Abstract
Human lungs single-cell RNA sequencing data from healthy donors (elderly and young; GEO accession no. GSE122960) were analyzed to isolate and specifically study gene expression in alveolar type II cells. Colocalization of angiotensin-converting enzyme 2 (ACE2) and TMPRSS2 enables severe acute respiratory syndrome coronavirus 2 (SARS-CoV 2) to enter the cells. Expression levels of these genes in the alveolar type II cells of elderly and young patients were comparable and, therefore, do not seem to be responsible for worse outcomes observed in coronavirus disease 2019 (COVID-19) affected elderly. In cells from the elderly, 263 genes were downregulated and 95 upregulated. Superoxide dismutase 3 (SOD3) was identified as the top-ranked gene that was most downregulated in the elderly. Other redox-active genes that were also downregulated in cells from the elderly included activating transcription factor 4 (ATF4) and metallothionein 2A (M2TA). ATF4 is an endoplasmic reticulum stress sensor that defends lungs via induction of heme oxygenase 1. The study of downstream factors known to be induced by ATF4, according to Ingenuity Pathway Analysis™, identified 24 candidates. Twenty-one of these were significantly downregulated in the cells from the elderly. These downregulated candidates were subjected to enrichment using the Reactome Database identifying that in the elderly, the ability to respond to heme deficiency and the ATF4-dependent ability to respond to endoplasmic reticulum stress is significantly compromised. SOD3-based therapeutic strategies have provided beneficial results in treating lung disorders including fibrosis. The findings of this study propose the hypotheses that lung-specific delivery of SOD3/ATF4-related antioxidants will work in synergy with promising antiviral drugs such as remdesivir to further improve COVID-19 outcomes in the elderly.
Innovation
For the first time, single-cell RNA sequencing data of the human lungs have been studied as a function of age to reveal that age-related weakening of specific components of the antioxidant defense system of the alveolar type II cells should be tested for a mechanistic connection of COVID-19 severity outcomes.
Introduction
T
Reactive oxygen species (ROS) and reactive nitrogen species (RNS) are known to be produced by cells of the innate immune system and others in response to viral infection (11). ROS/RNS are directly implicated in lung fibrosis and declining lung function (5). Antioxidant enzymes such lecithinized superoxide dismutase (SOD) have proven to be useful in patients suffering from lung fibrosis (25). There is substantial literature demonstrating a causative role of ROS/RNS in the development of lung fibrosis (21). In virus-induced lung disease, antioxidant treatment attenuated lung inflammation and airway hyper-reactivity (3). In this study, we utilized single-cell sequencing data from elderly and younger humans to specifically study alveolar type II cells to identify genes that are most profoundly affected as a function of age. Such effort was intended at developing novel hypotheses to understand why lungs of the elderly are more severely affected in COVID-19.
Results
Human lungs single-cell RNA sequencing (scRNA-seq) data from four healthy donors (9783 cells from old-age group, 57 and 63 years old, and 8501 cells from young-age group, 22 and 29 years old; GEO accession no. GSE122960) were analyzed (20). Analysis of scRNA-seq data generated 14 clusters visualized using t-distributed stochastic neighbor embedding (t-SNE). Such clustering was based on a nonlinear dimensionality reduction technique for embedding high-dimensional data with the objective of visualization in low-dimensional space. As depicted in Figure 1A, these clusters include epithelial cells (alveolar type II cells), macrophages in different states, epithelial cells (alveolar type I cells), monocytes, B cells, T cells/NK cells, endothelial cells, and stem cells. SingleR package in R (1), in conjunction with the Human Primary Cell Atlas (HPCA) data set, was used to identify alveolar type II cells from the total lung cell population. These cells heavily expressed the characteristic surfactant protein C gene (Supplementary Fig. 1A, B). Genes of this particular alveolar type II cell cluster were analyzed for differential expression as a function of aging (Fig. 1B). Colocalization of ACE2 and TMPRSS2 enables SARS-CoV 2 to enter the cells (8). Expression levels of these genes in the alveolar type II cells of old and young patients were comparable (Fig. 1C) and, therefore, do not seem to be responsible for worse outcomes in COVID-19-affected elderly.
