Abstract
A plant production system called Veggie was launched to the International Space Station (ISS) in 2014. In late 2015, during the growth of Zinnia hybrida cv. ‘Profusion’ in the Veggie hardware, plants developed chlorosis, leaf curling, fungal growth that damaged leaves and stems, and eventually necrosis. The development of symptoms was correlated to reduced air flow leading to a significant buildup of water enveloping the leaves and stems in microgravity. Symptomatic tissues were returned to Earth on 18 May 2016 and were immediately processed to determine the primary causal agent of the disease. The presumptive pathogen was identified as Fusarium oxysporum by morphological features of microconidia and conidiophores on symptomatic tissues; that is, by epifluorescent microscopy (EFM), scanning electron microscopy (SEM), metabolic microarrays, and ITS sequencing. Both EFM and SEM imaging of infected tissues showed that germinating conidia were capable of stomatal penetration and thus acted as the primary method for infecting host tissues. A series of ground-based pathogenicity assays were conducted with healthy Z. hybrida plants that were exposed to reduced-airflow and high-water stress (i.e., encased in sealed bags) or were kept in an unstressed configuration. Koch's postulates were successfully completed with Z. hybrida plants in the lab, but symptoms only matched ISS-flown symptomatic tissues when the plants were stressed with high-water exposure. Unstressed plants grown under similar lab conditions failed to develop the symptoms observed with plants on board the ISS. The overall results of the pathogenicity tests imply that F. oxysporum acted as an opportunistic pathogen on severely high-water stressed plants. The source of the opportunistic pathogen is not known, but virulent strains of F. oxysporum were not recovered from unused materials in the Veggie plant pillow growth units assayed after the flight.
1. Introduction
Bioregenerative life-support systems (BLSS) may be developed for future human habitats on the Moon and Mars (Ming and Henninger, 1989; Wamelink et al., 2014; Wheeler, 2017). The advantage of BLSS habitats that utilize higher plants versus physiochemical (i.e., nonbiological) life-support systems is that the former can provide for the regeneration of water, oxygen, and food stocks with minimal resupply from Earth. Both algae and traditional agronomic and horticultural crops have been proposed for BLSS (see reviews by Eckart, 1996; Massa et al., 2013; Wheeler, 2017), but the final configuration of such a system is still under development. However, as missions to the Moon and Mars become realistic over the next 10 years, the deployment of smaller plant-growth systems will provide fresh produce for the astronauts during Mars transit or in cis-lunar space. It will be these small plant-biomass production systems that will pioneer the development of BLSS modules for more expansive bases on the Moon and Mars over the next 30 years.
Currently, plant biomass production systems in low Earth orbit (LEO) have been limited to small plant growing systems that have been designed for either targeted plant physiology and metabolism studies or technology demonstration activities (see review by Zabel et al., 2016). One of the latter such systems is the Veggie plant growth system that was launched in 2014 to the International Space Station (ISS) (Morrow and Remiker, 2009; Massa et al., 2013, 2017a, 2017b). Veggie was designed as a small “salad machine” for producing fresh leafy greens for astronauts on the ISS. Initial production in 2014 (VEG-01A) and 2015 (VEG-01B) were both used to grow crops of red-romaine lettuce (Lactuca sativa cv. ‘Outredgeous’) leading to consumption of a portion of the VEG-01B crop by ISS astronauts (Massa et al., 2017a).
During the course of the VEG-01C crop between 16-Nov-2015 and 14-Feb-2016, several of the stressed Zinnia hybrida plants developed a foliar disease that advanced rapidly down the stems and caused significant leaf chlorosis, necrosis, and eventual wilting of plant canopies (Massa et al., 2017a). In addition, symptomatic and wilted leaves exhibited a significant amount of aerial mycelium suggesting that the causal agent of the disease was a fungus. However, no diagnostic equipment or protocols were available on the ISS during the disease outbreak in late 2015, so moribund leaf and root tissues were harvested on 22-Dec-2015 and 08-Jan-2016 (36 and 53 days after initiating the crop of zinnia plants, respectively), frozen at -80°C, and returned to Earth for diagnosis of the presumptive biological pathogen. Frozen zinnia tissues arrived at the Kennedy Space Center (KSC), Florida, labs on 18-May-2016, and subsequent, diagnostic assays immediately started.
Historically, only two previous studies have been published that examined the development of plant diseases in microgravity (μg) that occurred in small plant-growth systems on board the shuttle (Bishop et al., 1997; Ryba-White et al., 2001). First, a wheat crop (Triticum aestivum cv. ‘Super Dwarf’) was damaged by growth of the endophytic fungus Neotyphodium chilense, leading to a loss of approximately 50% of the crop during an 8-day shuttle mission (Bishop et al., 1997). Next, the development of the fungal root pathogen Phytophthora sojae on soybean plants (Glycine max cv. ‘Williams 82’) was examined during a directed plant-disease development experiment on shuttle flight STS-87 called the Collaborative Ukrainian Experiment (Ryba-White et al., 2001). In both disease outbreaks, restricted airflow within the plant-growth systems was linked to the development of severe symptoms.
The primary goals of the current study were to identify the causal agent of disease on the zinnia plants during the VEG-01C experiment and to determine whether specific environmental conditions contributed to the development of severe symptoms in μg. None of the diagnostic procedures were possible on board the ISS. The following procedure is presented in a linear fashion to highlight the sequence of events that were required to determine the relationship between zinnia plants and the microbial causal agent of the disease.
2. Methods
2.1. Veggie plant growing system
The Vegetable Production System (i.e., Veggie) plant growth chamber (Fig. 1) was launched to the ISS in April 2014 along with the plant pillows for the VEG-01C experiment (Massa et al., 2016, 2017a; Khodadad et al., 2020). Veggie is a small plant growth facility that uses red, green, and blue LED lights and a single fan to circulate ISS ambient air through the growing volume. The designed airflow could be adjusted between 1.2 and 4.2 cubic feet per minute. Transparent, extensible bellows contain the growing plants and ensure bottom-to-top air flow. For the VEG-01C experiments, plants were grown in plant pillows (Stutte et al., 2011). Plant pillows are small growing bags that interact with a root mat water reservoir on the Veggie baseplate (Massa et al., 2017b). Pillows contained a porous ceramic substrate, called arcillite, mixed with controlled-release fertilizer and wicks for seed attachment. Plant pillows were packaged and handled according to Massa et al. (2017b). The wicks are inserted through a silicon foam matrix (called gasket foam) that securely holds the seeds and wicks in the tops of the pillows. Surface sanitized seeds (Massa et al., 2017b) were glued with a 1% Guar solution onto the plant pillow wicks, and pillows were packaged for flight under sterile airflow in a biosafety cabinet.
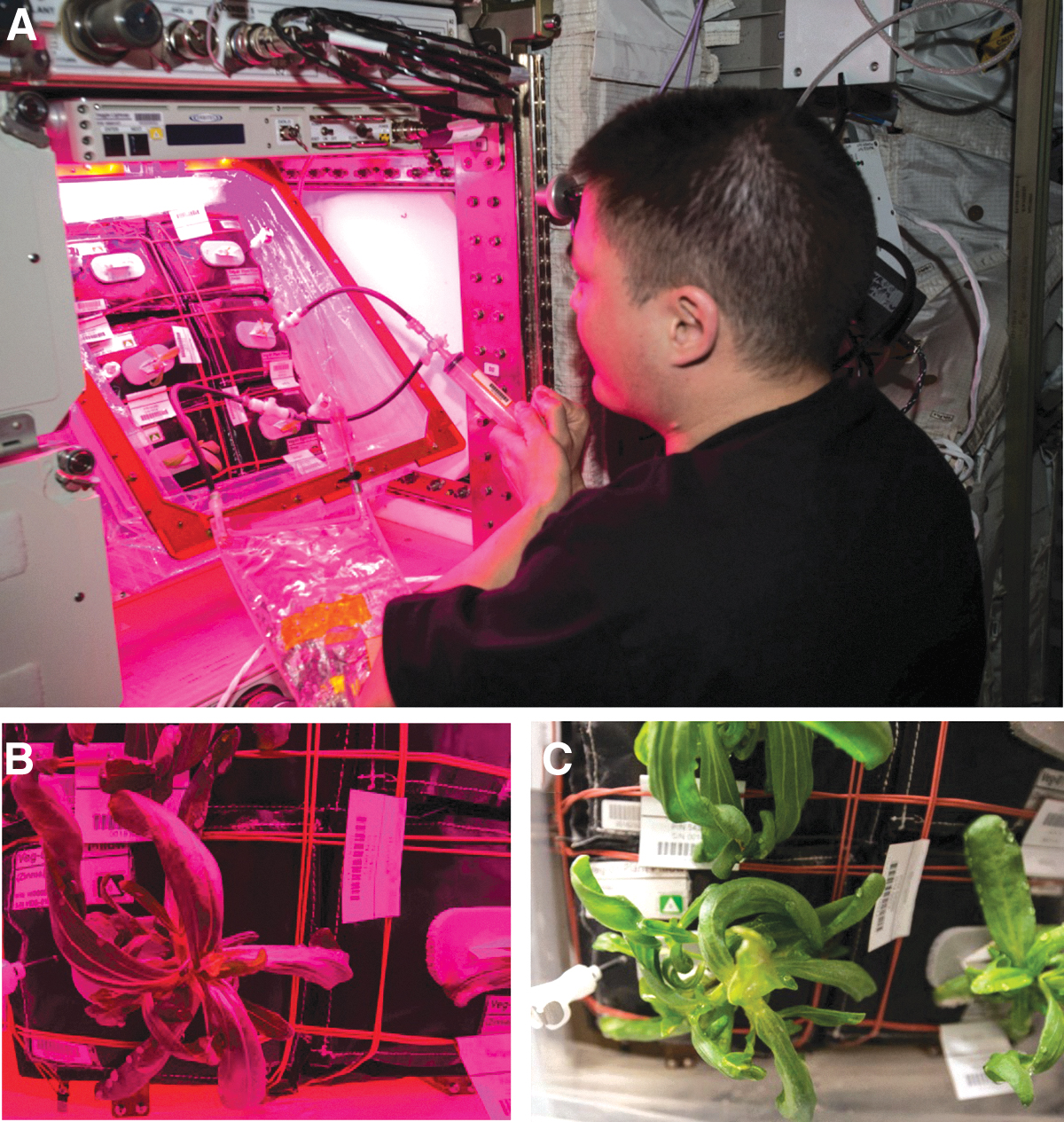

The red/blue LED images were less useful than white-light flash-photographs in diagnosing the development of the disease outbreak on Zinnia hybrida. (
The Veggie chamber was installed in the European laboratory, called Columbus, in May of 2014, and the first crops of lettuce were grown in the chamber between May–June 2014 and July–August 2015. In the third Veggie crop (VEG-01C), Zinnia hybrida cv. ‘Profusion’ (Johnny's Selected Seeds, Winslow, ME, USA) was grown between 16-Nov-2015 and 14-Feb-2016. Zinnia seeds had been sanitized and planted in the plant pillows in December 2013 according to the work of Massa et al. (2017b). Pillows were sealed in gas-impermeable Tedlar bags (SKC Inc., Eighty Four, PA, USA) under sterile laminar-flow hood air conditions to reduce any impact by the ambient volatile organic carbon compounds on board the ISS during storage. Zinnia plants were grown under a 10 h light/14 h dark photoperiod with red, blue, and green LEDs set to provide ∼200 μmol m2 s−1 of photosynthetically active radiation at the surface of the pillows with the Veggie baseplate set to a height of 30.5–31.75 cm below the lights (Massa et al., 2017a). The ISS temperature in the 90-day experiment averaged 23.1 ± 0.2°C, relative humidity averaged 40.3 ± 2.1%, and CO2 levels averaged 3156 ± 451 μmol mol−1 (Table 1).
Environmental Data from All Experimental Locations
Abbreviations: Temp = temperature; RH = relative humidity; CO2 = carbon dioxide; PAR = photosynthetically active radiation (400–700 nm); VEG-01C = Veggie experiment on board the International Space Station; LED = light-emitting diodes; PathLab = pathology lab in which Koch's postulate assays were completed; FL = fluorescent; na = not applicable.
Difficulty was encountered in maintaining optimal moisture for the germinating seedlings during the VEG-01C experiment. Power was inadvertently shut off to the Veggie hardware 3 days after initiation (i.e., on 19-Nov-2015) due to an unrelated issue. It was restored after a temporary (∼3 h) outage, and the lights and systems appeared to resume nominal operations when manually reset. No impact was assumed because plants had only just started to germinate, with the first seedlings just visible at 3 days after initiation when wicks were separated to assist seedling emergence. However, the Veggie ground team later determined in ground tests that when power was removed and restored to Veggie, although the Veggie LCD controller screen indicated that the fan speed was set to low, it was in fact off. Thus, in the absence of gravity and buoyancy-driven convection, water films accumulated within the chamber and around plant tissues. Plant leaves, stems, and internodes became water-soaked (Fig. 2) after approximately 7 days following the power-restart event.
