Abstract
Significance:
Mitochondria are eukaryotic organelles with various essential functions. They are both the source and the targets of reactive oxygen species (ROS). Different branches of a mitochondrial quality control system (mQCS), such as ROS balancing, degradation of damaged proteins, or whole mitochondria, can mitigate the adverse effects of ROS stress. However, the capacity of mQCS is limited. Overwhelming this capacity leads to dysfunctions and aging. Strategies to interfere into mitochondria-dependent human aging with the aim to increase the healthy period of life, the health span, rely on the precise knowledge of mitochondrial functions. Experimental models such as Podospora anserina, a filamentous fungus with a clear mitochondrial aging etiology, proved to be instrumental to reach this goal.
Recent Advances:
Investigations of the P. anserina mQCS revealed that it is constituted by a complex network of different branches. Moreover, mitochondrial architecture and lipid homeostasis emerged to affect aging.
Critical Issues:
The regulation of the mQCS is only incompletely understood. Details about the involved signaling molecules and interacting pathways remain to be elucidated. Moreover, most of the currently generated experimental data were generated in well-controlled experiments that do not reflect the constantly changing natural life conditions and bear the danger to miss relevant aspects leading to incorrect conclusions.
Future Directions:
In P. anserina, the precise impact of redox signaling as well as of molecular damaging for aging remains to be defined. Moreover, natural fluctuation of environmental conditions needs to be considered to generate a realistic picture of aging mechanisms as they developed during evolution.
Introduction
Mitochondria are semiautonomous eukaryotic organelles: their biogenesis and function are controlled by both the genetic information in the nucleus and in mitochondria. The vast majority of the more than 1000 proteins required for a functional mitochondrion are encoded by the nuclear deoxyribonucleic acid (nDNA), synthesized in the cytoplasm and delivered to different mitochondrial locations. Only a small number of essential proteins, 13 in mammals, and a set of ribosomal ribonucleic acid (rRNA)'s and transfer ribonucleic acid (tRNA)'s are encoded by the mitochondrial DNA (mtDNA).
The central soluble space of the organelle, the mitochondrial matrix, is surrounded by a phospholipid bilayer, the inner mitochondrial membrane (IMM), which is typically invaginated and forms the so-called cristae reaching out into the matrix space. In this way, the surface of the IMM is largely increased and provides sufficient space for the incorporation of proteins essential for the architecture, biogenesis, and function of the organelle. The IMM can be divided into two regions: the cristal membrane (CM) separating the intracristal space (ICS) from the matrix and the inner boundary membrane (IBM), which runs in parallel with the outer mitochondrial membrane (OMM). This phospholipid bilayer separates the organelle from the cytoplasm. The space between the IBM and OMM represents the intermembrane space (IMS). The IMS and the ICS are connected by small slits, the cristae junctions (CJ).
Both the IMM and the OMM control the exchange of molecules into and out of the organelle. In contrast to the IMM, which is an impermeable barrier for charged molecules, the OMM contains voltage-dependent anion channels that allow the transfer of such molecules across this membrane.
Mitochondria are not synthesized de novo but are derived from existing organelles. “Growth” and propagation of mitochondria, processes that are essential for the development of organisms, proceed via the incorporation of components into existing mitochondria. Depending on physiologic conditions, mitochondria can fuse or divide by fission. In this way, different morphotypes are formed such as small spherical units, filaments, and even networks of connected filaments. The underlying dynamic processes are controlled by nuclear genes (Shaw and Nunnari, 2002; Westermann, 2010; Westermann, 2008).
Mitochondria fulfill a number of essential functions, including iron/sulfur cluster formation, phospholipid synthesis, the control of programmed cell death (PCD), the generation of reactive oxygen species (ROS), and the formation of the cellular energy unit adenosine triphosphate (ATP). The latter function is why mitochondria are generally called the “energy plants” of the cell. While a smaller fraction of mitochondria-generated ATP results from substrate phosphorylation utilizing enzymes of the Krebs cycle, most of the cellular ATP is generated by oxidative phosphorylation (OXPHOS) at the IMM. The latter depends on electron transfer and the transport of protons across the IMM by which an electromotive force is generated that drives the phosphorylation of adenosine diphosphate (ADP) to ATP. The whole process utilizes large protein complexes, the respiratory chain (RC), and the F1Fo-ATP-synthase complex, which are embedded in the membrane.
Apart from the standard RC utilizing protein complexes I–IV, in many organisms such as filamentous fungi, plants, or protozoa, alternative routes are used (Hamann and Osiewacz, 2018). This flexibility allows to deal with impairments of individual components of the RC or with nutritional stress situations. For instance, impairments of complex IV lead to the induction of an alternative oxidase (AOX) acting as the terminal RC oxidase (Borghouts et al., 2001; Dufour et al., 2000; Gredilla et al., 2006). Mutants respiring via such alternative routes provided valuable insight into the aging process of Podospora anserina.
In this review, I summarize work, integrating recent data, and draw an updated picture that demonstrates a key role of mitochondria in aging and life span control of P. anserina. I emphasize different aspects of mitochondrial biology that have a strong impact on aging. These studies identified effects of molecular and nutritional interventions on organismic aging with strong relevance for the development of strategies aimed to increase the health span of biological systems. For more details of earlier work and on the biology of P. anserina, the reader is referred to a number of published review articles (Esser and Tudzynski, 1980; Osiewacz, 2002; Osiewacz, 1997; Osiewacz and Scheckhuber, 2006; Silar et al., 2001).
For a detailed discussion of different aspects of mitochondrial biology and physiology, the reader is referred to a number of excellent reviews (Anand et al., 2021; Brand, 2016; Cobine et al., 2021; Colina-Tenorio et al., 2020; Jezek, 2021; Jezek et al., 2023; Rampelt et al., 2017). For recent comprehensive reviews on the role of mitochondria in mammalian aging, see Batalha et al. (2022), Harrington et al. (2023), and Kauppila et al. (2017).
Age-Related mtDNA Reorganization in P. anserina
P. anserina is a filamentous fungus with a limited life span for which a paramount role of mitochondria was demonstrated soon after the development of the first methods of molecular biology. The life cycle of P. anserina starts with a single ascospore, the product of sexual reproduction (Fig. 1). This single cell germinates and forms a multicellular, macroscopically visible vegetation body (thallus), a mycelium, which consists of branched, filamentous cells (hyphae). These cells grow with a constant linear growth rate at their tips until they reach a strain-specific “senescence point.” At this point, the growth rate slows down, the pigmentation of the hyphae increases, and finally, the thallus dies at the periphery (Rizet, 1953). At the microscopic level, branching of senescent hyphae is abnormal, and finally, the hyphal tips swell and burst (Delay, 1963; Esser and Tudzynski, 1980).
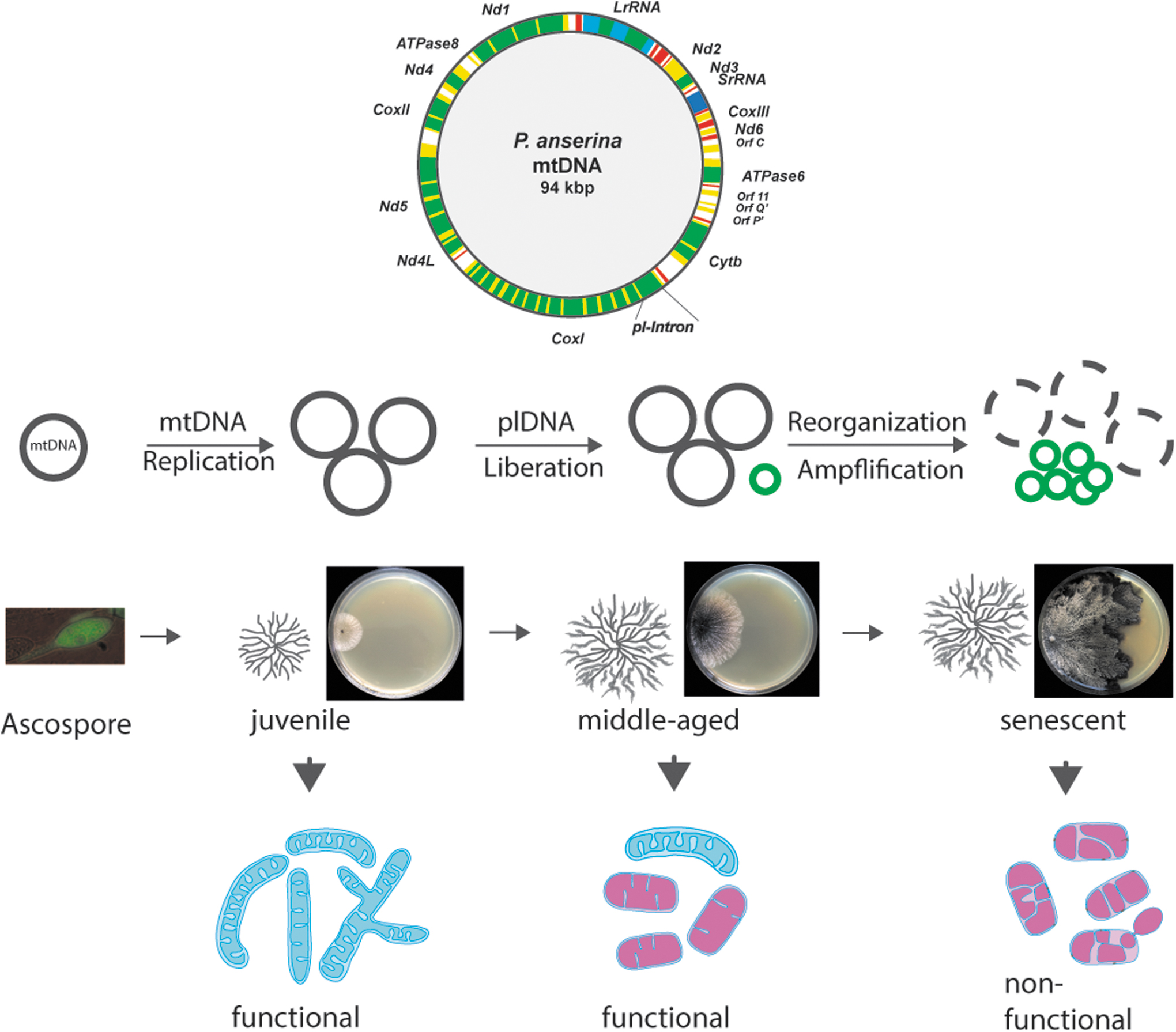
The time until this stage is reached takes a few weeks and depends on the strain isolated from different geographical regions. For instance, the life span of the wild-type “s,” that is commonly used in the laboratory, is about 25 days on a standard complete growth medium when cultivated at 27°C. On other media, and at other temperatures, the life span differs. Many specific mutants were generated from this wild-type strain that display altered life spans. These mutants provide substantial insights into the genetic mechanisms of aging (Table 1).
