a feature (e.g., a stromatolite) at the time of its development from the microscopic to the macroscopic scales. Deciphering whether morphological features whose characteristic size range/distribution, spatial distribution, shape, orientation, or texture/fabric can be indicative of biological synthesis or biological influence will be enhanced when multiple morphological attributes are considered in combination and when structural attributes are combined with chemical and activity attributes (Section 4). Structural attributes of potential biosignatures are especially useful when they are obtained in an environmental context. There are five measurable properties in the Structure category.
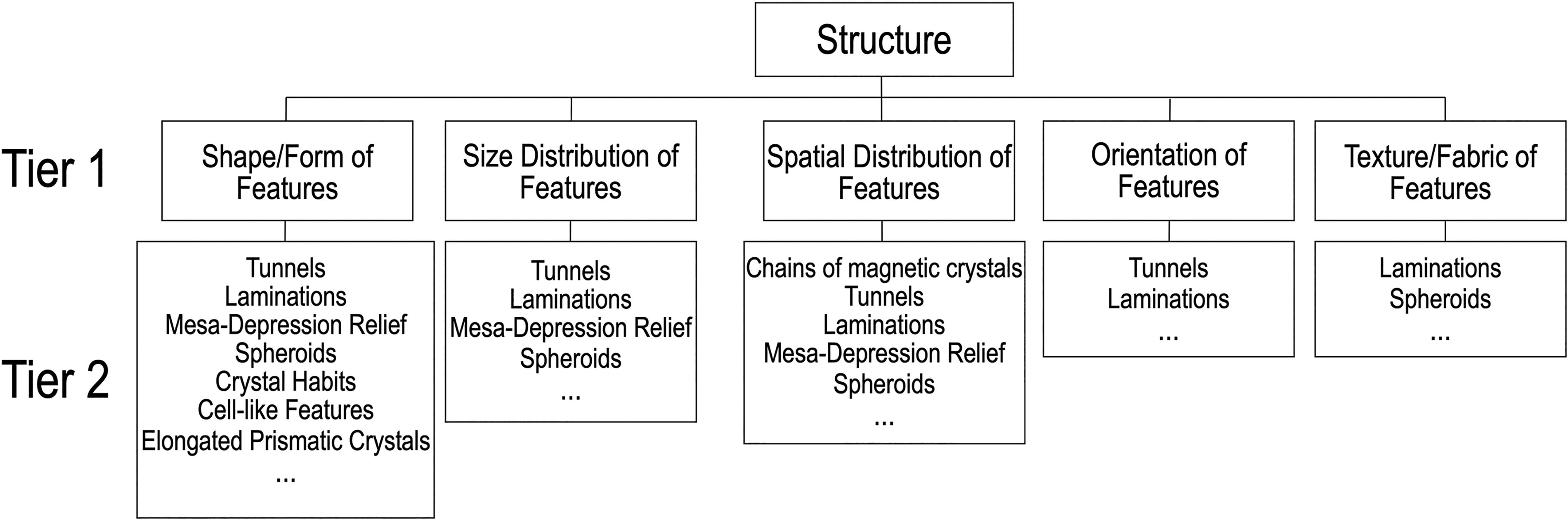
Taxonomic elements of the Structure category. Tier 1 and Tier 2 contents can be added without changing the overall structure of the taxonomy.
(1) Shape/Form of features: Shape/form is the 3D geometric shape or habit of a volumetric structural feature (e.g., crystals) or the 2D or 3D relief or topographic patterning of a potential biogenic object or feature on a surface. For example, cocci (i.e., spherical cells) represent a morphotype of unicellular organisms that can be distinguished from other cellular morphotypes, such as rods and filaments. Upon burial and compaction, coccoidal microfossils may undergo deformation to resemble ruptured or deflated (e.g., raisin-like) spheres, or they may become flattened. Abiotic processes commonly produce objects with spherical morphologies that, like microbial cells, maximize their surface area/volume ratio, such as the “blueberries” on Mars (Squyres et al., 2004). Abiotic processes can also produce curved and sinuous features whose morphologies resemble cells and their fossilized equivalents (e.g., García-Ruiz et al., 2017; McMahon and Cosmidis, 2022). Interestingly, such objects can display many biomimetic attributes, including physical (e.g., helical twisting, hollow or porous tubules, segmented and pinching/swelling ends of filaments) and chemical features (e.g., organic films and membranes) (cf. McMahon and Cosmidis, 2021).
