Abstract
In recent years, strong evidence has emerged indicating the potential habitability of the subsurface of Mars. Occasional discharge events that bring subsurface fluids to the surface may carry with them the biological traces of subsurface organisms. Similar events are known to take place on Earth and are frequently associated with long-term mineralogical preservation of organic material, including DNA. Taking advantage of this process may allow for the development of life-detection strategies targeting biosignatures from the more habitable subsurface environment without the need for direct subsurface exploration. To test the potential for this approach to life-detection, we adapted a protocol to extract microbial DNA preserved in carbonate rocks and tested its efficacy in detecting subsurface organisms at a Mars analog site in southeastern Utah, USA, using samples from ancient and modern carbonate deposits associated with natural and artificial springs. Our results indicated that DNA from deep-subsurface organisms preserved in carbonate deposits can remain recoverable for up to 100,000 years, supporting life-detection strategies based on the detection of deep-subsurface biosignatures in surface-exposed rocks on Mars.
Introduction
The deep subsurface of Mars has been identified as a priority for life-detection efforts targeting both extant and extinct life. Mounting evidence supports the high potential habitability of the martian subsurface where, in contrast to the inhospitable surface, conditions are likely to be more favorable to life. In particular, the martian subsurface may offer protection from the ionizing radiation and freezing conditions present on the surface (Horne et al., 2022).
Perhaps most importantly, current estimates suggest that liquid water may be present in the subsurface at kilometer-scale depths (Carrier et al., 2020). Where water is present in liquid form in the subsurface, water-rock interactions may provide an energy source for microbial rock-hosted life (Schrenk et al., 2013; Lau et al., 2016). Finally, the deep subsurface of Mars is likely to have a considerable longevity, thereby creating a persistent and stable environment where life, if present, may have survived to the present day (Michalski et al., 2013).
Growing interest in the martian subsurface is fueled, in part, by recent advances in our understanding of terrestrial subsurface biodiversity. In recent years, it has become clear that the deep subsurface of the Earth harbors a great diversity and abundance of microbial life (Edwards et al., 2012). Subsurface lifeforms have been discovered multiple kilometers into the subsurface, and estimates suggest that the deep biosphere may contain a significant portion of the Earth's total biomass (McMahon and Parnell, 2014; Purkamo et al., 2020). While the terrestrial subsurface biosphere is sustained in part through chemical energy derived from the surface, underground water-rock interactions generate an energy source for chemolithotrophic microbial primary producers that are not dependent on the surface biosphere. For example, the interaction of liquid water and ultramafic rocks, known as serpentinization, produces hydrocarbons that can be used in subsurface microbial metabolism (Schrenk et al., 2013). Evidence suggests that this process likely occurred on Mars in the past and may continue in the present (Vance and Melwani Daswani, 2020).
The capacity of the martian subsurface for meeting key habitability requirements and its potential similarity to the subsurface of Earth make it a top priority for life-detection efforts. Indeed, a number of mission concepts and exploration strategies targeting this environment are currently proposed or underway, including strategies focused on drilling directly into subsurface environments to search for evidence of extant life, as well as strategies targeting surface-exposed rocks that may preserve evidence of past life (Onstott et al., 2019; Carrier et al., 2020). The latter strategy has several important advantages. Surface-exposed rocks are directly accessible, negating the need for complex drilling operations. Habitable conditions in the subsurface are likely to be vertically stratified, with the most habitable conditions occurring kilometers beneath the surface where liquid water may be present. While future drilling missions may be capable of sampling this depth, they would also be limited in the number of sites that could be sampled over the mission lifetime.
By targeting surface-exposed rocks, candidate sites from many locations across a broad spatial scale can be identified, characterized, and prioritized before sampling, both in situ through surface exploration and from orbital observations (Onstott et al., 2019). Because habitability may be heterogenous in the subsurface, diversifying the number of target sites is a significant advantage. Habitability may also be heterogenous over time, such that environments may have been habitable in the past but are no longer capable of supporting life, or potentially have had sporadic bursts of habitability throughout time. Therefore, targeting sites where recorded evidence of past life may have accumulated over time is also a priority.
Ancient subsurface-associated rocks dating to millions or billions of years ago, which may preserve fossilized records of ancient life, can become exposed at the surface through different processes such as erosion, meteoric impact, or faulting (Hays et al., 2017; Onstott et al., 2019). However, more recently formed rocks may also occur at or near the surface in conjunction with mineral precipitation associated with groundwater flow to the surface. Although the extent of groundwater flow to the surface on present day Mars is an area of active debate, such events may occur sporadically in association with fracture networks, impact events, or recurring slope lineae (Osinski et al., 2013; Ojha et al., 2015). Terrestrial examples of this phenomenon are common and include hot and cold spring systems, where groundwater is driven to the surface along fracture networks by hydrothermal processes or the buildup of deeply sourced gasses.
Mineral precipitates that form as a result of groundwater outflow can trap and entomb organisms present in the subsurface fluids, preserving biosignatures in the form of morphologic structures or biomolecules (Bosak et al., 2021). If recent precipitates formed in this manner can be located on Mars, it may be possible to extract biomolecules or genetic material from any preserved subsurface organisms directly from the surface-exposed rock, offering an alternative to directly targeting subsurface environments.
Previous studies provide substantial evidence for the preservation of subsurface biosignatures such as morphological structures, lipids, and isotope composition, which are recorded in the geological record of Earth dating back billions of years (Onstott et al., 2019; Bosak et al., 2021). However, complex biomolecules such as DNA may be preserved and remain detectable in more recently formed deposits.
DNA is a top priority target for biosignature detection because it is ubiquitous within terrestrial organisms and would provide unambiguous evidence of life (Parnell et al., 2007). Therefore, recently formed mineral precipitates associated with subsurface outflow deserve attention as potential targets for astrobiological investigations. To assess the utility of this approach, it is critical to evaluate whether genetic material from subsurface organisms can be preserved and detected in surface-exposed rocks at terrestrial sites analogous to those that might be found on Mars.
In the present study, we investigated the potential for detecting genetic material from subsurface organisms preserved in surface-exposed carbonate precipitates dating back 100,000 years. Our study evaluated three questions relating to DNA trapped within carbonate precipitates: (1) Can DNA be extracted from the protected interiors of carbonates? (2) Can DNA be preserved inside the carbonates over timescales spanning at least 100,000 years? (3) Is DNA at least partially derived from microorganisms transported from subsurface environments? We evaluated these questions using a series of meteogene travertine deposits in southeastern Utah, USA, located at a Mars-analog site called Crystal Geyser. The carbonaceous travertine deposits result from subsurface fluid discharge events at different time points over the past ∼100,000 years (Burnside et al., 2013). The subsurface fluid originates from a system of stratified subsurface aquifers, including both shallow and deeply sourced subsurface fluids rich in dissolved salts and carbon dioxide, analogous to that which might be found on Mars.
