Abstract
Aims:
The MarR/DUF24-family QsrR and YodB repressors control quinone detoxification pathways in Staphylococcus aureus and Bacillus subtilis. In S. aureus, the QsrR regulon also confers resistance to antimicrobial compounds with quinone-like elements, such as rifampicin, ciprofloxacin, and pyocyanin. Although QsrR was shown to be inhibited by thiol-S-alkylation of its conserved Cys4 residue by 1,4-benzoquinone, YodB senses quinones and diamide by the formation of reversible intermolecular disulfides. In this study, we aimed at further investigating the redox-regulation of QsrR and the role of its Cys4, Cys29, and Cys32 residues under quinone and oxidative stress in S. aureus.
Results:
The QsrR regulon was strongly induced by quinones and oxidants, such as diamide, allicin, hypochlorous acid (HOCl), and AGXX® in S. aureus. Transcriptional induction of catE2 by quinones and oxidants required Cys4 and either Cys29′ or Cys32′ of QsrR for redox sensing in vivo. DNA-binding assays revealed that QsrR is reversibly inactivated by quinones and oxidants, depending on Cys4. Using mass spectrometry, QsrR was shown to sense diamide by an intermolecular thiol-disulfide switch, involving Cys4 and Cys29′ of opposing subunits in vitro. In contrast, allicin caused S-thioallylation of all three Cys residues in QsrR, leading to its dissociation from the operator sequence. Further, the QsrR regulon confers resistance against quinones and oxidants, depending on Cys4 and either Cys29′ or Cys32′.
Conclusion and Innovation:
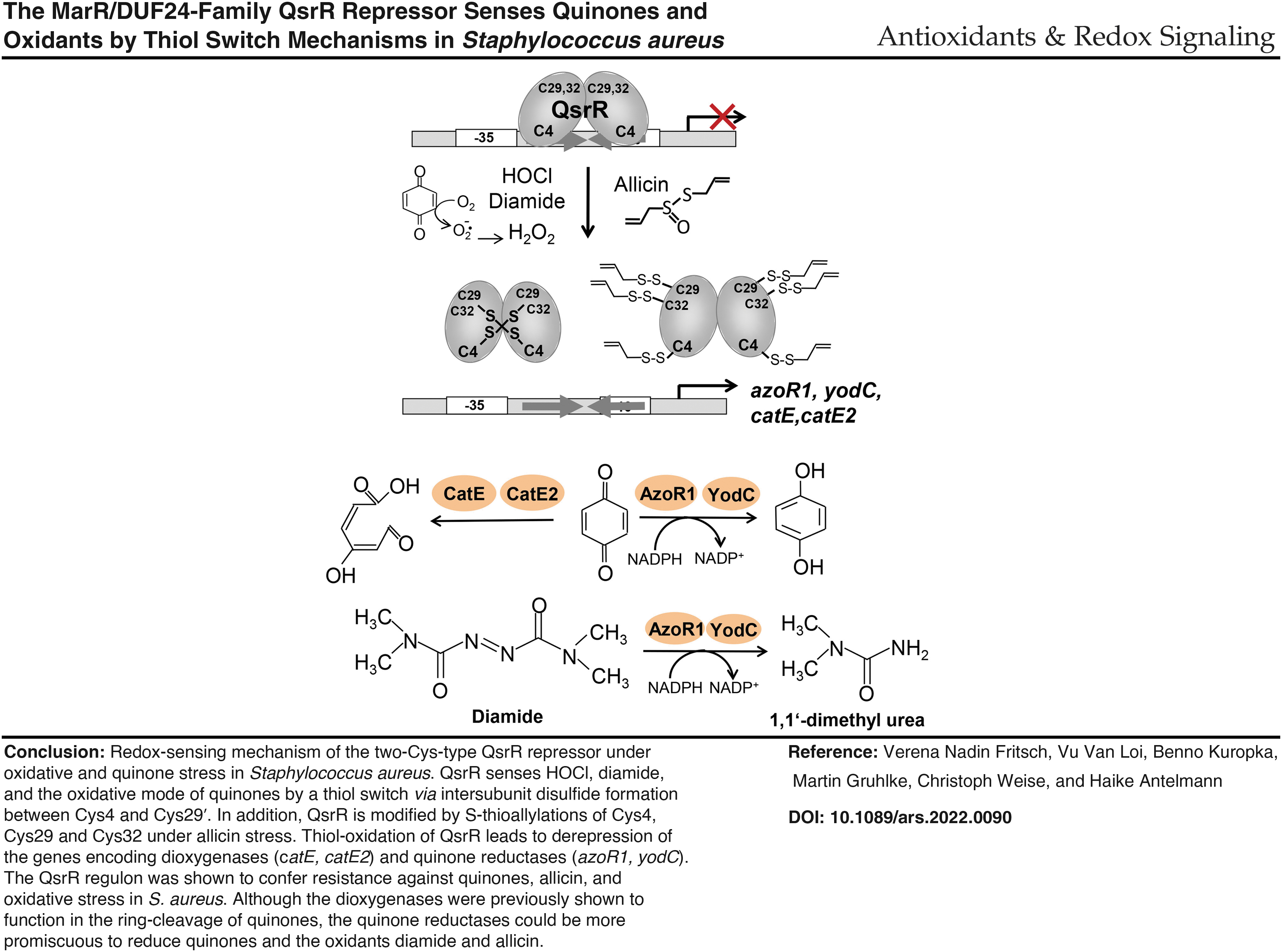
QsrR was characterized as a two-Cys-type redox-sensing regulator, which senses the oxidative mode of quinones and strong oxidants, such as diamide, HOCl, and the antimicrobial compound allicin via different thiol switch mechanisms.
Introduction
S
During infections, cellular metabolism, and antibiotic treatments, S. aureus has to cope with reactive oxygen species (ROS), reactive electrophile species (RES), and reactive chlorine species (Linzner et al, 2021; Loi et al, 2015). S. aureus encounters endogenous ROS, such as superoxide anions, hydrogen peroxide (H2O2), and hydroxyl radicals, which are produced as byproducts in the aerobic respiratory chain (Imlay, 2008). After phagocytosis of S. aureus cells, activated macrophages and neutrophils release the strong oxidant hypochlorous acid (HOCl) as the most potent killing agent against the invading bacteria (Ulfig and Leichert, 2021).
Innovation
Staphylococcus aureus is a major human pathogen, which has to cope with oxidative and electrophile stress as well as redox-active antibiotics during infections. In this work, we have shown that the MarR/DUF24-family regulator QsrR senses and responds not only to quinones and quinone-like antimicrobials, but also to strong oxidants, such as diamide, hypochlorous acid, and allicin stress. Using transcriptional studies, DNA binding assays, and mass spectrometry, QsrR was shown to sense disulfide stress by diamide and allicin via different thiol-switches, which involve Cys4 and Cys29′. Thus, QsrR allows the adaptation of S. aureus toward quinones, antimicrobials, and strong oxidants via reversible thiol-switch mechanisms.
The ROS and HOCl can damage all cellular macromolecules, including proteins, DNA bases, and carbohydrates, leading to bacterial killing by the immune cells. The Cys thiol group is the most susceptible target for oxidation by ROS and HOCl, leading to reversible thiol switches or irreversible thiol-oxidation products (Linzner et al, 2021). The oxidation of cellular metabolites, such as glucose, unsaturated fatty acids, and amino acids by ROS, further generates RES as secondary reactive species (Jacobs and Marnett, 2010; Marnett et al, 2003).
The RES include quinones and aldehydes, which have electron-deficient carbon centers and can react with the nucleophilic Cys thiol group (Linzner et al, 2021). S. aureus not only synthesizes endogenous menaquinone as an electron carrier but has also to cope with external quinone-like antimicrobial compounds, such as pyocyanin, ciprofloxacin, and the naphthoquinone lapachol. Many fully substituted quinones, such as lapachol, act via the oxidative mode to generate semiquinone radicals, resulting in ROS formation and protein thiol-oxidation (Linzner et al, 2020; Monks et al, 1992; O'Brien, 1991).
In addition, unsubstituted quinones, such as 1,4-benzoquinone (BQ), act as oxidants and electrophiles via the thiol-S-alkylation chemistry with protein thiols, resulting in the aggregation and depletion of thiol-containing proteins in the proteome, especially at toxic concentrations (Liebeke et al, 2008; Loi et al, 2015; Monks et al, 1992; O'Brien, 1991). Apart from ROS and RES, S. aureus encounters other redox-active antimicrobials, such as the natural organosulfur compound allicin from garlic, which are used as alternative treatment option to combat methicillin-resistant S. aureus. Allicin has been shown to act as strong antimicrobial against bacteria, fungi, and parasites, including multi-drug-resistant strains.
The killing effect of allicin is mediated by S-thioallylation of low molecular weight (LMW) thiols and protein thiols as revealed in several bacteria (Borlinghaus et al, 2021; Borlinghaus et al, 2014; Loi et al, 2019; Müller et al, 2016).
In Bacillus subtilis and S. aureus, the MarR-type repressors MhqR and YodB (QsrR) sense and respond directly to quinones and regulate quinone detoxification pathways, including quinone reductases (AzoR1, YodC) and ring-cleavage dioxygenases (CatE, CatE2, MhqE), which confer resistance to quinones (Chi et al, 2010a; Fritsch et al, 2019; Ji et al, 2013; Leelakriangsak et al, 2008; Töwe et al, 2007). The quinone reductase (AzoR1) and nitroreductase (YodC) might function as NADPH-dependent flavoenzymes in the reduction of quinones to redox-stable hydroquinones. The thiol-dependent dioxygenases (CatE, CatE2, MhqE) are involved in the ring cleavage of quinones and catechol to the γ−hydroxymuconic semialdehyde as demonstrated for CatE in B. subtilis (Leelakriangsak et al, 2008; Tam le et al, 2006).
In S. aureus, the QsrR and MhqR regulons confer independent resistance to methylhydroquinone (MHQ), BQ, and quinone-like antimicrobials, such as pyocyanin, ciprofloxacin, and rifampicin (Fritsch et al, 2019; Ji et al, 2013; Noto et al, 2017). QsrR also mediates tolerance of S. aureus to photodynamic inactivation by the photosensitizer methylene blue (Snell et al, 2021). Thus, the dioxygenases and quinone reductases might function in the detoxification of phenolic antimicrobials and dyes as mechanisms of antimicrobial resistance.
Recently, the quinone-sensing Rrf2-family SifR repressor was characterized in Streptococcus pneumoniae (Zhang et al, 2022). SifR controls the Fe2+-dependent catechol-2,3-dioxygenase CatE and the NAD(P)H-dependent quinone reductase YwnB, which was capable of reducing the host-derived catecholamines norepinephrine and adrenochrome (Zhang et al, 2022). Thus, SifR is postulated to make iron available from catecholate-Fe3+ complexes, taken up via the PiuBCDA transporter. The DNA-binding activity of SifR was inhibited by S-alkylation of the redox-sensing Cys102 in vitro, whereas the redox-sensing mechanism of SifR remains to be investigated in vivo (Zhang et al, 2022).
Interestingly, the catDE operon of B. subtilis was shown to be controlled by two MarR/DUF24-type repressors (YodB, CatR) and the iron-sensing Fur repressor, providing a link between iron limitation and catechol degradation (Chi et al, 2010b; Pi and Helmann, 2018). CatE was shown to be involved in the degradation of the endogenously produced catecholate siderophore bacillibactin to avoid accumulation of toxic catechol derivatives during iron starvation in B. subtilis (Pi and Helmann, 2018). Thus, the quinone response is also important for removal and reduction of endogenous catecholate siderophores and utilization of host-derived catecholamine hormones in bacteria (Alghofaili et al, 2021; Pi and Helmann, 2018; Zhang et al, 2022).
In S. aureus, MhqR controls the mhqRED operon, encoding the dioxygenase MhqE and phospholipase/carboxylesterase MhqD (Fritsch et al, 2019). MhqR does not sense quinones via thiol-based redox switches or S-alkylation mechanisms. We hypothesize that quinones bind to a specific ligand-binding pocket, which is conserved in MarR-type regulators (Grove, 2017; Perera and Grove, 2010; Wilkinson and Grove, 2006), resulting in conformational changes of the DNA-binding helix-turn-helix (HTH) motifs and the dissociation of MhqR from the promoter DNA (Linzner et al, 2021). The MarR/DUF24-family QsrR repressor of S. aureus resembles a two-Cys-type redox-sensing regulator, which has three Cys residues in positions 4, 29, and 32 (numbering based on the S. aureus COL QsrR sequence).