(2) Size range/distribution of features: This includes the size range of individual features as well as the distribution of the sizes of a feature. For laminations, for example, this can refer to the range of laminae thicknesses, measured in cross-section parallel to the lamina growth direction, and the distribution of those thicknesses in a particular volume of sediment. For tunnels, this can refer to the range of their lengths and the distribution of the range of tunnel diameters and lengths for chemically distinct rocks.
(3) Spatial distribution of features: Spatial distribution refers to the distribution of a physical object in three dimensions (3D), which can be expressed in a two-dimensional (2D) cross section as the proportion of a surface covered by a feature within an area of interest. This includes patterns on surfaces and the spatial frequency (including repetitions) of distinctive features that intersect rock/sediment surfaces. Observable features in this group include crystalline (minerals) and non-crystalline (mineraloids) aqueous/amorphous precipitates, microbial-like objects, sedimentary structures that resemble biofilm surfaces, and so on. For example, in basalt, the spatial distribution of tunnels can be along fractures and start at voids, which provide an opening for water and/or microorganisms to enter a host rock.
(4) Orientation of features: Orientation is the position of a feature relative to its surroundings or another parameter in its depositional environment. Orientation includes linear arrangements of features, minerals, cell-sized objects, and 3D morphological spatial arrangements across scales. One example is tunnels in igneous and metamorphic rock, which can be positioned relative to fractures, vesicles (bubbles), minerals, or other tunnels. As an example, tunnels that reside at the margins of a vesicle mine the glass for energy and material since tunnel orientation allows the cells to continually encounter fresh glass. Abiotic tunnels around olivine follow lines of stress created by the differential thermal shrinkage of olivine and glass (McLoughin et al., 2010). Another example includes filamentous cell-like objects that have a vertical or horizontal orientation relative to a planar surface, like the orientation of viable cells that produce a benthic “carpet” in a microbial mat.
(5) Texture/fabric of features: The primary texture of sedimentary rocks (detrital and chemical) refers to the attributes that arise from grain (or aqueous precipitate for chemical sediments) size, grain shape (form, roundness, and surface texture/microrelief), and fabric, which refers to grain orientation, grain-to-grain relationships, and grain packing (Boggs, 2011). An understanding of the primary texture of sedimentary rock is essential for proper interpretation of its depositional environment and transport history. Though genetic interpretations of primary sedimentary textures remain challenging, they are often a defining characteristic of a deposit. Fabric is the spatial and geometric configuration of all elements and features of a sedimentary rock, that when formed in the presence of a microbial community can include grains, minerals, aqueous precipitates, microbial fossils, cellular and extracellular relics, organics, and other rock fragments. For example, biogenic stromatolites can display distinctive fabrics and textures, and the combination of such component parts may reveal attributes and visual patterns that can be diagnostic of a biological origin or influence (Grey and Awramik, 2020). The antithesis of these visual features would be layering that may look like a typical stromatolite but is not part of a photosynthetic or other biological process.
The Activity category reflects time-dependent processes that are the result of life's exchange of matter and energy with the environment (Fig. 4). This is manifested in measurable, time-dependent chemical and physical phenomena (e.g., movement of an object). In addition, biological systems can efficiently catalyze specific chemical reactions at rates significantly higher than abiotic processes, and the activity of biological systems can be modulated by environmental factors in ways that are predictable (e.g., seasonal changes) or within well-defined margins (e.g., temperature ranges). The boundaries of the Activity category may appear ambiguous since biological activity can be manifested in the form of chemical compounds or structural features. This category encompasses changes that a substance or structure undergoes over time, but not the substance (Chemistry category) or structure (Structure category) itself. There are currently five physical/chemical processes in the Activity category.