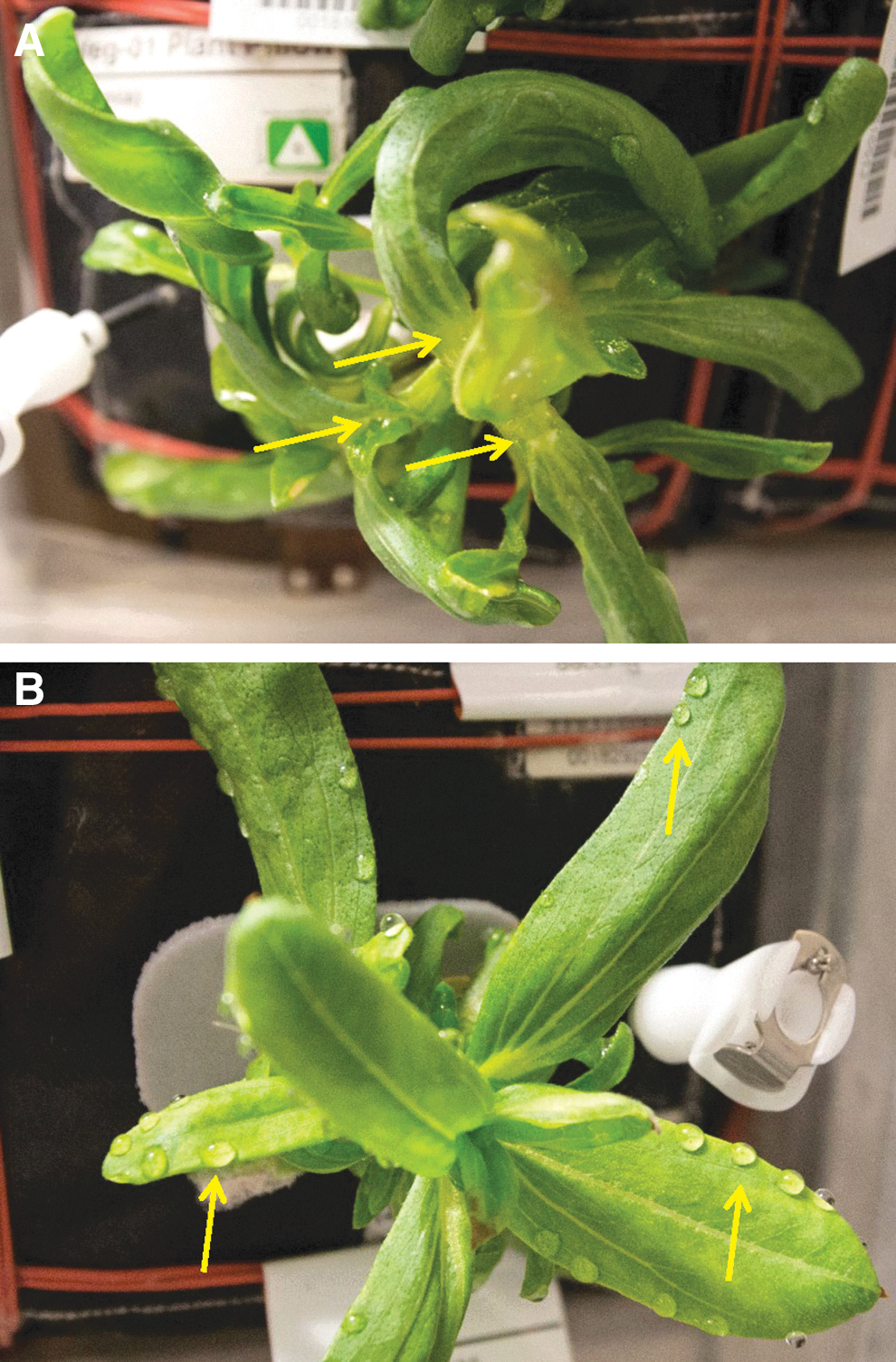

Zinnia hybrida plants grown on the ISS during the VEG-01C experiment; images collected on 15-Dec-2015 (29 days after initiation). Developing plants exhibited significant amounts of stunting, epinasty, and chlorotic leaves prior to the observation of aerial mycelium. (
Plants were watered on a preset schedule by the astronauts for the first 36 days of the experiment. The first water application was at 7 days after initiation when seedlings were thinned from two plants to one plant per pillow. Watering continued approximately every 3 days but varied with crew activities. Water was assumed to have accumulated inside plant pillows, and some free water was observed on the bellows of the Veggie facility. Due to a lack of atmospheric convection (i.e., the circulating fan was off), the normal transpiration loss of water was inhibited. By 29 days after initiation, some of the plants were stunted, chlorotic, and water-saturated (Fig. 2). Some plants exhibited abnormal growth with leaf epinasty and distorted morphologies (Fig. 2A). Water also visibly coated some leaf surfaces, and plants that were not otherwise wet showed distinct guttation droplets along the edges of leaves (Fig. 2B). The guttation prompted the team to schedule on-orbit crew activities to program the Veggie fan to the high setting in order to dry out the plant-growth compartment. However, ISS operations unrelated to the Veggie experiment prohibited the fan reprogramming prior to crew members notifying the ground Payload Operations team of suspected microbial growth on two of the plants. The Veggie fans were off for approximately 33 days.
Visible fungal growth on plants was observed 36 days after initiation on 22-Dec-2015 (Fig. 3A), and cream-colored mycelium was observed on the sewn seamed edges of several of the plant pillows (Fig. 3B). The crew member caring for the plants attempted to mitigate this by removing visibly infected tissues, wiping plants and hardware with sanitizing wipes, and setting the Veggie fans to high. Plants were wiped with custom-generated citric-acid based wipes (KIMTECH PURE W4 Dry Wipers; Kimberly-Clark Professional, Roswell, GA, USA, saturated with 1% Pro-San [Microcide Inc., Sterling Heights, MI, USA]). Hardware was wiped with commercial 0.4% benzalkonium chloride wipes (BZK Antiseptic Towelettes, Nice-Pak/PDI Inc., Orangeburg, NY, USA) available on the ISS. Consequently, after an additional 36 h with the Veggie fan on high, it appeared that the fan was drying leaf canopies out too much. Plants were watered, and the fan speed was switched to low speed on 24-Dec-2015.
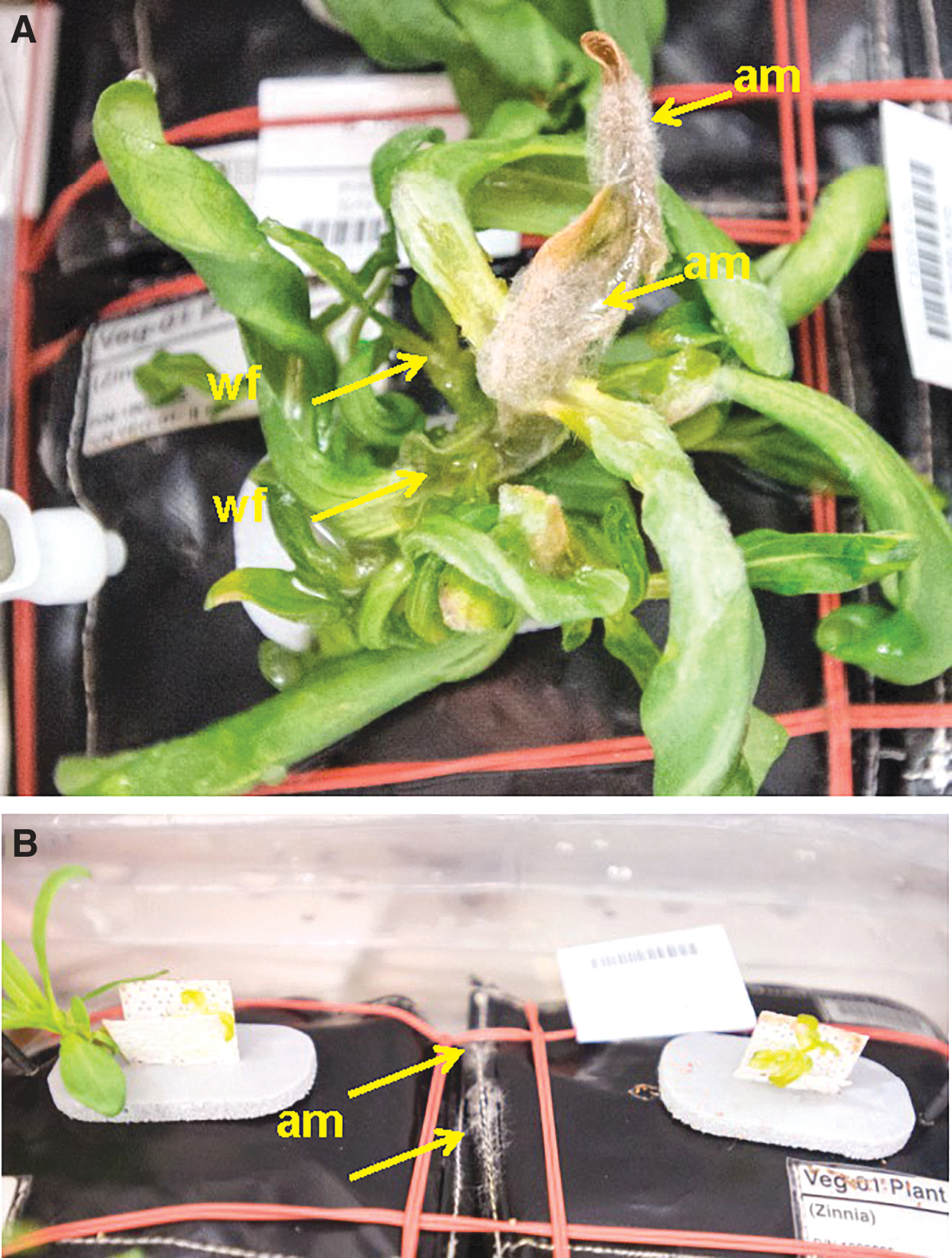

Signs of an active disease outbreak on Zinnia hybrida tissues during the VEG-01C experiment on board the ISS. (
At this point the plant care was switched from scheduled activities to crew-determined activities. Although plant care by the crew continued, the first two visibly infected plants that previously had been trimmed appeared wilted, necrotic, and in an advanced state of decay on 08-Jan-2016, 53 days after initiation (Fig. 4). Over the course of the VEG-01C experiment, one plant remained symptom free (upper left plant; Fig. 4), two plants failed to thrive after germination (upper right and middle right plants; Fig. 4), one plant developed symptoms but recovered (middle left plant; Fig. 4), and two plants were killed by the pathogen (lower two plants; Fig. 4). Interestingly, the symptom-free plant (upper left) was notable in that it experienced delayed germination. All other seedlings were apparent at 5 days after initiation, but the upper left plant did not appear until 9 days after initiation, after the second seedlings in all other plant pillows had already been removed through thinning.