Molecular Pathways Defined by Longevity Mutants Involved in the Control of Life Span and Aging of Podospora anserina
Genes and encoded products: PaAif2: autophagy inducing factor 2, PaAmid1: AIF homologous mitochondrion inducer of death 1, PaAmid2: AIF homologous mitochondrion inducer of death 2; PaAtg1: autophagy gene 1, PaAtg23: autophagy gene 24; PaAtpe, PaAtpg: ATP-synthase subunit E, G; PaCox1: cytochrome c oxidase subunit I, PaCox17: cytochrome c oxidase copper chaperone COX17, PaClpP: CLP protease, PaCrd1: Cardiolipin synthase, PaCypD: cyclophilin D, PaDnm1: Dynamin1, PaIap: i-AAA protease, PaLon: LON protease, PaMth1: S-adenosylmethyl-dependent methyltransferase, PaSod2: superoxide dismutase 2, PaSod3: superoxide dismutase 3, PaMic 10, PaMic26, PaMic19, PaMic60: subunits 10, 26, 19, 60 of the “mitochondrial contact site and cristae organization system,” PaMca1: metacaspase1, PaMca2: metacaspase2.
Δ: strains in which the indicated gene is deleted by gene replacement.
Mean life span compared with that of the P. anserina wild type from which the mutant strains are derived.
Molecular pathway affected by the genetic intervention.
ex1/ex2, two spontaneous mutants in which large parts of PaCox1 are deleted; grisea, gene coding for the copper-regulated transcription factor GRISEA; n.a., not analyzed; OEx, strains that overexpress the indicated gene.
ADCD, autophagy-dependent cell death; AIF, apoptosis inducing factor; ATP, adenosine triphosphate; COX, cytochrome c oxidase; H2O2, hydrogen peroxide; RC, respiratory chain; ROS, reactive oxygen species.
Early formal genetic experiments suggested genetic traits both inside the nucleus and in the cytoplasm (extranuclear) to be involved in aging and life span control (Marcou, 1961; Tudzynski and Esser, 1979). An extrachromosomal element was subsequently identified to accumulate in mitochondria of senescent mycelia (Fig. 1). This element consists of a series of autonomous covalently closed circular DNAs and thus resembles the structure of bacterial plasmids. It was termed plasmid-like DNA (plDNA) (Stahl et al., 1978) or α-sen DNA (Cummings et al., 1979). In juvenile cultures, plDNA is an integral part of the mtDNA, which is a circular molecule of 94 kbp and thus is considerably larger than the human mtDNA with a size of ∼16 kbp. This size difference is not due to a significant difference in the coding potential.
Both the mtDNA of P. anserina and of humans encode a set of mitochondrial tRNAs, 2 rRNAs, and 13 proteins of the RC (Cummings et al., 1990). However, in P. anserina, several of these genes contain a number of introns that are completely missing in the mtDNA of humans. One of these introns, the pl-intron, is the first intron of the gene coding for subunit one of the cytochrome c oxidase I (COXI), which gives rise to the formation of plDNA (Osiewacz and Esser, 1984). It acts as a mutator element which reintegrates into the standard mtDNA leading to the generation of large-sequence duplications (Sellem et al., 1993). Homologous recombination between these sequences leads to large reorganizations of the mtDNA and the deletion of sequences coding for essential components of the mitochondrial gene expression system (tRNAs, rRNAs) and of RC proteins (Kück et al., 1985).
Consequently, after passing the “senescence point,” neither the damaged mtDNA-encoded proteins of the RC can be replaced by newly synthesized new ones nor can functional mitochondria increase in number and size at the growing periphery of a mycelium. Different longevity mutants, in which the mtDNA is stabilized by different means, emphasize the key impact of mtDNA instability for P. anserina aging (Scheck-huber and Osiewacz, 2008). One example is a mutant that contains a linear plasmid as an autonomous element and a copy of it integrated in the standard mtDNA. In this strain, the amplification of plDNA and the reorganization of mtDNA are delayed and the life span is increased (Hermanns and Osiewacz, 1996; Hermanns et al., 1994; Osiewacz et al., 1989).
Other examples are the ex and mex mutants in which parts of the mtDNA are deleted (Belcour and Vierny-Jamet, 1986; Schulte et al., 1989; Schulte et al., 1988) (see chapter 2, section 1).
After the identification of mtDNA instabilities in P. anserina as a cause of aging, work with different Neurospora species, closely related filamentous ascomycetes of P. anserina, was reported (Bertrand et al., 1986; Court et al., 1991; Griffiths, 1992; Griffiths and Bertrand, 1984; Osiewacz, 1990). Basically, in these cases, the disruption of the coding sequence of the essential mitochondrial rRNA gene results from the integration of linear mitochondrial plasmids. The defective mtDNA becomes suppressive and accumulates leading to senescence. Subsequently, mitochondrial mutations, gross deletions as well as subtle point mutations, were demonstrated to occur during aging in diverse organisms, including organisms such as Caenorhabditis elegans, Drosophila spec, as well as mice, rats, and also in humans (Batalha et al., 2022; Cortopassi et al., 1992; Kadenbach and Müller-Höcker, 1990; Linnane et al., 1989; Melov et al., 1994; Piko et al., 1988; Vandiver et al., 2023; Wallace, 2005).
Respiration, ROS Generation, and ROS Balancing
The mtDNA encodes a few genes for components of the RC, the F1Fo-ATP-synthase, and of RNAs of the mitochondrial gene expression machinery. In P. anserina, the age-associated mtDNA instability leads to impairments of the energy metabolism and to senescence and death (Fig. 1). Detailed studies of the underlying molecular processes provided evidence for an important role of ROS. For instance, studies of mutants such as ex1 and ex2 revealed that the stabilization of the mtDNA and a reduction in ROS generation at the RC lead to a pronounced effect on life span (Kück et al., 1985; Schulte et al., 1989; Schulte et al., 1988). These and similar mutants appear to be immortal. They contain large deletions of the CoxI gene, including the mobile pl-intron. Consequently, the RC is compromised due to the lack of RC complex IV.
This lack is compensated by the induction of the gene coding for an AOX, which becomes integrated into the IMM and accepts electrons from ubiquinol and transfers them to oxygen (Fig. 2). In this way, the RC is shortened and electrons are not traveling via complex III, a major site of superoxide generation to the matrix and the ICS, and complex IV (Brand, 2016; Brand, 2010; Jezek et al., 2023). As a consequence, less superoxide is generated and, since the two RC complexes pump protons from the matrix space to the IMS, the membrane potential is lowered and less ATP is generated than in strains using the standard cytochrome c-dependent RC. The corresponding mutants grow slower than the wild-type strain and are affected in energy-consuming processes such as sexual reproduction.