Of note, the QsrR protein of S. aureus Newman (NWMN_2027) is annotated with an additional N-terminal Met residue, leading to the renumbering of Cys5, Cys30, and Cys33 in the previous study (Ji et al, 2013). For unification, we refer to the S. aureus COL numbering of Cys4, Cys29, and Cys32 regarding QsrR regulation throughout the article. The N-terminal redox-sensing Cys is conserved across the MarR/DUF24-family of regulators, including YodB, CatR, and HypR of B. subtilis and QsrR of S. aureus (Antelmann and Helmann, 2011; Ji et al, 2013; Linzner et al, 2021; Yurimoto et al, 2005).
These MarR/DUF24 proteins harbor the HxlR-type winged HTH domain (IPR002577) of 90–100 aa (
According to our previous RNA-seq analyses, the QsrR regulon was strongly induced under oxidative and electrophile stress, such as MHQ, lapachol, HOCl, allicin, and AGXX® stress in S. aureus (Fritsch et al, 2019; Linzner et al, 2021; Linzner et al, 2020; Loi et al, 2019; Loi et al, 2018a, Loi et al, 2018b). Thus, we hypothesize that QsrR might sense ROS, generated in the oxidative mode of quinones, and disulfide stress by strong oxidants and antimicrobials due to intersubunit disulfide formation between the conserved Cys4 and the Cys29′ or Cys32′ residues of opposing subunits, as revealed for the homologous YodB repressor of B. subtilis (Chi et al, 2010a; Linzner et al, 2021).
In this work, we have investigated the regulatory mechanisms of the QsrR repressor and the role of its three Cys residues for DNA-binding activity and redox sensing under quinone and oxidative stress in S. aureus. Using transcriptional analyses of the QsrR Cys4, Cys29, and Cys32 mutants, we showed that Cys4 and either Cys29 or Cys32 residues are involved in the redox sensing of quinones and oxidants in S. aureus. Mass spectrometry (MS) revealed that QsrR senses diamide stress by intermolecular disulfides between Cys4 and Cys29′, whereas allicin resulted in S-thioallylation of all three Cys residues of QsrR. Mutational phenotype analyses further supported that Cys4 and either Cys29′ or Cys32′ are required for redox regulation to confer resistance toward MHQ, HOCl, and H2O2 stress in S. aureus. Altogether, this work has unraveled the thiol switch mechanisms of QsrR for adaptation toward oxidative and electrophile stress as well as redox-active antimicrobials in S. aureus.
Results
Transcriptional induction of the QsrR regulon by quinone and oxidants requires the redox-sensing Cys4 and either Cys29 or Cys32 in S. aureus
Previous RNAseq analyses showed a strong upregulation of the QsrR regulon by quinones (MHQ, lapachol) and strong oxidants (HOCl, AGXX, and allicin) in the transcriptome of S. aureus (Fritsch et al, 2019; Linzner et al, 2020; Loi et al, 2019; Loi et al, 2018a, Loi et al, 2018b). The QsrR regulon genes azoR1, catE, catE2, yodC, and SACOL0409 were most strongly induced (14- to 278-fold) under allicin, AGXX, and MHQ stress (Supplementary Table S1). In addition, the MhqR and QsrR regulons were previously shown to confer independent resistance to antimicrobials with quinone-like elements, such as ciprofloxacin, rifampicin, and pyocyanin (Fritsch et al, 2019; Noto et al, 2017).
Thus, we analyzed transcription of the catE2 and qsrR genes upon exposure to different thiol-reactive compounds and antibiotics. Northern blot analyses verified the strong transcriptional induction of catE2 (25- to 48-fold) and qsrR (3- to 10-fold) under MHQ, diamide, AGXX, and allicin stress in S. aureus (Fig. 1A, B). The catE2 gene was only weakly ∼2-fold upregulated by different antibiotics, formaldehyde and methylglyoxal stress in S. aureus, whereas erythromycin and rifampicin caused a 10.7- and 4.1-fold transcriptional induction of the qsrR gene (Fig. 1A, B). These data suggest that QsrR might potentially sense and respond to quinones and strong oxidants, which cause disulfide stress in S. aureus.
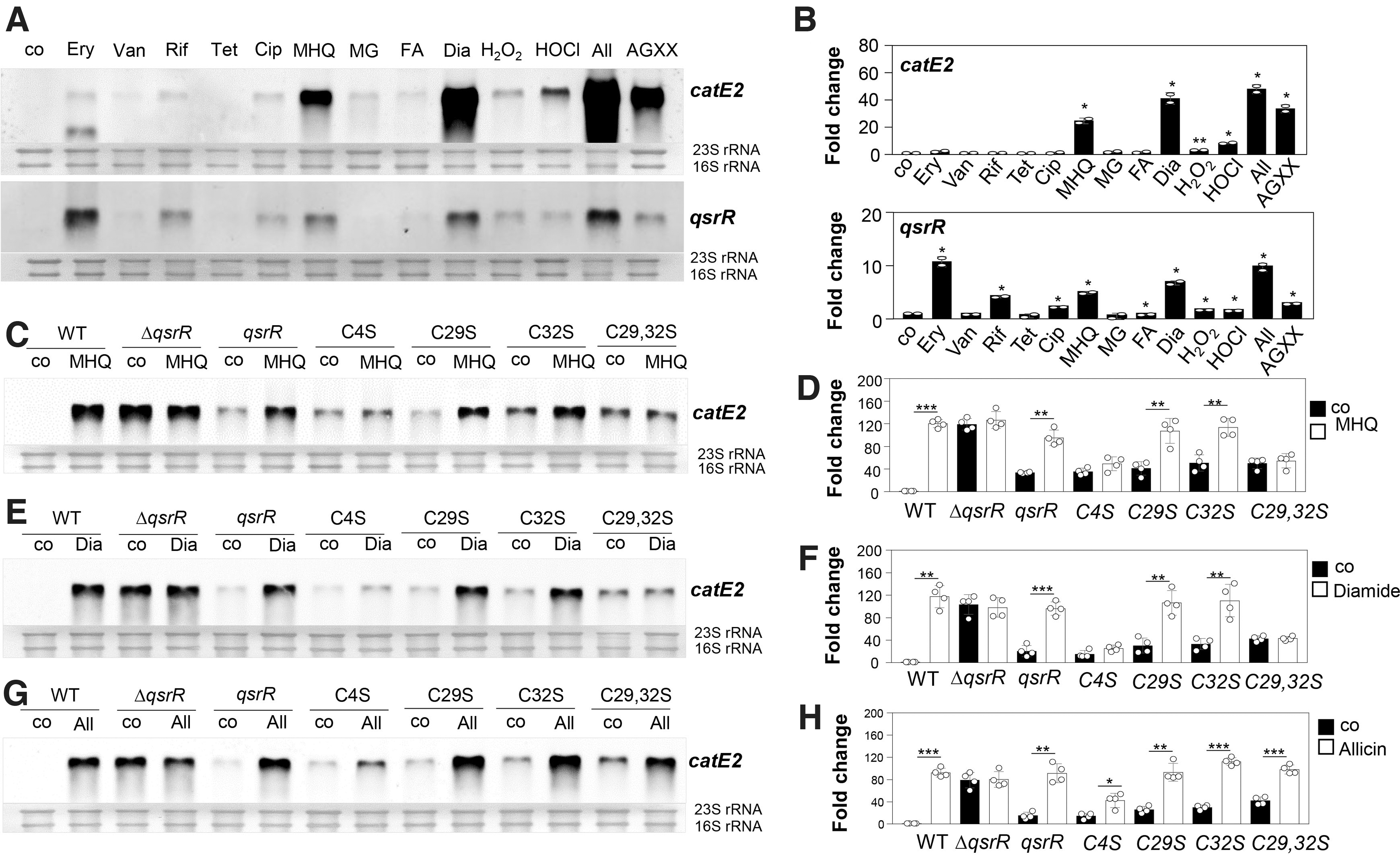
Next, we used quantitative Northern blot analyses to investigate the roles of the three Cys residues (Cys4, Cys29, and Cys32) of QsrR for redox sensing and DNA-binding activity under quinone and disulfide stress in vivo (Fig. 1C–H). Transcription of catE2 was analyzed in the qsrR mutant complemented with qsrR and the qsrR C4S, C29S, and C32S mutant alleles. We further confirmed that QsrR and the QsrR Cys mutant proteins are expressed in S. aureus (Supplementary Fig. S1).
The Northern blot analysis revealed that catE2 is fully >80-fold derepressed in the qsrR mutant, and repression could be restored in the qsrR complemented strain (Fig. 1C–H). Due to the strong autoregulation of qsrR expression in the WT after thiol stress, the basal level of catE2 transcription was higher in the qsrR complemented strain (Fig. 1C–H). Although catE2 transcription was fully inducible in the qsrR complemented strain after MHQ, diamide, and allicin stress, the C4S mutant failed to respond significantly to MHQ and diamide stress (Fig. 1C–F). There was only a weak threefold induction of catE2 transcription in the C4S mutant under allicin stress (Fig. 1G, H).
Thus, Cys4 was clearly identified as redox-sensing Cys, which is required for the responsiveness of QsrR to quinones and disulfide stress in S. aureus in vivo (Fig. 1C–F). However, both C29S and C32S mutants showed a similar strong response to MHQ and diamide as the WT protein, whereas the C29,32S double mutant was unresponsive and did not show increased catE2 transcription under thiol stress (Fig. 1C–F). This indicates that Cys4 and either Cys29 or Cys32 are required for redox sensing of QsrR and transcriptional induction of the QsrR regulon under MHQ and diamide stress in vivo.
In contrast, a significant induction of catE2 transcription was measured in the C29S, C32S, and C29,32S mutants after allicin stress, supporting that the allicin response requires only Cys4 (Fig. 1G, H). Thus, the redox-sensing mechanisms of QsrR differ between MHQ, diamide, and allicin stress. In addition, the C29S, Cys32S, and C29,32S mutants were slightly impaired in DNA binding as revealed by the higher basal level of catE2 transcription (Fig. 1C–H).
The Cys residues do not affect the DNA-binding activity of QsrR to the specific QsrR operator sequence in vitro
Gel shift assays were used to investigate the DNA-binding activity of QsrR and the single C4S, C29S, and C32S mutant proteins to the palindromic QsrR operator in the catE2 and qsrR promoter regions as previously defined with the consensus sequence GTATA-N5-TATAC (Ji et al, 2013). QsrR was shown to bind with high affinity to the catE2 and qsrR upstream promoter regions (Fig. 2A, B). The dissociation constants (KD) of QsrR were calculated as 68.3 nM for the catE2 promoter and as 112.4 nM for the qsrR promoter (Fig. 2C, D). The C4S, C29S, and C32S mutant proteins showed similar DNA-binding affinities and their KD values for the catE2 promoter were determined as 61.5, 66.5, and 61.1 nM and for the qsrR promoter as 103.7, 105.4, and 105.5 nM, respectively.