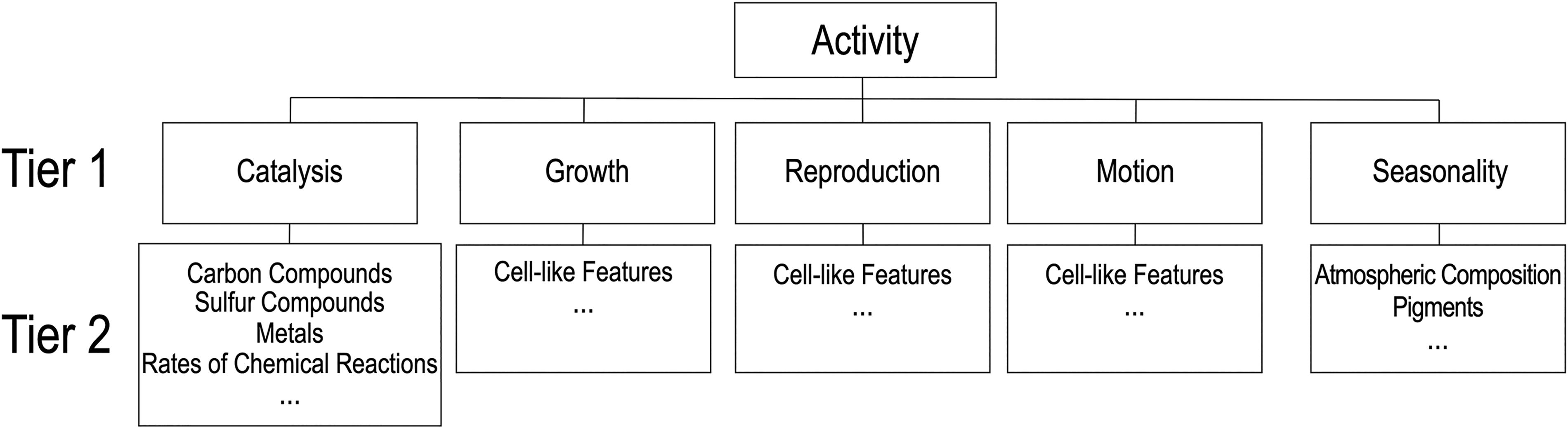
Taxonomic elements of the Activity category. Tier 1 and Tier 2 contents can be added without changing the overall structure of the taxonomy.
(1) Catalysis: Catalysis describes a hastening of chemical reactions leading to a lesser amount of time needed to reach chemical equilibrium between reactants and products at given environmental conditions. For example, reduction of CO2 to CH4, even if the latter is the thermodynamically stable species, can be a protracted process in the absence of life and at normal temperature (e.g., Etiope and Sherwood Lollar, 2013) but is much faster if living organisms deriving energy from this reaction use enzymes to accelerate its completion. Catalysis also includes the production of compounds whose synthesis requires energy input, hence moving away from chemical equilibrium in ways that are not observed in abiotic environments.
(2) Growth: Growth describes the rate of change in size of a physical object. Organisms, or their structural components, typically change in size during a natural life cycle. Populations of organisms (e.g., a biofilm, a forest…) can also change in size at rates that can be predicted based on physical and chemical conditions. Again, the boundaries between growth and reproduction can be fuzzy (or even interchangeable as in microbiology), and for certain organisms, changes in size can be significant during certain stages of the life cycle, also leading to changes in morphology, like in the case of reproduction.
(3) Reproduction: Reproduction describes the rate of increase in numbers of a physical object. The most immediate biological example is cell division, which can occur at rates that are predictable based on physical and chemical conditions (e.g., temperature, pH, nutrient availability). Sometimes, biological reproduction results in distinct morphologies such as that observed during different reproductive stages and modes, like binary fission or budding. Measurements of those morphologies would fall under the Structure category, despite their link to reproduction. This highlights the fact that the boundaries between taxonomic entries are not always sharp and that combinations of potential biosignatures (i.e., change in numbers plus morphology) often increase their diagnostic power (Section 4).
(4) Motion: Motion describes the change in position of a physical object. The changes in direction of motile organisms appear purposeful and are inconsistent with drift or Brownian motion (Nadeau et al., 2018). Biological motion can be interrupted by changes in temperature, radiation, or addition of chemical compounds. Certain organisms navigate in response to various physicochemical parameters (taxis) including light, magnetic fields, or chemical gradients, which are testable means to differentiate biological activity from abiotic phenomena.
(5) Seasonality: Seasonality describes changes in the properties of an environment in response to cyclic or periodic changes in environmental conditions, for example, seasonal changes in the wavelength-dependent reflectance of a planetary surface or in its atmospheric composition as a function of the periodic variation in the distance to its star or the latitude of its subsolar point (e.g., Olson et al., 2018). Measurements of seasonality (and growth) are often applied to the search for evidence of life at planetary scales (e.g., exoplanets) by means of remote sensing methods.
The taxonomy meets several important functional requirements. One of the key requirements is to facilitate the infusion of life-detection knowledge into the design and implementation of spacecraft missions (Section 5). Four additional qualities were considered during the development of the taxonomy: (1) Durability, (2) Flexibility, (3) Granularity, and (4) Neutrality.
Durability: The taxonomy must be stable and resilient against significant reconstructions over periods of time relevant to the life cycle of spaceflight missions, which might encompass several decades from inception to execution. This trait is important to ensure continuity as the scheme evolves through community input, research advances, and the challenges faced with each potential discovery of ancient terrestrial life or extraterrestrial life.
Flexibility: While stability is needed for durability, the taxonomy must still be able to absorb new knowledge derived from the study of life on Earth and new potential biosignatures that become accessible through technology development, without altering the underlying architecture.