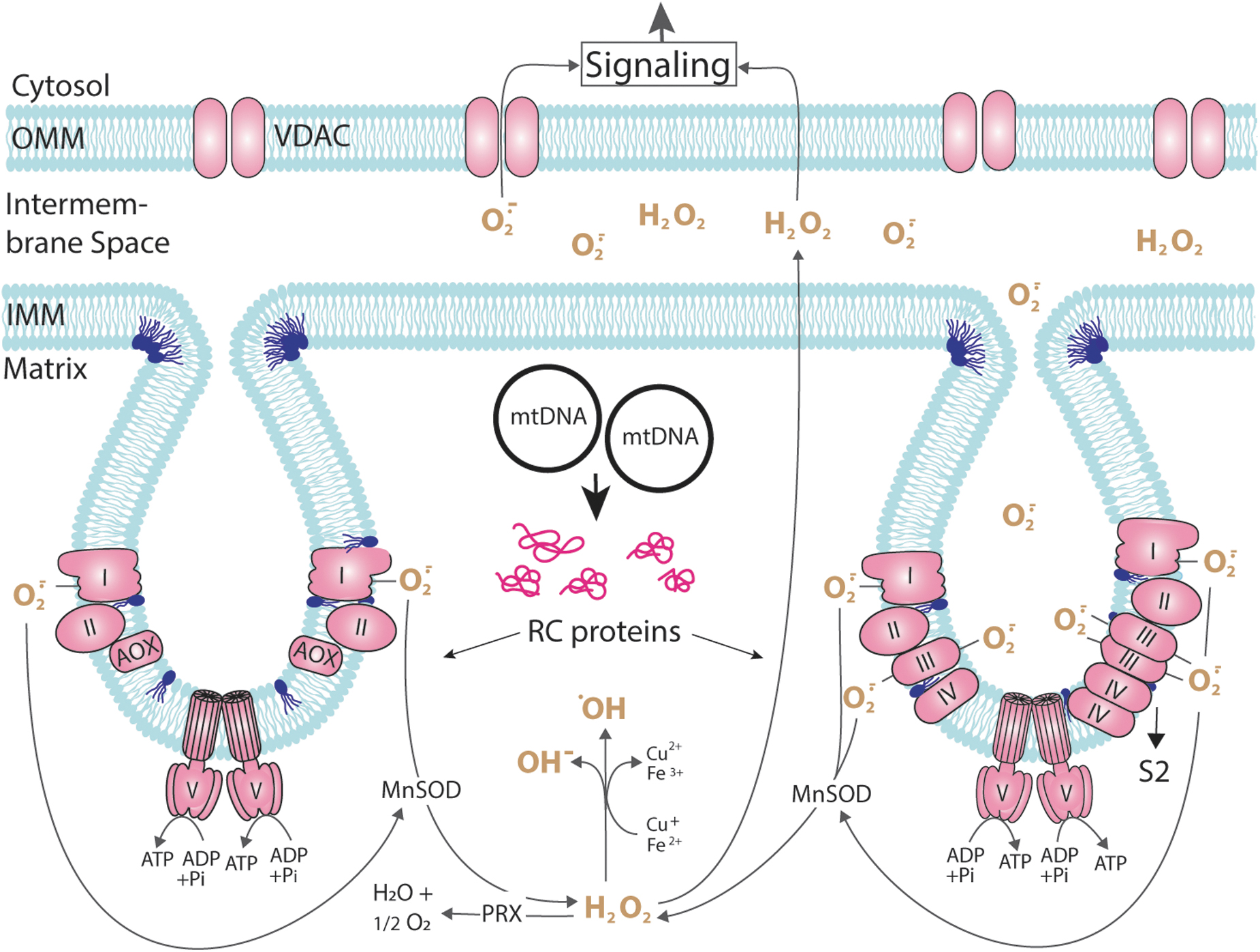
It is important to note that the life span extending effect of PaAox expression appears under well-controlled conditions using endogenous regulators. The experimental, unphysiological overexpression of PaAox under control of a strong promoter affected long-lived respiratory P. anserina mutants for different reasons (e.g., altering ROS levels) and therefore leads to life span reduction (Lorin et al., 2001). Similarly, the unphysiological, heterologous expression of the Aox from the tunicate Ciona intestinalis in a long-lived complex I Drosophila melanogaster mutant was found to decrease the life span possibly due to changes in electron transport in the presence of AOX and altered ROS levels (Scialo et al., 2020; Scialo et al., 2016). By these alterations, the beneficial effect of mild oxidative stress found in the mutants may be lost (for more details see chapter 7, section 9).
Superoxide is a charged molecule and basically does not cross the phospholipid bilayer of membranes. At the RC, it is released into the mitochondrial matrix as well as to the IMS (Fig. 2). Superoxide from the IMS may be released to the cytoplasm via protein pores in the OMM (Han et al., 2003). It also can be converted to hydrogen peroxide, a secondary ROS, which is uncharged and can pass biomembranes. Moreover, hydrogen peroxide can be detoxified by peroxidases or catalase to oxygen and water. On the contrary, in the presence of copper (I) and iron (II), hydrogen peroxide can be oxidized to form the hydroxyl radical, a free radical that is highly reactive for which no detoxification system exists.
Another long-lived mutant respiring via the alternative pathway is mutant grisea (Table 1), in which a nuclear gene coding for the transcription factor GRISEA is mutated. GRISEA senses cellular copper levels and regulates the uptake of copper from the growth medium. The mutation at the first nucleotide of the single intron in the Grisea gene leads the ablation of the GRISEA transcription factor (Osiewacz and Nuber, 1996). In the mutant, cellular copper is depleted because the expression of GRISEA target gene PaCtr3, which codes for a high-affinity copper transporter, is affected. In mitochondria, copper is an essential cofactor of RC complex IV (Borghouts and Osiewacz, 1998; Borghouts et al., 2002; Borghouts et al., 1997). Therefore, also in the grisea mutant, respiration utilizes the alternative respiration pathway with the AOX as terminal oxidase, which uses iron instead of copper as a cofactor.
In the grisea mutant, the generation of superoxide is strongly reduced (Gredilla et al., 2006). Moreover, the amplification of plDNA and the reorganization of the mtDNA are delayed (Borghouts et al., 2000; Borghouts et al., 1997).
The relevance of copper and respiration for the observed life span extension is further supported by a mutant, ΔPaCox17, in which copper is specifically excluded from complex IV of the RC. In this case, while cellular copper levels are basically similar to those in the wild type, the delivery of copper to complex IV is specifically inhibited by the ablation of the PaCOX17 copper chaperone (Stumpferl et al., 2004). The mutant displays an increased life span, which is much longer than that of the grisea mutant (mean life span: 375 days vs. 39 days). In contrast to the grisea mutant in which the life span increase results from a general cellular copper depletion, in the PaCox17 deletion mutant, only the activity of COX is impaired. Importantly, superoxide dismutase 1 (SOD1), which also requires copper as a cofactor, is inactive in the grisea mutant, but active in the PaCox17-deletion strain.
SODs are enzymes of the enzymatic ROS scavenging system, which are localized in different cellular locations (Fridovich, 1978). They convert superoxide into molecular oxygen and hydrogen peroxide. In P. anserina, three isoforms have been identified. SOD1 is the isoform that is predominantly localized in the cytoplasm. Similar to in yeast, a small fraction of the enzyme is likely to be active in the mitochondrial IMS (Sturtz et al., 2001; Zintel et al., 2010). The protein uses copper and zinc as cofactors. SOD2 colocalizes with the endoplasmic reticulum and appears to be a secreted manganese SOD isoform (Zintel et al., 2010). A third SOD (PaSOD3) has been identified in isolated mitochondria from P. anserina, the abundance of which declines during aging (Groebe et al., 2007). It is a manganese-dependent enzyme. The analysis of the three SODs supports the role of ROS in aging and development.
Unexpectedly, while the deletion of the gene coding for the secreted PaSOD2 leads to a slight life span extension, the deletion of PaSod1 and PaSod3 does not affect the life span. Moreover, the overexpression of PaSod3 results in a decrease in mean life span by about 20%, linked to differences in the abundance and profile of the mitochondrial proteases PaLON, PaCLPP, the heat shock protein 60 (PaHSP60), and the hydrogen peroxide detoxification peroxiredoxin (PaPRX1) enzyme (Zintel et al., 2010). Mathematical modeling of experimental data revealed a model incorporating superoxide as part of a feedback loop with components of the antioxidant system (PaSOD3, PaPRX1) and of the mitochondrial matrix protease PaCLPP (Kowald et al., 2012).
Yet another component of the system involved in mitochondrial ROS balancing is PaMTH1, an S-adenosylmethyl-dependent methyltransferase (Table 1). This protein accumulates during aging in both the cytosol and the mitochondrial matrix of the P. anserina wild type. It was identified to protect the fungus against the destructive potential of flavonoid polyphenols such as quercetin, which contain vicinal dihydroxyl groups. They are commonly found in herbivorous dung, the natural growth substrate of the fungus (Averbeck et al., 2000; Chatterjee et al., 2015; Kunstmann and Osiewacz, 2009). In the presence of copper (I) or iron (II) ions, they give rise to the formation of the highly toxic hydroxyl radical. PaMTH1 is able to detoxify the polyphenol by methylation of hydroxyl groups (Knab and Osiewacz, 2010; Kunstmann and Osiewacz, 2008).
Consequently, compared with the wild type, protein carbonylation is decreased in PaMth1 overexpressing strains and mean life span is extended by 115%. In contrast, the deletion of PaMth1 leads to a decreased resistance against exogenous oxidative stress and a mean life span increase of 18% (Kunstmann and Osiewacz, 2009; Kunstmann and Osiewacz, 2008).
The cellular ROS load depends on the generation and the detoxification of ROS, processes that are depending on physiological conditions (Brand et al., 2004; Dröse and Brandt, 2008; Jezek, 2021). While low cellular ROS levels are essential for proper development, too high levels lead to molecular damage and degeneration (Fig. 3). The damaging role of mitochondrial ROS is the basis of the “mitochondrial free radical theory of aging,” which states that aging is caused by the damaging effect of mitochondrial ROS, the levels of which accumulate during aging (Harman, 1972). More recently, this causative impact of ROS became critically challenged (Lapointe and Hekimi, 2010). However, it is current consent that, while not the exclusive cause, ROS strongly contribute to life span control and aging. Both damaging and signaling functions of ROS appear to be effective (Barja, 2019; Hekimi et al., 2011; Lapointe and Hekimi, 2010; Scialo et al., 2016). The data from P. anserina fully support this scenario.