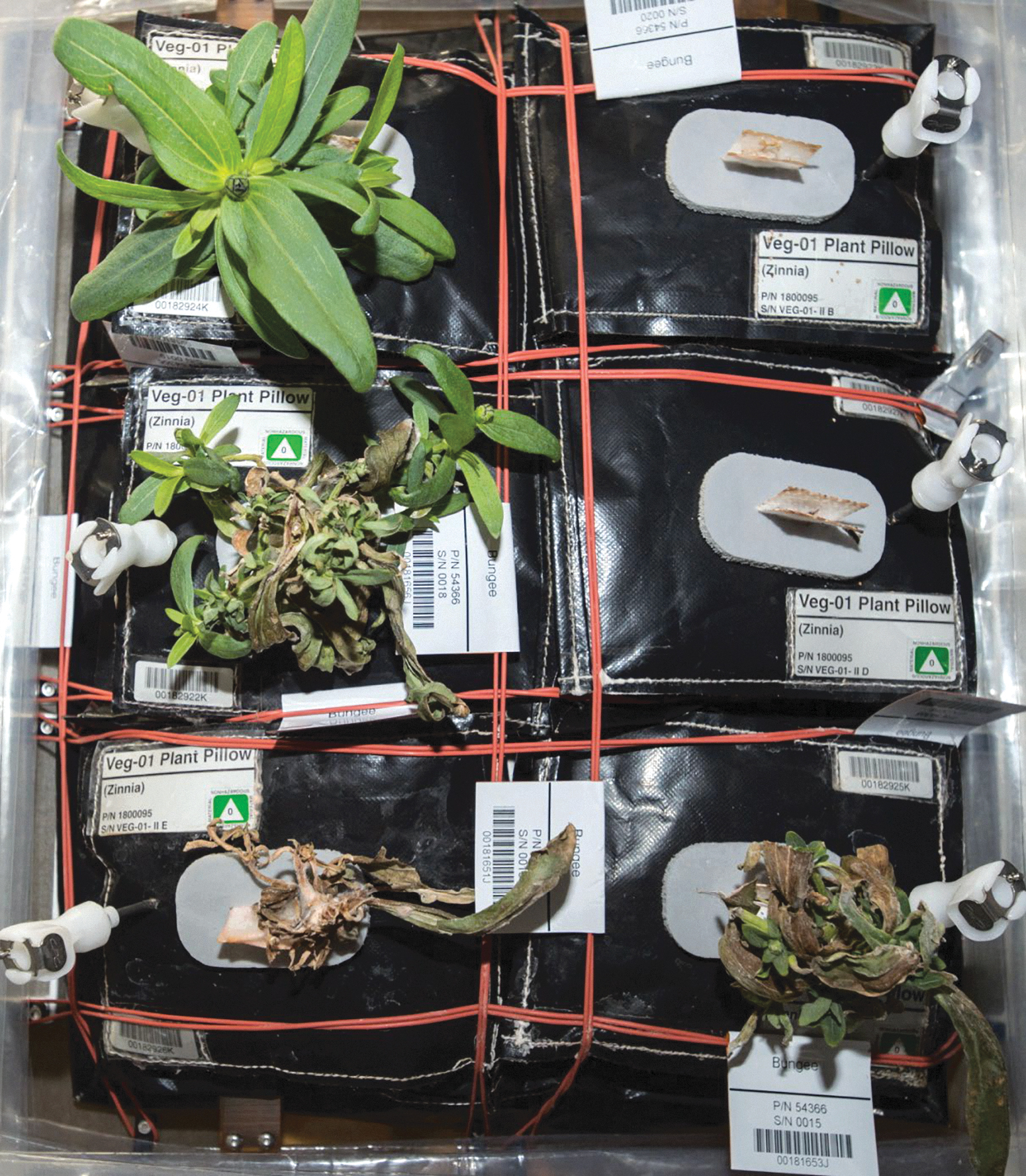

Flash-photography of the VEG-01C plants on 08-Jan-2016. Plant pillows are designated A (upper left) through F (lower right) moving in a left–right and top–down direction. Plant seedlings in pillows A, B, and D exhibited delayed seed germination, with plants in pillows B and D failing to develop, but plants in pillow A eventually developed into the only non-infected zinnia plant in the experiment. The first occurrence of the disease (e.g., Fig. 3A) appeared on plant stems and leaves in pillows E and F on 22-Dec-2015. The image shown here and taken on 08-Jan-2016 (i.e., 17 days after first symptoms) exhibits healthy (pillow A), diseased (pillow C), and dead (pillows B, D, E, and F) plants. (Image courtesy of NASA; iss046e005913.) Figure 4 is approximately 60 cm (high) by 30 cm (wide).
After 53 days after initiation (08-Jan-2016), two wilted and moribund plants (lower two plants in Fig. 4) were harvested and frozen at -80°C and later returned to Earth for diagnosis. Increased airflow and conscientious care by the crew allowed the two surviving plants to flourish. By 61 days of growth (16-Jan-2016), flowers on both plants had opened. Plants continued to flower; and after 90 days of growth, 28 flowers were harvested, with one being removed prior to this for photographic opportunities in the ISS Cupola (cover image of this issue). Both non-symptomatic plants and associated pillows were frozen at -80°C and returned for analysis. Half of the harvested flowers were returned at ambient temperature for seed testing. The remainder of the flowers were preserved by the crew as souvenirs.
2.2. Ground PathLab and MicroLab analyses
VEG-01C diseased tissues were frozen at -80°C on board the ISS on day-53 and returned to Earth on 14-May-2016 on board the SpaceX-CRS8 capsule. The frozen tissues arrived at the KSC microbiology labs on 18-May-2016 and were divided between a University of Florida plant pathology lab (PathLab) and the KSC microbiology lab (MicroLab). The following protocols were started on 23-May-2016 and are presented in sequential order to describe the process utilized to diagnose the causal agent of the ISS zinnia disease.
In the KSC MicroLab, symptomatic tissues were processed by first weighing and washing them in sterile phosphate-buffered saline (PBS) to extract surface microbes. Samples were shaken in tubes with sterile buffer and glass beads for 2 min. Each sample extract was plated onto inhibitory mold agar (Becton Dickenson, Franklin Lakes, NJ, USA) for fungal and trypticase soy agar (TSA) (Becton Dickenson) for aerobic bacterial counts. After plating and the appropriate incubation periods, colonies of fungi or bacteria were counted and individually isolated. Pure cultures of fungal isolates were identified by using the tetrazolium dye–based metabolic microarray system called Biolog Micro ID System Filamentous Fungi plate (Biolog, Hayward, CA, USA; e.g., Pinzari et al., 2016) and confirmed by the MicroSEQ Fungal Identification System (ThermoFisher, Waltham, MA, USA) following the manufacturer's protocols. Deoxyribonucleic acid (DNA) from each culture was extracted with the PREPMAN DNA isolation media (Invitrogen, Grand Island, NY, USA) following manufacturer's protocol. The DNA was processed through the FAST MicroSEQ D2 LSU DNA Fungal polymerase chain reaction (PCR) and Sequencing kits (Invitrogen) and were sequenced on the ABI 3130 Genetic Analyzer. FASTA sequences were analyzed on the ABI 3130 Fungal Library 2.2 and then imported into BLAST for corroboration (see below).
In the PathLab, symptomatic tissues were aseptically dissected and placed on potato dextrose agar (PDA; Fisher Scientific, Pittsburg, PA, USA) for fungal isolation and on TSA for bacterial isolation. Other symptomatic tissues were divided into asymptomatic and symptomatic tissues and processed for epifluorescent microscopy (EFM), scanning electron microscopy (SEM), and recovery of the putative causal agent of disease.
First, tissues were stained with 0.1% aqueous Calcofluor (Fluorescent Brightner-28, Sigma-Aldrich, St. Louis, MO, USA), aseptically dissected for longitudinal and cross sections, mounted on glass slides, and viewed with a Zeiss Axioskop 40 EFM microscope (Zeiss, White Plains, NY, USA). Separate tissue samples were placed on the slides for observing both internal and external surfaces of all sections collected. The EFM process required approximately 20% of the available tissues.
For SEM processing, similar longitudinal and cross sections of asymptomatic and symptomatic tissues were excised and fixed in 2% glutaraldehyde in 1 × PBS for 24–36 h at 4°C. The samples were then triple rinsed in sterile and prechilled PBS (4°C) and postfixed in 1% PBS-buffered osmium tetroxide at 4°C. All samples were then washed three times in PBS and then sterile deionized water (SDIW) at 4°C, allowed to warm to 24°C, and then passed through a graded ethanol series. The final steps were to triple-rinse the samples in 100% ethanol, critical-point-dry the samples in liquid CO2 (Autosamdri-815, Tousimis Research Corp., Rockville, MD, USA), sputter-coat the mounted samples with gold-palladium (model Desk-V, Denton Vacuum, Moorestown, NJ, USA), and view them in a Hitachi SEM (model SU5000 FE-SEM, Hitachi High-Technologies America, Schaumburg, IL, USA). The SEM process required approximately 20% of the available tissues.
Third, asymptomatic and symptomatic tissues were aseptically dissected and placed on either TSA (bacteria), potato dextrose agar alone (PDA; fungi), or PDA supplemented with Tergitol NP-10 (Sigma-Aldrich, St. Louis, MO, USA) and 50 ppm chlortetracycline hydrochloride (Sigma-Aldrich) (PDATC; for fast growing fungi). The Tergitol NP-10 was used to slow down the growth of rapidly growing fungal strains, and the chlortetracycline hydrochloride suppressed bacterial growth. Because the causal agent of the disease was believed to be an airborne fungus, the combination of the three media would provide a method of determining the frequency of bacteria and/or fungi in the tissues. The isolation and recovery process required approximately 40% of the available tissues. The remaining 20% of the tissues were archived at -80°C in PBS.
In an attempt to determine the origin of the pathogen, a diversity of materials used in the Veggie plant production pillows (Massa et al., 2017a, 2017b) and Z. hybrida seeds were assayed for fungal contamination. The materials were arcillite, gasket foam, Nomex thread used to sew the pillows together, Nutricote controlled-released fertilizer, Teflon-Coated-Kevlar (TCK) used as the primary walls of the pillows, and Z. hybrida seed. All materials were placed on PDA or PDATC and incubated at 25°C for 7 days. All unique fungal colony morphotypes were subcultured and purified by hyphal tipping. All fungi were then identified with ITS sequencing, as described below.
2.3. 16S and ITS sequencing of bacteria and fungi from ISS zinnia plants
Deoxyribonucleic acid was extracted from all bacterial and fungal isolates recovered in the PathLab with the UltraClean Microbial DNA Isolation Kit (12224-50, MoBio Laboratories Inc., Carlsbad, CA, USA) and PCR amplified according to the protocols from the work of Benardini et al. (2003). The PCR amplicons were cleaned with the QIAquick PCR Purification Kit (model 28104, QIAGEN Sciences, Valencia, CA, USA). All bacteria were sequenced with the universal 16S primers B27F (5'-GAGTTTGATCMTGGCTCAG-3') and B1512R (5'-AAGGAGGTGATCCA NCCRCA-3') as described previously (Lueders et al., 2004). All fungal isolates were sequenced with the ITS1F (5'-CTTGGTCTTTAGAGGAAGTAA-3') and ITS4 (5'-TCCTCCGCTTAT TGATATGC-3') primers, as described previously (Toju et al., 2012). Purified PCR amplicons were Sanger sequenced at the Interdisciplinary Center for Biotechnology Research (ICBR) at the University of Florida (Gainesville, FL, USA).
Taxonomic affinities were based on the National Center for Biotechnology Information (NCBI) BLAST function 1 . Isolates were identified to species based on affinities ≥97.5% (Stackebrandt et al., 2002). Bacterial isolates were archived at -80°C in 25% (final concentration) sterile aqueous glycerol solutions, and fungal isolates were archived at −80°C in sterile PBS buffer with 10% (final concentration) sterile aqueous glycerol. All 16S and ITS sequences were uploaded to GenBank 2 (Anonymous, 2011), and accession numbers are given in the data tables described below.
2.4. Multi-locus sequence typing on Fusarium isolates
2.4.1. Isolates
Additional multi-loci sequencing (see below) of eight key Fusarium spp. isolates used in the pathogenicity tests (Section 2.5) was completed to improve the identification of the presumptive pathogen strains derived from only ITS sequencing (Section 2.3). Eight F. oxysporum isolates from infected ISS Z. hybrida plants (ASF-151, ASF-154, ASF-159, ASF-162, ASF-164, ASF-167, ISS-F3, and ISS-F4) obtained from the ISS were used for the analysis. The ISS-F3 and ISS-F4 were recovered from the ISS galley and characterized by personnel at the Jet Propulsion Laboratory (Pasadena, CA, USA; see Urbaniak et al., 2019) and were graciously provided by JPL for this study. The F. oxysporum isolates were grown on 90 mm diameter Petri plates with PDA and layers of permeable GelAir cellophane on the agar surfaces (Bio-Rad Laboratories Inc., Hercules, CA, USA). Once isolates reached the edges of plates, mycelia were scraped and stored in PBS at −80°C for DNA extraction
2.4.2. DNA sequencing
Genomic DNA from all fusaria strains was extracted by using a DNeasy Plant Mini Kit (Qiagen, Valencia, CA, USA) according to the manufacturer's protocol. Multi-locus sequence typing was carried out employing sequences of the nuclear ribosomal DNA intergenic spacer region (IGS-rDNA), translation elongation factor 1α gene (TEF), and β-tubulin gene (BT). Previous studies on various F. oxysporum formae speciales have shown that the above three loci are better candidates in characterizing the genetic diversity F. oxysporum species complex (O'Donnell et al., 2009; Laurence et al., 2014; Cañizares et al., 2015). Published reports (Ramdial et al., 2017) as well as our experience indicate ITS sequences of Fusarium isolates are less informative and therefore not included here.
Three overlapping primer pairs were used (iNL11/iNLr; NLa/CNSa; CNS2/iCNS1) to sequence IGS region (O'Donnell et al., 2009) as it was more than 2000 bp long. These contigs were aligned together to form a single contig with Geneious 9.1 software (Biomatters Inc., Newark, NJ, USA). Partial sequencing of TEF (∼690 bp) and BT (∼600 bp) regions was carried out by primer pairs EF1/EF2 and T1/T2, respectively (O'Donnell and Cigelnik, 1997; O'Donnell et al., 1998). PCR was performed as described by O'Donnell and Cigelnik (1997) with Phusion High-Fidelity DNA Polymerase (New England BioLabs Inc., Ipswich, MA, USA) in an Eppendorf Mastercycler thermocycler (Hamburg, Germany). The following program was used for amplifying BT locus: 1 cycle of 150 s at 98°C; 32 cycles of 10 s at 98°C, 50 s at 52°C, and 55 s at 72°C; followed by 1 cycle of 10 min at 72°C and a 4°C soak. The TEF primers, and iNL11/iNLr and CNS2/iCNS1 of IGS, had an annealing temperature of 55°C while NLa/CNSa primer pair was amplified by using an annealing temperature of 65°C. The conditions of denaturing and extension steps for TEF and IGS primer pairs were the same as BT. All PCR fragments were visualized and verified for correct size range by electrophoresis on 1.2% agarose gels stained with Sybr Green I Nucleic Acid Gel Stain (Invitrogen, Waltham, MA, USA) using 1 × TEA buffer. The PCR products were purified with a Wizard SV Gel and PCR Clean-UP System kit (Promega Corporation, Madison, WI, USA) and shipped to Eurofins Genomics, LLC (Louisville, KY, USA) for direct sequencing with both forward and reverse primers.