Due to the presence of active microbial communities on rock surfaces, we developed a methodology for evaluating whether DNA can be preserved within carbonates that involves the addition of a surface DNA tracer as a control, followed by the sterilization and removal of surface DNA. We used this sterilization method on carbonate samples collected from both modern and ancient travertine deposits and then extracted DNA preserved within the minerals to evaluate the longevity of preserved DNA in this system.
Importantly, terrestrial travertine deposits are expected to contain organisms originating from both surface and subsurface environments. We used our procedure to evaluate whether subsurface organisms can be detected in carbonates precipitated at the surface, and in this context, our approach may have applications to life-detection strategies for Mars.
Materials and Methods
Study site and sampling
Samples were collected from the area surrounding Crystal Geyser in southeastern Utah (38.938304, −110.135467, ±500 m). The site consists of both active and inactive spring-associated travertine deposits, connected to a system of stratified shallow (200–450 m) and deep (>1500 m) subsurface aquifers by fault lines and artificial bore holes (Shipton et al., 2004; Wilkinson et al., 2009). The inactive deposits result from natural eruptions occurring over the past 400,000+ years.
The active deposit, Crystal Geyser, is situated directly on top of one of the inactive deposits (L2) and resulted from drilling activity in 1935 (Baer and Rigby, 1978; Burnside et al., 2013). A total of eight rock samples were collected from four separate travertine deposits. These included five “modern” rock samples from the active deposit Crystal Geyser (labeled S1–S5), and three “ancient” rock samples from the inactive deposits (labeled L2–L4).
The ancient deposits had the following ages as estimated by Burnside et al. (2013): L2: 5.4 ka; L3: 50.8 ka; and L4: 103.1–113.9 ka (Fig. 1). Rock samples were collected with a rock hammer, placed into sterile bags, and stored on ice during transportation followed by storage at −80°C on returning to the university.
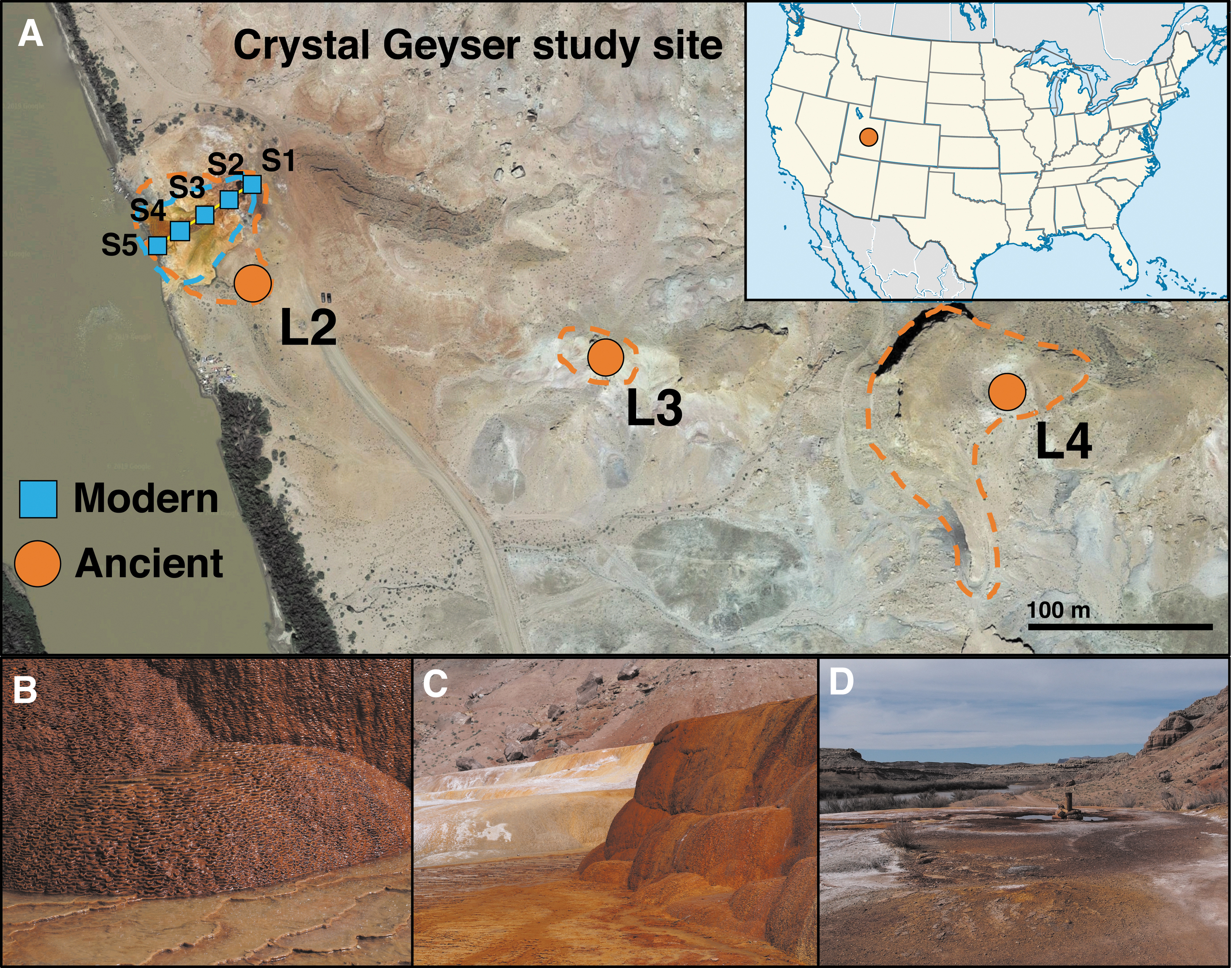
Crystal Geyser study site, in southeastern Utah.
All samples were subjected to a two-by-two factorial approach involving a combination of DNA spiking and a sterilization procedure (described below). Rock samples were subdivided into four subsamples, each receiving one of four treatments: (1) non-spiked, sterilized (“sterilized”); (2) non-spiked, non-sterilized (“control”); (3) spiked, sterilized (“spiked-sterilized”); and (4) spiked, non-sterilized (“spiked”) (Fig. 2).