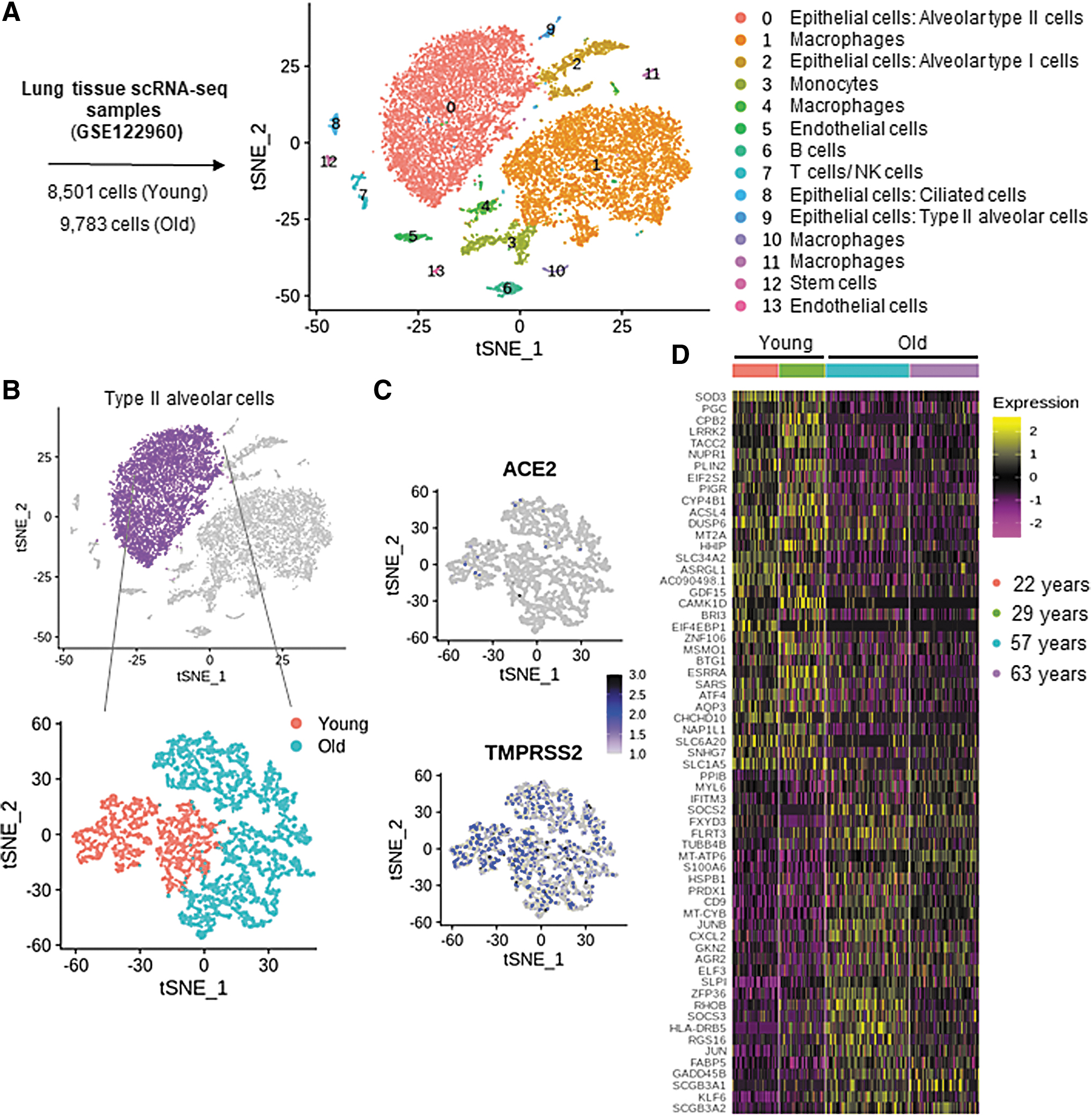
Compared with that in alveolar type II cells from younger donors, in cells of older donors, 263 genes were differentially downregulated and 95 genes were upregulated (adjusted p-value <0.05 and 10% log fold change; Fig. 1D, Supplementary Table S1). SOD3 was identified as the top-ranked (by log of fold change) gene that was most downregulated in alveolar type II cells (Supplementary Table S1). This observation piqued our interest in other genes with known redox functions. Other genes with known redox-based functions that were also downregulated in alveolar type II cells of elderly lung donors included activating transcription factor 4 (ATF4) and metallothionein 2A (M2TA) (Fig. 2A–C). Additional studies to look for SOD3-interacting genes, as predicted by String database (String version 11.0), recognized the following: SOD2, catalase (CAT), glutathione peroxidase 1 (GPX1), GPX2, GPX3, GPX5, GPX7, GPX8, antioxidant 1 copper chaperone (ATOX1), and ATPase copper transporting alpha (ATP7A) (Supplementary Fig. 2A, B). Among these candidates, GPX1 was the only gene that was differentially low in expression in cells from elderly donors (Supplementary Fig. S2B, Supplementary Table S1). Viral infection is known to employ endoplasmic reticulum (ER) stress to cause lung fibrosis (12). ATF4, an ER stress sensor that can defend lungs via induction of heme oxygenase 1, was downregulated in alveolar type II cells of the elderly (Fig. 2B). Study of downstream factors that are known to be induced by ATF4, according to Ingenuity Pathway Analysis™ (IPA), identified 24 candidates. Twenty-one of these 24 were significantly downregulated in the alveolar type II cells of the elderly (Fig. 2C). The downregulated candidates were subjected to pathway enrichment using the Reactome Database. These analyses identified that in the elderly, the ability to respond to heme deficiency and the ATF4-dependent ability to respond to endoplasmic reticulum stress are significantly compromised (Fig. 2D, Table 1).