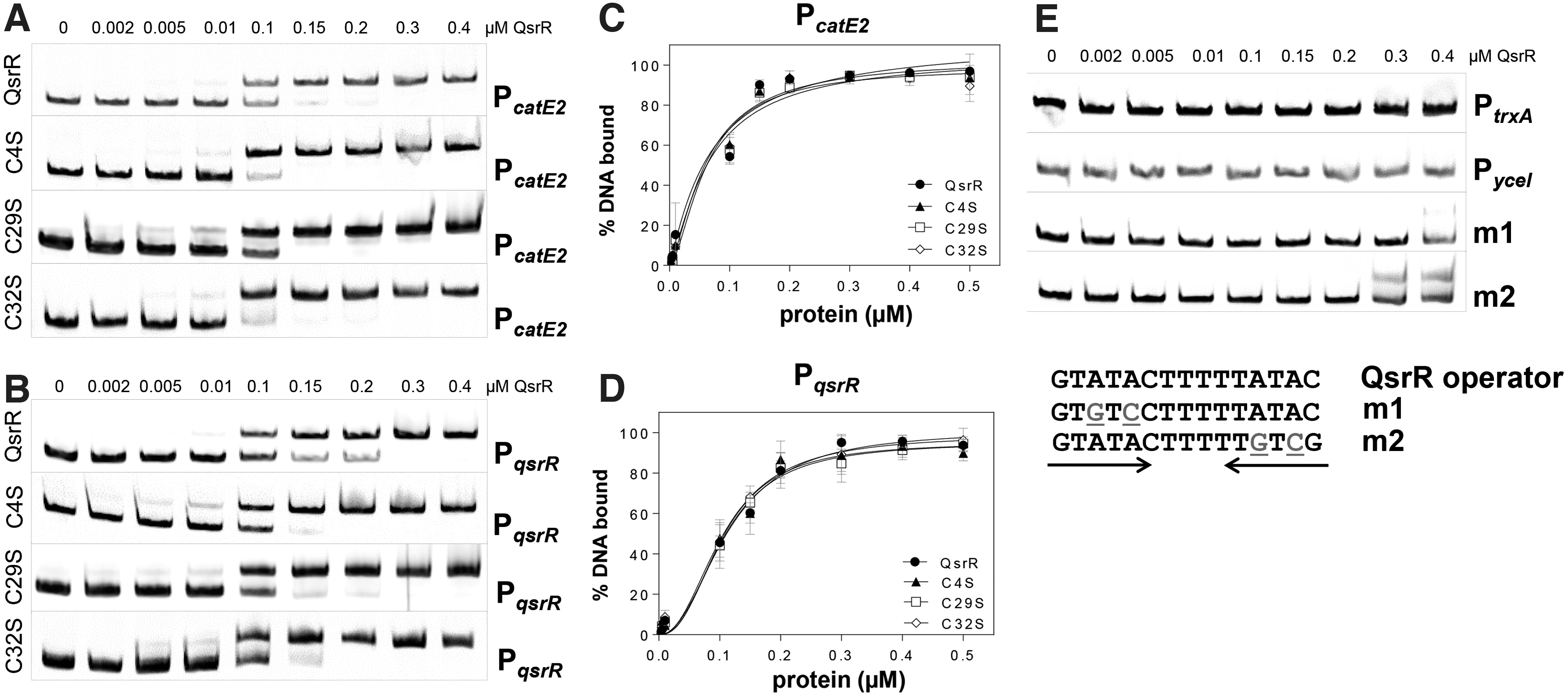
Thus, the single Cys mutations do not affect the DNA-binding activity of QsrR in vitro (Fig. 2A–D). However, the Cys29,32S double mutant might reveal the role of the Cys residues in DNA binding in vitro as was shown by the transcriptional assays in vivo (Fig. 1C–H).
The specificity of QsrR binding to the catE2 and qsrR promoter regions was verified using the upstream promoter region of the unrelated trxA and yceI genes and mutated qsrR promoter probes (m1, m2), with two base substitutions in each half of the inverted repeat sequence (Fig. 2E). QsrR was unable to bind to the trxA and yceI promoters or to the mutated qsrR promoter probes, supporting the specific DNA-binding activity to the QsrR operator sequence.
Quinones, diamide, and HOCl lead to reversible inhibition of the DNA-binding activity of QsrR in vitro, which depends on Cys4 and Cys29
Next, we used electrophoretic mobility shift assays (EMSAs) to study the role of the three Cys residues of QsrR for redox sensing of quinones, diamide, and HOCl stress in vitro. Previously, BQ was shown to inhibit the DNA-binding activity of the QsrRC29,32S mutant protein in vitro due to thiol-S-alkylation of the redox-sensing Cys4 (Ji et al, 2013). However, the reversibility of the effect of quinones on the DNA-binding activity of the QsrR WT protein and the three Cys mutants has not been analyzed in the previous study (Ji et al, 2013).
Treatment of QsrR with increasing concentrations of MHQ (0.25–20 μM) resulted in the dissociation of QsrR from the catE2 and qsrR promoter regions (Fig. 3A, B). DNA binding of MHQ-treated QsrR could be almost restored after treatment with 10 mM dithiothreitol (DTT), indicating that MHQ acts mainly via the oxidative mode, leading to reversible thiol-oxidation and inactivation of QsrR (Fig. 3C, D). DNA-binding assays of the three Cys mutants in the presence of MHQ revealed that Cys4 is essential for redox sensing of QsrR, since the DNA-binding activity of the C4S mutant to the catE2 and qsrR operators was not inhibited by MHQ (Fig. 3A, B).
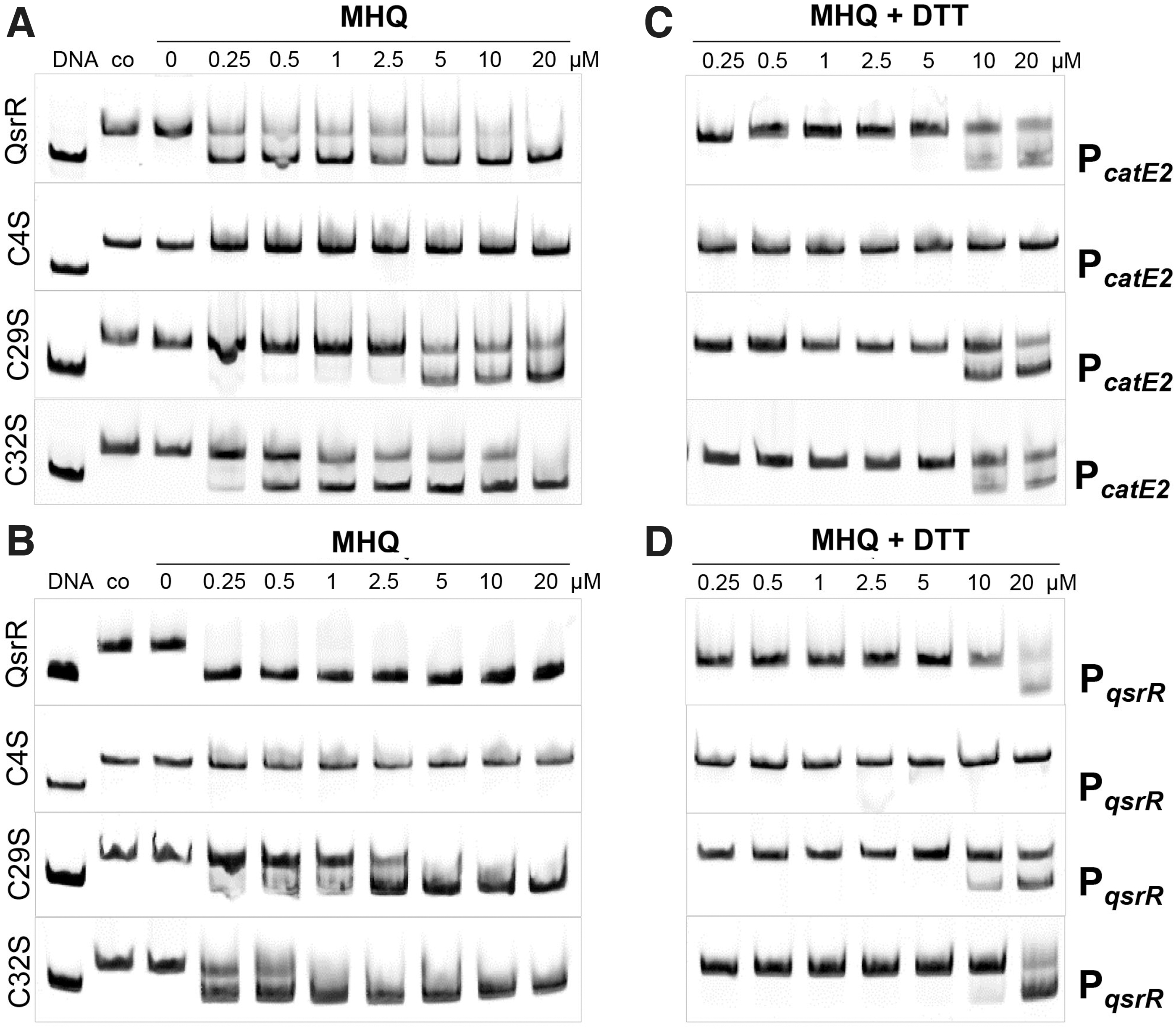
The C32S mutant showed a similar response to quinones as the QsrR WT protein, whereas the C29S mutant was less sensitive to inactivation by MHQ, indicating that Cys29 is partly involved in quinone sensing (Fig. 3A, B). Further, the DNA-binding activities of the quinone-treated C29S and C32S mutant proteins could be almost restored after DTT reduction, supporting that QsrR, the C29S and C32S mutants are reversibly oxidized by MHQ in vitro (Fig. 3C, D). In conclusion, Cys4 is essential for MHQ sensing, whereas the non-conserved Cys29 also functions in redox regulation of QsrR in vitro, involving a reversible thiol switch mechanism.
Since the QsrR regulon was strongly induced under disulfide stress (Fig. 1A, B), we analyzed the effect of diamide and HOCl on the DNA-binding activity of QsrR and its Cys mutants. Treatment of QsrR with 25–50 μM diamide resulted in its partial release from the catE2 promoter region (Fig. 4A). DNA-binding activity of diamide-treated QsrR could be restored by DTT reduction, indicating that QsrR is reversibly oxidized and inactivated by a thiol switch mechanism in response to diamide stress (Fig. 4A).
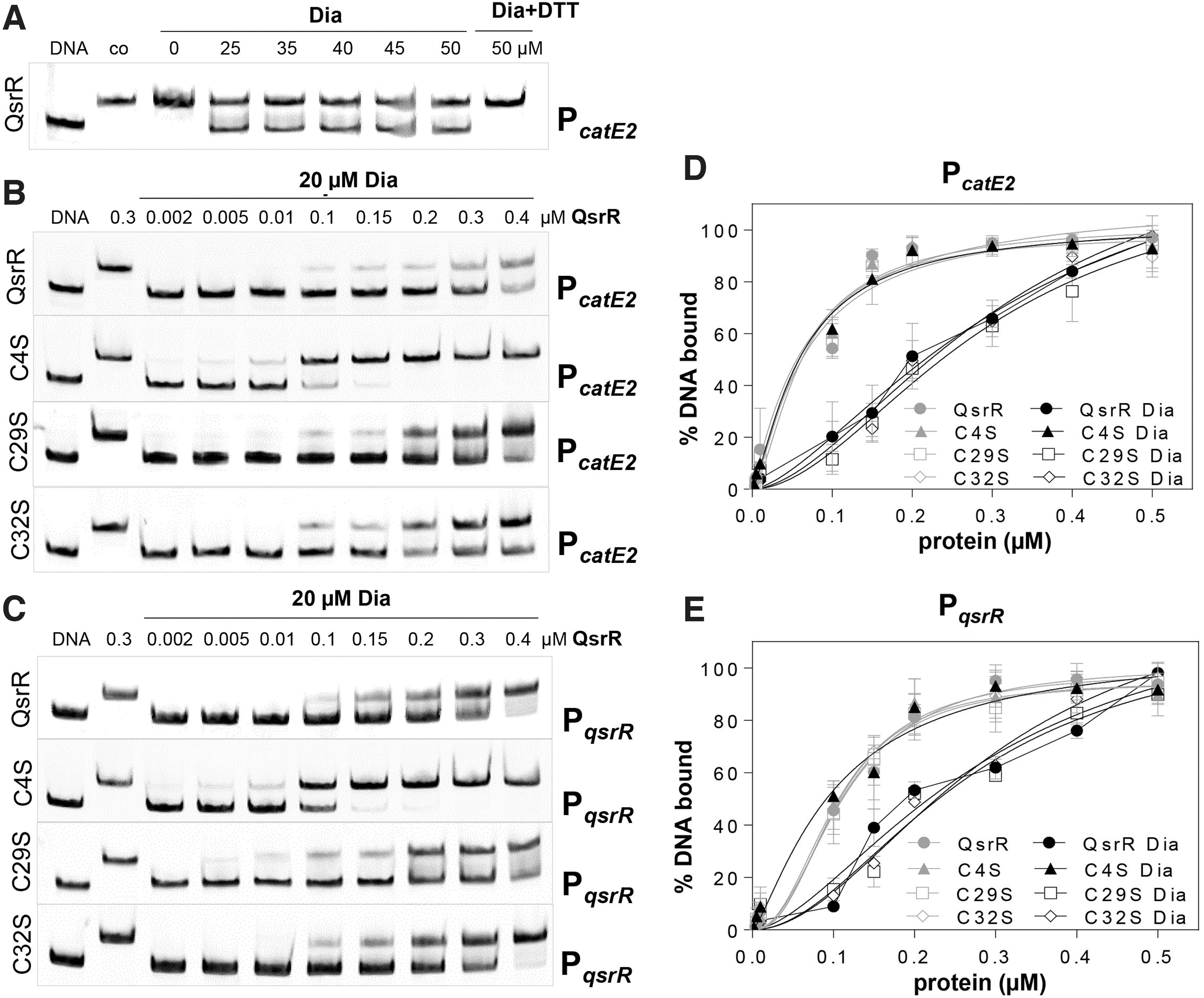
The comparison of the DNA-binding activities of QsrR and the three Cys mutants to the catE2 and qsrR promoters after diamide treatment revealed that Cys4 is required for redox sensing of diamide, since the C4S mutant did not respond to disulfide stress (Fig. 4B–E). However, the DNA-binding activities of the C29S and C32S mutants were inhibited by diamide stress, indicating that both Cys residues are not essential or can replace each other for redox sensing of disulfide stress (Fig. 4B–E). The unresponsiveness of the C4S mutant to diamide was reflected by the KD values, which were determined for the catE and qsrR promoters as 61.1 and 102.9 nM, respectively.