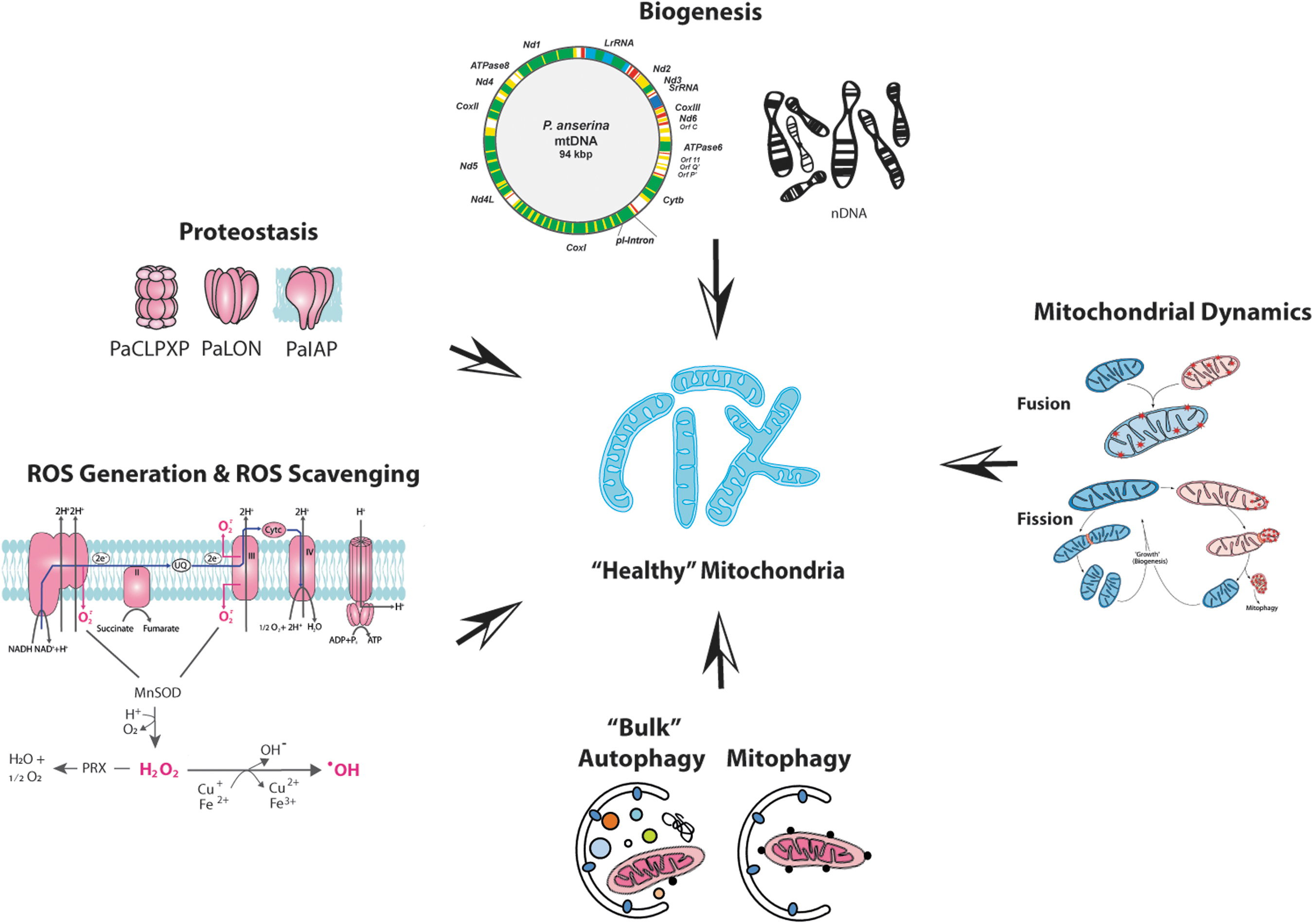
Due to the dual opposing role, balancing of cellular ROS levels is of key importance. Nonenzymatic and, as mentioned above, enzymatic processes are active in the control of proper cellular ROS levels. However, the capacity of ROS-balancing pathways as part of a complex mitochondrial quality control system (mQCS) is limited (Fischer et al., 2012; Tatsuta and Langer, 2008). If the system is overwhelmed, molecular damage becomes manifested. In this case, different higher order branches of the mQCS, including those that are involved in repair, degradation, secretion, and biogenesis of new functional components, are induced and active in keeping a “healthy” population of mitochondria (Fig. 3). The different processes of the mQCS and their regulation are very complex and genetically controlled (Fischer et al., 2012; Hamon et al., 2015; Luce et al., 2010; Tatsuta and Langer, 2008).
The detailed knowledge of the components of the mQCS and their interactions provides a key for interventions to keep mitochondria functional over longer periods of time in the lifetime of organisms.
Mitochondrial Proteostasis
In mitochondria, a number of soluble and membrane-bound proteases are involved in the control of mitochondrial homeostasis (Friguet et al., 2008; Luce et al., 2010; Quiros et al., 2015). They are active in both balancing the correct quantity of proteins, which often are part of multiprotein complexes, and in controlling protein quality under changing physiological situations. These basic functions are of paramount importance for the growth and development of organisms to adjust the mass of functional mitochondria (number and size) to any given situation. For both processes, compounds need to be synthesized and correctly processed. Proteins (i.e., enzymes), which themselves are prone to molecular damage, are key regulators of these processes. They are essential for the degradation of damaged or surplus compounds as well as their biosynthesis and their assembly to functional units.
For mitochondria, the latter requires a delicate system of nuclear/mitochondrial interactions since three cellular compartments—the nucleus, the cytoplasm, and the mitochondrial matrix—are involved in the biogenesis of individual components, which have to be delivered to the site of their action and have to be correctly assembled.
In P. anserina, mitochondrial proteases were shown to strongly impact aging and life span (Fig. 4). Compared with the wild type, mean life span was increased by 73% in a mutant in which the mitochondrial soluble matrix protease PaLON, a homo-oligomeric complex with each subunit containing a chaperone and protease domain, was constitutively overexpressed (Table 1). In this mutant, levels of carbonylated aconitase and carboxymethylated proteins were found to be reduced in mitochondria and hydrogen peroxide secretion, and resistance against exogenous oxidative stress was lowered. The vital functions of the mutants, including their growth rate and fertility, were unaffected. Strikingly, oxygen consumption of isolated mitochondria was increased in 18-day-old strains of the mutant, an age that corresponds to old age in the wild type (Luce and Osiewacz, 2009).
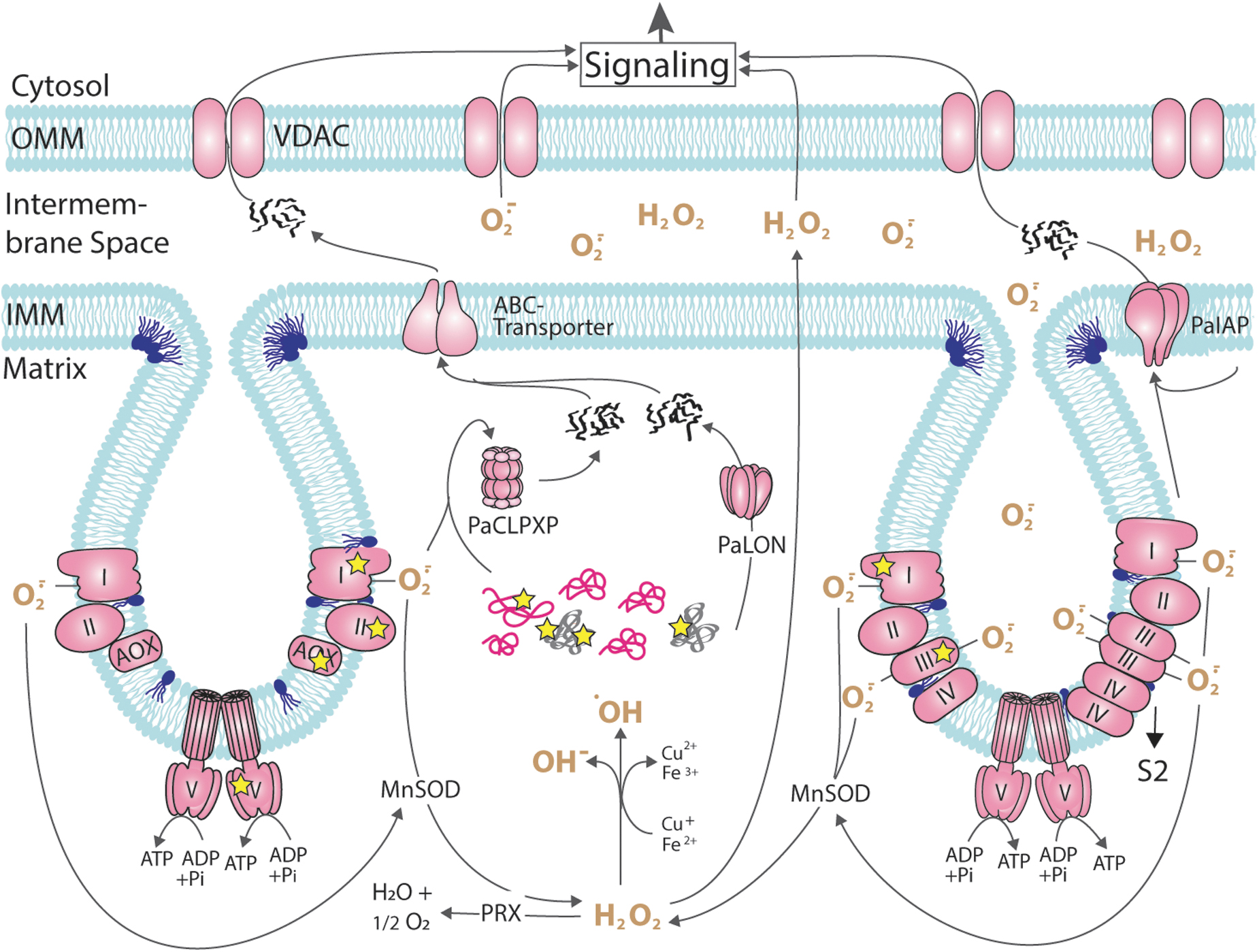
Overall, these data demonstrate that increasing the capacity of the mitochondrial protein QCS has a positive effect on health span. This role was further underlined by studying a PaLon deletion mutant (Table 1), which is viable under standard growth conditions at 27°C incubation temperature (Adam et al., 2012). In contrast, growth at higher (36°C) and lower (18°C, 11°C) incubation temperatures, respectively, is lethal. Significantly, the deletion strain showed an increased sensitivity to hydrogen peroxide in the growth medium. Reintroduction of a wild-type copy of PaLon suppressed this effect supporting the results of the PaLon overexpression study.