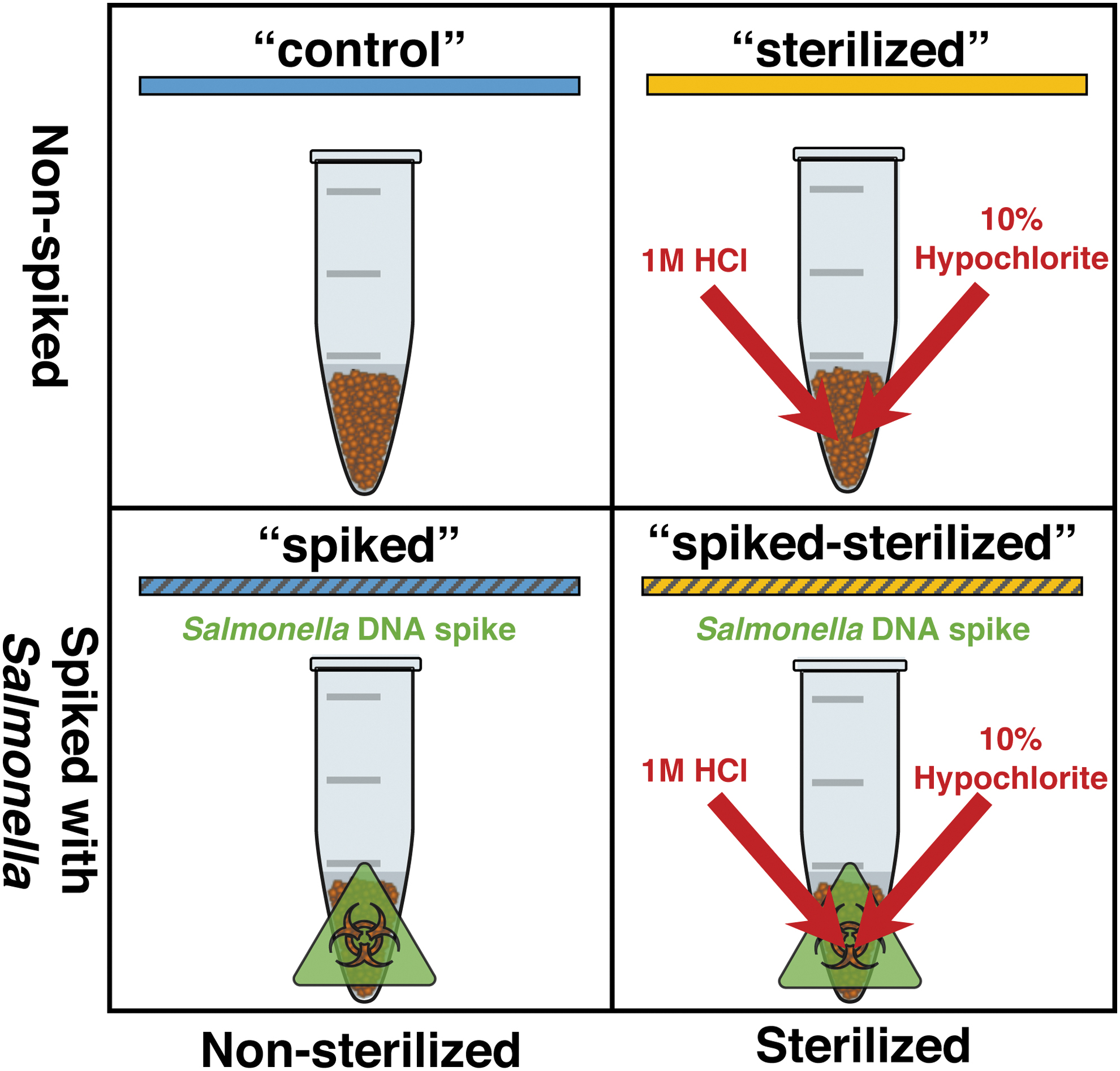
Experimental approach, depicting each of the four separate treatments: “control,” “sterilized,” “spiked,” and “spiked-sterilized.”
Samples were beaten with a rock hammer inside a sterile bag until roughly pea-sized, and they were then ground using a flame-sterilized mortar and pestle until fine to coarse grained. One gram of crushed sample was then loaded into a 2 mL tube and washed using filter-sterilized water. The washing step is intended to facilitate the removal of the finest grained sediment and dust, leaving only small rock particles ∼0.5–1 mm in diameter. The water was then removed using a 1 mL pipette.
We utilized DNA spiking to evaluate the efficacy of the sterilization procedure. In two of the treatment types (“spiked” and “spiked-sterilized”), 1000 ng of Salmonella DNA was diluted in ultrapure water to a volume of 500 μL and added to the rock samples. In the non-spiked treatments (“sterilized” and “control”), 500 μL of ultrapure water was added. This step was done following pulverization, but before sterilization. All sample treatments were then vortexed and allowed to incubate at 4°C for 24 h, to ensure sufficient time for the spiked DNA fragments to penetrate and bind to rock particles.
We predicted that, if the sterilization procedure was not effective and/or organisms trapped in the rock interior were not sufficiently isolated from the surface, fragments of spiked Salmonella DNA would reach these locations and would be detected following sequencing. Following incubation, the fluid within the tube was removed using a 1 mL pipette.
Sterilization procedure
The sterilization procedure, adapted from the work of Sankaranarayanan et al. (2011), was designed to remove all organisms and their genetic material from the rock surface. We hypothesized that any genetic material located sufficiently deep within rock particles to escape the sterilization fluids and be detected post-sterilization would represent organisms that were trapped at the time of deposition. These organisms may have been localized within fluid inclusions or trapped within the mineral matrix. The following steps were applied to two of the treatment types (“sterilized” and “spiked-sterilized”); the other two treatment types (“control” and “spiked”) were left aside until the DNA extraction steps.
First, 600 μL of 1 M HCl was mixed with the pulverized sample by pipetting. This was allowed to sit for 5 min before centrifuging at 5000g for 30 s and removing the acid wash. This step was repeated for a total of three acid washes. Following treatment with acid, the samples were washed with ultrapure water, and they were then treated to a bleach wash using 1 mL of 10% hypochlorite solution. The tube was placed in a rotator and left for 20 min. Following the bleach wash, the sample was vortexed and then washed with filter-sterilized water five times to ensure complete removal of bleach. The sample was then considered to consist of surface-sterilized particles and ready for the extraction of DNA from the interiors of the particles.
DNA extraction
Our protocol was modified from a method for DNA extraction from low-biomass carbonate rock (Barton et al., 2006). Six hundred microliters of buffer AE was added to rock samples followed by 20 μL of poly-dIdC to boost DNA yield. The poly-dIdC coats the carbonate minerals, which have an affinity for binding DNA, replacing native DNA strands.
