Abstract
Significance:
Cytochrome bd is a ubiquinol:oxygen oxidoreductase of many prokaryotic respiratory chains with a unique structure and functional characteristics. Its primary role is to couple the reduction of molecular oxygen, even at submicromolar concentrations, to water with the generation of a proton motive force used for adenosine triphosphate production. Cytochrome bd is found in many bacterial pathogens and, surprisingly, in bacteria formally denoted as anaerobes. It endows bacteria with resistance to various stressors and is a potential drug target.
Recent Advances:
We summarize recent advances in the biochemistry, structure, and physiological functions of cytochrome bd in the light of exciting new three-dimensional structures of the oxidase. The newly discovered roles of cytochrome bd in contributing to bacterial protection against hydrogen peroxide, nitric oxide, peroxynitrite, and hydrogen sulfide are assessed.
Critical Issues:
Fundamental questions remain regarding the precise delineation of electron flow within this multihaem oxidase and how the extraordinarily high affinity for oxygen is accomplished, while endowing bacteria with resistance to other small ligands.
Future Directions:
It is clear that cytochrome bd is unique in its ability to confer resistance to toxic small molecules, a property that is significant for understanding the propensity of pathogens to possess this oxidase. Since cytochrome bd is a uniquely bacterial enzyme, future research should focus on harnessing fundamental knowledge of its structure and function to the development of novel and effective antibacterial agents.
Table of Contents
Introduction
Distribution, Evolution, and Regulation of Gene Expression Distribution Phylogeny CydX and CydH subunits CydX CydH Regulation of expression
Structure and Assembly Structure Assembly—the role of CydDC The cydDC genes The CydDC proteins: structure and function Structural investigations into the CydDC complex Physiological impacts of CydDC function Conclusions
Spectral and Redox Properties
Catalytic Cycle
Physiological Functions Respiratory protection of nitrogenase An oxygen-reactive oxidase in anaerobes? Environmental stressors and their relationships with cytochrome bd
Peroxide NO and ONOO−
Sulfide Chromate
Antibiotics and Antimicrobial Agents Cationic amphiphilic peptides with antimicrobial activities Gramicidin S Cathelicidin LL-37 Microcin J25 Other antimicrobial compounds Bedaquiline Clofazimine Isoniazid Cytochrome bd protection of mycobacteria from compounds targeting cytochrome bcc
Cytochrome bd As a Potential Therapeutic Target
Possible Biotechnological Applications
Conclusions
I. Introduction
Cytochrome bd, first described in 1928 [(345), see also Cook and Poole (75) and references therein], is a respiratory terminal oxidase thus far uniquely identified in the electron transport chain of prokaryotic organisms (36, 48, 151, 163, 252). As it promotes virulence in a number of pathogenic bacteria, it is currently recognized as a prospective drug target for the development of new antibacterial drugs.
Phylogenetically unrelated to the more extensively investigated haem/copper oxidases, such as mitochondrial cytochrome aa 3 (53, 58, 59, 243, 274, 301, 336), cytochrome bd catalyzes the reduction of O2 to H2O with remarkably high affinity, using quinols as physiological reducing substrates (55, 163). The reaction, although not associated with a proton pumping activity (267), is electrogenic (19, 20, 38, 51, 155, 267) and thus contributes to generating a proton motive force (PMF) and sustaining bacterial energy conservation.
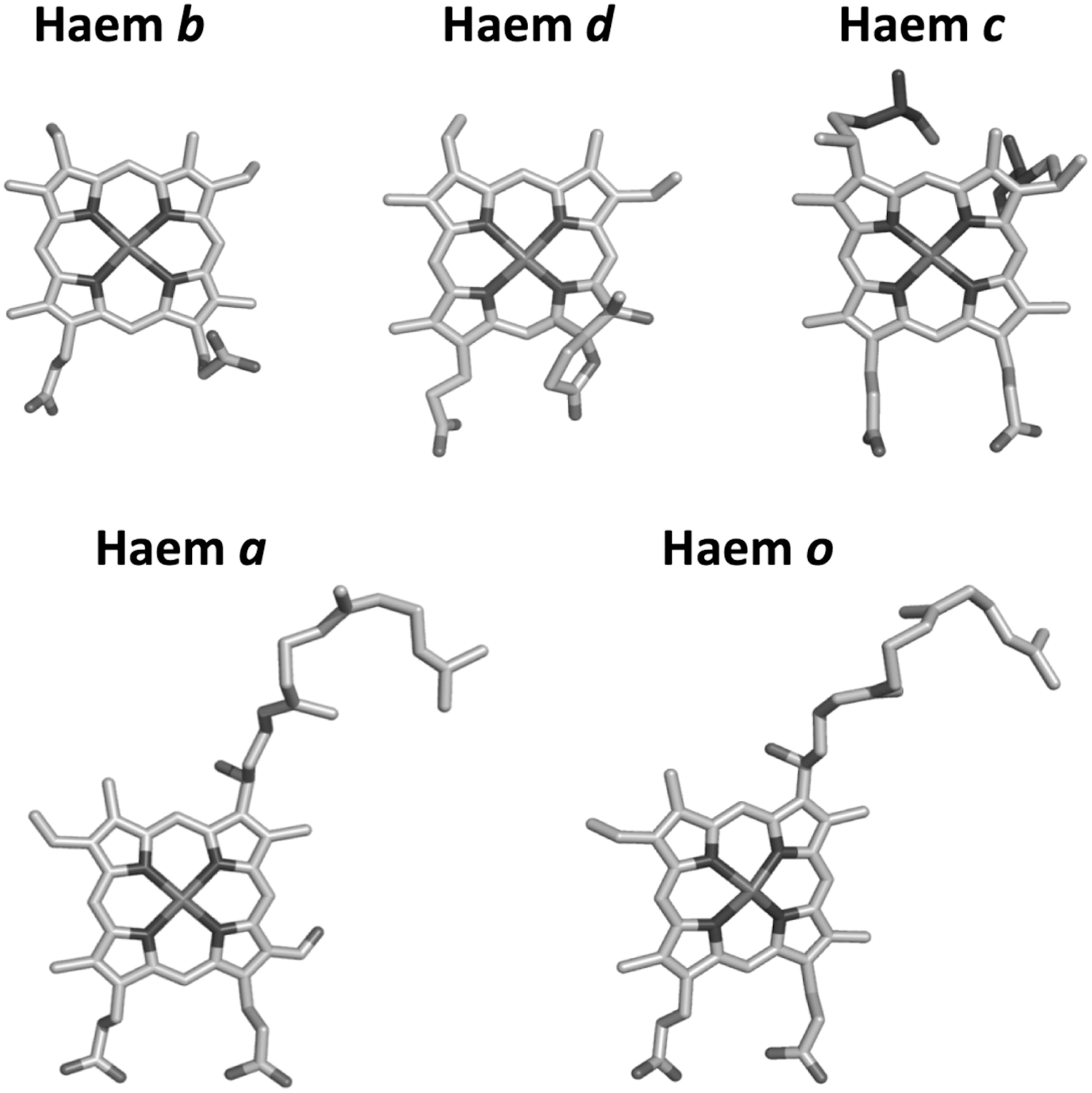
Cytochrome bd displays distinct biochemical features with respect to haem/copper oxidases, including a d-type O2-reactive haem and no copper atoms, which, on the contrary, are invariantly present in haem/copper oxidases (see Fig. 1 for haem types). The three-dimensional structure of cytochrome bd has remained a mystery for a long time, but it was recently solved for two members of this protein family, the enzymes from Geobacillus thermodenitrificans (281) and, more recently, from Escherichia coli (280, 317), revealing unexpected structural differences between the two proteins, most likely of evolutionary significance.