LON is considered to be the main protease in the mitochondrial matrix, which is active in the degradation of oxidized proteins. Enzymes of the RC (Hori et al., 2002) or the Krebs cycle (Bota and Davies, 2002; Bulteau et al., 2006; Luce and Osiewacz, 2009) have been identified as substrates of the protease. During aging of mice, a decline of LON transcripts, and of protein levels, was observed (Bota et al., 2002; Lee et al., 1999). In liver samples of senescent rats, decreased LON activity was reported (Bakala et al., 2003). These and other data underline the relevance of the protein in the aging process also in mammals.
While the data of the studies with PaLON were rather as expected, those of the analysis of another mitochondrial serine protease, PaCLPP, were completely unexpected and counterintuitive. This protease is part of a hetero-oligomeric protein complex consisting of individual PaCLPX chaperone and PaCLPP protease subunits (Fig. 4). The identification of potential mitochondrial interaction partners and substrates by stringent tandem affinity purification and differential proteomics identified predominantly components associated with mitochondrial metabolic pathways, including proteins of the pyruvate dehydrogenase complex, the Krebs cycle, as well as subunits of the RC complex I. This interactor/substrate profile strongly suggests a central role of PaCLPP in the control of the energy metabolism, a role that seems to be conserved from bacteria to humans (Fischer et al., 2015; Huang et al., 2020; Petereit et al., 2020; Seiferling et al., 2016; Szczepanowska et al., 2020).
CLPP was found to play a key role in aging, human acute myeloid leukemia, and Perrault syndrome, a human hearing loss disease (Cole et al., 2015; Fischer et al., 2013; Gispert et al., 2013; Seiferling et al., 2016).
Surprisingly, compared with the P. anserina wild type, a PaClpP deletion mutant (Table 1) shows a life span increase by 71%. Life span can be reverted to the life span of the wild type by the heterologous expression of a human ClpP gene copy, indicating a conserved function of the fungal and the human protein (Fischer et al., 2013). This partial conservation is supported by the potential CLPP substrates. Most significantly, homologues of mitochondrial RC complex I proteins were identified as putative substrates in the fungus P. anserina, the plant Arabidopsis thaliana, and in humans (Huang et al., 2020; Petereit et al., 2020; Szczepanowska et al., 2020).
Another mitochondrial protease is PaIAP, which is a protein in the IMM (Fig. 4). Ablation of this protein let to a mean life span increase of 76% at standard growth conditions at 27°C growth temperature. Unexpectedly, at increased temperature (37°C), the life span was decreased by 20% below the life span of the wild type, and spore germination and fruiting body development were affected (Weil et al., 2011). Significantly, PaIAP abundance in the wild type was found to be strongly increased. In the PaIap deletion, strain levels of PaCLPP and of heat shock protein 60 (HSP60) were reduced at 37°C growth temperature (Table 1). In addition, deletion of PaIap was found to affect the mitochondrial RC.
At 27°C, the abundance of complex I and IV monomers and complex V dimers was lower in the mutant than in wild type and the abundance of RC supercomplexes S1 and S2, consisting of one complex I and IV and two complex III units (I1III2IV1), and of one complex I and two complex III and IV (I1III2IV2) subunits, respectively. While the total abundance of the OXPHOS complexes was unchanged in wild type and the mutant, a shift toward the formation of stable RC supercomplexes and of complex V dimers was observed (Weil et al., 2011). Significantly, the observed alterations in the architecture of the membrane-bound RC are linked to a slight overall increase in the abundance of cardiolipin (CL), a cone-shaped phospholipid that is involved in the stabilization of IMM complexes, including the RC complexes and supercomplexes.
Mitochondrial Membrane Lipids
RC complexes and supercomplexes are embedded in the phospholipid bilayer of the IMM. At this membrane, the chemiosmotic gradient generated at the RC drives the synthesis of ATP at the F1Fo-ATP-synthase complex. CL is a signature phospholipid of the IMM that contains four fatty acid moieties, which may differ in individual CL molecules and consequently in the number of carbon atoms and double bonds in the fatty acid residues.
In the P. anserina wild type, an initial lipidome analysis revealed age-associated alterations. In particular, phosphatidylethanolamine (PE) and phosphatidylcholine (PC) levels were found to increase significantly, while CL levels remained constant (Löser et al., 2021). A more detailed analysis identified the IMM protease PaIAP as an adaptive short-term regulator involved in the adjustment of phospholipid biosynthesis in response to energetic demands and natural temperature fluctuations. Deletion of PaIap leads to a shift in the fraction of CLs toward CL72:8, a tetra-linoleoyl containing CL species with four C18 acyl residues and eight double bonds, and leads to a strong increase of unsaturation in the fraction of IMM phospholipids (Löser et al., 2021), which is known to result in increased membrane fluidity (Ernst et al., 2016).
Higher amounts of CL72:8 appear to be positive for mitochondrial functions. For instance, increased levels of C72:8 in cardiac mitochondria of mammals were previously linked to a higher respiratory activity (Stefanyk et al., 2010). In rats with severe heart problems, elevated CL72:8 amounts improved mitochondrial functions (Chicco et al., 2008; Mulligan et al., 2012). Since it is known that CL, and probably in particular CL72:8, is effective in the stabilization of RC complexes and supercomplexes, the increase in active CL appears to be a key for increasing the functionality of mitochondria in the PaIap1 deletion mutant at 27°C growth temperature. At higher temperature, the alteration of the mitochondrial membrane may be negatively affected leading to the observed adverse effect on life span.
The reason for the observed alteration in CL and specifically CL72:8 is certainly due to an alteration in CL biosynthesis and remodeling, which occurs in the IMM. In yeast, YME1, the homologue of PaIAP, is known to be involved in the turnover of different components of the mitochondrial lipid metabolism, including cardiolipin synthase (CRD1) and the CL remodeling acyl transferase TAZ1 (Stefanyk et al., 2010). In P. anserina, a strong increase of PaCRD1 occurs in the PaIap1 deletion mutant although transcript levels of PaCrd1 of the wild type and the mutant essentially do not differ indicating that PaCRD1 levels are most likely accumulating due to the lack of degradation of the protein by its protease (Löser et al., 2021).
Further evidence for the impact of CL on P. anserina aging comes from a PaCrd1 deletion mutant. This mutant lacks CL, while the abundance of phosphatidylglycerol (PG) and PE is increased. The latter is another cone-shaped phospholipid and involved in the induction of negative membrane curvature and leads to membrane stiffening (Cullis and de Kruijff, 1979; Escriba et al., 1997). Its increase leads to a change in the ratio of PE and PC from 1.0 to 1.7 and may, such as in mammalian cells (Tasseva et al., 2013), cause the observed changes in mitochondrial characteristics and functions. Compared with the P. anserina wild type, the life span of the mutant is decreased by 18% (Löser et al., 2021).
The unexpected life span extension of the PaIap deletion mutant raised the question about the evolutionary benefit of this protease. The life span extension was observed at a constant temperature of 27°C at which P. anserina is usually cultivated in the laboratory. At increased temperatures (37°C), the life span is shorter than that of the wild type indicating a role of PaIAP1 as an adaptive regulator of temperature fluctuation as it occurs normally at the natural habitat of P. anserina. This effect is achieved by proteolytic adjustments of phospholipid (PL) biosynthesis contributing to adaptations in membrane plasticity and mitochondrial physiology (Löser et al., 2021).
Mitochondrial Morphology
The morphology of mitochondria is dynamic depending on physiological conditions and linked to genetically controlled fission and fusion events, the proportion of which leads to the characteristic morphotypes (ellipsoid, filamentous, networks) at a given time point.