Tubes were then vortexed and stored at 4° C overnight to allow the poly-dIdC to bind the carbonate particles. Following incubation, the tubes were vortexed and 100 μL of 10% sodium pyrophosphate was added as a chelating agent. Samples were then incubated in a dry bath at 65° C for 60 min to promote cell lysis, followed by centrifugation for 2 min at 5000g. Finally, the supernatant was removed and subjected to a standard phenol/chloroform DNA extraction protocol, followed by cleanup with magnetic beads (Rohland and Reich, 2012) and quantification with Qubit (Thermo Fisher).
DNA sequencing
Bacterial 16S rRNA gene amplicon sequencing was conducted by the Michigan State University genomics core facility. The V4 region of the bacterial 16S rRNA gene was amplified with dual-indexed Illumina fusion primers with the 515F/806R primers (Kozich et al., 2013). Amplicon concentrations were normalized and pooled using an Invitrogen SequalPrep DNA Normalization Plate. After library QC and quantitation, the pool was loaded on an Illumina MiSeq v2 flow cell and sequenced using a standard 500 cycle reagent kit. Base calling was performed by Illumina Real Time Analysis (RTA) software v1.18.54. The output of RTA was demultiplexed and converted to fastq files using Illumina Bcl2fastq v1.8.4.
Quality control and processing of sequence data was based on the work of Kozich et al. (2013), using the software platform mothur (Schloss et al., 2009) and the algorithms USEARCH8 (Edgar, 2010) and UCHIME (Edgar et al., 2011). In brief, overlapping paired-end reads were merged and filtered by quality score, chimeric sequences, and sequences with ambiguous bases or homopolymers longer than eight base pairs were removed, identical sequences were merged, and finally sequences were pre-clustered using a difference threshold of one.
These steps resulted in a set of 87,488 unique, quality-controlled sequences (operational taxonomic units [OTUs]) derived from an original set of 971,864 raw sequencing reads. These refined sequences were OTUs and used for all subsequent analyses. No further similarity clustering was done to avoid the unnecessary loss of data. Sequences were also re-processed with DADA2 (Callahan et al., 2016) at a later stage of the project with similar results. Representative sequences of each OTU were then given taxonomic assignments using the SILVA reference alignment in mothur (Pruesse et al., 2012).
Sequence data analysis
Comparisons of diversity indices, including observed OTU richness, Shannon, and inverse Simpson statistics, as well as total sequence abundance and Salmonella sequence abundance, were performed, and significant differences between mean values of treatments were tested with analysis of variance (ANOVA). The R package phyloseq (McMurdie and Holmes, 2013) was used to calculate Bray-Curtis community composition dissimilarities and to make ordination plots that visualize the dissimilarities.
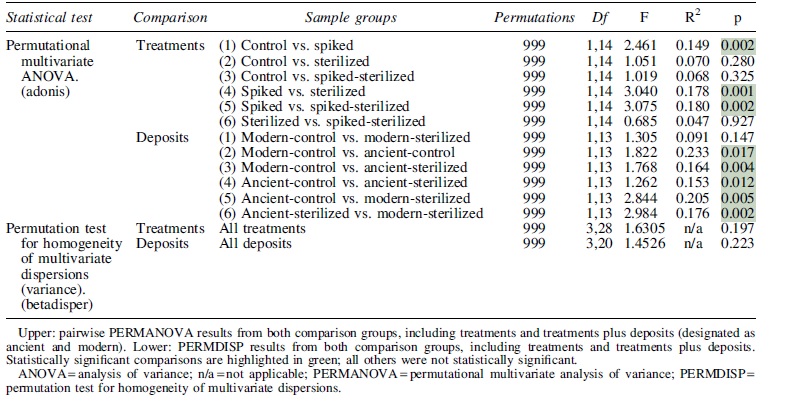
Statistical significances of Bray-Curtis dissimilarities between groups of samples were calculated with permutational multivariate analysis of variance and permutation test for homogeneity of multivariate dispersions as implemented by the R package vegan (Oksanen et al., 2019). The differential abundance of OTUs was calculated using the R package edgeR (Robinson et al., 2010) and measured as log2-fold change between sample types.
The R package iNEXT (Hsieh et al., 2016) was used to calculate asymptotic diversity estimates for ancient samples. To identify subsurface bacteria within travertine deposits, a blastn search was conducted with the 16S rRNA gene amplicon sequences from our study against the metagenomic data from (Probst et al., 2018).
Each assembly downloaded from NCBI BioProjects PRJNA362739, PRJNA349044, and PRJNA29758 was categorized as being enriched in the “major,” “minor,” or “recovery” eruptions as indicated in Supplementary Table 5 of Probst et al. (2018). Blast hits were filtered through these categories to identify those representing organisms enriched in the groundwater.
Data availability
All DNA sequences from this study are available via the NCBI SRA BioProject PRJNA672135. The full table of OTUs and their counts among all samples is provided as Supplementary Table S1.
Results
Sterilization procedure
We obtained a total of 306,514 sequences of the 16S rRNA gene from all control samples (
A total of 87,488 OTUs were clustered from the total set of 776,769 quality-controlled amplicon sequences. Total abundance, OTU observed richness, Shannon diversity, and inverse Simpson diversity were all significantly lower in sterilized samples compared with control samples (Fig. 3).