These KD were similar under oxidized and reduced conditions (Figs. 2C, D and 4D, E). In contrast, the KD values of QsrR and the QsrRC32S mutant protein increased after diamide treatment to a similar extent for the catE2 (337.5 and 325.5 nM) and qsrR promoters (291.9 and 295.6 nM), respectively, indicating that Cys32 is not involved in diamide sensing (Fig. 4D, E). A slightly decreased KD value of the QsrRC29S mutant protein was estimated for the catE2 (296.3 nM) and qsrR promoters (266.2 nM) in response to diamide treatment (Fig. 4D, E). These results support that Cys4 and Cys29 of QsrR are involved in redox sensing of strong oxidants, such as diamide stress in vitro.
Similarly, HOCl treatment resulted in decreased DNA-binding activity of QsrR and the C29S and C32S mutant proteins to the catE2 and qsrR promoters, indicating that Cys29 and Cys32 are dispensable or can replace each other for redox sensing of HOCl stress in vitro (Fig. 5A, B). The DNA-binding activities of the diamide-treated QsrR, the C29S and C32S mutants could be restored with DTT, supporting that QsrR responds via thiol switch mechanisms (Fig. 5C, D). In contrast, the C4S mutant was impaired in redox sensing and did not respond to HOCl stress in the DNA-binding assays in vitro (Fig. 5A, B, D, E).
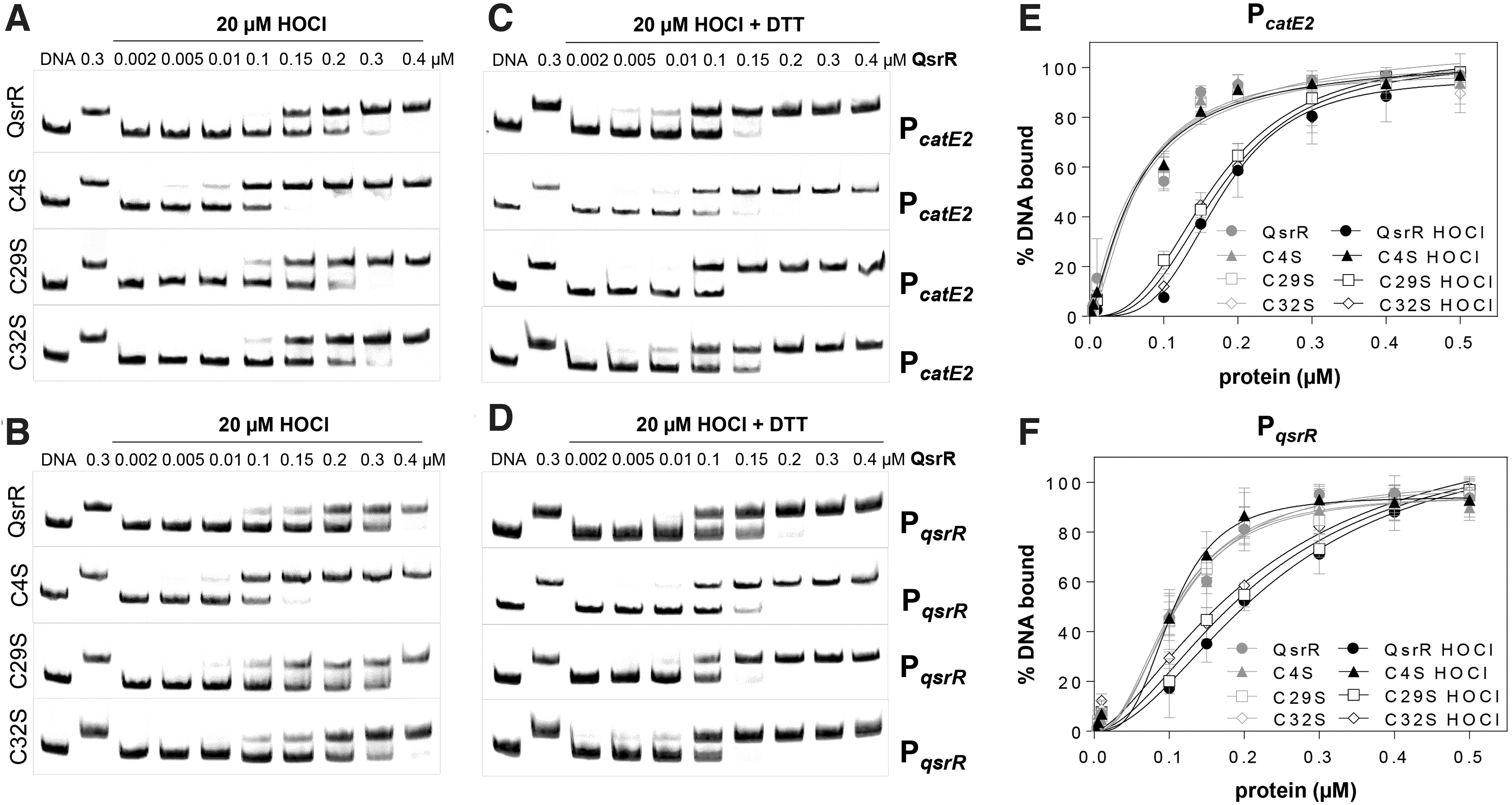
The unresponsiveness of the C4S mutant toward HOCl stress was evident also by the low KD values, which were determined as 63.3 and 103 nM for the catE2 and qsrR promoters, respectively (Fig. 5E, F). In contrast, the KD values of the QsrR, QsrRC29S, and QsrRC32S proteins were increased after HOCl treatment at similar levels for the catE2 (176.3, 167.8, and 175.5 nM) and the qsrR promoters (239.9, 222, and 221 nM), respectively (Fig. 5E, F). Altogether, these results revealed that QsrR is reversibly inactivated under quinone and disulfide stress in vitro, which requires Cys4 and in part Cys29 for redox sensing in vitro.
QsrR senses disulfide stress and quinones by intermolecular disulfides between Cys4 and Cys29′ in vitro and in vivo
To reveal the redox-sensing mechanism of QsrR under quinone and disulfide stress in vitro, thiol-oxidations of QsrR and the QsrR Cys mutant proteins were analyzed by non-reducing sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) (Fig. 6). Exposure to diamide and HOCl resulted in QsrR oxidation and formation of an intermolecular disulfide band with the size of ∼30 kDa, which was reversible with DTT (Fig. 6A, B). These QsrR intermolecular disulfides were also visible after MHQ treatment, although less pronounced compared with diamide-treated QsrR (Fig. 6A, C).
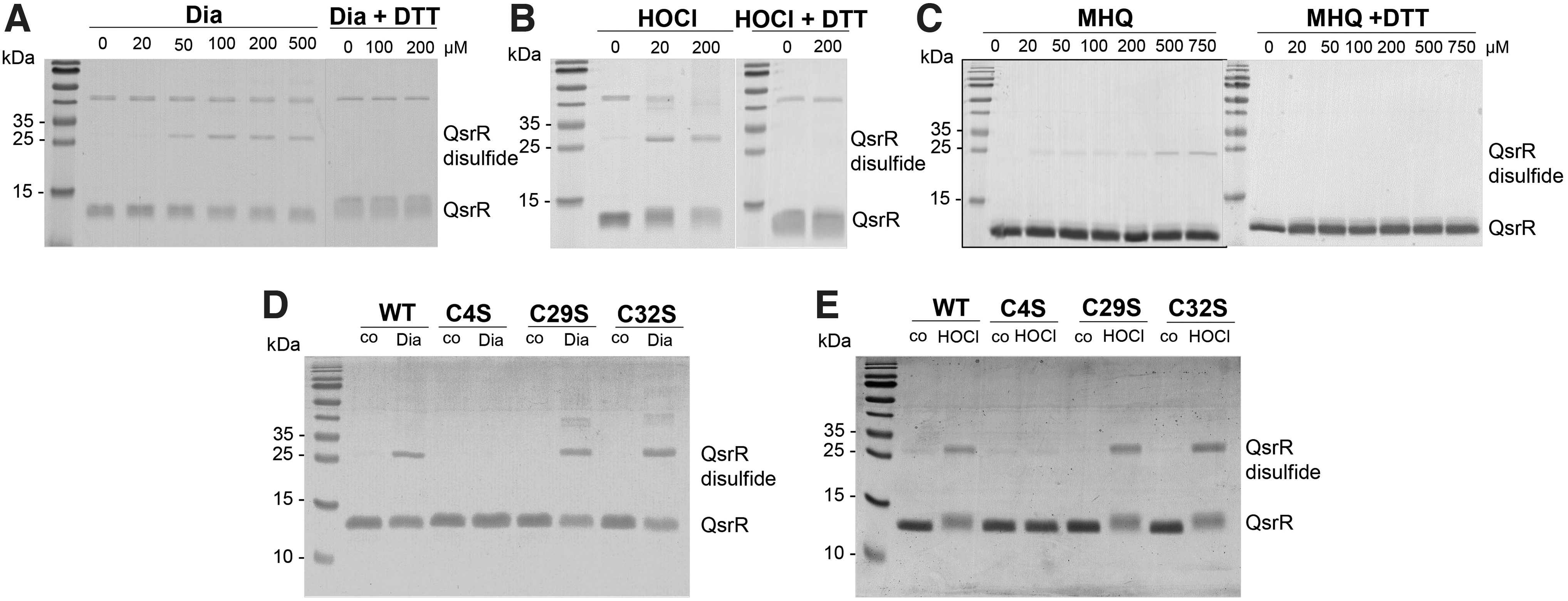
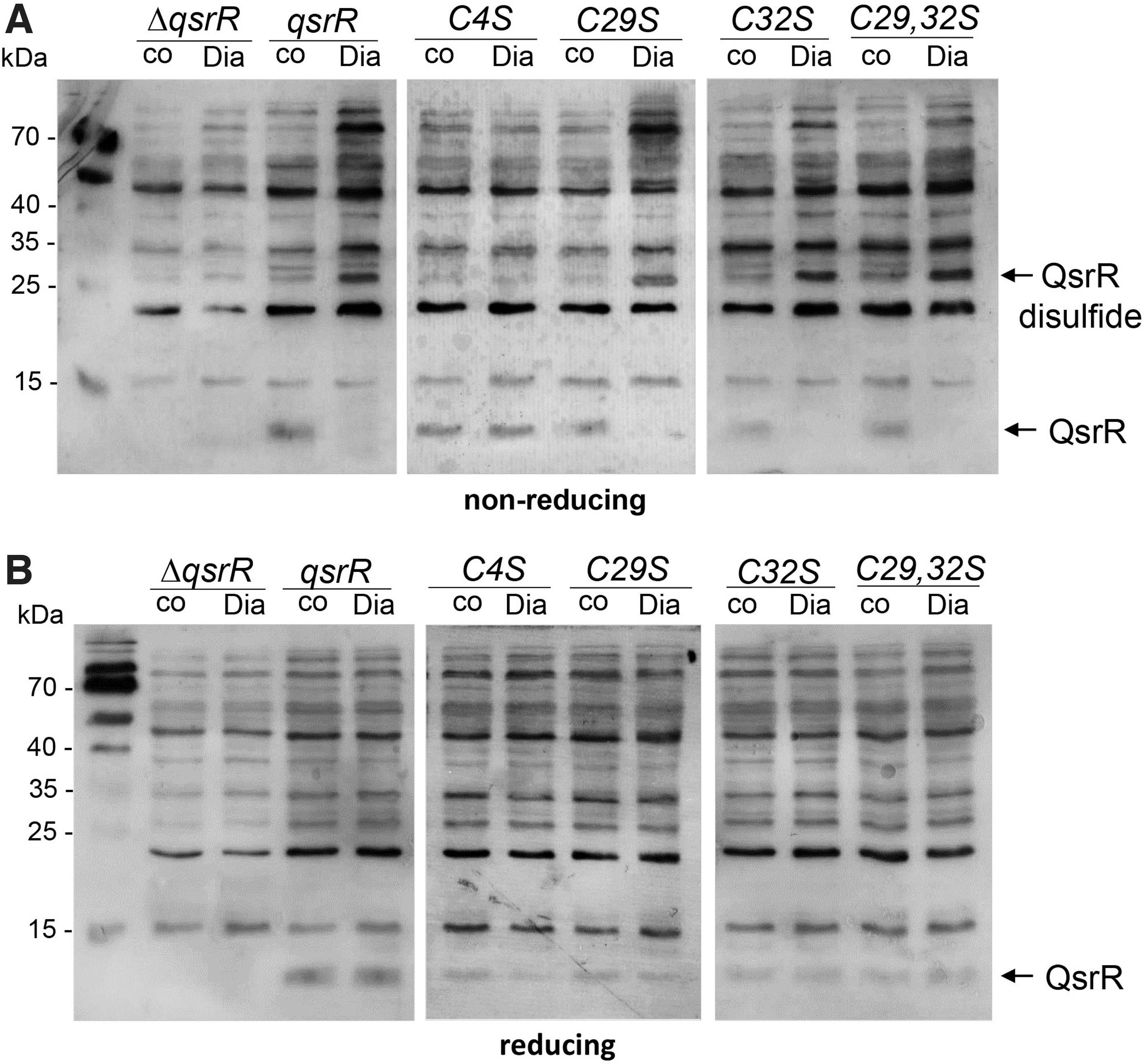
The oxidation to intermolecular disulfides was observed also in the C29S and C32S mutants after diamide and HOCl treatment, but not in the C4S mutant, supporting that Cys4 is the redox-sensing Cys and involved in intersubunit disulfide formation in vitro (Fig. 6D, E). To monitor QsrR oxidation to intermolecular disulfides in vivo, we performed non-reducing Western blot analyses of S. aureus complemented strains expressing the His-tagged QsrR, C4S, C29S, C32S, and C29,32S proteins. The protein extracts were harvested after alkylation with iodoacetamide (IAM) in the dark to block reduced thiols in S. aureus cells.