A large body of evidence suggests that cytochrome bd enables bacterial O2 consumption, either for bioenergetic purposes, to drive specific metabolic pathways and/or to afford protection from O2 toxicity and a variety of stress conditions, including hypoxia, medium alkalinization, high temperature, and exposure to toxic compounds, such as uncouplers, antibiotics, or classical respiratory inhibitors, for example, nitric oxide (NO), cyanide, and hydrogen sulfide (H2S) [see Borisov et al. (46), Borisov et al. (48), Forte et al. (115), Giuffre et al. (124), Giuffre et al. (125), Korshunov et al. (192), Poole and Cook (252) and references therein].
Furthermore, cytochrome bd was shown to facilitate degradation of harmful reactive oxygen and nitrogen species (ROS and RNS), such as hydrogen peroxide (H2O2) or peroxynitrite (ONOO−) [reviewed in Forte et al. (117)], produced by the host immune system to control microbial infections.
The growing attention that is being paid by the scientific community toward bd-type bacterial terminal oxidases as potential drug targets prompted us to present here a comprehensive update on the information currently available about these enzymes and their impact on bacterial physiology.
A prelude on nomenclature is in order. The accepted name for the subject of this review is “quinol oxidase (electrogenic, proton-motive force generating), EC 7.1.1.7′, product of the cydAB (and other) genes.” It is generally called cytochrome bd and, less logically, “cytochrome bd oxidase.” The latter is unfortunate because the complex does not oxidize cytochrome bd but is cytochrome bd! However, there have been more uncertainty and debate on the correct nomenclature of the other major oxidase in E. coli and many other bacteria, to which we refer, namely cytochrome bo 3, bo oxidase, or bo′. The last name was proposed (251) because some years ago the IUB Enzyme Nomenclature “rules” proposed that ligand-binding haems should be designated with the suffix ′. Hence aa′ (for cytochrome aa 3) and bo′ appeared correct.
The term cytochrome bo 3, emphasising the correlation with aa 3 did not, and still does not, appear logical, because there is, and never was, a bo 1 or bo 2, whereas the names a 1 and a 2 were both once common currency (for cytochromes b 595 and d, respectively). However, bo 3 has been so widely adopted to describe EC 7.1.1.3 ubiquinol oxidase (H+-transporting), product of cyoABCD in E. coli (219), that we have adopted it here.
II. Distribution, Evolution, and Regulation of Gene Expression
A. Distribution
Cytochrome bd is a membrane-integrated complex comprising two to four subunits (see later, sections II.B and II.C). The two largest and omnipresent subunits are subunits I and II, which are encoded by the chromosomal cydA and cydB genes. In E. coli, two different cytochrome bd-type oxidases have been described, the best studied cytochrome bd-I and the more recently discovered cytochrome bd-II (93). Whereas bd-I is encoded by the cydAB operon, bd-II is encoded by the cyxAB operon (55). The bd-family of oxygen reductases has a very broad taxonomic distribution with orthologs found in various bacterial phyla. These include diverse groups of Eubacteria, from gram-positive Firmicutes and Actinomycetes to the whole phylum of Proteobacteria.
A number of Archaea also encode bd-family homologues, with members of the family found in Crenarchaeota (Thermoproeus tenax, Pyrobaculum neutrophilum, Vulcanisaeta moutnovskia), Euryarchaeota (Methanosarcina barkeri, Archaeoglobus sulfaticallidus) (57), and Korarchaeota (Korarchaeum cryptofilum) (96). Cytochrome bd-type oxygen reductases are very common in some phyla, such as the Proteobacteria and Actinobacteria, and sporadically distributed in others. Intriguingly, homologues of cytochrome bd have been detected in many species described as strict anaerobes such as Methanosarcina barkeri, Methanosarcina acetivorans (57), Bacteroides fragilis (13), Desulfovibrio gigas (202), Desulfovibrio vulgaris Hildenborough (288), Geobacter metallireducens (141), Moorella thermoacetica (92), and Chlorobaculum tepidum (204).
A more recent survey of cytochrome bd sequences highlighted the diversity of molecular forms of cytochrome bd (96). Although no clear pattern could be discerned in the distribution of the two types of cytochrome bd among the classes of Alpha-, Beta-, and Gammaproteobacteria, Epsilonproteobacteria possess only the bd-I type. In contrast, the majority of Deltaproteobacteria has short CydA and CydB proteins that are evidently related to the catalytic subunits of ancestral cytochromes bd of Bacillus subtilis (337).
In addition, a phylogenetically isolated group of Deltaproteobacteria such as Desulfobulbus propionicus contains cydA sequences that are longer than those of the rest of the class, due to Q loop extensions similar to those of bd-I type oxidases. Furthermore, the genomes of over 50 Deltaproteobacteria contain atypical chimeric forms of cydA that are fused with genes encoding multiple c-type cytochromes but not associated with a cydB gene.