2.4.3. Bioinformatics and phylogenetic analysis
The forward and reverse sequences of the IGS, TEF, and BT fusaria genes in all isolates were aligned, and consensus sequences were made using Geneious Version 9.1. Sequences and accession numbers of all isolates were deposited in the GenBank for reference (Supplementary Table S1). GenBank-deposited Fusarium oxysporum sequences that were closely related to the above eight isolates were determined by BLAST search and incorporated in the analysis. Multiple alignments of sequences were created for each gene using MUSCLE algorithm (Edgar, 2004) in MEGA 7 software (Kumar et al., 2016) for each locus. The missing sequence data at each end of the multiple alignments were trimmed to remove ambiguous regions; then phylogenetic trees were constructed after combining IGS, TEF, and BT alignments together. Phylogenetic analysis was done in MEGA 7 to generate a neighbor-joining (Saitou and Nei, 1987) tree with Kimura-2-parameter (K2P) (Kimura, 1980). Fusarium sibiricum isolate NRRL 53421 was used as the outgroup (see Table S1 for GenBank accessions). To ascertain integrity of clades, bootstrap tests were performed with 1000 replicates per run (Felsenstein, 1985).
2.5. Koch's postulates with Fusarium isolates
Preliminary results (May 2016) from the EFM, SEM, and IR processes suggested that the causal agent of disease (henceforth, pathogen) was a strain of Fusarium sp. (see Results for more details). To confirm whether the Fusarium sp. isolates were responsible for inciting disease on the ISS Z. hybrida plants, a series of Koch's postulates (KP) (Agrios, 2005) assays were conducted with six Fusarium oxysporum strains isolated from symptomatic Z. hybrida tissues and two Fusarium oxysporum strains obtained from the galley in the living quarters on the ISS (Urbaniak et al., 2019).
Zinnia hybrida (cv. ‘Profusion’) plants were sown in hydroponic-grade rockwool (FloraFlex, Los Angeles, CA, USA) for 7 days and then transplanted into separate containers with 250 cc of arcillite (Turface Proleague, Profile Products, LLC, Buffalo Grove, IL, USA), supplemented with 7 g/1000 cc of controlled-release fertilizer pellets called Nutricote (18-6-8, type 180, Florikan, CRF, Sarasota, FL, USA) (see Massa et al., 2016, 2017a, 2017b). The seedlings were irrigated twice on day-1 and day-7 after transplanting with a modified Hoagland's solution (Schuerger and Hammer, 2009) and thereafter with SDIW. Zinnia plants were grown in separate 250 cc Magenta Vessels (Sigma-Aldrich) placed in separate dishes to prevent cross-contamination via irrigation drainage.
Supplementary Figure S1A shows the plant-growth system setup on a bench top in the PathLab with a fluorescent (FL) lamp as the primary light source (model Sun Blaze, T5HO-44, Sunlight Supply Inc., Vancouver, WA, USA) bordered by reflective aluminum shields on the back and front sides of the plant production area. On either side of the FL lamp, HOBO dataloggers (model U12-012, ONSET Computer Corp., Bourne, MA, USA) were positioned to monitor the temperature and relative humidity of the plant production area (henceforth called the KP zone), and small muffle fans were driven by 12-V DC batteries for circulating the air within the KP zone. Environmental conditions reported for the VEG-01C experiment on board the ISS and for the KP assays are given in Table 1.
Four-week-old plants were inoculated in one of two ways to study the effects of the F. oxysporum strains on leaves, stems, and roots. First, cultures of the F. oxysporum strains were grown on fresh PDA for 7 days at 25°C. Plant stems were superficially wounded to a depth of approximately 0.5 mm with sterile scalpels at the apical buds and at three proximal stem nodes. Mycelium was harvested on the ends of sterile scalpels (∼1 mg) and placed directly on the wounded stem tissues. Plants were immediately bagged with 1 L polypropylene bags (Fig. S1) and placed under the FL lamp in the KP zone (henceforth, Protocol-1). Second, 40 mg of mycelium and microconidia were macerated in 1 mL of 0.5 × PBS with separate microfuge tube mortars and pestles (DWK Life Sciences Kimble Kontes Pellet Pestle systems; model K749515, Fisher Scientific). Two 1 mL aliquots of macerated fusaria cell debris were injected into two opposite sides of the plant root zones. Plants were again immediately bagged and placed under the FL lamp (henceforth Protocol-2). The two inoculation protocols were completed on separate plants. Both protocols were conducted twice on healthy Z. hybrida plants over the course of approximately 4 months of KP assays. The bagged plants were kept under the FL lamps for 14 days to induce a reduced-airflow high-water stress environment, as was observed with the Z. hybrida plants on the ISS. Symptoms and signs of the disease and pathogen, respectively, were observed daily for 14 days. And lastly, a similar set of plants were inoculated as described above but were bagged for only 1 day followed by 13 days of normal growth under the FL lamps. Plants were irrigated as required over the 14-day assays.
3. Results
3.1. Symptoms and signs of disease
Initial symptoms of stressed Z. hybrida plants on the ISS were epinastic curling of leaf margins and twisting of leaves in canopies (Fig. 2A), followed by leaf guttation (Fig. 2B) and water-soaking of apical buds (Fig. 2A). Excess water was attributed to a temporary failure of the circulating fans in the Veggie unit (Massa et al., 2017a), which lasted for 6 days (precise time unknown) from 16-Dec-2015 through 22-Dec-15. On 22-Dec-2015, the apical buds showed signs of fungal infections in which aerial mycelium and leaf margin necrosis were observed on most plant canopies (Fig. 3A). At this time, aerial mycelium was also observed colonizing the Nomex thread used to sew the plant irrigation pillows together (Fig. 3B, arrows). It was believed at the time (late December 2015) that the symptoms and signs represented an active and aggressive fungal pathogen attacking Z. hybrida plants because of the combination of aerial mycelium, leaf margin necrosis followed by whole-leaf necrosis, and eventual wilting of plant canopies (Figs. 3A and 4). By 08-Jan-2016, two of six plants failed to establish healthy canopies during germination, two plants were dead, one plant was severely damaged by the disease, and only one plant appeared fully healthy (Fig. 4). Plant tissues from the bottom two plants in Fig. 4 were harvested on 10-Jan-2016, frozen at -80°C, and returned to Earth for diagnosis of the pathogen.
3.2. Diagnosis process with infected zinnia tissues from the ISS
Infected and healthy tissues from the ISS-grown Z. hybrida plants returned to Earth on 14-May-2016. Tissues arrived at KSC on 18-May-2016 and were available for diagnostic processing on 23-May-2016. The following diagnostic protocol is presented in sequential order to demonstrate the process and time involved in diagnosing the pathogen on Z. hybrida plants.
First, on 23-May-2016, symptomatic tissues from ISS Z. hybrida plants were photographed with a stereo microscope (model Stemi 2000-C, Zeiss) fitted with a digital camera (model ProgRes Speed XT, Jenoptik Optical Systems, Jena, Germany) (Fig. 5). Symptomatic stem, leaf, and root tissues were aseptically dissected and placed in SDIW for visual examination. In general, all longitudinal stem sections exhibited an advanced stage of decay with vascular bundles, cortex, and apical meristems (Fig. 5A) discolored, moribund, and with obvious loss of tissue integrity. Aerial mycelium was obvious on all external surfaces of stems and leaves (Figs. 5B and 5C). Root sections were also moribund, discolored, and in an advanced state of decay (Fig. 5D).
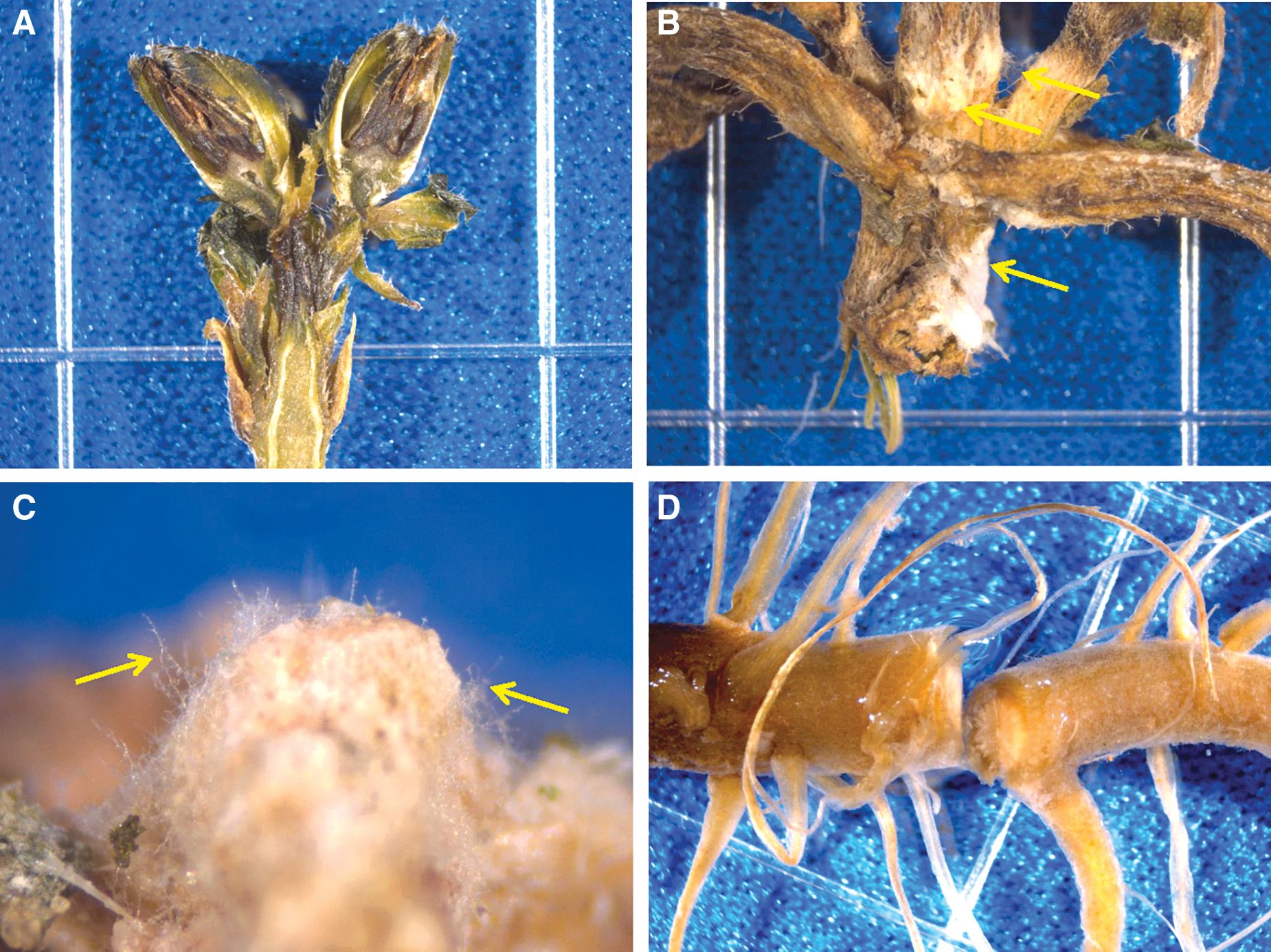
All diseased plants from the ISS were returned to Kennedy Space Center, FL, on 18-May-2016. Plant tissues were cut into longitudinal, transverse, and cross sections and then processed for recovery of the presumptive causal agent of the disease beginning on 23-May-2016. (
Second, aerial mycelium on leaves, apical buds, stems, and roots were aseptically excised from tissues, stained with 0.1% Calcofluor, and examined under EFM. On all tissues, microconidia were observed attached to the distal ends of short monophialides (i.e., conidiophores) (Fig. 6A), with the conidiophores periodically developing into more complex polyphialides (data not shown). However, no macroconidia or chlamydospores (Nelson et al., 1983) were observed on any tissues examined. The structure, abundance, and organization of the microconidia and monophialides suggested that the aerial mycelium was derived from fungi in the genus Fusarium (based on Nelson et al., 1983; Leslie et al., 2006). However, with no macroconidia present on diseased zinnia tissues, no further speciation was possible at the time. And lastly, individual conidia were observed with germtubes penetrating stomates on the abaxial surfaces of leaves (Fig. 6B) and thus were suspected as being the primary propagule for plant infection.