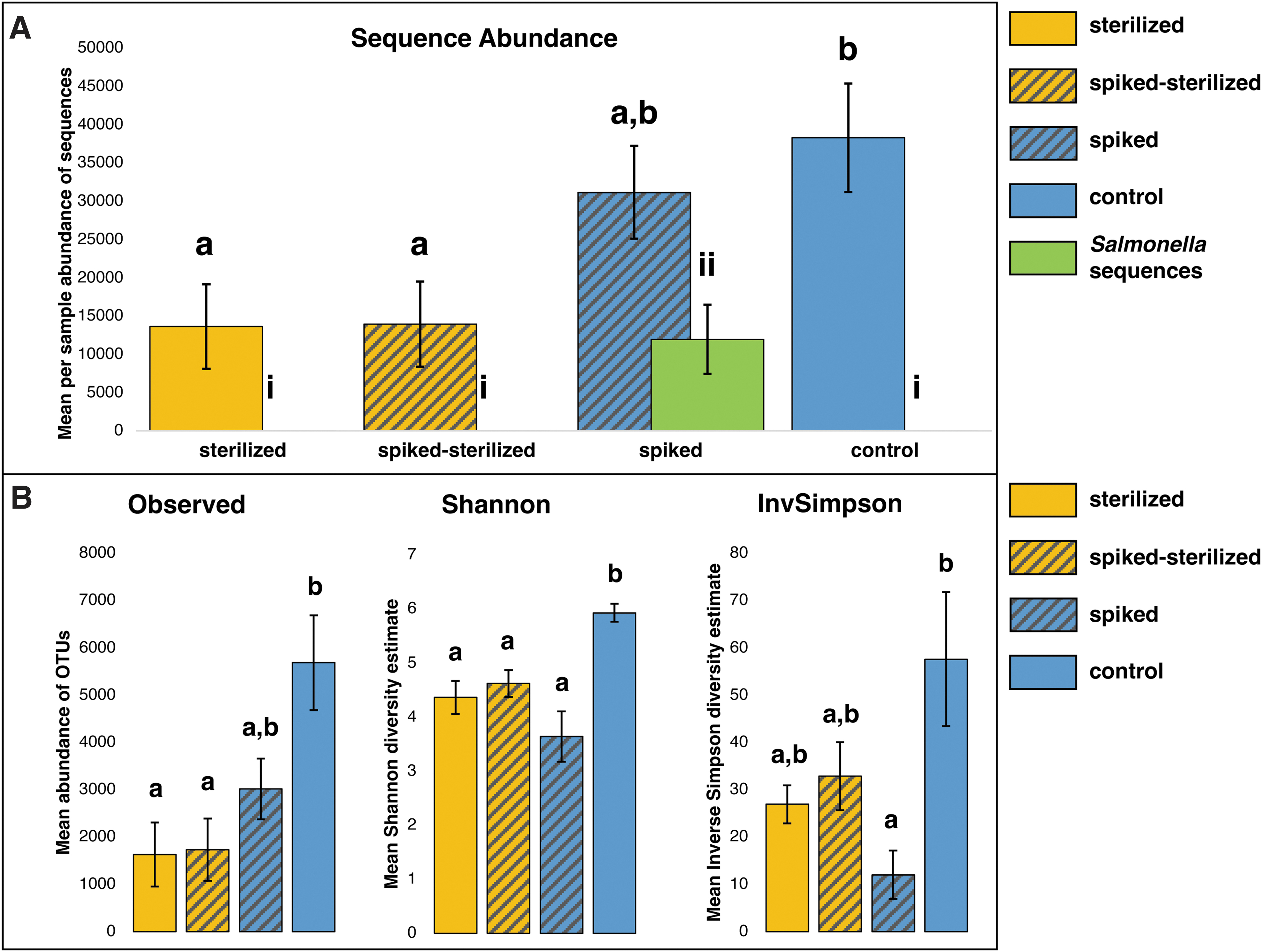
Bar charts of diversity indices, comparing treatments.
“Spiked” samples, as expected, contained very high Salmonella sequence abundance, with a total of 95,822 sequence counts among all spiked samples (n = 8), and a mean of 11,978 sequences (38.42% of total) per sample. Subsequent sterilization of spiked samples (i.e., “spiked-sterilized” samples) almost entirely removed Salmonella sequences, with a total count of 18 sequences among all spiked-sterilized samples (n = 8), for a mean of two sequences (0.016% of total) per sample. Spiked-sterilized samples did not differ significantly from non-spiked sterilized samples in total sequence counts or OTU diversity (Fig. 3).
The microbial community composition of modern travertine deposits (i.e., those collected from active springs) was not affected by sterilization (modern-control vs. modern-sterilized in Table 1). Ancient travertine deposits (ages of ∼5000 to ∼100,000 years), in contrast, showed significant shifts in community composition after sterilization (ancient-control vs. ancient-sterilized in Table 1). These patterns in community composition dissimilarity are visualized in Fig. 4, where ancient, sterilized samples appear to be highly variable and distinct from all other samples.
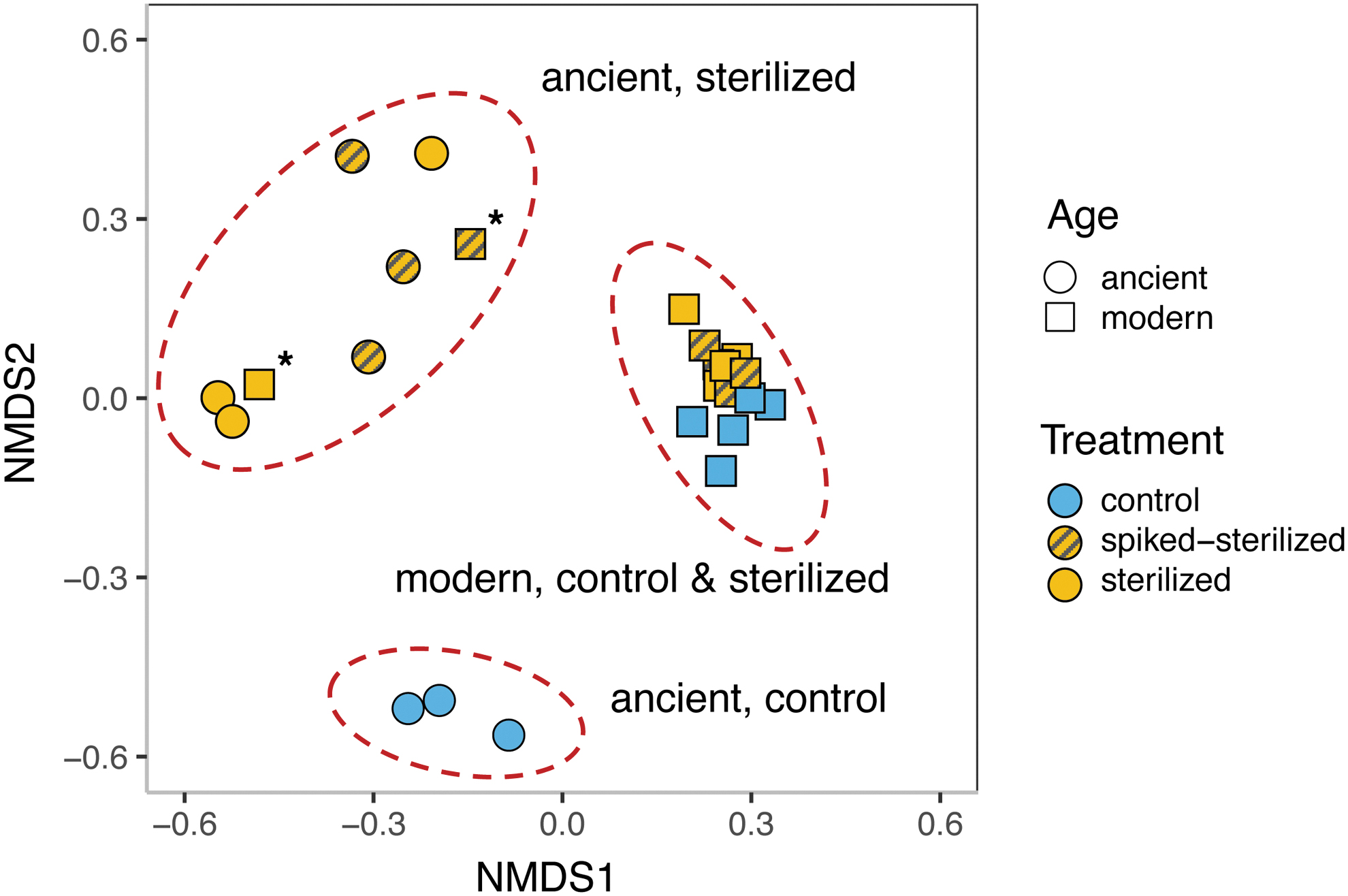
NMDS ordination plot comparing modern and ancient rock samples, including control, sterilized, and spiked-sterilized treatments. Dashed-ovals represent statistical significance (p < 0.05 as measured by PERMANOVA). Asterisks indicate sterilized and spiked-sterilized treatments of modern outlier sample S1. PERMANOVA, permutational multivariate analysis of variance.