Although the Western blot results revealed the oxidation of the QsrR WT and C29S, C32S, and C29,32S mutant proteins to the intersubunit disulfides upon diamide stress in S. aureus cells, the C4S mutant was unable to form intermolecular disulfides upon diamide stress in vivo (Fig. 7A). The intermolecular QsrR disulfides were reversible in the reducing Western blot analysis (Fig. 7B). These results support the critical role of Cys4 in the redox sensing of disulfide stress by an intersubunit thiol switch mechanism in S. aureus.
Next, we were interested in identifying the cross-linked Cys residues in the QsrR disulfide after diamide stress in vitro. The reduced QsrR and oxidized QsrR intermolecular disulfide bands were tryptic in-gel digested, and the peptides were subjected to LC-MS/MS analysis (Fig. 8). The Mascot search results identified two main disulfide cross-linked peptides (Fig. 8A, B), which were present in the oxidized QsrR disulfide band but not in the reduced QsrR sample as revealed by the extracted ion chromatogram (Fig. 8C, D).
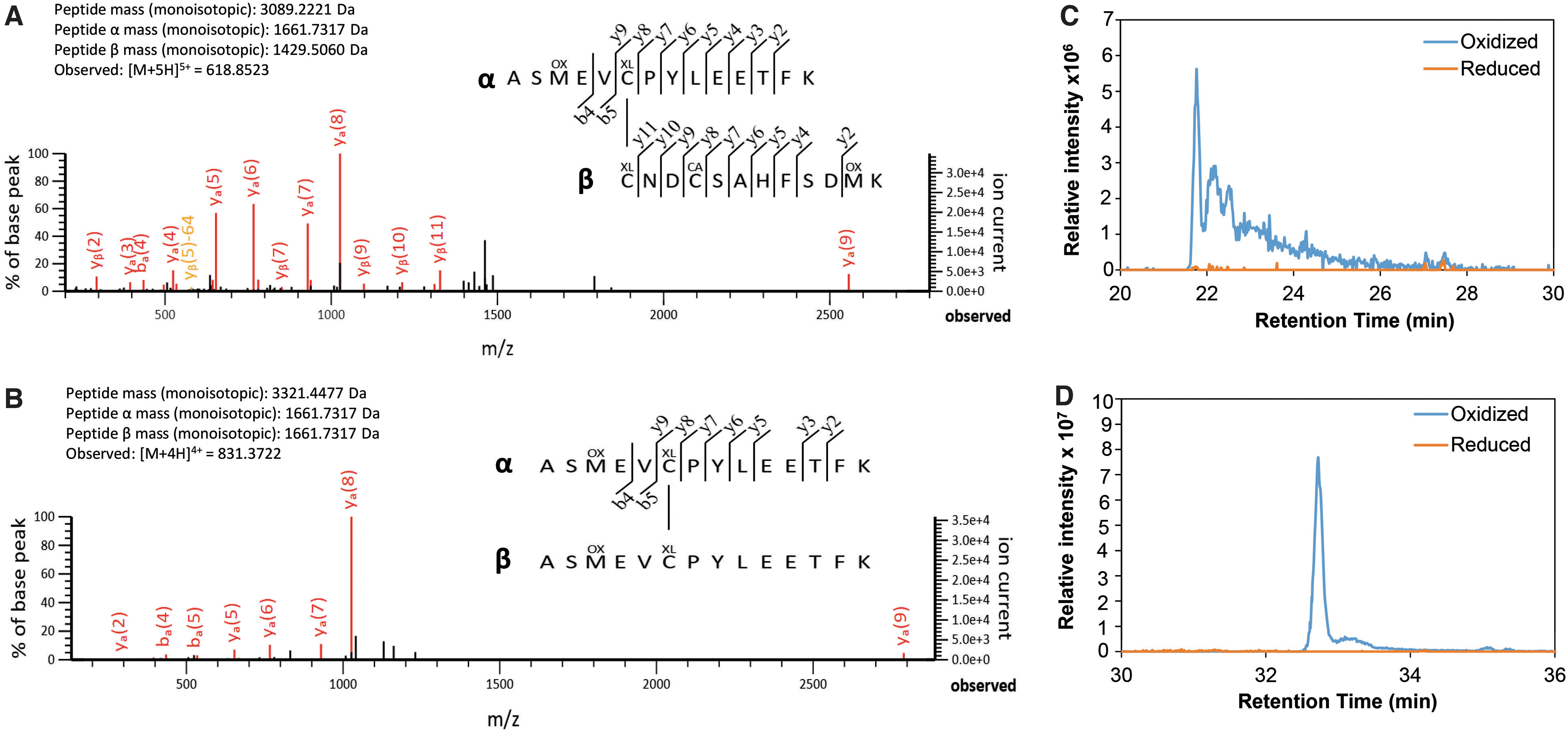
These oxidized disulfide peptides include the Cys4-Cys29′ disulfide with the monoisotopic mass of 3089.2221 Da (Fig. 8A) and the Cys4-Cys4′ intermolecular disulfide with the monoisotopic mass of 3321.4477 Da (Fig. 8B) as revealed by the MS/MS spectra. The Cys4-Cys32′ disulfide peptide was not detected in the diamide-treated QsrR sample. According to the previously characterized YodB mechanism (Chi et al, 2010a; Lee et al, 2016), QsrR is most likely oxidized to Cys4-Cys29′ intermolecular disulfides, which have been reshuffled to Cys4-Cys4′ disulfides in vitro due to the high reactivity of the redox-sensing Cys4.
These results are in agreement with our Northern blots, indicating that Cys4 and either Cys29′ or Cys32′ are required for the redox sensing of diamide and MHQ in vivo (Fig. 1C–F). Thus, QsrR senses quinones and oxidants by Cys4-Cys29′ intersubunit disulfide formation, leading to the derepression of transcription of the QsrR regulon.
Allicin causes S-thioallylation of QsrR, leading to inhibition of DNA binding in vitro
Transcription of the QsrR regulon was strongly upregulated by the organosulfur compound allicin (Fig. 1A, B and Supplementary Table S1). The mode of action of allicin involves widespread S-thioallylations of protein and LMW thiols in bacteria and human cells (Chi et al, 2019; Gruhlke et al, 2019; Loi et al, 2019). Thus, we were interested in whether QsrR is redox-controlled by the formation of intersubunit disulfides or S-thioallylations after allicin treatment. In gel-shift assays, the DNA-binding activity of QsrR to the catE2 and qsrR promoters was reversibly inhibited after allicin treatment (Fig. 9A).
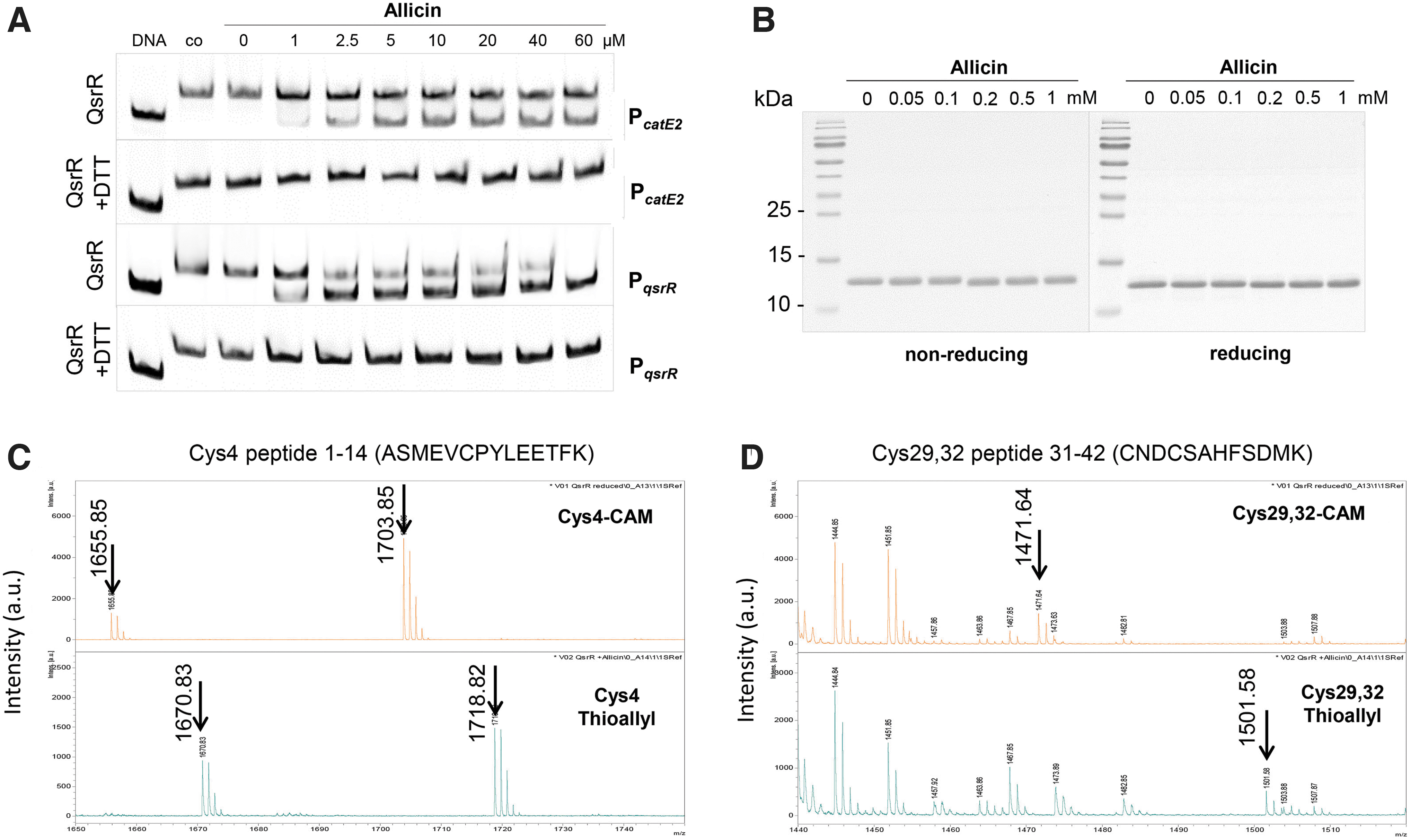
The non-reducing SDS-PAGE analyses revealed no formation of the intermolecular disulfide in the QsrR protein after exposure to allicin (Fig. 9B). Thus, we hypothesized that allicin might cause S-thioallylation of the Cys residues of QsrR, leading to its inactivation and relief of repression. The thiol-modifications of untreated and allicin-treated QsrR bands were investigated by matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF-MS) analysis after in-gel tryptic digestion.