The recent discovery of the small subunit CydX (68, 102, 144, 145, 314, 324) provided the opportunity for multiple taxonomic coverage studies. Using the previously characterized E. coli CydX protein as a query sequence, Allen et al. conducted a survey for orthologs using multiple homology-based in silico tools in completed genomes from 1095 taxa spanning the major Eubacterial divisions (111). Their comprehensive approach resulted in the identification of over 300 CydX homologues that are restricted to the phylum of Proteobacteria and more specifically to its Alpha, Beta, and Gamma classes (3). Only two orthologs were identified using refined methods in the species Leptospirillum ferrooxidans C2–3 and Leptospirillum ferriphilum ML-04, both members of the Nitrospiraceae family in the phylum Nitrospirae. A more recent analysis (96) identified a class of remote orthologs of CydX, termed CydX-like proteins. Their function remains currently unknown.
B. Phylogeny
Previous studies suggested that the bd-family of oxygen reductases is an ancient innovation, already present in the ancestor of both Bacteria and Archaea (63). However, it has been reported that the family may have originated in Bacteria and was later acquired by Archaea via lateral gene transfer (57, 135). Phylogenetic reconstructions of the bd-family showed that lateral gene transfer plays a substantial role in the distribution of the family, with many phyla acquiring cytochrome bd genes multiple times independently. A representative phylogenetic tree is shown in Figure 2. Sequence analysis has demonstrated that CydA and CydB have different rates of evolution, with CydB evolving 1.2 times faster than CydA (135). The biological relevance of this asymmetrical evolution remains essentially unknown.
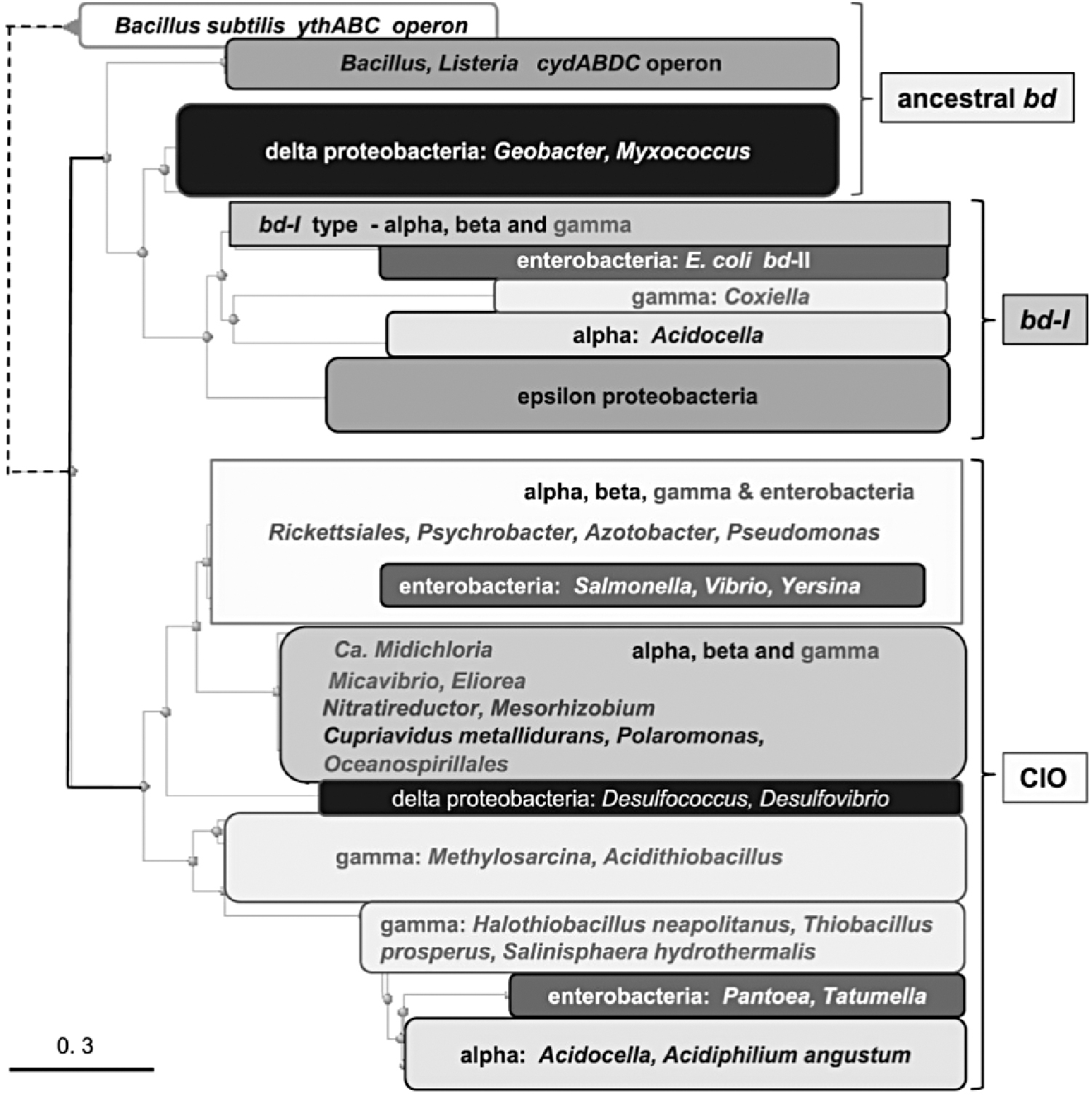
More recently, Degli Esposti et al. specifically studied proteobacterial cytochromes bd using integrated approaches of genomic and protein analysis (96). Their work generated a molecular classification of diverse types of cytochrome bd, allowing reinterpretation of their evolution and substantiating the occurrence of multiple lateral gene transfer events. Specifically, their findings provided insights on basal taxa of Alphaproteobacteria from which the Gammaproteobacterial lineage probably emerged. A duplication of the original gene cluster of a cytochrome bd might have occurred in the ancestors of extant Alphaproteobacteria of the Rhodospirillales order, such as Acidocella, from which the bd-I type of oxidase might have diffused to other proteobacterial lineages.