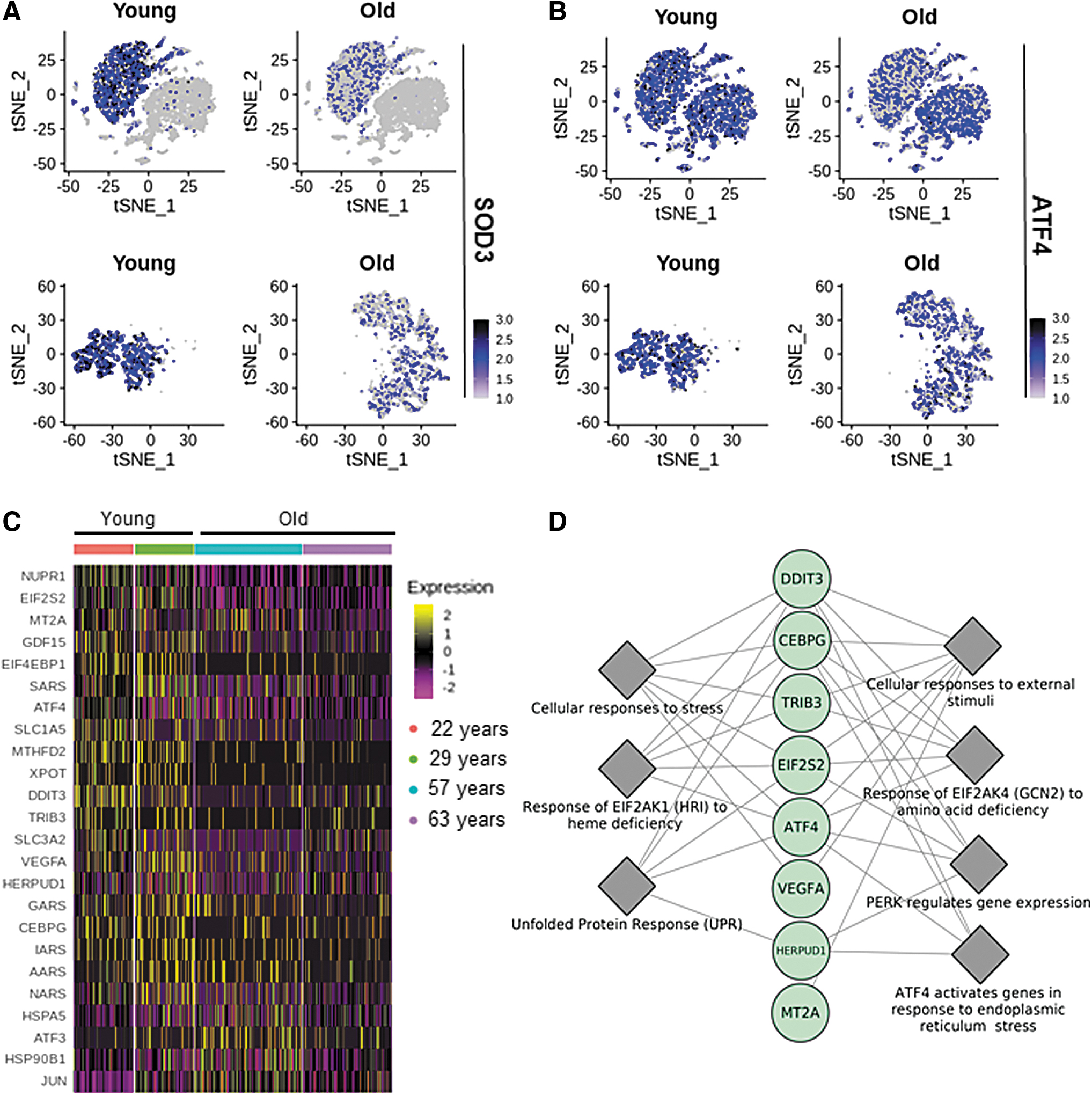
Reactome Pathways Enrichment for the Downregulated Genes Among ATF4 Downstream Targets in Alveolar Type II Cells from Elderly
Pathways with adjusted p-value <2E-6 are shown.
EIF2AK1, eukaryotic translation initiation factor 2 alpha kinase 4; FDR, false discovery rate; GCN2, general control nonderepressible 2; HRI, heme-regulated inhibitor.
Discussion
At the time of this reporting, of 13,130 COVID-19 deaths (of reported death from all causes 582,565 on April 19, 2020) in the United States as reported by the Center for Disease Control, 13,001 decedents were of age 35 years or above, representing 99% of all COVID-19 deaths. Decedents aged 55 years or above account for 91% of all COVID-19 deaths (19). The younger donors of this study were both <30 years, whereas the elderly donors were both >55 years (19). In brief, the SARS-CoV 2 virion contains four proteins: spike, envelope, membrane, and nucleocapsid, and a single-stranded RNA. The virion binds to ACE2 receptor located on alveolar type II cells making these cells a target for infection. Once SARS-CoV 2 has attached to these receptors, the TMPRSS2 protease cleaves the spike protein to expose a fusion peptide that helps the virus enter the cell (8). This enables the virions to release their RNA into infected cells. Coexpression of ACE2 and TMPRSS2 on type II cells makes them a preferred site for SARS-CoV 2 attachment, entry, and replication (16). In late stages of the severe form of COVID-19, cytokine storm has been evident (17). Under these conditions, when the inflammatory system has gone awry in COVID-19–unrelated pathologies, there is ample evidence of oxidative stress (15). Findings of this study, involving an unbiased query of differentially expressed genes specifically in alveolar type II cells