Mitochondria dynamics impacts mitochondrial function and as such aging (Fig. 5). In P. anserina it was early found that a gene coding for the mitochondrial dynamin-like protein PaDNM1 (in mammals DRP1) is differentially expressed. Compared with juvenile cultures, transcripts of this gene were increased in abundance in senescent cultures (Scheck-huber et al., 2007). This suggested an increase of fission activity to occur during aging. Also, indeed, this conclusion was supported by the demonstration of punctate mitochondria in senescent cultures, whereas mitochondria in juvenile strains were of filamentous or of network structure. Deletion of the PaDnm1 gene resulted in a pronounced increase in life span from 22 days of mean life span of the wild type to 244 days in the mutant, in which fragmented mitochondria occur just before death (Table 1). Fitness of the mutant and the wild type (i.e., growth rate, fertility) was identical on standard growth medium.
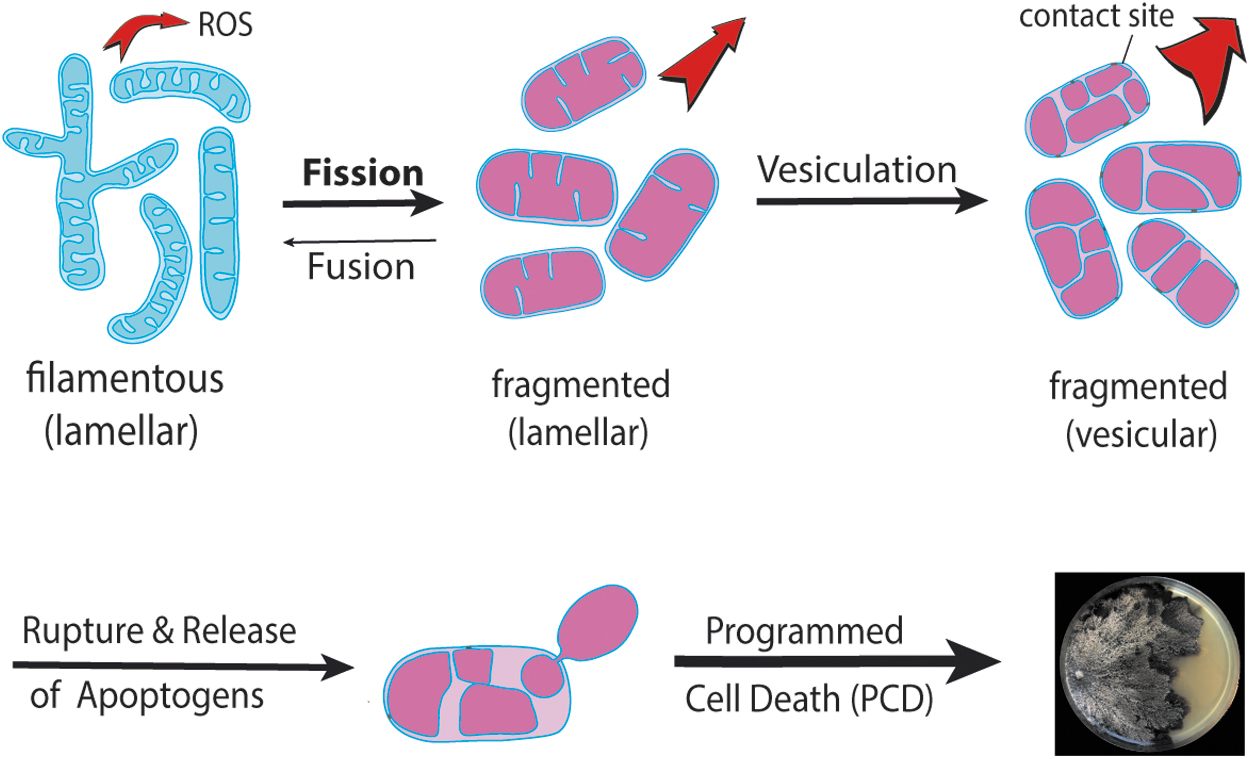
At the molecular level, the generation of cellular ROS and the release of hydrogen peroxide into the growth medium were delayed in the PaDnm1 deletion strain. Further experiments utilizing specific apoptosis mutants and the apoptosis elicitor etoposide revealed experimental evidence for an increased resistance of the mutant to the induction of apoptosis. Significantly, a shift from filamentous to fragmented mitochondria and an increase in cellular hydrogen peroxide were also observed to occur during replicative aging of yeast Dnm1 deletion strains (Scheckhuber et al., 2007). In both, P. anserina and Saccharomyces cerevisiae, deletion of Dnm1 leads to a DNM1-independent fragmentation of mitochondria in late age and the induction of an apoptotic cell death program.
Mitochondrial Ultrastructure
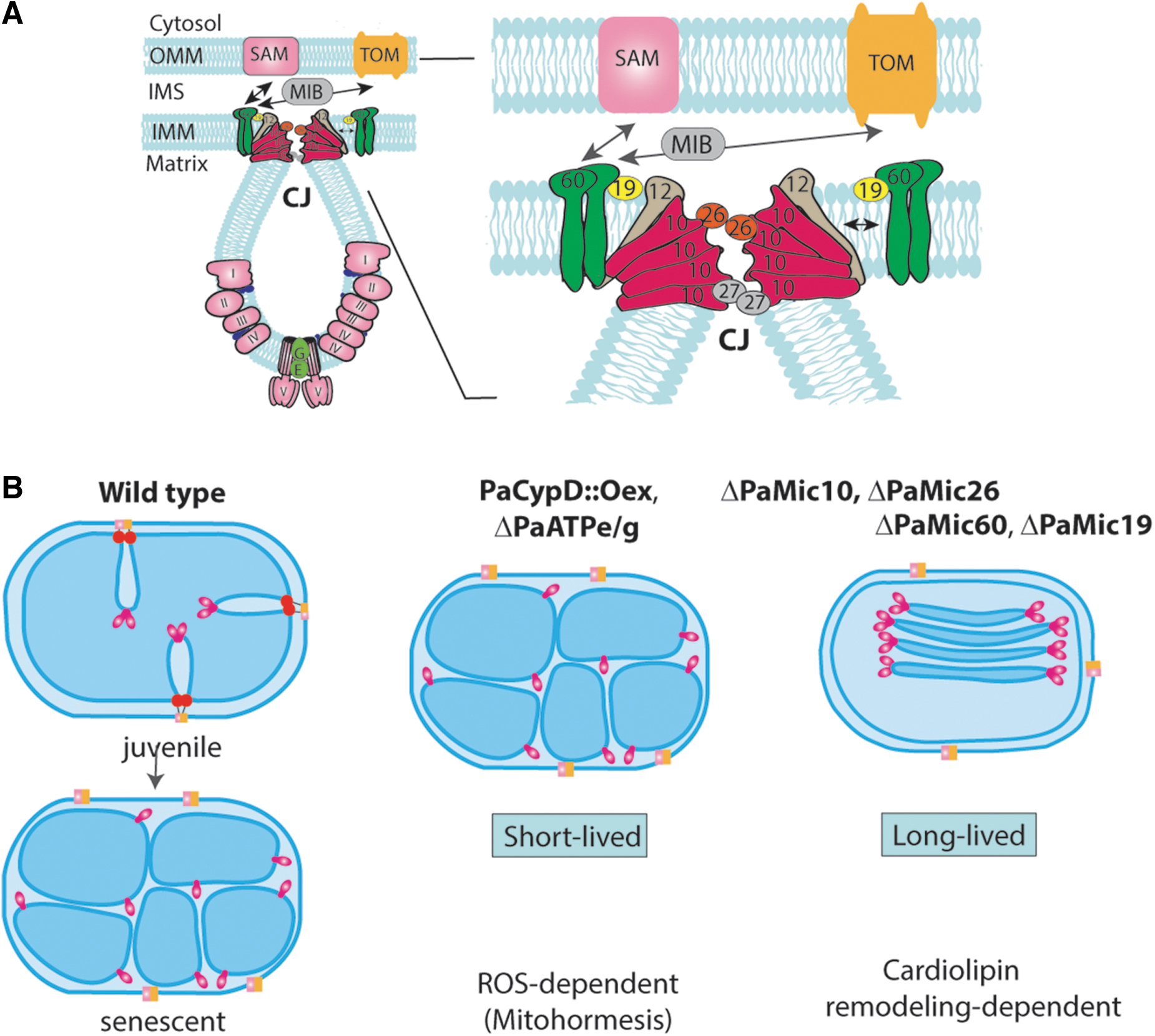
The IMM forms invaginations increasing the surface of this membrane, which is the main site of RC complexes and supercomplexes. Cristae shape varies from organism to organism and is dynamic depending on metabolic conditions (Colina-Tenorio et al., 2020). It is constituted by protein complexes and their interactions with the nonbilayer phospholipids, CL and PE. Both contribute to the characteristic architecture of the organelle and the dynamic alterations. For instance, conical PLs such as CL and PE are involved in negative curvature formation of the IMM. Rows of F1Fo-ATP-synthase dimers are leading to the positive curvature at the cristae tips (Davies et al., 2012; Paumard et al., 2002; Strauss et al., 2008). At CJ, in addition to CL, large protein complexes, the “mitochondrial contact site and cristae organization system” (MICOS), generate a negative curvature of the IMM (Fig. 6) (Eydt et al., 2017; Rampelt et al., 2017).