This conclusion derives from the robust clustering of the Gammaproteobacterium Salinisphaera hydrothermalis with Acidocella sp. MX-AZ02 and Acidocella facilis. As Acidocella lives in strongly acidic environments of wetlands and lakes, a transfer of the cytochrome bd operon from Acidocella to the halophilic Salinisphaera must have occurred in ancient evolutionary times, when both organisms shared common marine environments, or ancestors. The cyanide-insensitive oxidase type, on the contrary, may have differentiated into recognizable subtypes after another gene cluster duplication. These subtypes are widespread in the genomes of Alpha-, Beta-, and Gammaproteobacteria, with occasional instances of lateral gene transfer.
The small size of CydX proteins makes it difficult to robustly resolve phylogenetic inferences and requires critical interpretation. Similar challenges have been encountered with relatively small globin proteins, particularly in invertebrates (147) and simpler species (325). The comparative bioinformatic approaches by Allen et al. (3) to examine the distribution of the CydX protein across bacterial species resulted in a predominant association with the Proteobacter phylum and identified two orthologs in the Nitrospiraceae family. Given the phylogenetic distance between these Leptospirillum species and the other CydX-containing species, of which all are contained in the Proteobacter phylum, it is likely that these bacteria gained the cydABX operon through horizontal gene transfer.
Furthermore, in contrast to the CydA and CydB genes, with orthologs in a broad range of phyla, the CydX-containing species were found to be members of the Alpha, Beta, and Gamma classes of the Proteobacteria. This distribution difference is consistent with a model in which CydA and CydB evolved earlier than the CydX small subunit. Since the Alpha, Beta, and Gamma classes have been suggested to be diverged after the earlier branching of Delta- and Epsilonproteobacteria (70), the distribution of CydX suggests that it may have evolved in association with the cydAB operon in a progenitor of the Alpha-, Beta-, and Gammaproteobacteria clades (3).
Intriguingly, the early diverging group, lacking CydX orthologs, contains anaerobic organisms, whereas the other includes predominantly aerobic and facultative anaerobic organisms. Additional evidence for horizontal gene transfer of CydX genes was found by superimposing a phylogenetic tree based on CydX on a reference phylogenetic tree of all investigated taxa, suggesting that Rhodospirillum photometricum DSM 122 gained its cydABX operon through horizontal gene transfer (3). It is very likely that the cydABX operon has been transferred numerous times between closely related and divergent bacterial species.
C. CydX and CydH subunits
1. CydX
Until recently, it was generally accepted that cytochrome bd consists of two subunits. These are CydA (∼52–57 kDa) and CydB (∼40–43 kDa) encoded by cydA and cydB genes, respectively (131, 183, 222, 283). CydA carries the quinol oxidation site and all haems, b 558, b 595, and d (see section III). However, recent studies (68, 102, 144, 145, 314, 324) showed that in Proteobacteria, including E. coli, Brucella abortus, Shewanella oneidensis, and Salmonella enterica serovar Typhimurium, there is a short gene, cydX, located at the 3′-end of the cydAB operons that encodes a small protein essential for the function of the oxidase (Fig. 3A).
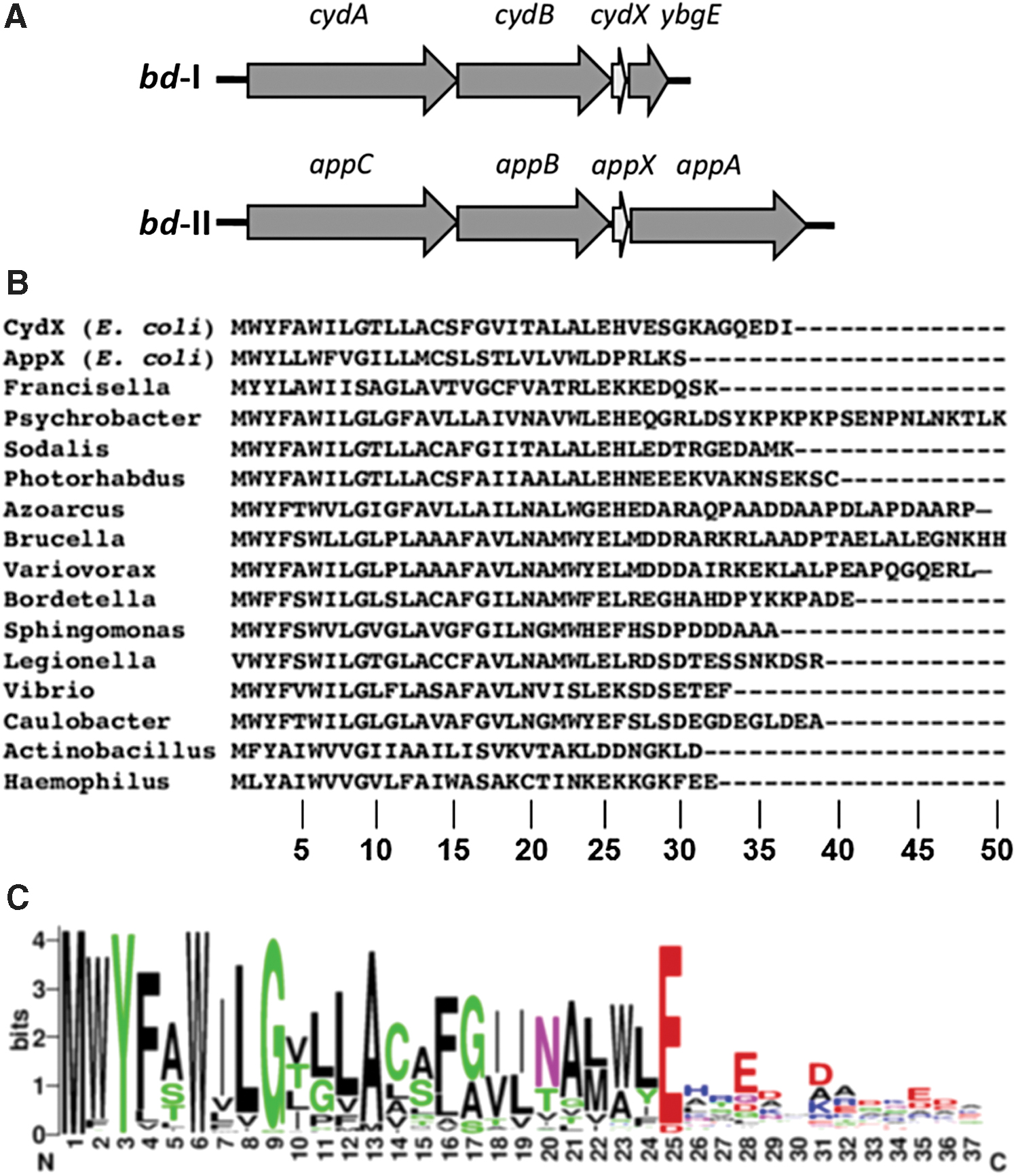
The CydX protein appeared to be a third subunit of cytochrome bd. Deletion of cydX in B. abortus leads to impaired intracellular growth, loss of viability in stationary phase, increased sensitivity to H2O2, and to the combination of the respiratory chain inhibitor sodium azide and the uncoupling agent nickel sulfate (314). Accordingly, ΔcydX E. coli mutants also reveal phenotypes associated with reduced cytochrome bd-I activity (324). Phenotypes of ΔcydX E. coli cells show slow growth in liquid culture, mixed-colony formation, and sensitivity to β-mercaptoethanol. Membrane extracts from ΔcydX E. coli cells have reduced N,N,N′,N′-tetramethyl-p-phenylenediamine oxidase activity. Interestingly, overexpression of appX, paralog of cydX, compensates the cydX deletion in E. coli (324). Upon isolation of E. coli cytochrome bd-I, CydX copurifies with CydAB (145).