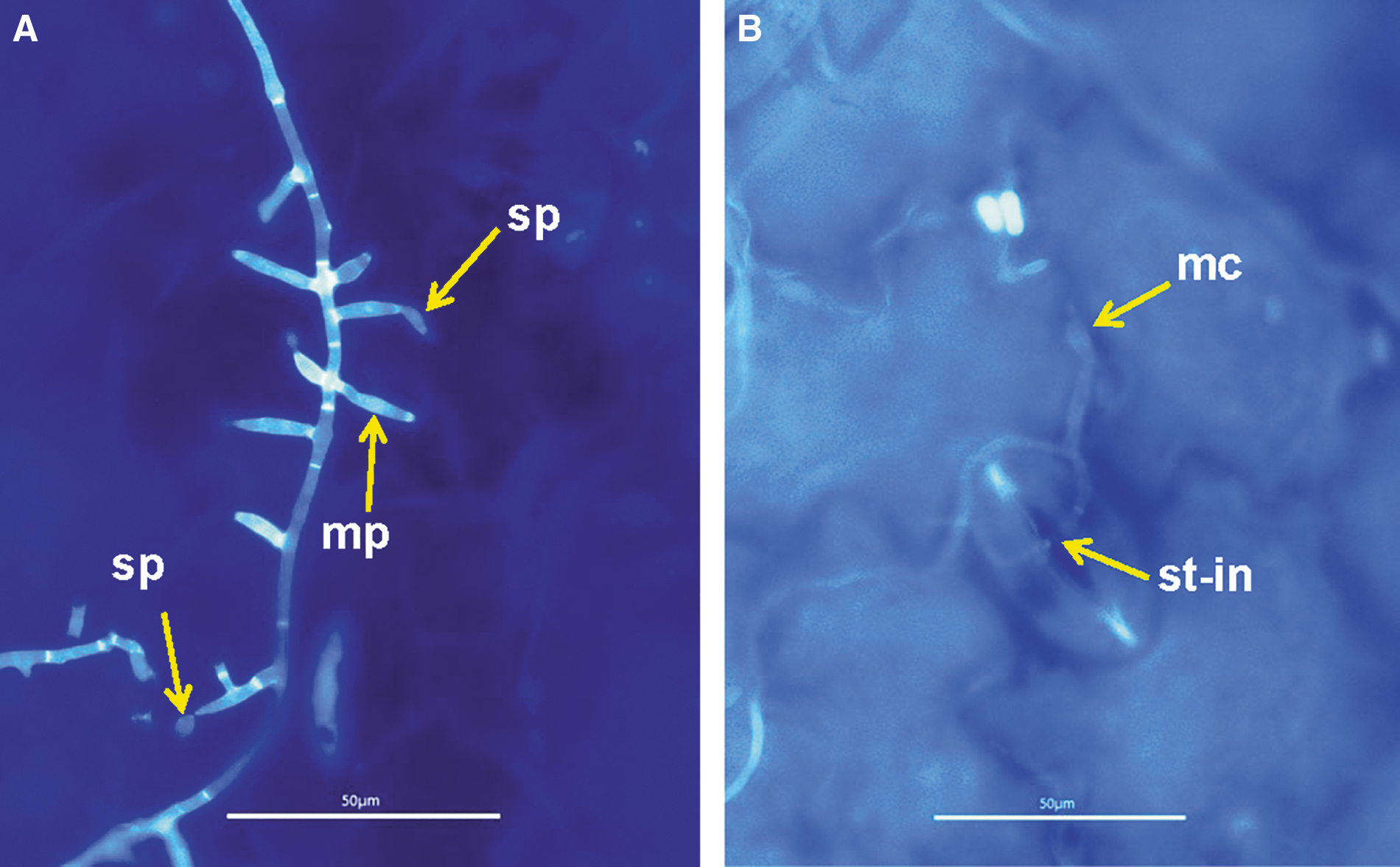
Fluorescent stained (0.1% Calcofluor) cells and aerial mycelium for Fusarium oxysporum on ISS Zinnia hybrida tissues imaged on 23-May-2016 in the lab. (
Third, unstained tissues were aseptically dissected from symptomatic leaves and plated on TSA, PDA, and PDATC, as described above. The cultures were examined daily for 7 days, and bacterial and fungal colonies with unique colony-morphologies were either streak purified (bacteria) on TSA or hyphal-tipped purified (fungi) on PDA. The isolation of diverse bacteria from symptomatic tissues yielded no consistent morphotypes. However, >95% of the fungal isolates from PDA or PDATC yielded a single colony morphotype composed of purple-pink aerial mycelium (Fig. S1B) with abundant microconidia. Again, no macroconidia or chlamydospores were observed on pure cultures of the presumptive fusarium pathogen.
Fourth, purified cultures of fungal mycelium were processed within 72 h for taxonomic affiliation with the Biolog Fungal ID system. All tested fusarium-like cultures were identified as Fusarium oxysporum. Other non-fusarium fungal isolates were subsequently identified through ITS sequencing (described below). Thus, within 3 days of the diagnostic processing on the ground at KSC, we had a strong candidate species for the causal agent of the ISS zinnia disease—namely, F. oxysporum. However, at this point the recovered presumptive pathogen was still only suspected of being the causal agent of disease because F. oxysporum is a ubiquitous species with numerous nonpathogenic and pathogenic strains described (Nelson et al., 1983).
Fifth, a similar cohort of the zinnia tissues—as described above for the visual light microscopy with Calcofluor—were fixed and processed for SEM imaging. The SEM protocols took approximately 1 week to complete. Abundant monophialides and microconidia were observed on symptomatic leaf, stem, and root tissues (Fig. 7A). In addition, individual microconidia were observed to form germtubes that subsequently infected host tissues only through stomates (Fig. 7B). Infection sites were not observed for stem or root epidermal cells. However, intracellular and intercellular mycelia were observed in cross-sections of vascular bundles in the roots and stems (Fig. 7C). And lastly, abundant conidiophores were observed emerging from leaf stomates (Fig. 7D), indicating that stomates served as the sites for both infection and for the formation of dispersal conidiophores. The isolation and characterization protocols for the fusaria outlined above took approximately 3 months to complete.
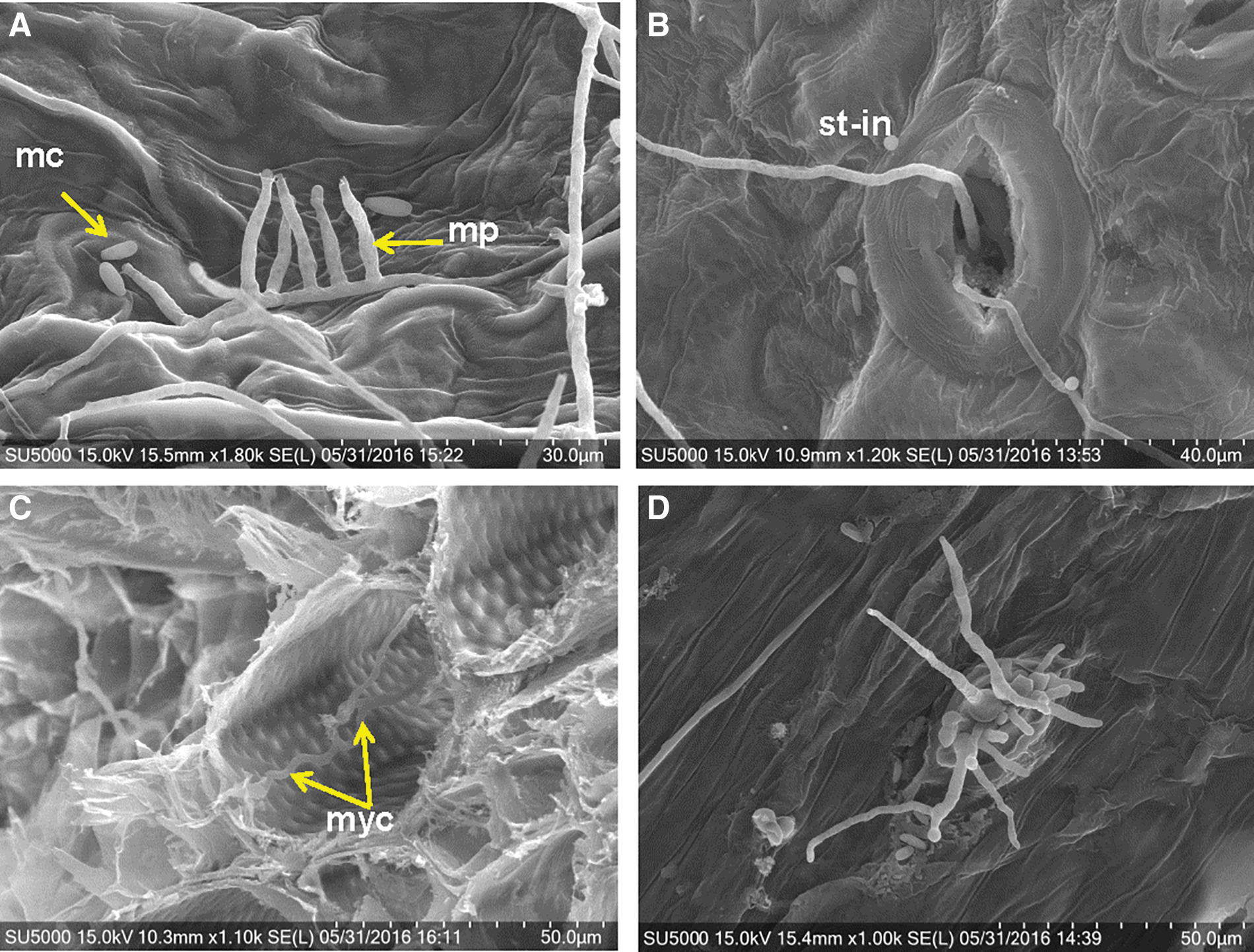
Scanning electron microscope images taken of fixed Zinnia hybrida tissues returned from the ISS and infected with Fusarium oxysporum. (
3.3. Koch's postulates
The process of diagnosing a causal agent of disease (i.e., a pathogen) in a manner consistent with historical norms is to move through a 4-step process called Koch's postulates (KP) (Agrios, 2005). The following steps must be completed successfully in order to conclude that a presumptive pathogen is in fact the actual pathogen. Step-1: the presumptive pathogen must be consistently isolated from symptomatic tissues. Step-2: the microorganism from diseased tissues must be separated from other microbes and grown in pure culture. Step-3: the pure cultures of the possible pathogen must cause disease when inoculated into healthy host tissues. Step-4: the presumptive pathogen must be re-isolated from newly diseased tissues and grown in pure culture. Failing to complete all four steps renders the diagnosis of the pathogen equivocal.
Step-1 and Step-2 were described above in Section 3.2. Eight pure cultures of F. oxysporum (strains ASF-151, ASF-154, ASF-159, ASF-162, ASF-164, ASF-167; and ISS-F3, ISS-F4) were tested individually against healthy Z. hybrida plants in the lab setup outlined above. Two types of plant inoculations were conducted. For KP Protocol-1, all inoculated plants incubated for 24 h in the polyethylene bags, and then uncovered for 13 additional days, developed no symptoms at the points of inoculation for all strains tested. In sharp contrast, all inoculated plants incubated 14 days in the polyethylene bags under the FL lamps (Fig. S1) developed symptoms (Fig. 8). No symptoms or signs of F. oxysporum were observed in non-inoculated control plants maintained for 14 days in the polyethylene bags. Longitudinal sections exhibited diseased vascular bundles and cortical tissues proximal to the inoculated apical buds (Fig. 8A). The inoculated apical buds developed stem rot symptoms advancing proximally from the inoculation points, with leaf necrosis and finally leaf/stem wilt (Fig. 8B). Inoculated lateral nodes initially developed leaf chlorosis and vascular rot moving away from the inoculation points (Fig. 8C). And finally, cream-colored aerial mycelia were observed on moribund leaves (Fig. 8D) and apical buds (Fig. 8B). The symptoms in the high-water stressed plants inoculated with strains of the presumptive F. oxysporum pathogen developed symptoms and signs that were nearly identical to zinnia plants grown on the ISS (Fig. 5).