Granularity: Potential biosignatures are defined at a similar level of detail using the same general language “[physical/chemical parameter] + [object/substance]” (Section 2.2). This conformity allows for equitable evaluations of the likelihood of false-positive and false-negative signals in a particular environment, and for meaningful comparisons, even between potential biosignatures that are seemingly unrelated. Defining each potential biosignature with the same level of detail also allows assessment of the combinatorial power of different potential biosignatures (Section 4).
Neutrality: The overall taxonomy must not imply an intrinsic hierarchy that could denote a preference toward specific potential biosignatures, desired measurement outcomes (i.e., biotic vs. abiotic interpretations), or some other perceived or unconscious bias (e.g., technical feasibility, historical treatment, dominant field, natural abundances). Thus, each potential biosignature is evaluated exclusively in terms of a specific set of criteria chosen to assess the likelihood of a false-positive or false-negative signal in a particular environment (Parenteau et al., 2025 in this issue).
Combinatorial Power of Potential Biosignatures
A search for evidence of life is rarely confined to measurements of one physical or chemical attribute of a substance or an object. Rather, multiple, independent lines of evidence must be combined to reliably establish or rule out biogenicity (NASEM, 2019; Perl et al., 2021; Shkolyar et al., 2025 in this issue). Because potential biosignatures captured in the taxonomy are defined with a similar level of detail (granularity), the LDKB framework can be used to evaluate the combinatorial power of different potential biosignatures and to determine how combinations thereof affect interpretations of biotic or abiotic phenomena or modify the likelihood of false positives or false negatives. In that sense, potential biosignatures are like atoms that interact to form the molecules that represent unambiguous evidence of life. Like in the case of atoms, there might be potential biosignatures that interact and combine with greater affinity, perhaps due to their biological relatedness (e.g., sedimentary structures combined with organic content). The LDKB framework can highlight known affinities or help establish new ones.
For example, enantiomeric ratios can be diagnostic of the origin of amino acids, but there is little consensus on where abiotic ratios end and biotic ratios begin; indeed, they can overlap. Certain meteorites are known to contain amino acids that show high enantiomeric imbalance (Glavin et al., 2010; Pizzarello et al., 2012), and both biotic and abiotic amino acids can racemize over time, eroding or completely erasing any original enantiomeric excess. For those reasons, independent measurements of several measurable attributes are needed to establish the origin of amino acids, including their abundance distributions and their isotopic compositions and ratios (12C, 14N, and H; Glavin et al., 2020). Each of these measurements represents one potential biosignature in the LDKB framework, and all three measurements are captured in the Chemistry category (see Fig. 2). Their combination strengthens the interpretation of the results and lowers the likelihood of false negatives and false positives.
Some entries in the Structure category are particularly amenable to combinatorial solutions, because structural information alone is rarely definitively diagnostic of biogenic origin or input (e.g., Cady et al., 2003). Structural features must often be complemented with chemical information (Chemistry category) and time-dependent information (Activity category) whenever such multiple lines of evidence can be obtained. For example, establishing the origin of sedimentary textures and/or fabrics might require a series of analyses of their environmental context and chemical composition; attributes of their laminated structure; the presence, type, and distribution of organic matter; the depositional context; and the degree of secondary or taphonomic alteration.
Combinations of potential biosignatures inform the types of technologies required to conduct more complete and robust life-detection experiments. They provide guidance on the types of instruments that must be developed and how they must interface with each other. This may be trivial when conducting life-detection experiments on Earth, where samples can be analyzed in multiple laboratories taking advantage of the full range of sample preparation methods and analytical techniques. But the search for evidence of life beyond Earth is different in that robotic spaceflight missions are handicapped by stringent limits on payload and resources (e.g., power, data), and once a mission launches, there is limited flexibility to respond to unexpected results or discoveries. The need to conform to the fundamental principles and practices that apply to all spaceflight projects is one of the main challenges in streamlining life-detection knowledge for implementation in planetary exploration. The taxonomy seeks to alleviate this challenge, as discussed in the next section.
Taxonomy of Potential Biosignatures and Science Traceability
Before a robotic spacecraft launches to another planet seeking evidence of life, mission planners must establish a clear and direct relationship between the types of measurements needed to achieve the mission goal(s), the instrumentation required to perform those measurements, and the type and quantity of data that must be obtained. In robotic spaceflight projects, the increasing detail in the requirement flow-down, from goals to data, is captured in the form of a science traceability matrix (STM) (Weiss et al., 2005).