Results from Statistical Analyses
All OTUs were classified with the SILVA reference database, reported in Supplementary Table S1. In general, unclassified Cyanobacteria, Piscirickettsiaceae (Proteobacteria), FamilyII (Cyanobacteria), Flavobacteriaceae (Bacteroidetes), Shewanellaceae (Proteobacteria), and Gallionellaceae (Proteobacteria) were the most abundant groups in all travertine deposits.
None of the taxa that were abundant in sterilized samples have been detected in our laboratory air or in various extraction blanks that our lab has conducted for other projects (Motamedi et al., 2020). To examine whether some taxa were more abundant in sterilized samples compared with controls, we calculated the differential abundance of each OTU in sterilized compared with control samples (Fig. 5).
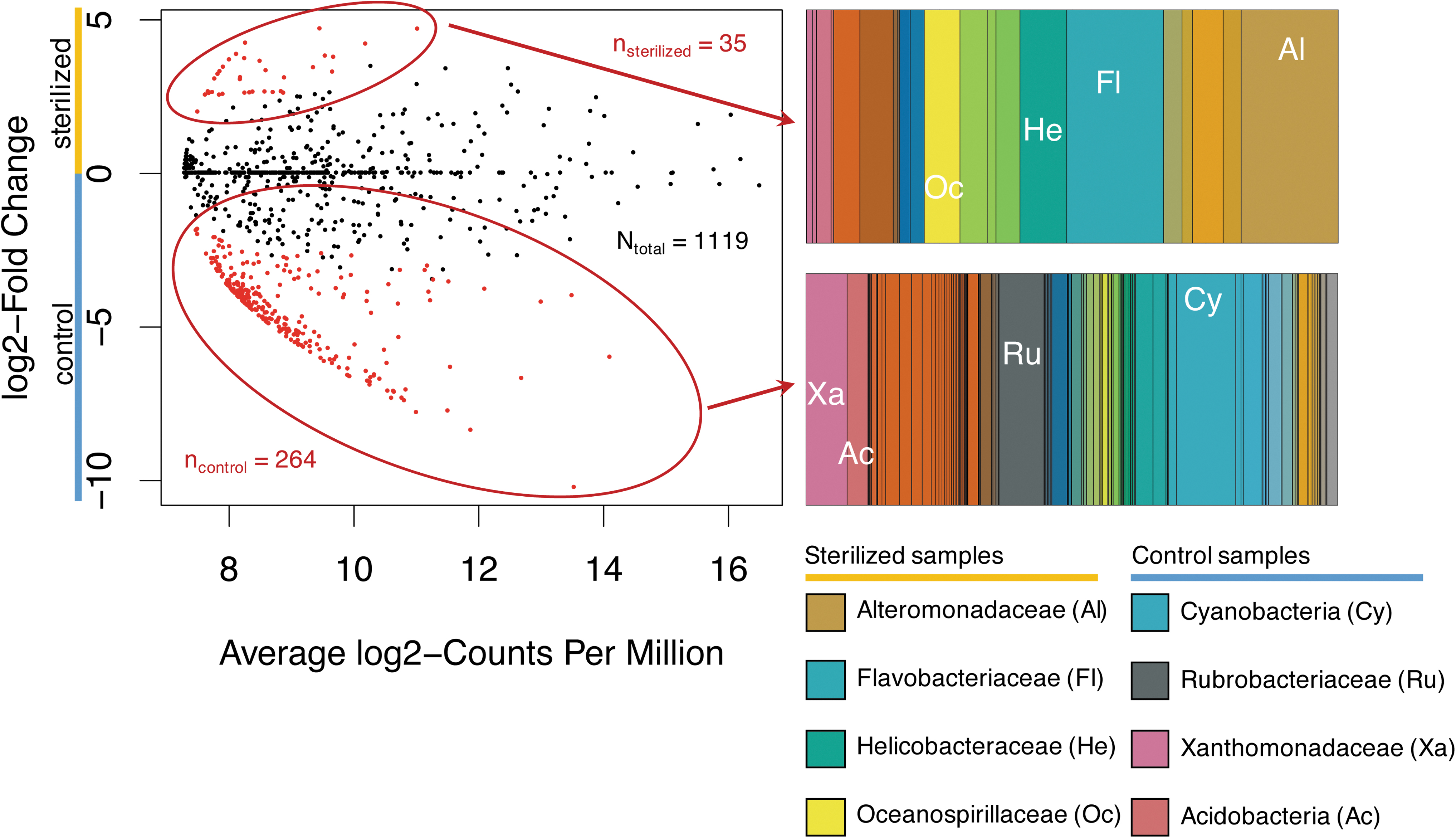
edgeR results, depicting statistically significant enriched sequences (red dots; n
sterilized = 35, n
control = 264, N
total = 1119) in sterilized and control treatment groups, and their respective family-level taxonomic breakdown within those treatment groups. The x-axis shows log-base two
The most abundant taxa in control samples were unclassified Cyanobacteria, Family II (Cyanobacteria), Xanthomonadaceae (Proteobacteria), and Flavobacteriaceae (Bacteroidetes), whereas unclassified Cyanobacteria, Piscirickettsiaceae (Proteobacteria), Shewanellaceae (Proteobacteria), and Oceanospirillaceae (Proteobacteria) were the most abundant taxa in sterilized samples.