The lack of CydX correlates with the absence of the high-spin haems b 595 and d, while the low-spin haem b 558 is retained (Fig. 4). The loss of the high-spin haems results in the loss of enzymatic activity. Hoeser et al. (145) suggested that CydX is essential for the assembly and/or the stability of the high-spin haem site. In contrast to CydX in E. coli, its counterpart in S. oneidensis does not seem to be essential to the oxidase activity (68). Although markedly impaired in function, CydX-lacking cytochrome bd from S. oneidensis can still confer a high level of resistance to nitrite.
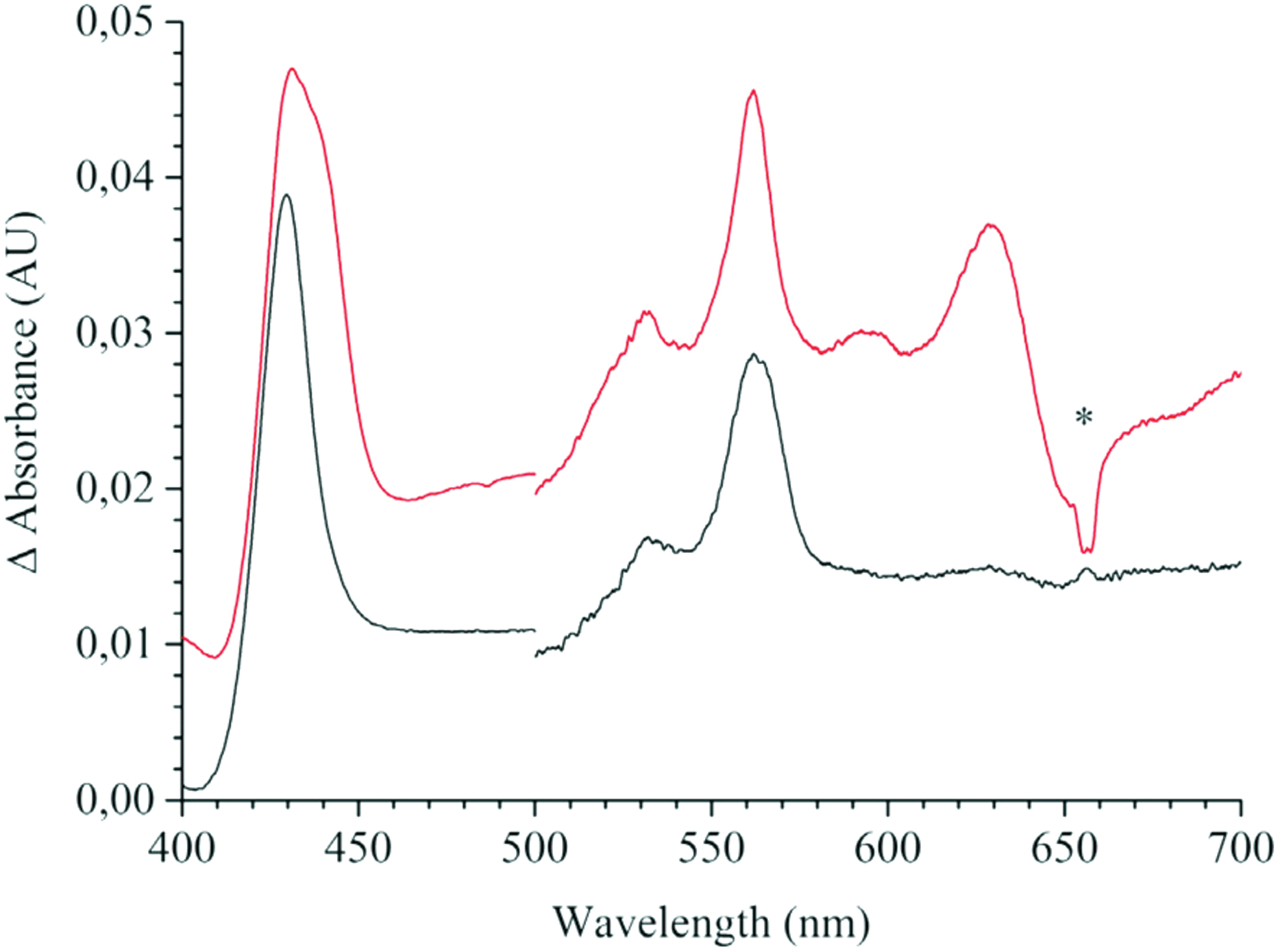
This suggests that a functional protein complex can be assembled by the two large subunits only. Chen et al. (68) concluded that complexation of CydA and CydB is independent of CydX. Furthermore, CydX does not rely on the CydA-CydB complex for its translocation and integration into the membrane. Nonetheless, as in E. coli, CydX in S. oneidensis is apparently critical to positioning and stabilization of the haems, especially haem d.