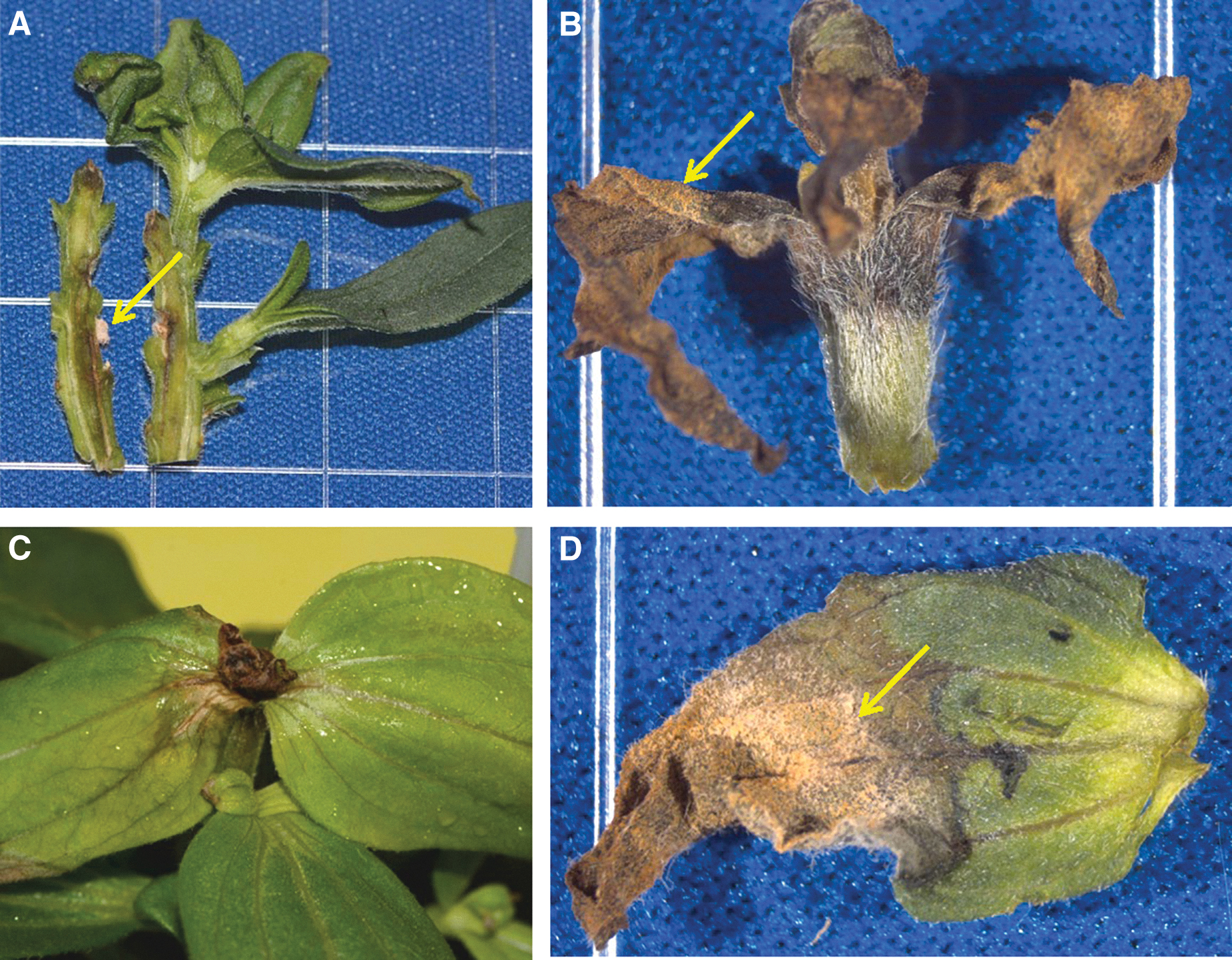

Ground-based symptomatic tissues from Zinnia hybrida confirming Koch's postulates. All leaf and stem tissues inoculated with Fusarium oxysporum strains and incubated under water-stress-inducing conditions developed symptoms that mimicked those observed during the VEG-01C experiment on the ISS (see Fig. 5). (
For KP Protocol-2, macerated mycelia in PBS buffer were used to inoculate roots of both water-stressed and non-water-stressed plants. In all cases, root inoculations failed to incite any root, stem, or leaf symptoms; thus, root infections were not believed to be the primary mode of infecting Z. hybrida plants in the ISS Veggie system.
And finally, symptomatic tissues from the high-water-stressed (i.e., bagged) plants were aseptically assayed for the presence of the presumptive F. oxysporum pathogen by placing symptomatic tissues on both PDA and PDATC. In all cases, symptomatic tissues from ground KP assays (Fig. S1C) yielded cultures of F. oxysporum identical to the colony morphotypes from ISS zinnia plants (Fig. S1B). Thus, all four steps of Koch's postulates were successfully completed for strains of F. oxysporum initially recovered from Veggie-grown Z. hybrida plants on the ISS. However, because disease only appeared on high-water-stressed plants, F. oxysporum was labeled as an opportunistic pathogen of Z. hybrida during the VEG-01C experiment.
3.4. Isolation of fungi from Veggie materials
To determine whether the opportunistic pathogen F. oxysporum was present on materials within the Veggie pillows, several materials were assayed for fusaria during the KP assays. The materials included small samples from the preflight archives of (1) Teflon-Coated-Kevlar (TCK) used as the bulk outer walls of the propagation pillows, (2) Nomex thread used to sew the walls of the pillows together, (3) gasket foam used to secure the wicks into the tops of the pillows, (4) arcillite propagation medium, (5) Nutricote T180 slow-release fertilizer granules, and (6) Z. hybrida seeds. All materials are described previously (Morrow and Remiker, 2009; Massa et al., 2017a, 2017b).
The focus for isolating and identifying microorganisms from pillow materials and zinnia seeds was to recover all unique fungal colony morphotypes but not to isolate all bacteria that might be present. Table 2 presents the fungi that were recovered from all materials assayed on PDA and PDATC (see also Fig. S2). No F. oxysporum strains were recovered from any of the assay plates (total of 30 plates). However, a F. sporotrichioides strain was recovered from the gasket foam. Other common fungi recovered from the Veggie materials were Alternaria alternata, Aspergillus niger, Cladosporium cladosporioides, Penicillium spp., and Perconia sp. (Table 2; Fig. S2). TSA was not used as an assay medium, so few bacteria were observed on the PDA or PDATC plates. And lastly, the Z. hybrida seeds were remarkably free of bacteria and fungi, with the exception of recovering a single isolate of Penicillium sumatrense from one of 30 seeds assayed.
Fungal Isolates from Materials Used in the Veggie Plant Propagation Pillows Assayed for the Presence of Fusarium oxysporum
See Figure S2 for visual details of the fungi recovered from these materials on PDA.
Based on these results, we concluded that the F. oxysporum opportunistic pathogen on ISS Z. hybrida plants was not present in the materials that were used for the Veggie VEG-01C plant pillows and zinnia seed. Furthermore, for the actual VEG-01C flight experiment, these materials were sterilized prior to launch. We understand that culture-based assays may miss viable but non-germinating cells. Based on these data and considerations, we conclude that inoculum of the opportunistic pathogen—F. oxysporum, that incited disease in Z. hybrida on board the ISS—was present in the ISS environment prior to the start of the VEG-01C experiment.
3.5. Isolation and KP assays for F. sporotrichioides isolated from gasket foam material
Of the fungi recovered from the Veggie materials, KP assays were conducted for the single strain of F. sporotrichioides recovered from gasket foam. The KP assays were conducted as described above for the ISS/Z. hybrida strains of F. oxysporum. Results were surprising and indicated that F. sporotrichioides was a more aggressive foliar pathogen than the F. oxysporum strains tested (compare Figs. 8 and 9). Aerial mycelia of F. sporotrichioides were observed on all inoculated plants within 24 h (Fig. 9A), followed by rapid development of disease, necrosis of stem and leaf tissues, and eventual wilting of plant canopies after 7 days (Fig. 9B). However, no root symptoms developed with F. sporotrichioides after 14 days, similar to the KP assays for the F. oxysporum strains.
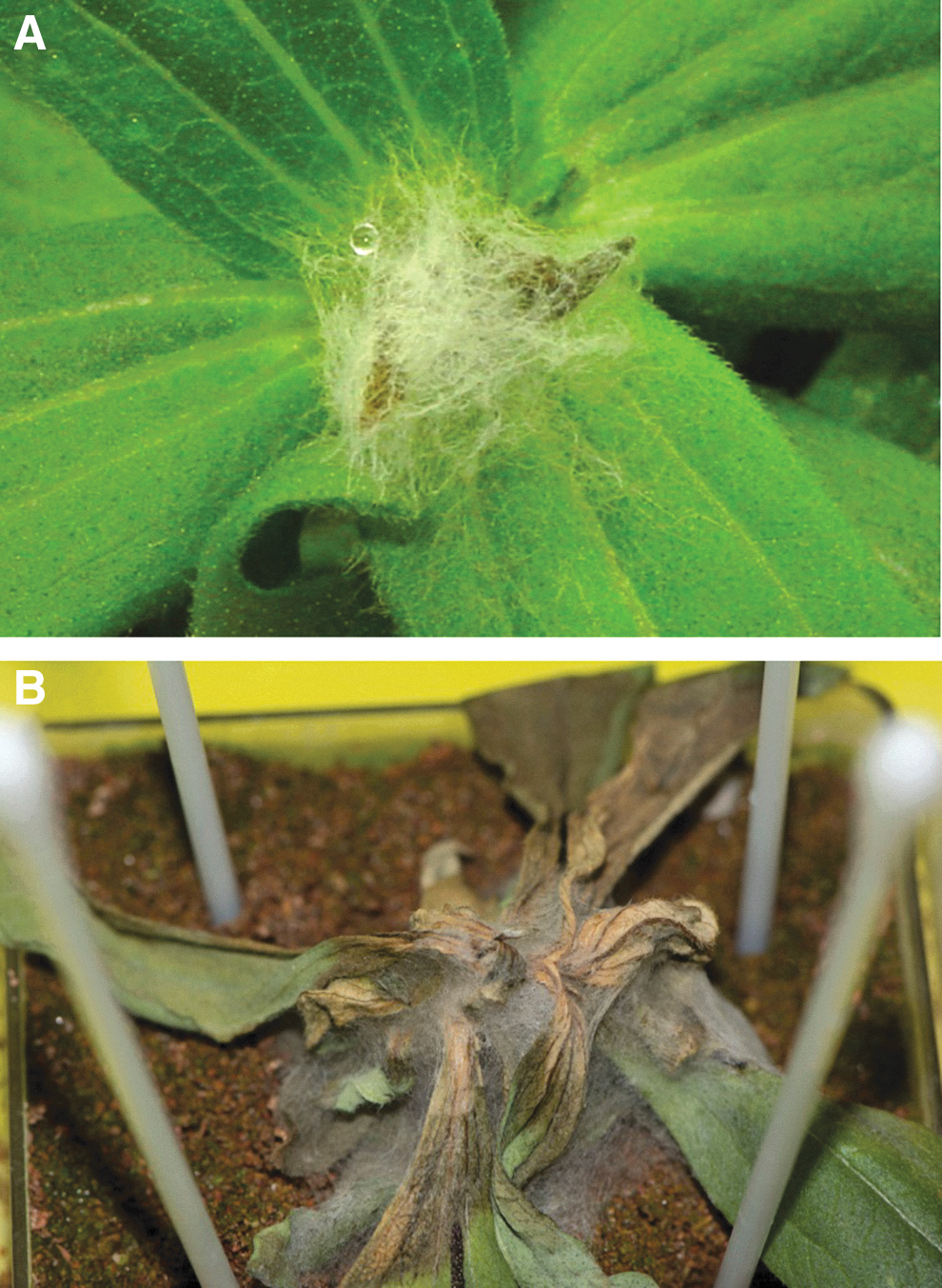

Fusarium sporotrichioides was isolated from gasket foam during ground-based assays of materials used in the Veggie plant pillows, but not from flight samples of infected Zinnia hybrida tissues. Zinnia plants were inoculated with F. sporotrichioides and incubated as described in the text. Ground-based symptoms in inoculated plants were much more severe for F. sporotrichioides shown here than for the ISS strains of F. oxysporum (Fig. 8). (
3.6. Phylogeny of flight F. oxysporum compared to other type-strains of Fusarium
The phylogenetic analysis was carried out with eight F. oxysporum isolates obtained from the ISS and twelve different closely related formae speciales of the same species. The phylogenetic tree (Fig. 10) clearly separates ISS-F3 and ISS-F4 from the six ASF isolates obtained from symptomatic zinnia plants. According to a previous analysis, ISS-F3 and ISS-F4 grouped closely with F. oxysporum f. sp. lycopersici and radicis-lycopersici (Urbaniak et al., 2019). The present study corroborates these previous results and indicates that the zinnia isolates showed clear distinction from ISS-F3 and ISS-F4 isolates by grouping closely together but separately within the tree. The average number of base differences per sequence between the zinnia and ISS groups was 61.8 bp. The multiple sequence alignment of three gene regions (IGS, TEF, and BT) of the eight isolates consisted of 3250 bp with gaps. There were 61 parsimony informative sites with two variants between zinnia and ISS-F3/-F4 isolates. The combined data set had unique sequences for all isolates except for ASF-159 and ASF-162, which were similar.

The phylogenetic relationships of Fusarium oxysporum isolates obtained from the ISS were inferred by constructing a neighbor-joining (NJ) tree using combined sequences of translation elongation factor (TEF), β-tubulin (BT), and intergenic spacer (IGS) regions. The bootstrap values from 1000 replications are shown next to the branches. The analysis also included several F. oxysporum sequences closely related to isolates under investigation. Fusarium sibiricum was used as the outgroup.