The MS1 scans of the peptides in the reduced QsrR sample showed the carbamidomethylated Cys4 peptide with the mass of 1703.85 Da and the carbamidomethylated Cys29,32 peptide with the mass of 1471.64 Da, indicating that Cys4, Cys29, and Cys32 are reduced in the QsrR control sample (Fig. 9C, D, upper panel). In the allicin-treated QsrR protein sample, all three Cys residues were found to be S-thioallylated as revealed by the mass shift of 72 Da for C3H5S1 at Cys residues.
In the MS1 spectra, the S-thioallylated Cys4 peptide was identified with the mass of 1718.82 Da (1646.7440 + 72 Da) and the Cys29,32 peptide with the mass of 1501.58 Da (1357.4970 + 144 Da) (Fig. 9C, D, lower panel). Of note, the Cys4 peptide was also identified with a dethiomethylated methionine sulfoxide in the reduced and S-thioallylated samples, as labeled in the MS1 spectra with the mass peaks of 1655.85 Da and 1670.83 Da, respectively. Overall, these data clearly confirm that allicin causes S-thioallylation of all three Cys residues in QsrR, leading to its inactivation and the strong induction of the QsrR regulon in allicin-treated S. aureus cells (Fig. 1A, B). Moreover, the S-thioallylation at Cys4 alone in the C29,32S mutant might be sufficient to cause conformational and structural changes, leading to derepression of catE2 transcription (Fig. 1G, H).
The QsrR regulon confers resistance toward quinones and oxidants, which depends on Cys4 and either Cys29′ or Cys32′
Previously, the qsrR deletion mutant has been shown to confer resistance toward quinone stress in S. aureus (Fritsch et al, 2019; Ji et al, 2013). Since QsrR responds to quinone and disulfide stress by different thiol switches, we investigated the growth and survival phenotypes of the qsrR mutant and Cys mutant strains under MHQ, H2O2, HOCl, and allicin stress in S. aureus. Consistent with previous data (Fritsch et al, 2019; Ji et al, 2013), the qsrR mutant was not affected in growth after 100 μM MHQ stress and showed a 3- to 17-fold increased survival after treatment with 250 μM MHQ, supporting that the QsrR regulon contributes most strongly to the quinone resistance in S. aureus (Fig. 10A, E).
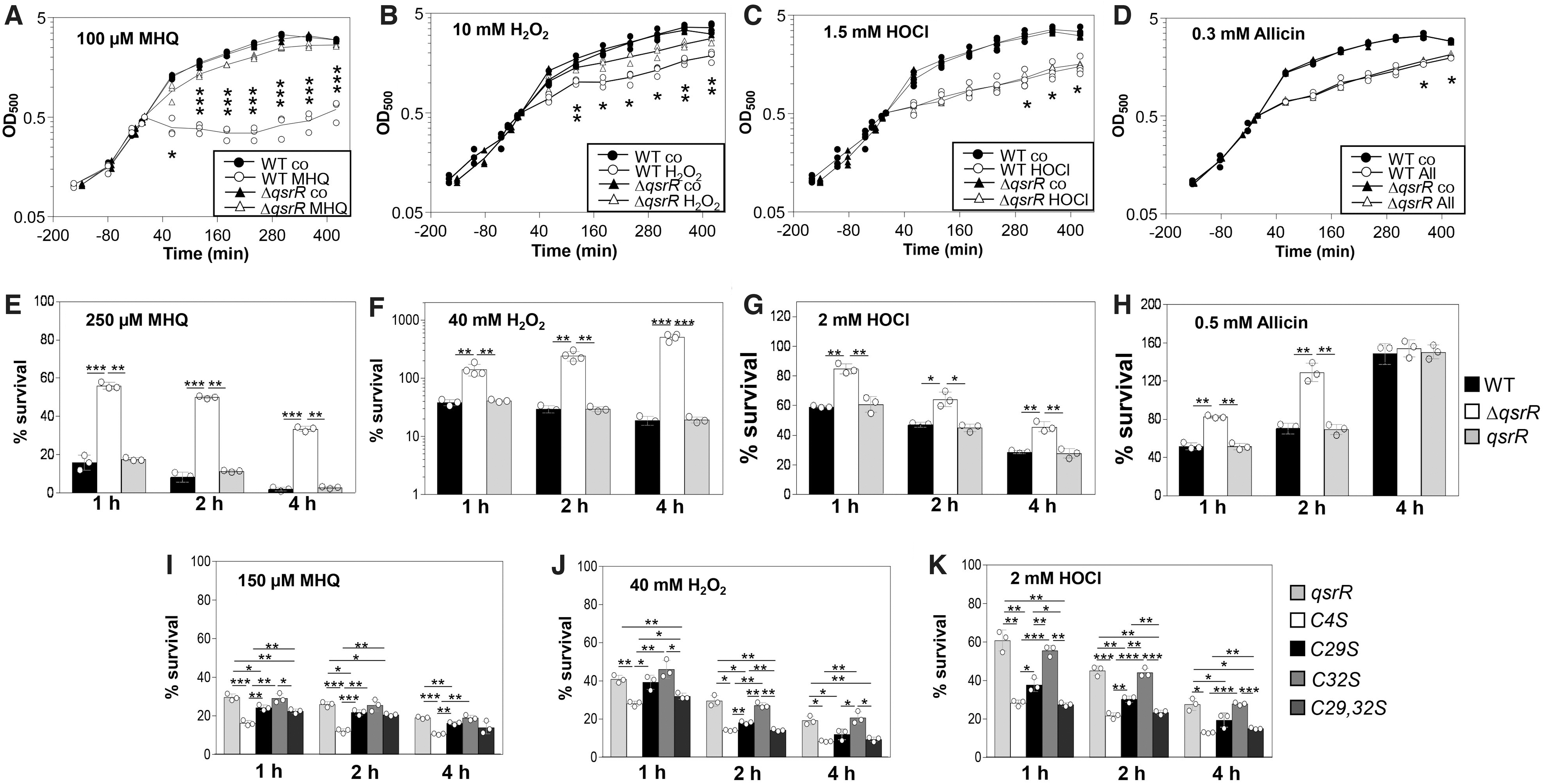
In growth curves, the qsrR mutant was only slightly more resistant after exposure to sublethal doses of 10 mM H2O2, 1.5 mM HOCl, and 0.3 mM allicin stress as compared with the WT (Fig. 10B–D). However, in survival assays, the qsrR mutant was significantly more resistant to the oxidants than the WT and showed an increased survival after treatment with lethal doses of 40 mM H2O2, 2 mM HOCl, and 0.5 mM allicin (Fig. 10F–H). All growth and survival phenotypes could be reversed to WT levels in the qsrR complemented strain (Fig. 10E–H and Supplementary Fig. S3A–D).
The phenotypes of the QsrR Cys mutants were analyzed regarding the resistance against MHQ, H2O2, and HOCl stress in comparison to the qsrR complemented strain (Fig. 10I–K and Supplementary Fig. S4). The QsrRC4S mutant showed a significantly decreased survival after MHQ, H2O2, and HOCl exposure as compared with the qsrR complemented strain, supporting its redox-sensing function under quinone and oxidative stress. In addition, the C32S mutant showed similar survival rates as the qsrR complemented strain after exposure to these stressors, indicating that Cys32 is not involved in redox sensing.
The C29S mutant showed a slightly reduced survival after MHQ, H2O2, and HOCl stress than the qsrR complemented strain, whereas the Cys29,32S double mutant was similarly sensitive as the C4S mutant to the oxidants (Fig. 10I–K). These data support that both Cys4 and Cys29 residues are required for redox regulation of QsrR by Cys4-Cys29′ intersubunit disulfide formation in vivo.
Discussion
In this article, we have shown that the quinone-sensing QsrR repressor is redox-controlled by different thiol switch mechanisms to sense quinones and disulfide stress in S. aureus. QsrR belongs to the MarR/DUF24 family of winged HTH transcriptional regulators, which possess the highly conserved HxlR-type HTH domain (IPR002577) and are widely distributed across firmicutes, actinobacteria, bacteroidetes, proteobacteria, and euryarchaeota (Supplementary Fig. S6). All HxlR-type HTH domain proteins share the conserved N-terminal Cys4 residue of QsrR, which was shown to be essential for redox sensing in the MarR/DUF24-homologs HxlR, YodB, CatR, and HypR of B. subtilis and QsrR of S. aureus (Supplementary Fig. S7) (Chi et al, 2010a, Chi et al, 2010b; Ji et al, 2013; Leelakriangsak et al, 2008; Palm et al, 2012; Yurimoto et al, 2005).
In addition, most MarR/DUF24-family regulators have one or two Cys residues at non-conserved positions, suggesting that these could also function as two-Cys regulators. Interestingly, Cys29 of QsrR was found to be widely conserved in homologs across Staphylococcus species, whereas Cys101 of YodB showed some conservation in homologs of Bacillus species, supporting the common regulatory model of QsrR and YodB by intersubunit disulfide formation in response to oxidative stress (Supplementary Fig. S7). However, there are other MarR/DUF24 homologs, such as HxlR, which harbor only the conserved Cys and might use monothiol mechanisms for redox sensing (Yurimoto et al, 2005).
QsrR was previously shown to respond to BQ by thiol-S-alkylation of the redox-sensing Cys4 to control the ring-cleavage dioxygenases (CatE, CatE2) and quinone reductases (AzoR1, YodC) in S. aureus (Ji et al, 2013). However, the S-alkylation model has been shown for the C29,32S mutant protein in vitro, but not for the QsrR WT protein containing three Cys residues in vivo (Ji et al, 2013). In addition, the reversibility of the QsrR response to quinones and other thiol-reactive compounds was not investigated in the previous study (Ji et al, 2013).
In our earlier RNAseq transcriptome data and the Northern blot results of this article, the QsrR regulon was found to respond most strongly to MHQ and strong oxidants, including diamide, HOCl, allicin, and AGXX, which cause disulfide stress in S. aureus (Fritsch et al, 2019; Loi et al, 2019; Loi et al, 2018a, Loi et al, 2018b). Similarly, the QsrR-homologue YodB was shown to sense and respond to quinones and diamide by a thiol switch mechanism in B. subtilis (Chi et al, 2010a, Chi et al, 2010b), indicating that QsrR might be not directly alkylated by quinones, but rather senses the oxidative mode of quinones in vivo.