In P. anserina, in the final stage of the life cycle, the induction of PCD leads to death of the peripheral hyphae of the vegetation body. This kind of PCD differs from that in mammals. For instance, in contrast to mammals, P. anserina does not utilize caspases but, similar to other lower organisms, metacaspases (Hamann et al., 2008; Hamann et al., 2007; Madeo et al., 2002; Minina et al., 2020; Strobel and Osiewacz, 2013). Significantly, proteomic work revealed, compared with mitochondrial samples from juvenile cultures, an increase of cyclophilin D (PaCYPD) abundance in mitochondria from senescent strains (Groebe et al., 2007). CYPD is a mitochondrial peptidyl prolyl-cis,trans-isomerase, which is involved in the regulation of the mitochondrial permeability transition pore (mPTP), a large IMM protein complex of yet unsolved biochemical structure (Azzolin et al., 2010; Bernardi et al., 2023; Halestrap, 2005).
The inhibition of PaCYPD by cyclosporin A leads to life span extension, the overexpression of PaCypD (Table 1) to accelerated fragmentation of mitochondria, and a decreased life span in two independent mutants by 61% and 50%, respectively. Remarkedly, strains from mitochondria of 4-day-old cultures (juvenile stage in wild type) of the mutant displayed a vesicular (reticulate) ultrastructure similar to that found in mitochondria of the 20-day-old wild type (Brust et al., 2010a). This kind of age-associated alteration of the mitochondrial ultrastructure was confirmed in a subsequent study in which the transition from the tubular to the vesicular ultrastructure was linked to the dissociation of F1Fo-ATP-synthase dimers, the formation of contact sites of the OMM and IMM, rupture of the OMM, and the release of apoptogens (Daum et al., 2013; Davies et al., 2011).
This role of F1Fo-ATP-synthase dimers in the formation of typical cristae was finally demonstrated in a study in which either of the two assembly factors (PaATPe, PaATPg) of these dimers was ablated (Fig. 5). The corresponding strains (Table 1) were affected in mitochondrial functions, accumulated senescence markers, including vesicular mitochondrial ultrastructure already in 3-day-old cultures. The mean life span of the corresponding strains was strongly reduced by 33% and 42%, respectively (Rampello et al., 2018).
A molecular model explaining ultrastructural changes culminating in the rupture of the OMM and the release of apoptogens during aging postulated specific changes to occur at the IMM at the CJ (Daum et al., 2013) at the MICOS protein complex. In the genome of P. anserina, five homologues of the six MICOS subunits of yeast are encoded (Warnsmann et al., 2022). These are as follows: MIC10, and the regulatory proteins MIC26, MIC27, and MIC12, forms the MIC10-subcomplex. Oligomerization of MIC10 monomers leads to curvature of the IMM at CJ (Fig. 6). P. anserina seems to miss an MIC12 homologue but encodes the core subunit PaMIC60 and its regulatory subunit PaMIC19. In yeast and mammals, it is known that this subcomplex forms contact sites between the IMM and OMM.
In P. anserina, the deletion of genes PaMic60, PaMic19, PaMic10, and PaMic26, encoding the PaMIC60 and PaMIC10 subcomplexes, respectively, resulted in a changed mitochondrial ultrastructure, and affected mitochondrial functions and life span (Warnsmann et al., 2022). In the mutants, the majority of mitochondria contain floating stacks of tubular cristae, which are not connected to the boundary membrane (Table 1). An exception is the PaMic12 deletion strain in which cristae were wild-type-like. The life span of this strain was similar to that of the wild type. In sharp contrast, the mean life span of the PaMic60, PaMic19, PaMic10, and PaMic26 deletion strains was increased by 88%, 100%, 56%, and 108%, respectively (Warnsmann et al., 2022). The individual vesicles in the stacks contain two tips, one on each site. It is yet an open question how these tips are formed.
Although not shown yet, it is tempting to speculate that dimers of FoF1-ATP-synthase are involved in curvature formation of these membranes similar to that was shown at cristae tips in wild-type mitochondria. In the case of the long-lived Mic mutants, one can speculate that this occurs on both the opposite sites of the individual membrane sacks. This may in fact have bioenergetic consequences, which can explain the observed effect on life span. Further analysis certainly will provide experimental evidence toward a better understanding of the newly generated mutants.
Simultaneous deletion of one subunit of the PaMIC60- and PaMIC10-subcomplex led to a synergistically life span increase, suggesting that the two MIC-subcomplexes affect independent molecular pathways (Warnsmann et al., 2022). Challenging the different mutants with the ROS-generator paraquat in the growth medium revealed that, compared with the wild type, the life span of the MIC10-subcomplex mutants PaMic10 and PaMic26 was decreased. In contrast, paraquat did not decrease the life span beyond the life span of the wild type of the MIC60-subcomplex mutants. These data suggested that the life span increasing effect of MIC10-subcomplex mutants may result from a mild increase in ROS stress, which positively effects mitochondrial functions and increases life span. Such adaptive responses by mild mitochondrial stress are well known from different systems and are collectively termed mitochondrial hormesis or mitohormesis (Barcena et al., 2018; Palmeira et al., 2019; Ristow and Zarse, 2010; Yun and Finkel, 2014).
Increasing oxidative stress in PaMic10 deletion mutants by exposure to exogenous paraquat increases cellular oxidative stress beyond beneficial threshold levels, leading to the induction of cell death and a decreased life span. The mitohormetic effect in the PaMic10 and PaMic26 deletion mutants is further supported by the reduction of life span of these mutants by ROS scavenging using ascorbic acid added to the growth medium. This treatment let to a decrease in life span, while life span of the two MIC60 subunit mutants remained increased compared with the life span of the wild type (Warnsmann et al., 2022).
Overall, these data indicate that longevity of the two PaMic10-subcomplex deletion mutants is triggered by ROS-induced mitohormesis, while life span extension of the two PaMic60-subcomplex mutants is due to another mechanism. This mechanism was found to be linked to phospholipid homeostasis (Marschall et al., 2022). In particular, compared with the wild type, the long-lived phenotype of a PaMic60 deletion mutant was linked to alterations in the phospholipid profile. The abundance of PaCRD1 and CL was found to be increased. In addition, the acyl composition of CL, which is under the control of acyl transferases such as tafazzin, is changed. In particular, the proportion of CL72:8 is increased.
A closer look at the synthesis of CL in the IMM identified an important impact of the remodeling of premature CL (pCL) that is generated by CRD1 from PG. Subsequently, pCL is converted to monolysocardiolin by phospholipases, followed by the generation of mature CL (mCL) by acyl transferases. Tafazzin is the major, but likely not the only, enzyme catalyzing this reaction. Deletion of the single gene coding for PaTAZ1 had no effect on life span. However, most strikingly, deletion of PaTaz1 in the PaMic60 deletion strain resulted in a wild-type-like life span indicating that the life span increase of the PaMic60 mutant is PaTAZ1 dependent. Compared with the wild type and the PaMic60 single mutant, the proportion of CL72:8 was largely reduced. Most strikingly, the growth on medium supplemented with 0.8 mM linoleic acid reestablished the long-life phenotype of the ΔPaMic60 single mutant in the ΔPaMic60/ΔPaTaz1 double mutant.
Taken together, the data underline the importance of mitochondrial phospholipid homeostasis and in particular a strong impact of mCL linked to mitochondrial ultrastructure and health span of P. anserina (Marschall et al., 2022).
Vacuolar Degradation of Mitochondria, Hormesis, and “Autophagy-Induced Cell Death”
The capacity of pathways involved in repair/remodeling of functionally impaired mitochondrial components is limited and mitochondria become severely damaged over time. Such mitochondria or surplus mitochondria can be degraded in the cell by autophagy (Klionsky et al., 2016; Yang and Klionsky, 2010; Yin et al., 2016). In fungi, unlike in mammals, where degradation takes place in lysosomes, this process occurs in vacuoles (Knuppertz and Osiewacz, 2016). During nonselective macroautophagy (hereafter termed autophagy), whole mitochondria together with portions of the cytoplasm are enclosed by a newly formed double biomembrane leading to the formation of autophagosomes. In addition, mitochondria can be specifically selected as autophagosomal cargo. This type of autophagy is termed macromitophagy (hereafter mitophagy) (Knuppertz and Osiewacz, 2016; Wang and Klionsky, 2011). After fusion with the vacuole, mitochondria are degraded and the degradation products are made available for further degradation or the synthesis of new macromolecules (e.g., proteins). Alternatively, if autophagy is impaired, damaged mitochondria encapsulated in vesicles can be secreted and removed from the cell (Liang et al., 2023).
Research of the last decade revealed a wide impact of autophagy, which includes but is not limited to mitochondrial function, on aging in P. anserina. First, it was found that the autophagosome number strongly increases during aging. Moreover, in young cultures, which were grown under nitrogen-depleted condition, autophagosomes were found. Under such conditions, nonselective autophagy is induced (Knuppertz et al., 2014). Deletion of genes coding for proteins (e.g., PaAtg1) involved in autophagy revealed that autophagy is a longevity assurance mechanism, because, compared with their wild type, mutants defective in auto-phagy are short-lived. Mitophagy was found to be induced in a mutant in which the gene for the mitochondrial MnSOD3 was deleted. In this mutant, the induction of mitophagy compensates deficiency in mitochondrial superoxide scavenging and explains why this mutant is not affected in life span.