The species richness of sterilized travertine samples was examined by measuring the observed species richness (i.e., number of observed OTUs), by calculating the Shannon and Inverse Simpson alpha diversity indices, and by estimating asymptotic richness and diversity (Table 2).
Alpha-Diversity Values and Estimates for Sites, Reported Separately for Each Deposit, Using Only the “Sterilized” and “Spiked-Sterilized” Samples
Alpha-Diversity Values and Estimates for Sites, Reported Separately for Each Deposit, Using Only the “Sterilized” and “Spiked-Sterilized” Samples
Observed diversity values are reported first, followed by diversity estimates from iNEXT and their respective SE, and LCL and UCL. All values are rounded to the nearest whole number.
LCL = lower confidence level; OUT = operational taxonomic unit; SE = standard error; UCL = upper confidence level.
The ancient travertine deposit from site L2 (age ∼5000 years) contained the lowest observed richness, but this result is due to the poor sequencing success of these samples (∼100 total counts per sample). The Shannon and Inverse Simpson alpha diversity indices decreased with increasing deposit age, as did their asymptotic estimates (Table 2).
Treatment efficacy
Our results suggest that both the DNA-spiking and sterilization treatments were effective. Spiked samples contained massive amounts of Salmonella 16S rRNA gene sequences, which were, by far, the most abundant organism therein (Fig. 3) and had a significantly different community composition compared with all other treatment types (Table 1). For sterilized samples, there was nearly complete removal of the surface DNA tracer, indicating that the remaining DNA was protected from surface sterilization. This indicates that the 16S rRNA gene sequences in sterilized samples represent the “internal” community of carbonates and originate from the contemporary microbial community present at the time of carbonate deposition. This is consistent with previous work finding that carbonates, including those at Crystal Geyser, are known to trap organic material present during mineral deposition (Wolf et al., 2022).
Preservation of microbial DNA in carbonates
Consistent with our expectations, the composition of 16S rRNA gene sequences obtained from modern carbonate samples was not significantly affected by sterilization (Fig. 4; Table 1). Modern samples were taken directly from the active Crystal Geyser, which began forming in 1935 during oil and gas exploration activity, giving the carbonates an age range of 0–85 years (McKnight, 1940; Waltham, 2001).
Most organisms preserved during carbonate deposition are expected to be members of the environmentally structured surface community, which is unlikely to have changed substantially over the past 85 years. Conversely, the deposits from which ancient samples were collected range up to 105,000 years old (Burnside et al., 2013), and are situated in arid conditions no longer subject to active water flow, a vastly different situation compared with the water-saturated conditions under which they formed. Based on this, we predicted that the internal and external communities of ancient samples would be substantially different, reflecting shifts in microbial community composition due to environmental changes since their formation. Consistent with our prediction, the community composition of ancient samples varied significantly between sterilized and control samples (Fig. 4; Table 1).
In addition, the alpha diversity of sterilized samples decreased with age, consistent with long-term DNA degradation (Table 2). In contrast, no change in alpha diversity over time was observed for control samples, in which most of the DNA would be derived from the external community. Interestingly, the sterilized and spiked-sterilized treatments of one modern sample (S1) did cluster with those of the ancient samples (Fig. 4, asterisks), whereas the control samples of S1 clustered with those of the modern samples. We posit that sample S1 is part of a more ancient (∼5.4 Ka), naturally formed deposit underlying the active Crystal Geyser formation (L2, Fig. 1A; Burnside et al., 2013).
The area adjacent to the borehole of the active Crystal Geyser, from which sample S1 was taken, is frequently inundated with erupting fluids but does not have any obvious recent carbonate formation (Fig. 1D), which is visually striking several meters away from the borehole (e.g., Fig. 1B, C). In this case, the internal 16S rRNA gene sequences of sterilized treatments would represent an ancient community formed ∼5.4 Ka, whereas the external 16S rRNA gene sequences would be largely from the present-day community formed at the active spring. This is consistent with the observed and estimated alpha diversity measures for the sterilized and spiked-sterilized treatments of sample S1, which are significantly lower than all other modern samples but similar to those of the ancient samples (Table 2).
Detection of subsurface organisms
Most of the water erupting from the Crystal Geyser system is derived from shallow, meteorically recharged aquifers <1 km beneath the surface (Shipton et al., 2004). However, a substantial fraction of erupting water, 10–20%, originates more than 2 km below the surface from a hydrothermal saline aquifer (Wilkinson et al., 2009). We hypothesized that organisms inhabiting this deep-subsurface hydrothermal environment would be transported to the surface and preserved during carbonate formation, forming a component of the internal 16S rRNA gene sequences derived from carbonate samples.
We tested this by examining the taxonomic classifications of 16S rRNA gene sequences recovered from sterilized samples. Table 3 reports several examples of taxonomic groups known to be associated with subsurface environments and were detected across multiple sterilized samples from ancient deposits. Genera such as Dethiobacter, Hydrogenophaga, and Thiomicrospira typically inhabit oxygen-poor, hydrothermal environments rich in sulfur compounds and/or hydrogen gas, and they are very unlikely to inhabit the dry, oxygenated, and sun-drenched surfaces of ancient carbonate deposits. In addition, most of the sequences we recovered from carbonate deposits, both sterilized and not sterilized, matched 16S rRNA genes present in samples of erupting fluids from Crystal Geyser reported by Probst et al. (2018). Several of the abundant members of the deep groundwater community identified by Probst et al. were also abundant inside the sterilized samples of our carbonate deposits (e.g., Piscirickettsiaceae and Gallionellaceae), directly demonstrating that subsurface organisms are preserved within the carbonate deposits.
Bacterial Taxa Identified from “Ancient, Sterilized” Samples That Represent Potential Suburface Inhabitants
Bacterial Taxa Identified from “Ancient, Sterilized” Samples That Represent Potential Suburface Inhabitants
Taxonomic treatment is followed by notes concerning known or likely metabolic activity and/or habitat conditions, deposits the taxon was identified from, the proportion of individual samples that contained the taxon, and the abundance of the taxon as a percentage of total sequences from
The surface of Mars may contain sites where, like the Crystal Geyser system, mineral deposits have formed in the past several hundred thousand years in association with the discharge of subsurface water. This type of site has major advantages as a candidate for future life-detection missions.