The YodB-regulated CatE was active as catechol-2,3-dioxygenase in catechol detoxification to generate 2-hydroxymuconic semialdehyde, whereas the azoreductases AzoR1 and AzoR2 were involved in quinone and diamide reduction in B. subtilis (Antelmann et al, 2008; Tam le et al, 2006). In addition, the SifR-controlled catechol-2,3-dioxygenase CatE and NAD(P)H-dependent quinone reductase YwnB were shown to catalyze detoxification of catechol iron complexes and catecholamine stress hormones in S. pneumoniae (Zhang et al, 2022). Based on their induction by strong oxidants, the QsrR-controlled enzymes CatE, CatE2, YodC, and AzoR1 might function in quinone, diamide, and allicin detoxification in S. aureus.
Phenotype assays revealed that the QsrR regulon not only conferred the strongest protection against quinones, but also mediated significant resistance under H2O2, HOCl, and allicin stress. Our data further revealed that QsrR responds rather to the oxidative mode of quinones and strong oxidants, which cause disulfide stress in S. aureus. The toxicity of BQ was previously shown to be related to its S-alkylation and oxidative mode (Kumagai et al, 2002; Monks et al, 1992; O'Brien, 1991). In B. subtilis, we could confirm that BQ can act via both the alkylation and oxidative mode, depending on the quinone concentration and the capacity of cells for detoxification and reduction of quinones (Liebeke et al, 2008).
Sublethal doses of BQ caused the reversible thiol-oxidation of GapDH in the redox proteome of B. subtilis, supporting the oxidative mode of BQ, whereas higher lethal doses resulted in aggregation and depletion of thiol-containing proteins by the thiol-S-alkylation chemistry (Liebeke et al, 2008). However, the alkylation mode was always accompanied by cell death due to the irreversible protein damage and depletion of thiol-containing proteins and LMW thiols (Liebeke et al, 2008).
Thus, under physiological conditions, QsrR should sense rather the oxidative mode of sublethal doses of quinones to induce the quinone detoxification enzymes, which enables the recovery of the cells from quinone stress. In the alkylation mode, the quinone response would not be reversible since the S-alkylated proteins cannot be repaired, indicating no shutdown of the quinone response and no recovery of the cells from quinone stress, which would not be favored under in vivo conditions.
In this study, QsrR was shown to function as a typical two-Cys-type redox-sensing regulator, which uses different thiol switch mechanisms for redox sensing of quinones and disulfide stress. The transcriptional data showed that Cys4 and either Cys29 or Cys32 are important for redox sensing of MHQ and diamide, whereas only Cys4 is required for allicin sensing. This is in agreement with the different thiol switch models of QsrR as revealed by non-reducing SDS-PAGE, MS, and Western blot analysis. Diamide and HOCl stress lead to Cys4-Cys29′ intermolecular disulfide formation, whereas allicin causes S-thioallylations of all three Cys residues to inhibit the repressor activity of QsrR.
The DNA-binding assays confirmed that Cys4 is most important for redox sensing of QsrR on quinone, diamide, and HOCl stress in vitro. The KD values of the C4S mutant were unchanged under diamide and HOCl stress compared with the untreated QsrR, indicating the unresponsiveness of the C4S mutant toward thiol stress. In contrast, the QsrR WT protein and the C29S and C32S mutants showed similar increased KD values after quinone and disulfide stress, supporting their capability to respond to thiol stress. However, the inhibition of the QsrR DNA-binding activity by diamide, HOCl, and MHQ stress was reversible with DTT, providing confirmation that thiol switch models are involved in QsrR regulation.
The thiol switch mechanisms of the QsrR repressor resemble the redox-sensing mechanisms of the OHP-sensing OhrR repressors of B. subtilis and Xanthomonas campestris (Antelmann and Helmann, 2011; Hong et al, 2005; Lee et al, 2007; Mongkolsuk and Helmann, 2002; Newberry et al, 2007). The one-Cys-type OhrRBS protein of B. subtilis senses OHP and HOCl stress via its single Cys15 residue by S-bacillithiolation (Antelmann and Helmann, 2011; Chi et al, 2011; Lee et al, 2007), whereas the two-Cys-type OhrRXC homolog of X. campestris forms intermolecular disulfides between Cys22 and Cys127′ leading to conformational changes in the winged HTH motif and dissociation of OhrR from the operator DNA (Newberry et al, 2007; Panmanee et al, 2006).
Moreover, the one-Cys OhrR protein could be converted to a two-Cys-type redox sensor by the introduction of a C-terminal Cys residue by either G120C or Q124C mutations, resulting in intersubunit disulfide formation as an alternative thiol switch mechanism (Soonsanga et al, 2008). Similar to QsrR, the YodB repressor of B. subtilis was demonstrated to sense quinones and diamide by intersubunit disulfide formation between Cys6 and either of the C-terminal Cys101′ or Cys108′ residues in vitro and in vivo (Chi et al, 2010a; Linzner et al, 2021).
The structural mechanism of YodB inactivation by intersubunit disulfide formation under diamide treatment and by S-alkylation of Cys6 with methyl-p-benzoquinone has been resolved in vitro (Lee et al, 2016). Interestingly, S-alkylation of Cys6 does not cause major conformational changes in the HTH motifs of YodB, whereas the oxidation to the Cys6-Cys101′ intersubunit disulfides under diamide stress was accompanied by large structural rearrangements (Lee et al, 2016). The structural changes of S-alkylated QsrR with menadione involved steric clashes with the DNA backbone, a 10° rigid-body rotation, and 9 Å elongation of the two QsrR monomers, but no major conformational changes in the monomer structures (Ji et al, 2013).
Since Cys4 and Cys29′ of adjacent subunits are 15.2 Å apart in the structure of the QsrR dimer (Supplementary Fig. S5), formation of the Cys4-Cys29′ intersubunit disulfides will require structural rearrangements, which should be incompatible with DNA binding. Although Cys4 is located at the N-terminal α1 helix in the QsrR dimer interface, Cys29 and Cys32 are located in the loop region between the α2 and α3 helices. The α3 and α4 helices form together with the β1 and β2 sheets the winged HTH DNA-binding domains (Ji et al, 2013). The mutation of Cys29 or Cys32 showed only minor effects on the DNA-binding activity of QsrR in vivo, whereas the Cys4-Cys29′ intersubunit disulfide might cause conformational changes in the winged HTH motifs, leading to dissociation of the QsrR repressor from the operator DNA as revealed by the gel shift assays.
Apart from the Cys4-Cys29′ intermolecular disulfide, the MS data also identified Cys4-Cys4′ crosslinks, which are most likely in vitro artefacts due to reshuffling of Cys4-Cys29′ peptides. The distance between Cys4 and Cys4′ of both subunits is 27 Å (Supplementary Fig. S5), which is too far apart for disulfide bond formation. The Cys4-Cys4′ disulfide is also not physiologically relevant, because the Cys29,32S mutant did not respond to diamide and MHQ stress in transcriptional studies in vivo.
Moreover, in phenotype analyses, both Cys4 and Cys29,32S mutants were sensitive in survival assays after oxidative stress, supporting that Cys4 and either Cys29′ or Cys32′ are important for redox sensing and engaged in intersubunit disulfide formation in vivo. Similar intermolecular thiol switch models were revealed as common redox-sensing mechanisms in two-Cys-type MarR-family redox sensors, such as OhrRXC (Newberry et al, 2007), HypRBs (Palm et al, 2012), YodBBs (Chi et al, 2010a), and the Rrf2-family regulator HypRSa (Loi et al, 2018b), supporting the Cys4-Cys29′ intermolecular disulfide model of QsrR as revealed in this work.
In addition, we found that QsrR senses allicin by S-thioallyalation of all three Cys residues, leading to the reversible inactivation of the QsrR repressor. Since the C29,32S mutant still responded to allicin in transcriptional analyses in vivo, S-thioallylation of Cys4 might be sufficient for repressor inactivation, causing similar structural changes as observed for S-alkylation of Cys4 by quinones (Ji et al, 2013). The S-thioallylation of the redox-sensing Cys residues of YodB, HypR, and OhrRBS was previously found in B. subtilis cells under allicin stress in vivo, resulting in upregulation of the corresponding regulons in the transcriptome (Chi et al, 2019).
In addition, the single Cys OhrR repressor of B. subtilis was shown to be inactivated by S-bacillithiolation under HOCl and OHP stress, resulting in derepression of the ohrA gene, encoding a peroxiredoxin, which confers resistance toward OHP and HOCl stress (Chi et al, 2011; Lee et al, 2007). Thus, S-thioallylation of QsrR at the conserved Cys4 might cause similar structural changes as observed for the S-bacillithiolated OhrRBS or the menadione-alkylated monothiol QsrR repressor (Ji et al, 2013).
Finally, the quinone-sensing QsrR and MhqR regulons were previously shown to confer resistance to antimicrobials with quinone-like elements, including ciprofloxacin, rifampicin, and pyocyanin in S. aureus (Fritsch et al, 2019; Noto et al, 2017). Although QsrR conferred stronger resistance to MHQ, the MhqR regulon showed a better protection against the quinone-like antimicrobials pyocyanin, ciprofloxacin, norfloxacin, and rifampicin in S. aureus (Fritsch et al, 2019). Ciprofloxacin belongs to the fluoroquinolone antibiotics, which are frequently applied in the clinics for the treatment of patients with S. aureus infections.
Although the target of ciprofloxacin is the DNA gyrase, the bacterial killing has been associated with ROS generation (Dwyer et al, 2007; Phillips-Jones and Harding, 2018). Pyocyanin is a blue-colored redox-active phenazine antibiotic produced by the pathogen Pseudomonas aeruginosa (Price-Whelan et al, 2006), which causes cystic fibrosis and often co-occurs together with S. aureus during wound and respiratory tract infections in cystic fibrosis patients (Biswas and Gotz, 2021). Pyocyanin was shown to generate ROS as a toxicity mechanism (Mahajan-Miklos et al, 1999; Noto et al, 2017), but its redox-cycling activity further contributed to the proton motive force, ATP generation, redox homeostasis, and biofilm formation in P. aeruginosa (Dietrich et al, 2008; Glasser et al, 2014; Price-Whelan et al, 2007).
The resistance of the qsrR mutant to quinone-like antimicrobials might be caused either by diminishing ROS produced by the antibiotics or by the degradation or reduction of the compounds by the QsrR-controlled catechol-2,3-dioxygenases (CatE, CatE2) and quinone reductases (AzoR1, YodC) (Noto et al, 2017).
Taken together, our results revealed that two-Cys-type MarR-type redox sensors, such as QsrR, can sense different redox-active compounds via distinct thiol switches, including intersubunit disulfides or S-thiolations, which lead to inactivation of the repressor function and induction of detoxification pathways with broad specificities for thiol-reactive oxidants and electrophiles. The S-alkylation model might be relevant under toxic quinone concentrations, whereas QsrR might sense mainly the oxidative mode via thiol switches upon the exposure of cells to sublethal doses of quinones or quinone-like antimicrobials. Future studies should be directed to elucidate the thiol-disulfide reductase pathways responsible to regenerate reduced QsrR upon recovery from quinone and oxidative stress in S. aureus.
Materials and Methods
Bacterial strains, growth, and survival assays
Bacterial strains and primers are listed in Supplementary Tables S2 and S3. Escherichia coli strains were cultivated in Luria Bertani (LB) medium for plasmid construction and protein expression. For growth and survival assays, S. aureus strains were cultivated in RPMI medium with 0.75 μM FeCl3 and 2 mM glutamine as supplements as previously described (Dorries and Lalk, 2013). The strains were exposed to the thiol-reactive compounds during the exponential growth phase at an optical density at 500 nm (OD500) of 0.5. Survival assays were performed by plating 100 μL of serial dilutions of S. aureus strains onto LB agar plates for colony forming units counting.