The induction of external oxidative stress by growth of the mutant and the wild type on medium, to which 20 μM of the superoxide generator paraquat was added, had different effects on the two strains (Knuppertz et al., 2017). In the wild type, this mild induction of oxidative stress is beneficial and leads to an extended life span. In contrast, in the mutant, in which, due to the ablation of PaSOD3, the basic oxidative stress level is higher than in the wild type, added paraquat increases oxidative cellular stress beyond beneficial limits and decreases life span.
The study nicely demonstrates the two sides of autophagy as a “double-edged sword.” Low stress resulting from the ablation of PaSOD3 is beneficial (hormetic, “prosurvival”), while excessive stress induced by paraquat has a “prodeath” effect leading to the induction of “type II” PCD also termed “autophagy-dependent cell death” (ADCD) (Hamann et al., 2007; Knuppertz et al., 2017). This type of cell death is also induced in a short-lived strain, which constitutively overexpresses PaCypD, the gene encoding the mPTP regulator CYPD. In this mutant, accelerated aging is not dependent on the P. anserina metacaspases involved in the control of “apoptotic” or “type I” PCD, but linked to an increase in nonselective autophagy and mitophagy and thus ADCD (Kramer et al., 2016).
ADCD appears also to be the cause of life span reduction in a mutant in which the F1Fo-ATP-synthase assembly factor PaATPE is ablated (Rampello et al., 2018). In this mutant with a vesicular cristae ultrastructure, nonselective autophagy is slightly increased while there is a pronounced induction of mitophagy, which depends on the mPTP regulator PaCYPD (Warnsmann et al., 2021).
Also, the induction of accelerated aging by treatment of P. anserina with gossypol, a natural compound from cotton seeds, appears to be due to the induction of CYPD-mediated ADCD. Treatment of P. anserina with gossypol leads to a dose-dependent decrease of the life span, mitochondrial dysfunction, increased hydrogen peroxide abundance, and an induction of nonselective autophagy and mitophagy (Meyer et al., 2018; Warnsmann et al., 2018).
As mentioned above, the life span increasing effect of the deletion of the PaMic10 and PaMic26 genes coding for the PaMIC10-subcomplex was linked to mitohormesis. The deletion of these genes leads to mild oxidative stress and life span increase. Challenging these mutants with an additional 80 μM of paraquat results in a life span reduction. It is significant to note that in these mutants, mitophagy is increased about 1.5-fold compared with the wild type (Warnsmann et al., 2022). Interestingly, although life span and mitophagy of the two PaMIC60-subcomplex mutants ΔPaMic60 and ΔPaMic19 are also increased, treatment with 80 μM paraquat does not reduce life span but increases it. This effect is linked to alterations in phospholipid homeostasis (see chapter 5, section 2).
The counterintuitive life span extension observed in the different deletion strains of the PaCLPXP complex can be explained by a compensatory induction of autophagy. In these mutants, in which components of the mitochondrial protein QCS are ablated, nonselective autophagy is induced (Knuppertz and Osiewacz, 2017).
Yet another mutant is ΔPaAtg24 (Table 1), which is characterized by a reduction of the mean life span by 18%. PaAtg24 codes for a sorting nexin, active in cellular vesicle transport, membrane trafficking, and protein sorting. In the deletion strain, nonselective autophagy and selective autophagy of peroxisomes are almost completely blocked, while mitophagy is still induced during aging of the mutant demonstrating a PaATG24-independent form of mitophagy to occur in P. anserina (Henkel et al., 2020), which, however, is not efficient enough to ensure mitochondrial function at a level that is sufficient for normal survival. Moreover, vacuoles, the organelles in which all kinds of cellular compounds are degraded, are smaller in the PaAtg24 mutant than in the wild type (Henkel et al., 2020).
Vacuolar size can be reverted to wild-type size by growth of the mutant on a medium to which oleic acid is added. This fatty acid, which is exclusively metabolized in peroxisomes, leads to a severe alteration of the metabolism. In mitochondria, complexes I and II of the RC are bypassed and ROS production is decreased. The autophagy defects of the mutant are abrogated and life span is strongly increased (Schürmanns et al., 2022).
Apart from oleic acid, curcumin, a polyphenol from the rhizome of Curcuma longa, has a life span-extending effect, which depends on a functional autophagy machinery. Curcumin treatment increases mitochondrial respiration, SOD1 activity, hydrogen peroxide release from growing cultures, and induces autophagy (Warnsmann and Osiewacz, 2016).
Overall, these data emphasize a central role of autophagy in aging and life span control. Further investigations are likely to emphasize this importance and will unravel the details controlling the regulation of the different forms of autophagy and their interaction with other cellular pathways. At the current stage, ROS-associated signaling pathways appear to play a key role (Knuppertz et al., 2017). However, many details of these pathways and their interactions remain to be elucidated.
Apoptosis
In multicellular organisms such as C. elegans or mammals, apoptosis or “type I” PCD is a process that is essential for development and part of the cellular QCS to remove surplus or severely damaged cells that are at risk to propagate in an uncontrolled manner. In mammals, there are caspase-dependent and caspase-independent pathways active and mitochondria play an important role in the control of apoptosis.
In P. anserina, apoptosis, in addition to ADCD, is not part of the QCS but leads to death of the individual at the end of its life span (Hamann et al., 2008; Osiewacz, 2011). Instead of a set of caspases, there are two metacaspases and caspase-independent factors (Table 1) such as the mitochondrial oxidoreductase PaAIF and PaAMID, which are involved in the control of apoptotic cell death. During aging they are released from mitochondria (Fig. 6). Ablation of the latter two types of factors leads to increased tolerance to ROS and to life span extension (Brust et al., 2010b). The two metacaspases PaMCA1 and PaMCA2 are calcium-dependent enzymes, the activity of which increases during P. anserina aging (Hamann et al., 2008; Minina et al., 2020; Strobel and Osiewacz, 2013).
Both enzymes are involved in degradation of poly (ADP-ribose) polymerase, a well-known substrate of mammalian caspases and an enzyme with a wide impact on aging processes (Bürkle et al., 2005; Grube and Bürkle, 1992; Müller-Ohldach et al., 2011; Strobel and Osiewacz, 2013). PaCYPD, which accumulates during aging and regulates the mPTP, is involved in the regulation of both apototic PCD and ADCD (Kramer et al., 2016; Schneider, 2005).
Conclusions
Wild-type aging of P. anserina is caused by mtDNA instabilities, which lead to mitochondrial impairments and ultimately to cell death. This process is fast occurring within 3–4 weeks in wild-type “s” and linked to the short period of time nutrients are available for the fungus on its natural substrate, herbivorous manure. During its lifetime, P. anserina propagates efficiently generating fruiting bodies with a large number of ascospores as progeny. In the laboratory, under well-defined conditions with no nutrient limitation, the aging process can be easily analyzed. Such investigations allowed to unravel the importance of mitochondria in aging. Processes balancing ROS levels by reducing ROS generation, turning available ROS over into nontoxic compounds, repairing molecular damage of cellular components (e.g., DNA, proteins, lipids), and replacing them by newly synthesized ones, strongly impact life span and health span.
In some cases, results of specific experiments were counterintuitive owing to the fact that the different pathways of the mQCS interact with each other relying on specific signaling pathways. However, the experiments normally do not reflect the natural constantly changing conditions (i.e., temperature, nutrient supply). Adaption to these conditions is a key for survival for any organism in nature. The underlying mechanisms of aging, as they developed during evolution, are thus incompletely understood. In case of redox signaling, it appears that, due to its ability to pass biomembranes and its stability, hydrogen peroxide is the main ROS (Sies, 2017). The targets of ROS signaling need to be precisely identified. Such information will be important to specifically intervene in aging, for instance, by the controlled induction of mitohormesis that increases the health span of organisms.
More recent studies identified dynamic alterations of the mitochondrial ultrastructure during aging of P. anserina. These processes are linked to alterations of specific protein complexes, including dimers of the F1Fo-ATP-synthase at the cristae tips and MICOS at the basis of cristae, as well as of nonbilayer phospholipids. While experimental impairment of F1Fo-ATP-synthase dimerization shows adverse effects on life span linked to a vesicular structure of the IMM, ablation of four of the five identified P. anserina MICOS proteins is beneficial and linked to the formation of stacked tubular and floating vesicles. The mechanistic impact of this new observation on aging is only beginning to be elucidated. While ablation of PaMIC10 and PaMIC26 respectively, leads to induction of ROS-dependent mitohormesis, the ablation of the two PaMIC60 subcomplex proteins is linked to altered phospholipid homeostasis.
It remains to be demonstrated in how far the conclusions about their impact drawn in P. anserina can be translated to higher systems including humans. This translation could generate valuable data for the establishment of scientifically driven strategies (e.g., nutritional regimens) to intervene into age-related degenerative processes. However, experiments with this goal may not be easy, because what takes place in the undifferentiated cells of a fungal vegetation body may only occur in one or a few critical organs or even cell types of the complex human body and, although of great significance, may therefore escape easy detection.
Author's Contribution
H.D.O.: Conceptualization and writing.
Footnotes
Author Disclosure Statement
No competing financial interests exist.
Funding Information
The work of Heinz D. Osiewacz has continuously been supported by the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation)—Project-ID 25913077-SFB1177 and Os75/17-2.