4. Discussion
4.1. Fusarium oxysporum and Zinnia hybrida disease complex
Many plant species have been proposed for long-term space-based BLSS modules that include a wide diversity of agronomic and vegetable crops (e.g., wheat, sweet potato, potato, rice, salad greens, swiss chard, radishes, tomatoes) (see Hoff et al., 1982; Wheeler et al., 2004; Tako et al., 2010; Zabel et al., 2016). Many of these crops also have been grown in large-scale BLSS testbeds in Japan, Russia, and the United States (see reviews by Tako et al., 2010; Wheeler, 2017) in order to model plant growth in highly controlled hydroponic systems similar to those proposed for BLSS modules. More recently, a subset of these vegetable crops (e.g., salad greens, swiss chard, kale, mustard greens) have been tested in space within small-scale plant-growth chambers (e.g., Zabel et al., 2016; Massa et al., 2017a) to focus on developing hardware and screening cultivars to supply fresh salad crops to astronauts for near-term ISS and Moon missions. As new systems are launched and used over many crop cycles, it is possible that the microbiomes of the systems will increase in both diversity and biomass that could create opportunities for disease development on the crops. We report here on the first outbreak of an opportunistic Fusarium oxysporum/Zinnia hybrida disease complex (called a pathosystem) within the Veggie plant-growth chamber on the ISS.
The genus Fusarium is a cosmopolitan group of fungi capable of saprophytic growth in soil, urban, and aquatic ecosystems, as well possessing strains capable of causing disease in plants, humans, and animals (Nelson et al., 1981; Summerell et al., 2001). Regarding plant pathogens, Fusarium spp. are generally assigned species-level taxa based on morphological characteristics of conidia and conidiophores with subspecies designations (called formae speciales; abbrv. ‘f. sp.’) depending on DNA sequencing and pathogenicity assays (Summerell et al., 2001; van Dam et al., 2016). For example, F. oxysporum f. sp. lycopersici would be morphologically similar to most other F. oxysporum strains but be capable of causing wilt disease in many tomato cultivars. Fusarium spp. can produce microconidia, macroconidia, and chlamydospores. In most cases, microconidia (approximate size: 5–12 μm in length and 2.2–3.5 μm in width; Booth, 1971) are the primary airborne propagules for the dispersal of most species in the environment.
In late 2015, a crop of Z. hybrida was propagated in the Veggie VEG-01C experiment on board the ISS as a technology demonstration project. On 22-Dec-2015 visible signs (i.e., aerial mycelium) and symptoms (i.e., chlorosis, necrotic leaves) (Fig. 3A) were observed on two zinnia plants during a routine inspection by astronauts of the VEG-01C experiment. Based on the digital images returned to Earth, no definitive diagnosis of the microbial problem was possible at that time. Initially, it was thought that the aerial mycelium might be caused by the fungus Botrytis cinerea based solely on the color and structure of the aerial mycelium (Fig. 3A). Over time, the active disease spread to an adjacent plant (Fig. 4), causing severe necrosis of leaf canopies on three separate zinnia plants (Fig. 4). Immediately after this observation, the astronauts attempted to mitigate the disease by removing visibly infected tissues and wiping the remaining leaves with citric acid-based wipes saturated with 1% Pro-San. Hardware was wiped with commercial benzalkonium chloride wipes. These mitigation procedures likely delayed the disease development only slightly, and on 08-Jan-2016 new diseased tissues were observed, harvested (on 10-Oct-2016), frozen at -80°C, and returned to Earth on 18-May-2016.
As described above, a 9-month diagnostic effort was initiated on 23-May-2016 to characterize the causal agent of the disease and to identify environmental factors that may have promoted the development of severe symptoms on zinnia. Within 48 h of beginning the diagnostic process, and based on the morphologies of conidia and conidiophores on tissues with 0.1% Calcofluor staining (Fig. 6), the PathLab concluded that B. cinerea was not present and that the pathogen was likely caused by a Fusarium sp. The first positive indication at the species-level taxon was achieved within 1 week when purified fungal tissues from zinnia plants were tested with the Biolog Fungal ID system. Eventually, ITS sequencing (3–4 weeks) confirmed the identification of the putative pathogen as F. oxysporum. However, without completing Koch's postulates with strains isolated from diseased tissues, we could not claim with certainty that the F. oxysporum strains recovered from diseased Z. hybrida tissues were the actual causal agent of the disease. Pathogenicity tests were initiated on 20-Oct-2016 and required 6 months.
The timeframe required to officially designate F. oxysporum as an opportunistic pathogen of the ISS-grown Z. hybrida plants (approximately 9 months) is undesirable for maintaining plant-growth modules intended to supply edible produce for astronauts. Only a few plant diseases can be easily diagnosed with clear imagery of the symptoms or signs (e.g., powdery mildew on cucumbers, wheat rust). Most plant diseases require some diagnostic lab work to identify the pathogen and characterize the effects of environmental factors on disease expression. Events of the current study have been presented sequentially in order to demonstrate the time and resources that are usually required to officially designate a microorganism the causal agent of a new disease. Such an approach might only be plausible when symptomatic tissues can be returned to Earth from LEO or cis-lunar space. However, returning diseased tissues to Earth will not be possible during human Mars missions. We will shortly propose technologies and protocols that may significantly shorten this time lag.
There are a number of important traits of the ISS F. oxysporum/Z. hybrida pathosystem that are important to note. First, Koch's postulates were successfully completed with six strains of F. oxysporum recovered from ISS zinnia plants (i.e., compare Fig. 5 [ISS symptoms] to Fig. 8 [ground KP assays]). However, symptoms and signs could not be fully replicated in ground KP assays (Fig. 8) unless the plants were severely water-stressed by covering them with transparent plastic bags (Fig. S1A). Plants inoculated with F. oxysporum strains from the ISS did not induce symptoms if inoculated plants were allowed to develop outside of the plastic bags. Because films of water were observed on some surfaces of leaves and stems of the ISS zinnia plants, and these accumulated films of visible water were the sites of aerial mycelium only a few days later (Fig. 3A), we conclude that the typically nonvirulent F. oxysporum strains took advantage of the water-stressed plants in μg by infecting leaf, stem, and root tissues in the VEG-01C experiment. Once the water stress was corrected by engaging the Veggie circulation fan, and plant care changed from scheduled- to crew-determined frequencies, the disease incidence was slowed, and one of the three infected zinnia plants recovered.
Second, Urbaniak et al. (2019) reported F. oxysporum strains (i.e., ISS-F3 and ISS-F4) recovered from the galley table in the ISS module Columbus, keyed out into a separate clade from the F. oxysporum strains recovered from infected zinnia plants in the VEG-01C experiment. This is an intriguing finding because the ISS-F3 and ISS-F4 strains sequenced by Urbaniak et al. (2019) were also capable of inciting disease in Z. hybrida in the current study if the plants were incubated in plastic bags. Thus, the F. oxysporum strains from the current study (ASF-151, ASF-154, ASF-159, ASF-162, ASF-164, ASF-167; Table S1) and strains ISS-F3 and ISS-F4 from the work of Urbaniak et al. (2019) were all capable of causing disease on water-stressed zinnia plants in the KP assays, and thus all could be labeled as opportunistic pathogens of water-stressed ISS Z. hybrida in μg. However, in both the current study and that of Urbaniak et al. (2019), the DNA sequencing could not designate a formae speciales of the virulent F. oxysporum strains; thus indicating that the disease was a new expression of the capability of these F. oxysporum strains to incite disease in the unique μg environment found on the ISS. Furthermore, a second paper has been recently published (Avila-Herrera et al., 2020) that confirms the frequent occurrence of F. oxysporum species in the ISS atmosphere and on spacecraft internal surfaces. All three studies suggest that F. oxysporum strains in the ISS microbiome may be a continual source of opportunistic plant pathogens in near-term plant growing systems like Veggie.
Third, the likely source of the F. oxysporum pathogen could not be precisely demonstrated. Most of the materials used to assemble the Veggie plant pillows were assayed for the presence of F. oxysporum (including 30 zinnia seeds used in VEG-01C), and all were negative for the opportunistic pathogen. However, we did not assay these materials when the plant pillows were constructed in December 2013 (Massa et al., 2016, 2017) but instead assayed the materials during the summer of 2016. Thus, the microbial communities or assemblages on or within the plant pillows may have changed over time. Based on the recovery of F. oxysporum from within the ISS habitats (Urbaniak et al., 2019; Khodadad et al., 2020), we conclude that the most likely source of the inoculum for the disease was the endemic ISS or Veggie microbiomes. However, further ISS and Veggie microbial assays are recommended to acquire viable cultures of Fusarium spp. to thoroughly test this hypothesis.
Fourth, the red/blue LED illumination was inadequate to detect or monitor the development of the foliar and stem rots caused by F. oxysporum. Plant canopies generally appear as dark blue to black when illuminated by red/blue LEDs (Fig. 1B), even though a small amount of green light was present in Veggie. Similar problems with red/blue LED illumination systems were observed for hydroponically grown pepper and chard plants (Schuerger and Brown, 1997; Schuerger and Richards, 2006). White light illumination (Figs. 2, 3, and 4) best captured the signs and symptoms of the disease on ISS zinnia plants. This illumination problem suggests that plant-growth modules will either need white LEDs or separate electronic-flash image capturing systems to allow astronauts and ground-based plant biologists to better track the health of plants over time.
Fifth, although isolates of F. oxysporum were not recovered from any of the Veggie plant pillow materials tested (Tables 2 and S2), a strain of F. sporotrichioides was recovered from the gasket foam. Furthermore, F. sporotrichioides (Fig. 9) was apparently a more aggressive foliar rot/wilt pathogen on Z. hybrida than the F. oxysporum strains (Fig. 8) when both were tested in the KP assays. The occurrence of this new strain highlights the possibility of introducing new Fusarium spp. into plant-growth systems intended for flight to the ISS, and potentially on other deep-space missions.
And lastly, cream-colored mycelial strands were observed on the Nomex threads at the edges of plant-growth pillows during the VEG-01C experiment (Figs. 3B and 4) that were identified as F. oxysporum (Table S2). However, F. oxysporum was not isolated from stock supplies of Nomex threads during ground assays (Table 2). Thus, we conclude that it is likely that the F. oxysporum that colonized the Nomex threads during the VEG-01C experiment was derived from the extant microbiome on the ISS, or within Veggie. The presence of F. oxysporum on the flight Nomex threads highlights the possibility that materials used in spaceborne plant-growth systems might be able to support the colonization and eventual sporulation of microorganisms and possibly contribute to outbreaks of plant diseases in μg.
4.2. Plant diseases in spaceborne plant-growth systems
Although the current study is the first report of a Fusarium sp. causing a severe outbreak of a plant disease in μg, there is evidence that the genus Fusarium may be a long-term issue for spaceborne plant habitats. First, Fusarium spp. are known to cause numerous wilt and root rot diseases in hydroponically grown plants in the commercial greenhouse industry (Jarvis, 1992). Second, fusaria have been recovered from within human habitats on Apollo (Taylor et al., 1973), the Mir space station (Ott et al., 2004), and the ISS (Urbaniak et al., 2019; Khodadad et al., 2020) and likely will be present in future human spacecraft habitats because of the ease of aerial dispersal of infective propagules in this group (Nelson, et al., 1981; Summerell et al., 2001). Sources of diverse fusaria associated with spacecraft are likely from airborne propagules accumulated during prelaunch preparations, experimental payloads, equipment, consumables, and even the astronauts themselves. And third, as μg plant-growth habitats increase in size, complexity, and long-term use it is likely that their microbiomes will increase in diversity and abundance.
Historically, only two previous plant-disease complexes have been reported in μg plant habitats. Bishop et al. (1997) described a seedborne endophytic fungal contaminant that caused a 50% loss in experimental wheat plants in a shuttle plant-growth module called the Plant Growth Unit (PGU). The fungus was identified as Neotyphodium chilense and caused both root and stem rots of germinating seedlings. Second, the root pathogen Phytophthora sojae was applied to soybean roots just prior to launch on shuttle flight STS-87 (in 1997) and evaluated for causing root rots in soybean seedlings (Ryba-White et al., 2001). The experimental hardware was the Biological Research in Canisters (BRIC) system in which soybean seedlings and P. sojae infective propagules were placed in germination pouches and inserted into multiple BRIC units 24 h prior to launch. In both cases, the experimental systems were presterilized, closed, and tightly controlled systems. The infective propagules of N. chilense on wheat were located on internal tissues of the wheat seed and thus not removed by prelaunch ‘surface sanitation’ protocols with seeds. And the infective propagules of P. sojae were actively added to the experiment prior to launch. In both cases, plants were not exposed to adequate light and were exposed to high-humidity environments and μg conditions in LEO for up to 8 days, similar to the off-nominal conditions encountered by zinnia plants within Veggie. It is likely that stagnant air in and around canopies promoted disease development in all three cases during μg experiments.