Statistical analysis was performed using the Student's unpaired two-tailed t-test. The chemicals MHQ, H2O2, diamide, methylglyoxal, formaldehyde and the antibiotics were purchased from Sigma Aldrich. NaOCl was purchased from Honeywell Fluka. NaOCl dissociates in aqueous solution to HOCl and OCl− (Estrela et al, 2002). Thus, the concentration of HOCl was determined by absorbance measurements as previously described (Winter et al, 2008). Allicin and AGXX were generated as previously reported (Linzner and Antelmann, 2021; Loi et al, 2019).
Construction of the S. aureus COL ΔqsrR mutant as well as the qsrR, qsrRC4S, qsrRC29S, qsrRC32S, and qsrRC29,C32S complemented strains
The S. aureus COL ΔqsrR (SACOL2115) deletion mutant was previously constructed by allelic replacement via the pMAD E. coli/S. aureus shuttle vector (Fritsch et al, 2019). For construction of the His-tagged S. aureus qsrR, qsrRC4S, qsrRC29S, qsrRC32S, and qsrRC29,32S complemented strains, the pRB473 plasmid was used as described (Loi et al, 2018b). The qsrRC29S and qsrRC32S sequences were amplified from plasmids pET11b-qsrRC29S-His and pET11b-qsrRC32S-His. The qsrR and qsrRC4S sequences were amplified from S. aureus chromosomal DNA using the primers pRB-qsrR-for-BamHI or pRB-qsrRC4S-for-BamHI and pRB-qsrR-rev-KpnI (Supplementary Table S3). For construction of pRB473-qsrRC29,32S-His, two first-round polymerase chain reaction (PCR) products were performed using the primer pairs qsrRC29,32S-for and pRB-qsrR-rev-KpnI as well as qsrRC29,32S-rev and pRB-qsrR-for-BamHI, respectively.
The purified PCR products were fused and amplified by a second-round PCR using the primers pRB-qsrR-rev-KpnI and pRB-qsrR-for-BamHI. The PCR products were digested with BamHI and KpnI and inserted into pRB473. The plasmids were introduced into the ΔqsrR mutant via phage transduction as previously described (Loi et al, 2018b).
RNA isolation and Northern blot analysis
For RNA isolation, S. aureus strains were cultivated in RPMI medium and treated with various thiol-reactive compounds and antibiotics at an OD500 of 0.5 for 30 and 60 min, respectively, as indicated in the figure legends. Northern blot hybridizations were performed as described (Tam le et al, 2006; Wetzstein et al, 1992) with the digoxigenin-labeled catE2 and qsrR-specific antisense RNA probes, which were synthesized in vitro using the T7 RNA polymerase and the specific primer pairs as previously described (Busche et al, 2018; Imber et al, 2018) and in Supplementary Table S3.
Cloning, expression, and purification of His-tagged QsrR, QsrRC4S, QsrRC29S, and QsrRC32S proteins in E. coli
To construct the plasmids pET11b-qsrR-His and pET11b-qsrRC4S-His, the qsrR gene (SACOL2115) was amplified from chromosomal DNA of S. aureus COL using primers pET-qsrR-for-NheI or pET-qsrRC4S-for-NheI and pET-qsrR-rev-BamHI (Supplementary Table S3). The PCR products were digested with NheI and BamHI and inserted into plasmid pET11b (Novagen) (Supplementary Table S2). For the construction of the plasmids pET11b-qsrRC29S-His and pET11b-qsrRC32S-His, two first-round PCR products were fused and amplified by a second round of PCR as previously described (Loi et al, 2018b).
For the qsrRC29S mutant, the primer pairs pET-qsrRC29S-for-f2 and pET-qsrR-rev-BamHI as well as pET-qsrRC29S-rev-f1 and pET-qsrR-for-NheI were used. For the qsrRC32S mutant, the primers pET-qsrRC32S-for-f2 and pET-qsrR-rev-BamHI as well as pET-qsrRC32S-rev-f1 and pET-qsrR-for-NheI were applied (Supplementary Table S3). The second-round PCR products were digested and cloned into pET11b as described earlier. For expression and purification of His-tagged QsrR, QsrRC4S, QsrRC29S, and QsrRC32S proteins, E. coli BL21(DE3) plysS strains with plasmids pET11b-qsrR-His, pET11b-qsrRC4S-His, pET11b-qsrRC29S-His, and pET11b-qsrRC32S-His were cultivated in 1.5 L LB medium until the log phase at an OD600 of 0.7, followed by the addition of 1 mM iso-propyl-β-
Recombinant His-tagged proteins were purified using His Trap™ HP Ni-NTA columns and the ÄKTA purifier liquid chromatography (LC) system as described (Loi et al, 2018b).
EMSAs of QsrR and QsrR Cys mutant proteins
For EMSAs, the 210 and 185 bp DNA fragments containing the upstream regions of qsrR and catE2, respectively, were amplified by PCR. The DNA-binding reactions were performed with 15 ng of the promoter region and purified His-tagged QsrR, QsrRC4S, QsrRC29S, and QsrRC32S proteins for 45 min, as previously described (Loi et al, 2018b). MHQ, diamide, HOCl, and allicin were added to the DNA-QsrR-complex for 30 min to observe the dissociation of QsrR from the DNA. To analyze the reversibility of the QsrR inhibition by quinones and oxidants, DTT was added for 30 min as previously described (Loi et al, 2018b).
The percentage of the protein-DNA complex formation was determined according to the band intensities of three to four biological replicates of the EMSAs and quantified using ImageJ 1.52a. The KD values of the QsrR, C4S, C29S, and C32S mutant proteins for the catE2 and qsrR promoter were determined using the Graph prism software version 7.03. The detailed individual data of percentage of DNA-protein complex formation of the QsrR, QsrRC4S, QsrRC29S, and QsrRC32S proteins under control conditions, diamide, and HOCl stress are shown in Supplementary Tables S4–S6, respectively.
Analyses of thiol-oxidation of QsrR and the QsrR Cys mutant proteins in vitro
Purified QsrR and its Cys mutant proteins QsrRC4S, QsrRC29S, and QsrRC32S were first pre-reduced with 10 mM DTT for 15 min. Reduced QsrR proteins were treated with increasing amounts of MHQ, diamide, HOCl, and allicin, followed by alkylation with 50 mM IAM for 30 min in the dark due to the light sensitivity and instability of IAM. The reversible thiol-oxidation of QsrR to intermolecular disulfides was analyzed by separation of the reduced and oxidized QsrR and Cys mutant proteins by non-reducing and reducing SDS-PAGE.
MS for identification of the QsrR disulfides and S-thioallylations
To identify the post-translational thiol-modifications, 5 μg of the QsrR protein was treated with 320 μM diamide or 1 mM allicin for 30 min, alkylated with IAM, and separated by non-reducing SDS-PAGE. Reduced and oxidized QsrR bands were cut and in-gel tryptic digested as previously described (Loi et al, 2018b). The tryptic peptides were dissolved in 30 μL of 0.05% trifluoroacetic acid (TFA) with 5% acetonitrile, diluted 1:30, and 6 μL were analyzed by an Ultimate 3000 reverse-phase capillary nano LC system connected to an Orbitrap Q Exactive HF mass spectrometer (Thermo Fisher Scientific).
Samples were injected and concentrated on a trap column (PepMap100 C18, 3 μm, 100 Å, 75 μm i.d. × 2 cm; Thermo Fisher Scientific) equilibrated with 0.05% TFA in water. After switching the trap column in-line, LC separations were performed on a reverse-phase column (Acclaim PepMap100 C18, 2 μm, 100 Å, 75 μm i.d. × 25 cm; Thermo Fisher Scientific) at an eluent flow rate of 300 nL/min. The mobile phase A contained 0.1% formic acid in water, and mobile phase B contained 0.1% formic acid in 80% acetonitrile/20% water.
The column was pre-equilibrated with 5% mobile phase B, followed by an increase of 5%–44% mobile phase B in 35 min. Mass spectra were acquired in a data-dependent mode utilizing a single MS survey scan (m/z 200–2000) with a resolution of 60,000 in the Orbitrap, and MS/MS scans of the 5 most intense precursor ions with a resolution of 15,000 and a normalized collision energy of 27. All spectra were recorded in the profile mode. The isolation window of the quadrupole was set to 2.0 m/z. The dynamic exclusion time was set to 10 s and automatic gain control was set to 3 × 106 and 1 × 105 for MS and MS/MS scans, respectively.
Data processing and identification of proteins was performed using the Mascot software package (Mascot Server version 2.7, Mascot Distiller version 2.8, Mascot Daemon version 2.7; Matrix Science). During MS/MS processing, the maximum charge state was set to the precursor charge state and singly charged fragment ions were used as an output. Processed spectra were searched against the S. aureus COL QsrR protein sequence with the N-terminal extension (AS) and the C-terminal His6-tag. The pre-existing disulfide bond crosslinking method from Mascot was used, which allows for the identification of possible intramolecular (Cys29-Cys32) and intermolecular (Cys4-Cys29′, Cys4-Cys32′, or Cys4-Cys4′) disulfides in the oxidized QsrR protein.
A maximum of two missed cleavages was allowed and the mass tolerance of precursor and sequence ions was set to 10 ppm and 0.02 Da, respectively. Methionine oxidation (Met +15.994915 Da), methionine dethiomethylation (Met −48.003371), and cysteine carbamidomethylation (Cys +57.021465 Da) were set as variable modifications. A significance threshold of 0.05 was used as a cut-off, and the MS/MS spectra of the identified disulfide peptides were manually checked.
The peptides of the allicin-treated QsrR sample were analyzed by MALDI-TOF-MS using an Ultraflex-II TOF/TOF instrument (Bruker Daltonics, Bremen, Germany) equipped with a 200 Hz solid-state Smart beam™ laser. Alpha-Cyano-4-hydroxycinnamic acid was used as matrix substance, and the mass spectrometer was operated in the positive reflector mode. Mass spectra were acquired over an m/z range of 600–4000. Cysteine S-thioallylations by allicin were identified in the MS1 spectra by the mass shift of 72.00337 Da for C3H5S1 at Cys peptides using the Mascot software.
Western blot analyses
S. aureus COL cells were harvested before and 30 min after treatment with 5 mM diamide as described (Loi et al, 2018b). After washing, protein lysates were prepared in TE-buffer (pH 8.0) with 50 mM IAM using the ribolyzer. Protein lysates were separated using 18% non-reducing SDS-PAGE and subjected to Western blot analysis using anti-His6 monoclonal antibodies (Sigma) as previously described (Loi et al, 2021; Loi et al, 2018a).
Data Availability Statements
All data of this article are available in the figures and Supplementary Figures and Supplementary Tables. Raw data are made available on request to the corresponding author. An electronic laboratory notebook by a specific platform was not used for data collection.
Footnotes
Acknowledgments
For mass spectrometry performed by C.W. and B.K., the authors would like to acknowledge the assistance of the Core Facility BioSupraMol supported by the Deutsche Forschungsgemeinschaft.
Authors' Contributions
V.N.F. and H.A. designed the concept of this project. V.N.F performed the majority of the experiments and analyzed the data. V.V.L. and V.N.F. constructed the plasmids and mutants. B.K. and C.W. performed the mass spectrometry and data analysis. M.G. contributed to allicin synthesis. H.A. and V.N.F. wrote the article. H.A. provided funding and supervised the project. All authors have read, edited, and approved the final article.
Author Disclosure Statement
No competing financial interests exist.
Funding Information
This work was supported by an ERC Consolidator grant (GA 615585) MYCOTHIOLOME and grants from the Deutsche Forschungsgemeinschaft (AN746/4-1 and AN746/4-2) within the SPP1710 on “Thiol-based Redox switches,” by the SFB973 project C08 and TR84 project B06 to Haike Antelmann.
Supplementary Material
Supplementary Table S1
Supplementary Table S2
Supplementary Table S3
Supplementary Table S4
Supplementary Table S5
Supplementary Table S6
Supplementary Figure S1
Supplementary Figure S2
Supplementary Figure S3
Supplementary Figure S4
Supplementary Figure S5
Supplementary Figure S6
Supplementary Figure S7
Supplementary Figure S8
Supplementary Figure S9
Supplementary Figure S10
Supplementary Figure S11
Supplementary Figure S12
Supplementary Figure S13
Supplementary Figure S14
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