CydX is conserved in over 200 species of Proteobacteria (3, 96, 144) (Fig. 3B) (see also sections II.A and II.B). There are cydAB operons that encode CydX-like proteins denoted CydY and CydZ, possibly performing similar functions (3). CydX is a 30- to 50-amino-acid protein containing a single hydrophobic α-helix. The protein has the conserved N-terminal region and the less conserved C-terminal region (3, 68, 144) (Fig. 3C). The N-terminal region contains the completely conserved W6, and the highly conserved Y3 (all but one homologue), G9 (all but seven homologues), and E/D25 (all but one homologue). The analysis predicts that Y3, W6, and G9 are part of the transmembrane α-helix (3). The N-terminal region of CydX is probably important for function (68). Interaction of CydX with the cytochrome complex may be coordinated through a combination of interactions between multiple residues rather than being dependent on individual residues (144).
A third subunit, denoted CydS, was identified in the structure of cytochrome bd from G. thermodenitrificans K1041 and is thought to stabilize haem b 558 (281) (see section III). CydS is conserved in Bacillales but shows no sequence similarity to its proteobacterial counterparts. Surprisingly, although the solved structure of the E. coli cytochrome bd-I shows the similar location of CydX (280), the CydX-lacking bd-I protein still retains haem b 558 (145) (Fig. 4).
In summary, it seems likely that all cytochromes bd possess a third, small subunit. The CydX or CydX-like component could be overlooked in cytochrome bd of some microbes using standard biochemical and genomic methods. It is indeed difficult to identify such a small-size protein in cytochrome bd complex preparations, as well as the corresponding short gene out of many short open reading frames. One more possible reason for the apparent absence of cydX in some cydAB operons is that the gene could have migrated to another part of the genome. For instance, 4 of 20 CydS in Bacillales are encoded at gene loci far apart from their cydAB operons (281).
2. CydH
The structure of the E. coli cytochrome bd-I reveals a fourth, previously unknown subunit called CydH (280) [or CydY (317)]. Like CydX, CydH is a small noncatalytic accessory single transmembrane subunit (see section III). The subunit is encoded by the orphan gene ynhF, which is not part of the cydAB operon. Only members of the proteobacterial clade with cytochromes bd belonging to subfamily L (one of the two subfamilies of the bd enzymes, see section III) seem to have CydH homologues (280). The role of CydH appears to be more than just “structural.” The subunit blocks the O2 entry route to haem b 595 in the E. coli cytochrome bd-I (317). In the G. thermodenitrificans enzyme, CydH is absent, and therefore, this channel is open and provides O2 access to haem d located in place of haem b 595 at this site (281) (see also section III).
D. Regulation of expression
Regulatory mechanisms underpinning the expression of cytochrome bd have long been the subject of intense study, driven by the extraordinary features of this oxidase and its physiological significance. For example, the earliest studies on bacterial respiration revealed that cytochrome bd, recognizable initially by its distinctive optical properties, was maximally expressed in E. coli under conditions of low oxygen supply (252, 272). This is now considered to be a reflection of the very high oxygen affinity of this oxidase (88). Expression is also increased in the presence of cyanide (8), NO (217), carbon monoxide (CO) (332), and Ru- and Mn-containing CO-releasing molecules (CORMs) (94, 333).
By far, the best-studied systems are in E. coli, where regulation has proven to be extremely complex, intricate, and responsive. Although we focus on the mechanisms in E. coli, which involve multiple promoters and at least three classes of transcriptional regulators, the identity of the regulators and their modes of action differ among genera, but are outside the scope of the present review. For example, in Shewanella, the two cytochrome bd subunit genes are cotranscribed with a third gene, cydE, that encodes a GbsR regulator that represses cydAB expression (340).
In several gram-positive genera, the transcription factor Rex is implicated in the regulation of genes important for fermentation and growth at low oxygen tensions, sensing the cellular redox poise in the form of NADH/NAD+ ratios. Examples of the role of Rex in regulating cytochrome bd expression are found in Streptomyces (56, 207), Saccharopolyspora spinosa (348), and Bacillus subtilis (200, 266, 291, 331).
In Rickettsia conorii, a small regulatory RNA is implicated as a potential regulator of cydA (234). OxyR is a master regulator in a wide range of bacteria, but its role in regulating cytochrome bd appears limited: an OxyR binding site has been identified in the promoter region of cydA in Corynebacterium glutamicum (225, 316) and it plays a role in regulating electron flux to cytochrome bd in S. oneidensis (330).
Here we focus on the complex systems involved in regulating cytochrome bd-I of E. coli. Control of cytochrome bd expression in E. coli is achieved primarily through the combined and complex actions of “fumarate nitrate reduction” regulator (FNR) and the ArcB/ArcA two-component system. A regulatory complex of five promoters initiates the transcription of cydAB: four of them (P1–P4) are coordinately regulated by oxygen (via ArcA and FNR). ArcA binds at three sites at this promoter (214), but only one is essential for transcriptional activation (82). Also, two distinct sites for binding FNR are discernable (82): the downstream FNR-2 site is critical for FNR-mediated repression in vivo, whereas the upstream site (FNR-1) plays an ancillary role in regulation of the cydAB operon by oxygen (129). Detailed discussion of the biochemistry of these oxygen-sensing global regulators is beyond the scope of this review but a summary follows.
FNR is a protein of the CPR-FNR superfamily of transcription factors, which all possess an N-terminal sensory domain and a C-terminal DNA-binding domain (132). FNR regulates transcription of target genes (its regulon) under anaerobic conditions by assembly of an oxygen-labile [4Fe-4S] cluster into the N-terminus. Assembly of this cluster facilitates formation of an FNR dimer with enhanced DNA-binding activity. It targets specific nucleotide sequences in selected promoters, where it acts as an activator of “anaerobic genes” and a repressor of “aerobic genes.” As oxygen availability increases, the [4Fe-4S] cluster is degraded to a [2Fe-2S] cluster so that site-specific inhibition is inhibited. The details of how O2 interacts with FNR are described elsewhere (83) and the effects of NO on the cluster are given in Crack et al. (84) and Cruz-Ramos et al. (87).
In Azotobacter vinelandii, CydR is an FNR-like protein, encoded adjacent to cydAB (180), that represses cydAB expression (339). Interestingly, FNR is structurally and functionally related to the cAMP receptor protein (CRP) [for a review see Green et al. (132)]. In Mycobacterium smegmatis, CRP directly regulates the expression of cydAB in response to hypoxia (9). No CRP site is present in the M. smegmatis cydDC promoter.