First, mineral deposits formed in this manner allow indirect access to the deep subsurface, which has a high potential for habitability on present day Mars and is a top priority for life-detection. Liquid water is likely thermodynamically stable in the martian subsurface at kilometer-plus depths, but it may also be stable at shallower depths due to the presence of salty brines (Burt, 2003; Carrier et al., 2020).
Terrestrial subsurface aqueous systems, which are often highly saline, are known to have a high level of biomass and diverse microbial communities (McMahon and Parnell, 2014). The discharging waters at Crystal Geyser are sourced from deep, hypersaline, thermodynamically stable aquifers, and our results suggest that organisms inhabiting this environment are discharged at the surface. Putative subsurface organisms inhabiting the martian subsurface would potentially also be preserved in surface deposits during discharge events.
Second, the process of mineral deposition can preserve biosignatures over long time periods. Habitable conditions on Mars are likely to be heterogeneous over space and time. Environments may fluctuate in their capacity to harbor extant life, and life-detection missions targeting extant life might miss the window of opportunity during which life occurs at a particular site.
Sites where biosignature preservation can occur are particularly attractive, as searching for preserved biosignatures expands the window of opportunity for life-detection. We demonstrated that genetic material from organisms present during mineral deposition is preserved for at least 100,000 years, well within the estimated detection limit for bacterial DNA at between 0.4 and 1.5 million years (Willerslev et al., 2004). While the upper bounds for preservation of genetic material on Mars are uncertain, and likely to be context-dependent, we suggest that mineral deposits formed in the past million years may have the potential to preserve genetic material from putative subsurface lifeforms.
Crystal Geyser as a Mars analog site
Testing life-detection strategies at terrestrial Mars-analog sites is critical for evaluating their potential. Several factors make the Crystal Geyser system an excellent Mars-analog candidate. The vertically stratified subsurface system of Crystal Geyser involves groundwater from multiple aquifers at various depths, including shallower, low-temperature aquifers and deeper, high-temperature, hypersaline aquifers, broadly similar to conditions proposed to exist on Mars.
Crystal Geyser, which can be characterized as a carbon dioxide driven cold spring, has been suggested to be particularly relevant for carbonate deposit formation on Mars (Hays et al., 2017; Wolf et al., 2022). The Crystal Geyser site consists of multiple springs, both active and extinct, some of which were naturally formed along fracture networks and others that formed as the result of drilling activity (Dockrill and Shipton, 2010).
This combination of both naturally and artificially caused subsurface fluid discharge can be used to study the potential consequences of drilling into the martian subsurface. In addition, the network of active and extinct cold-spring carbonate deposits that make up the Crystal Geyser site has an age range of zero (actively forming) to upwards of 400,000 years (Burnside et al., 2013). The presence of deposits of different ages that formed from the same subsurface environment allows for studies into the preservation of biosignatures over time.
Challenges and limitations
In this study, we provided supporting evidence for selecting mineral deposits associated with subsurface fluid discharge as targets for life-detection missions. However, several important challenges and limitations to this approach exist. Currently, there is uncertainty regarding the existence of this type of site on Mars. While aqueous deposits likely do exist, whether any have formed during the past several hundred thousand years is currently unknown. In addition, although current and future technology may be used to identify and characterize mineral deposits from orbit and in situ, it would be difficult to ascertain the origin of the water in which they formed.
A further limitation is uncertainty regarding the longevity of biosignature preservation under martian conditions. Factors such as temperature, radiation, and oxidation have a significant impact on the preservation of biosignatures, and ionizing radiation is a particular concern given the high-radiation conditions of present-day Mars, currently estimated at 0.54–0.85 Gy per year (Dartnell et al., 2007). Although this level is well within the survivable range of living terrestrial bacteria, preserved genetic material would be more vulnerable as damage would no longer be actively repaired. Therefore, the detectable lifetime of genetic material would depend on the dose of radiation accumulated over time and the context in which it is preserved.
For example, interactions with minerals such as ferric iron and silicates can enhance the radiolytic degradation of organic material (dos Santos et al., 2016; Pavlov et al., 2022). Current studies suggest that radiolytic degradation would be a major problem for the preservation and detection of biosignatures over timescales of millions of years (Pavlov et al., 2012, 2022). However, at timescales of <1 million years, ionizing radiation is not likely to be a significant problem. For example, Dartnell et al. (2007) estimated that radioresistant organisms in a dormant state would need an exposure time of ∼450,000 years under martian conditions to receive a lethal dose of radiation. Even at biologically lethal doses of radiation, genetic material would remain detectable for some time.
Additional evidence suggests that the adsorption of DNA to rock surfaces can confer UVC-protection, supporting the use of DNA as a biosignature marker (Lyon et al., 2010). While further work regarding the longevity of DNA under ionizing conditions is required, we suggest that DNA is a promising biosignature candidate in the context of mineral preservation at sites with ages of several hundred thousand years.
Conclusions
Our results suggest that it is possible to detect ancient subsurface organisms preserved within surface-level carbonate deposits up to at least 100,000 years old, consistent with previous studies focused on extracting ancient microbial DNA preserved in minerals and recent advances in the field of ancient DNA (Sankaranarayanan et al., 2011; Green and Speller, 2017).
We suggest that mineral deposits with ages <1 million years and with the involvement of deep-subsurface water are excellent candidates for future life-detection missions. Advantages of this type of deposit include indirect access to the deep subsurface and long-term preservation of biosignatures. However, there are critical unknowns regarding the availability of this type of site on Mars and our ability to identify them.
While our study focused on detecting genetic material, other biomarkers would be similarly preserved and may have longer lifespans under martian conditions. Future research should focus on studying biosignature preservation in other mineral types, over a larger range of deposit ages and preservation conditions and detecting other biomolecule types that may originate in the deep subsurface.
Footnotes
Acknowledgments
The authors thank Brenda Bowen for helpful discussion and assistance in planning the research project and Marjorie Chan for the contribution of samples. They thank Lizethe Pendleton and Shahrzad Motamedi for valuable lab assistance, Alex Hyer and Chris Thornton for their help with data analysis, and all the members of the Brazelton Lab for helpful discussion and for their many contributions.
Authors' Contributions
M.P.P.: conceptualization, methodology, investigation, writing—original draft, and writing—review and editing.
W.J.B.: conceptualization, methodology, writing—review and editing, and supervision.
Author Disclosure Statement
No competing financial interests exist.
Funding Information
Funding was provided by the University of Utah and the NASA Astrobiology Institute (CAN-5) via the Rock-Powered Life team.
Supplementary Material
Supplementary Table S1
Abbreviations Used
Associate Editor: Don Cowan
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.