of the human lung, point toward specific elements of the antioxidant defense system that are weakened as a function of age. SOD3-deficient mice develop severe lung damage in the presence of normal oxygen tension along with marked inflammatory cell infiltration and alveolar hemorrhage (6). In the lungs, extracellular SOD3 is primarily expressed in bronchial and alveolar type II epithelial cells, alveolar macrophages, and pulmonary endothelial cells (4). Importantly, SOD3 mRNA expression in alveolar cells correlates with locally secreted enzyme activity, defending functional significance of observed changes in gene expression (27). With SOD3 as the top-ranked candidate, this study identifies specific component of the alveolar cell antioxidant defense systems that are weakened in response to aging. Evidence in the current literature shows that SOD administration can decrease the severity of respiratory illness (7). Intravenous SOD administration in rabbit models reversed allergic emphysema (2). SOD-based therapies have also shown encouraging results in managing infectious diseases by improving host immune responses. Melon SOD restored CD4+/CD8+ ratio in cats infected with feline immunodeficiency virus (28). Cu/Zn-SOD significantly inhibited HIV replication (13). Cu/Zn-SOD inhalation protected murine lungs against pulmonary emphysema by decreasing ROS levels and proinflammatory cytokines expression (26). A limited-scale prospective double-blinded controlled trial NCT04323228 is about to start recruiting to test the effect of an antioxidant oral nutrition supplement in SARS-CoV 2 positive cases. All of these mentioned studies are preliminary at best and may help lay the foundation for a more serious effort testing whether lung-specific delivery of SOD3-related antioxidants may work in synergy with promising antivirals such as remdesivir (NCT04323761) to further improve COVID-19 outcomes in the elderly.