In a practical way, the STM reflects the completeness of a robotic mission concept, and it typically adheres to a template provided in mission Announcements of Opportunity. The template is a relatively rigid framework in which to summarize the relationships between high-level objectives, measurement requirements, instrument requirements, and spacecraft requirements to expected data products and publications. All that information must be captured in a way that is succinct and precise (Table 1, top).
Top: Example Science Traceability Matrix from the 2018 Revision of the Standard Mission Announcement of Opportunity from NASA's Science Mission Directorate. Reproduced from SOMA (2018). Bottom: Simplified Example of a STM Highlighting the Entry Point for Content Captured in the Taxonomy (Gray Boxes).
Top: Example Science Traceability Matrix from the 2018 Revision of the Standard Mission Announcement of Opportunity from NASA's Science Mission Directorate. Reproduced from SOMA (2018). Bottom: Simplified Example of a STM Highlighting the Entry Point for Content Captured in the Taxonomy (Gray Boxes).
In the example STM, Science Goals (column 1) are broadly defined, long-term, and “high-value” and are quoted from relevant NASA and NASEM documents (e.g., determine whether life exists elsewhere in the Solar System). Science Objectives (column 2) are narrow, short-term, measurable actions to achieve the stated goals (e.g., search for evidence of life in martian sedimentary deposits). Scientific Measurement Requirements (column 3) are qualitative or quantitative analyses used to address the science objectives. Measurement requirements are split into Physical/Chemical Parameters (column 3A)—measurable attributes—and Observables (column 3B)—measurable signal that contains information on physical parameter properties. Because information in a STM must be traceable and flow logically, measurement requirements are typically defined in terms of one physical/chemical parameter and its corresponding observable (Table 1, bottom). This partitioning of measurement requirements into discrete STM entries is key to understanding the technical resources needed to address each objective and to evaluating the scientific consequences of any science descopes, for example due to cost, mass, or power constraints.
Transferring life-detection strategies to a STM format is far from straightforward. As explained in Section 4, evidence of life is rarely confined to measurements of one physical or chemical parameter and its corresponding observable. For example, recognizing a sedimentary feature as a biofabric (regardless of whether it is associated with a larger structure such as a stromatolite) often requires morphological characterizations, details of its internal structure (fabric), and chemical information (organic, isotopic, and mineralogical). Each one of these analyses must be bookkept in a STM as a separate measurement requirement (a physical/chemical parameter and an observable pair) so that the associated instrument requirements can be adequately captured. This approach makes it clear that measurements of morphology, internal structure, and chemical composition require distinct instrumentation (see Table 1, bottom).
The taxonomy of potential biosignatures naturally deconstructs life-detection strategies into single physical/chemical parameters (e.g., shape/form of sedimentary features; texture/fabric of sedimentary features; mineralogy of sedimentary features) that can be directly transferred to column 3A of a standard STM. This entry point is critical as it represents the transition from science requirements to instrument requirements. Each potential biosignature (or physical/chemical parameter) entry in the STM has an associated observable (column 3B) that is method-specific. The taxonomy does not prescribe observables, because the same physical/chemical parameter might be observed with different methods (e.g., the abundance distribution of amino acids and their enantiomer ratios can be observed with fluorescence intensity and with mass spectrometry). However, the taxonomy can help identify technology gaps in life-detection science. For instance, of the chemical parameters needed to establish the origin of amino acids, such as enantiomer ratios; abundance distributions; and C, N, and H isotopic compositions (Glavin et al., 2020), the first two can be observed with flight instruments (e.g., Creamer et al., 2017; Mora et al., 2020), but the technology to measure the isotopic composition of amino acids is still at a low readiness level. The taxonomy provides a template to quickly evaluate similar technological gaps.
We have described a taxonomy of potential biosignatures. This taxonomy comprises three overarching categories that correspond to hallmark attributes of life: Chemistry, Structure, and Activity, each including two subordinate tiers: (1) physical or chemical attributes or processes and (2) substances or objects. A tier 1 element plus a tier 2 element constitutes a potential biosignature. This organization allows one to (1) evaluate the likelihood of false-positive and false-negative results in a given environment (Pohorille and Sokolowska, 2020; Pohorille et al., 2025 in this issue), (2) evaluate the combinatorial power of different potential biosignatures, (3) carry out 1 and 2 without emphasizing the preconception that some potential biosignatures are intrinsically more valuable than others, (4) expand the taxonomy as new knowledge becomes available while maintaining its long-term consistency, and (5) assess the technical requirements needed to seek each potential biosignature. Ultimately, the taxonomy seeks to facilitate infusion of life-detection knowledge in the design of robotic missions.
Footnotes
Abbreviations Used
Associate Editor: Lewis Dartnell