While FNR is a direct sensor of oxygen (and NO), ArcBA is a representative of the two-component regulators and is an indirect oxygen sensor. The ArcB protein is a membrane-integral sensor kinase and ArcA is its cognate response regulator. Two key cysteine residues in the cytosolic domain of ArcB must be reduced for the protein to be functional. It is generally accepted that it is the redox poise of respiratory chain quinones, such as those transferring electrons to the oxidase itself that is key to Arc function. Both the ubiquinone and menaquinone (MQ) pools are thought to be involved in regulating ArcBA (15). Fermentation products act as allosteric effectors (121, 122).
When oxygen is available, ArcB transits from being an autokinase to a phosphatase, via formation of two intermolecular disulfide bonds, resulting in the formation of a covalently linked ArcB dimer. Ultimately [for references see Bettenbrock et al. (30)], a phosphate group is transferred to ArcA, which assumes the role of a sequence-specific DNA-binding protein. In this form, it controls expression of several target promoters including cydAB (284). The system is switched off by the phosphatase activity of the oxidized (disulfide) form of ArcB, via dephosphorylation of ArcA (120).
When the oxygen tension drops, ArcA is phosphorylated by ArcB and activates cydAB transcription (81, 153). In early experiments, cytochrome bd expression was shown to reach a maximum at <2% oxygen tension (320) but more deliberate control of oxygen availability (2) questions this. When oxygen is decreased further, FNR becomes active and represses cydAB transcription (82, 320). A unique feature of the cydAB operon is the fact that FNR repression requires the presence of a functional ArcA protein (81, 82, 130). FNR has been proposed to act as an antiactivator by counteracting ArcA-mediated activation rather than directly repressing transcription (82, 130).
A role for the histone-like protein H-NS has also been reported (130). H-NS binds to an extended region within the cydAB promoter element, including sequences upstream from, and overlapping, the four regulated promoters. Oxygen control of cydAB transcription is thought to be mediated by three alternative protein-DNA complexes that are assembled sequentially on the promoter region as the cells are shifted from aerobic to microaerobic and to anaerobic conditions.
A recent systems approach to the oxygen response of cytochrome bd-I and the two other oxidases of E. coli (30, 104) has adopted a rigorous chemostat methodology, transcriptome profiling, and powerful mathematical modeling tools (287). In chemostats poised at quantified and constant oxygen provision rates, 89 genes exhibited expression changes when oxygen availability was modulated experimentally (276). O2 availability in the chemostats was expressed as “% aerobiosis” (2), defined by the converse relationship between declining q acetate (a measure of “overflow” metabolism) and increasing oxygen transfer rate. Steady-state simulation results of the three oxidase activities in comparison with measurement data showed good correlations with measured oxidase levels (104).
Cytochrome bd-I levels were very low at the highest and lowest rates of O2 transport and highest at 56% aerobiosis. Expression of this operon was 6.4-fold enhanced compared with anoxia (where acetate flux is maximal).
The expression of the second cytochrome bd-type oxidase of E. coli, cytochrome bd-II, is subject to less extreme regulation and has been studied less intensively; the function of this oxidase is unclear. Our current knowledge of the structures of cytochromes bd-I and bd-II suggests that their functions should be similar and perhaps interchangeable, yet they appear to be differentially regulated. Whereas cydAB is maximally expressed at O2 tensions between microaerobic and oxygen-rich conditions (320), appCB, encoding cytochrome bd-II, is maximally expressed at 0% aerobiosis (using the acetate criterion). Spectroscopic assays of oxidase levels confirm these conclusions (276, 319). In contrast, the cyoABCDE operon, encoding cytochrome bo3 , was maximally induced under fully aerobic conditions.
Thus, shifts in oxidase expression are consistent with a progressive switch from fermentation to microaerobic respiration (where cytochrome bd-I acts as the dominant oxidase) and then to aerobic respiration (where cytochrome bo3 is the prevailing terminal oxidase. Posttranscriptional regulation appears not to play a major role in oxidase expression.
III. Structure and Assembly
A. Structure
To date, cytochrome bd structures from two bacteria have been reported. In 2016, the first crystal structure of cytochrome bd, from the thermophile G. thermodenitrificans K1041, was determined at 3.1 Å (Protein Data Bank [PDB]: 5DOQ) and 3.80 Å (PDB: 5IR6) resolutions (281). The structure shows that this is a three-subunit enzyme. Both main subunits, CydA (∼52 kDa) and CydB (∼40 kDa) (283), are integral membrane proteins, each consisting of nine transmembrane helices, with the N-terminus and the C-terminus in the periplasmic and cytoplasmic sites, respectively (Fig. 5A). CydA and CydB are encoded by paralogous genes, which resulted from a duplication of a single ancestral unit codifying for a homodimeric oxidase (75). The heterodimeric structure is stabilized by hydrophobic interactions involving residues coming from the paired α3, α4, and α9 of CydA and CydB. The CydA and CydB subunits are structurally similar (root mean square deviation [RMSD]: 3.1 Å). A search for structurally similar subunits in PDB yielded a relatively low similarity with Complex III (cytochrome bc 1) from Flavobacterium johnsoniae (PDB:6BTM; RMSD: 3.4 Å; Fig. 6A) (311) and Rhodothermus marinus (PDB:6F0K; RMSD: 3.4 Å; Fig. 6B) (307), and polysulfide reductase from Thermus thermophilus (PDB:2VPX; RMSD: 3.2 Å; Fig. 6C) (162). CydS, in turn, is a single transmembrane helix of 33 amino acid residues (∼4 kDa), which is positioned at the peripheral interface formed by helices α5 and α6 of CydA. CydS is proposed to stabilize haem b 558 during potential structural rearrangements of the CydA upon binding and oxidation of quinol (281).