Notes
Data acquisition
Primary single-cell RNA sequencing data were obtained from the GEO database (accession no. GSE122960). Authors of the published study performed single-cell RNA sequencing on eight healthy donor lungs and nine lungs from patients with pulmonary fibrosis (20). Four samples were chosen from the healthy donor lungs in which two of them were 57 and 63 years old and the other two samples were from donors who were 22 and 29 years old. All four donors were females, nonsmokers, and African American.
Processing of raw data and quality control
All analyses were performed using Seurat package (v.3.1.1) in R (v.3.3.5) (22). The initial data set contained 20,163 cells from four lung samples (two young and two elderly). Gene expression values were log normalized using 10,000 transcripts per cell as scaling factor. Canonical correlational analysis was performed to integrate all four samples to identify the shared cell types using the top 2000 highly variable genes. Cells expressing <200 or >5000 genes, as detected, were excluded. Further filtering was performed to exclude cells that contain >15% of their reads from mitochondrial encoded genes. Cells with total number of counts between 2000 and 25,000 were kept for downstream analysis. After the filtration step, 18,284 cells (9783 from elderly and 8501 from young donors) were maintained for downstream analysis. Principal component analyses were performed and the first 15 principal components were chosen for clustering.
Determination of cell type identity
To identify cluster identity, SingleR package in R was used to calculate the similarity between each cluster and the HPCA data set (1). In addition, differential gene expression was performed between each cluster and the rest of the cells to identify cluster markers and further identification of the cluster identity using known markers from the literature.
Differential gene expression analysis
Differential gene expression analysis was performed for alveolar type II cells (cluster zero) to compare between young- and old-age groups using Wilcox-Rank Sum test with adjusted p-value <0.05. To avoid detection of genes only altered in one sample of either group, four additional comparisons were performed (each sample with the two samples from the other group). The genes that were not simultaneously upregulated or downregulated and in at least three such comparisons were excluded.
Ingenuity upstream regulator analysis in IPA
IPA (23,24,29) was used to identify the cascade of transcriptional regulators that can explain the observed gene expression changes in the data set. This approach is based on prior knowledge of expected effects between transcriptional regulators and their target genes. Two statistical measures (an overlap p-value and an activation z-score) were computed for each potential transcriptional regulator. The activation z-score was used to infer likely activation states of regulators based on comparison with a model that assigns random regulation directions.
Pathways enrichment
Reactome database (10) was used for enrichment of ATF4 downstream targets that were found to be downregulated in alveolar type II cells in the cells from the elderly.
Footnotes
Authors' Contributions
A.S.A. and K.S. performed data analysis. A.S.A., K.S., H.M.E.A., and C.K.S wrote the article.
Funding Information
This work was supported in part by National Institutes of Health grants GM108014, NS042617 and NR015676.
Supplementary Material
Supplementary Table S1
Supplementary Figure S1
Supplementary Figure S2
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