Why have few plant-growth experiments on board the shuttle or ISS (see review by Zabel et al., 2016) experienced outbreaks of plant diseases over the last 40 years, and why did the F. oxysporum/Z. hybrida pathosystem occur recently? Most previous plant biology experiments in μg were conducted in sanitized and presterilized plant-growth modules that were reconditioned prior to each experiment (see review by Zabel et al., 2016). In contrast, the Veggie system was designed to serve as a plant-growth habitat for multiple cropping cycles for fresh produce for the astronauts (Massa et al., 2016, 2017a) and was open to the ISS environment (Fig. 1A). The former systems were sanitized for individual experiments; thus the microbiomes were significantly constrained prior to launch. Although the Veggie hardware was partially sanitized with Prosan wipes between crops, the Veggie system was designed to serve as a long-term salad machine on the ISS; thus its microbiome likely underwent a more dynamic successional process over time. However, keep in mind that the primary factor that created the off-nominal conditions in the Veggie VEG-01C experiment was the inadvertent shutdown of the air recirculation fan.
Why do symptoms appear more severe in μg than in ground-based assays? In the studies by Bishop et al. (1997), Ryba-White et al. (2001), and the current outbreak (2016), disease symptoms in all three pathosystems appeared more severe in μg than in ground-based assays. One common factor for all three disease outbreaks was that plants were in contained systems with high humidity and lacking buoyancy convection. Plants grown in μg have been shown to exhibit physiological changes due to low-convection and high-humidity environments (Kitaya et al., 2003); thus environmental factors alone might be the reason for increased severity of disease in μg.
Liquid water in μg is often dominated by strong surface tension forces on droplets (Bousman et al., 1996; Meseguer et al., 2014). In the VEG-01C experiment, water logging of plant canopies was observed on most plants during the phase of the disease outbreak associated with low air flow within Veggie. It is plausible that the water condensing on leaf and hardware surfaces formed multi-millimeter layers of standing water held in place by high surface tensions. Without gravity or strong air flow to disrupt these water films, the films became adhered to plant tissues, creating a barrier to gas diffusion and a niche for the easy germination of fusaria microconidia.
Furthermore, two other factors were not tested in the current experiments but should be explored in the future. Several studies have suggested that plant host-resistance mechanisms may be down-regulated in μg compared to 1g environments, including thinner cell walls, decreased lignin, altered genomics and transcriptomics profiles, constrained gas diffusion leading to the buildup of ethylene and epinasty, and root hypoxia (see reviews by Wolff et al., 2013; Ferl et al., 2015; Poulet et al., 2016; Paul et al., 2017). In addition, a number of papers describe increased microbial virulence in μg compared to Earth controls (see reviews by Klaus and Howard, 2006; Wilson et al., 2007; Rosenzweig et al., 2010). What has not yet been demonstrated is whether environmental factors alone (e.g., high humidity) can explain the increased severity of symptoms in microbial-challenged plants in μg or whether the combination of down-regulation of host-resistance genes coupled to the up-regulation of pathogen virulence genes is the primary μg process of increased disease development in space.
In summary, as spaceborne plant-growth modules are used for sequential cropping cycles, the complexity of maintaining crop health may become dynamic, requiring a diversity of mitigation protocols to keep the plants at peak performance. Although it is beyond the scope of the current study to develop a comprehensive Integrated Pest Management (IPM) program for spaceborne plant-growth modules, we suggest that long-term missions consider developing IPM protocols for plant production activities.
4.3. Integrated Pest Management (IPM) program for spaceborne plant-growth systems
Future IPM programs for spaceborne plant-growth modules (see reviews by Nelson, 1987; Schuerger, 1998, 2004) likely will be uniquely tailored for each payload, crop, and mission scenario. However, a few common concepts could be included in most mission scenarios.
First, significant risks can be obviated or mitigated by launching sanitized plant-growth modules, supplies, and consumables. Furthermore, the most aggressive plant pathogens like wheat rust, tomato mosaic virus, root-parasitic nematodes, and so on, can be excluded from spaceborne plant-growth modules by maintaining such sanitization programs for all equipment and supplies prior to launch.
Second, where possible, utilize crop cultivars that are genetically resistant to known microbial risks in space. This requires continued efforts to characterize the microbiomes within the ISS (e.g., Urbaniak et al., 2019) and plant-growth modules (e.g., Khodadad et al., 2020). For example, based on the current report of the F. oxysporum and Z. hybrida pathosystem in Veggie, host resistance should be preselected in plants (when available) against fusaria plant pathogens. Schuerger and Mitchell (Schuerger, 1991; Schuerger and Mitchell, 1992) were some of the first to suggest that Fusarium spp. are likely to comprise a group of plant pathogens in spaceborne plant-growth modules, and they recommended host resistance as one of the first lines of defense when mitigating against such outbreaks. In addition, other potential plant pathogens were recovered from the materials used for the Veggie plant pillows (Table 2), including Alternaria alternata, Cladosporium cladosporioides, and Aspergillus niger; all of these fungi can cause diseases on many vegetable and agronomic crops (Jarvis, 1992). Thus, developing an inventory of possible plant pathogens for likely crops to be grown on the ISS, in cis-lunar space, or on future Mars missions will inform plant scientists on the host-resistance genomes to be screened or developed for spaceborne plant-growth modules.
Third, the occurrence of N. chilense on wheat seed was unexpected because the wheat seed were surface sterilized prior to launch (Bishop et al., 1997). However, Bishop et al. developed a heating protocol to expose wheat seeds to 50°C for 1–6 h that completely inactivated the endophytic spores and mycelia of N. chilense. Thus, combinations of heat and surface sterilization protocols are likely to reduce seedborne pathogens more effectively than surface sterilization alone.
Fourth, automatic remote sensing capabilities in spaceborne plant-growth modules (e.g., Schuerger and Brown, 1997; Schuerger and Richards, 2006) will significantly enhance crop monitoring and disease diagnosis. However, such systems should utilize other illumination systems besides the red/blue LEDs used as the primary illumination source for the Veggie system. White light illuminated, in-focus, high-resolution images were the most helpful for monitoring crop growth and disease monitoring in the VEG-01C experiment described here.
Fifth, environmental manipulation (i.e., adjusting relative humidity in canopies, adjusting root or canopy temperatures, altering pH of nutrient solutions, etc.) can be used to control many plant diseases in hydroponic systems (e.g., Gold and Stanghellini, 1985; Jarvis, 1992; Schuerger and Mitchell, 1992). For the current disease outbreak, it is entirely possible that the leaf and stem rot on zinnia caused by the opportunistic pathogen F. oxysporum could have been completely prevented if the plant canopies would not have accumulated standing water on apical terminals and leaves (Fig. 2). Extending this concept, most foliar plant pathogens are unlikely to infect leaves or stems if the aerial humidities in canopies are held ≤70% relative humidity. Accumulating water films on zinnia leaves in the Veggie system in μg was a unique situation that was likely caused by the absence of air convection and the lack of gravity-induced settling of water on plant canopies (Kitaya et al., 2003).
In summary, complete IPM programs for specific mission scenarios will assist in the management of plant disease outbreaks in spaceborne plant-growth habitats or modules. Future IPM programs for spaceborne plants should approach the problem of disease management by first characterizing the microbiomes typically encountered in space habitats (e.g., Apollo, Mir, ISS; Taylor et al., 1973; Urbaniak et al., 2019; Khodadad et al., 2020) and looking for plausible pathogen species, as have been described for wheat (Bishop et al., 1997) and zinnia (current study). Prelaunch sanitation programs, crop diversity, use of resistant cultivars, and manipulation of environmental conditions in plant-growth habitats offer the best options to manage crop production and prevent disease outbreaks. Application of pesticides is considered a non-option in spaceborne small plant-growth modules and human BLSS systems due to bioaccumulation of toxic materials within closed life-support systems.
4.4. Technologies for rapid diagnosis of plant diseases in space
A rapid diagnostic system for spaceflight applications would have greatly accelerated the speed at which the Fusarium/Zinnia pathosystem was characterized. Instead of requiring materials returned to Earth for weeks to months of laboratory diagnostic assays, a rapid diagnostic system for the ISS or missions to the Moon or Mars should be developed that might provide the potential taxa of the most likely candidates for the causal agents of the disease. Two rapid testing systems that seem most applicable to the current discussion for space applications are the MinION DNA/RNA sequencing system (Oxford Nanopure Technologies; e.g., Castro-Wallace et al., 2017) and high-density microarrays of small-subunit rDNA (e.g., Wilson et al., 2002) or 16S rRNA (e.g., Brodie et al., 2007). The advantage of the MinION technology is that such a system is already being tested on the ISS (e.g., McIntyre et al., 2016; Castro-Wallace et al., 2017), it is currently being used for rapid field diagnosis of plant diseases (e.g., Donso and Valenzuela, 2018), and it provides complete DNA or RNA sequencing of the microbiomes on target materials. The disadvantage of the MinION is that the flow-cells are only stable for approximately 6 months, something that currently precludes the use of this technology for deep-space missions to Mars. The advantages of the microarrays are their easy use, the rapid bioinformatics associated with a completed assay, and the longer stability of the microarrays. However, microarrays have the disadvantage of only being able to detect what has been preloaded onto the microarray fluorescent chip surfaces.
Regardless of the technology selected, a rapid plant disease diagnostic system should be developed for use in near-term μg and planetary surface missions. Such a system could in theory reduce the diagnostic process from the 9 months described herein to a few hours on the ISS or in a Mars base. If disease diagnosis can be reduced to a few hours, it is plausible to manage disease outbreaks in real-time by a coordinated effort between astronauts and ground controllers.
4.5. Summary
The F. oxysporum/Z. hybrida disease outbreak during the VEG-01C experiment was caused by an operational issue (i.e., circulating fan left off for a period of time) that allowed the accumulation of significant water films on stems and leaves within the Veggie habitat. Endemic spores of F. oxysporum within the ISS or Veggie were likely the source of the opportunistic pathogen on Z. hybrida. Had a rapid diagnostic system been available on the ISS, the likely cause of the disease outbreak could have been established within a few hours to a day or two by a quick coordinated effort between astronauts and ground-control plant scientists. Instead, two zinnia plants were killed and one damaged by the outbreak of the F. oxysporum wilt/rot disease. Only a single zinnia plant was visibly unaffected.
Spaceborne plant pathology is a nascent discipline that requires significant work over the next few years to build a rapid diagnostic system and an IPM program for near-term plant-growth modules for space applications. Early success and validation of using plant biomass production systems in space—in order to provide fresh produce for astronauts—will inform the design and development of longer-term systems for Moon and Mars missions and eventually the development of large-scale BLSS modules. It is difficult to predict the precise list of plausible plant pathogens in future spaceborne plant habitats or BLSS modules due to the unique conditions in space (e.g., μg, increased ionizing radiation, closure, constrained microbial diversity, reduced plant resistance, and up-regulated microbial virulence; see reviews by Nickerson et al., 2004; Ott et al., 2012), but a comprehensive approach to disease mitigation through an IPM program offers the best way forward for using plant-based growth chambers like Veggie and larger-scale BLSS modules in future human missions to the Moon and Mars.
Footnotes
Acknowledgments
The authors would like to thank the National Aeronautics and Space Administration for providing the Zinnia hybrida tissues from the VEG-01C flight experiment for this study. The VEG-01C validation test flight was funded by NASA's Space Biology program. The flight experiment would not have been possible without help from the VEG and Veggie teams at KSC and SNC-ORBITEC, the ISS program, the Payload Operations and Integration Center and Marshall Space Flight Center, and the tireless efforts of the astronauts on the ISS. We also wish to thank K. Venkateswaran at the Jet Propulsion Laboratory, Pasadena, California, for graciously providing cultures of the ISS-F3 and ISS-F4 strains of Fusarium oxysporum for these studies; and C.S. Cockell at the University of Edinburgh, Scotland—and a second anonymous reviewer—for their helpful suggestions during the review process.
Author Contributions
A.C.S. envisioned the experiments discussed above, was the lead on preparing the paper for review, and worked to coordinate all aspects of the project with the other coauthors. G.D.M. was the lead scientist on the Veggie VEG-01C flight experiment and assisted in the writing and editing of the manuscript. B.S.A. and N.S.D. assisted in the genetic sequencing of Fusarium strains, worked on preparing sections of the paper, and assisted in the overall editing of the manuscript. M.E.H., J.T.R., and C.L.K. all assisted in the lab work for the KSC MicroLab and assisted with the editing of all sections in the manuscript. J.T.R. also assisted in the acquisition and organization of the ISS photographs used to characterize the dynamics of the disease complex. And finally, T.M.S. managed the flight coordination with the astronauts and KSC team organization during the VEG-01C mission.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Data Management Plan
Original and high-resolution images used throughout this paper will be posted in the University of Florida Institutional Repository (UFIR) at the website
All strains of Fusarium spp. reported here have been archived at the Northern Regional Research Laboratory (NRRL) Culture Collection, United States Department of Agriculture, 1815 N. University Street, Peoria, IL 61604, USA. The NRRL accession numbers for the following strains are given in parentheses: ASF-151 (66981), ASF-154 (66982), ASF-159 (66983), ASF-162 (66984), ASF-164 (66985), ASF-167 (66986), and ASF-171 (66987).
Supplementary Material
Supplementary Figure S1
Supplementary Figure S2
Supplementary Table S1
Supplementary Table S2
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