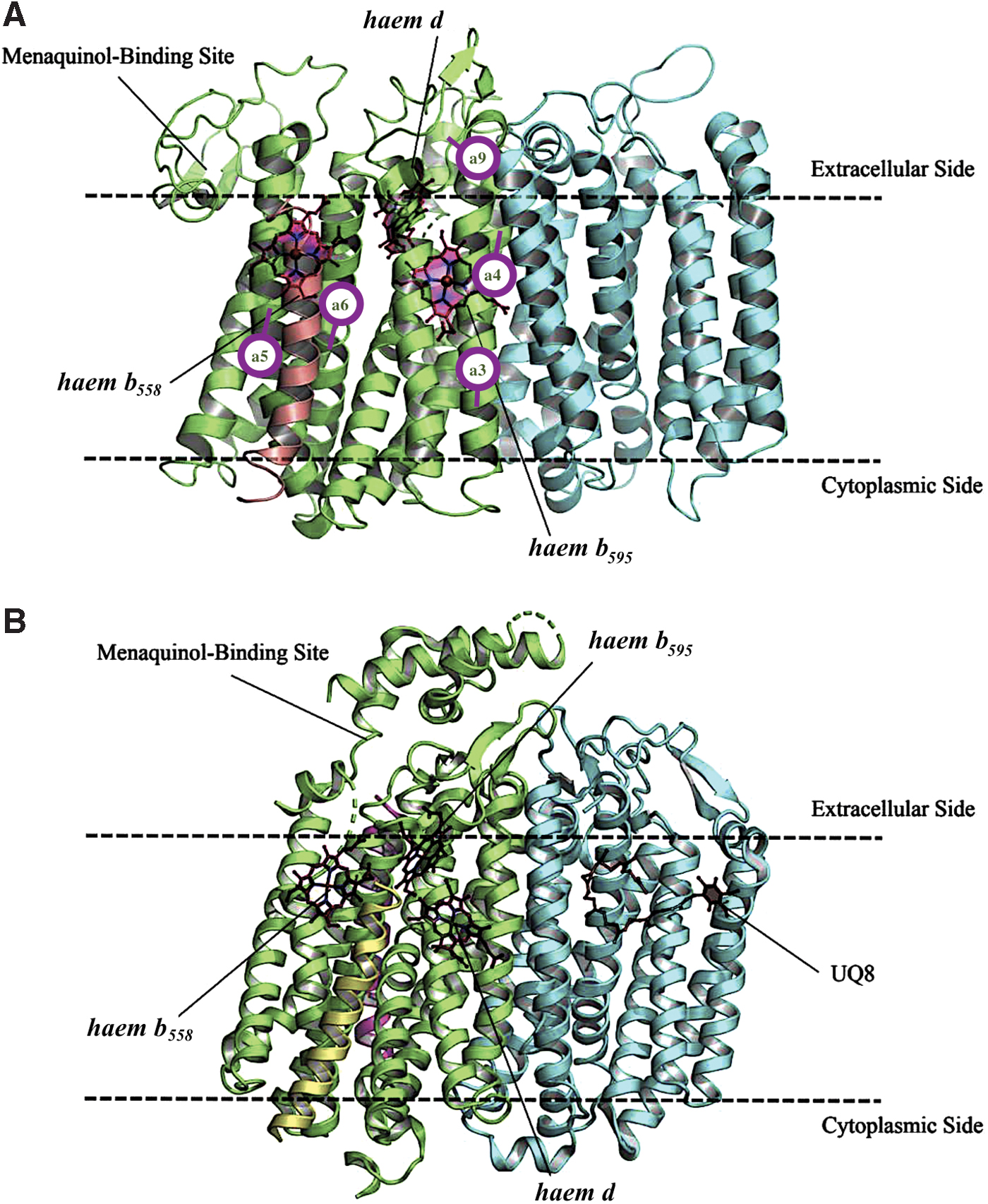
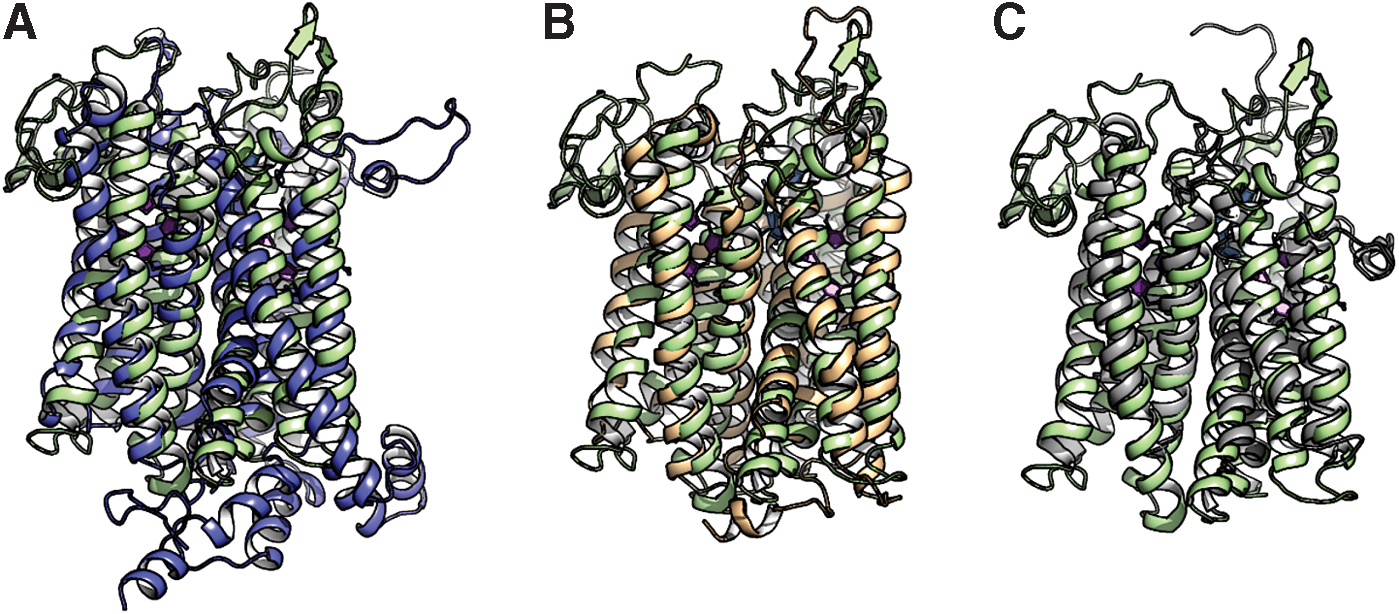
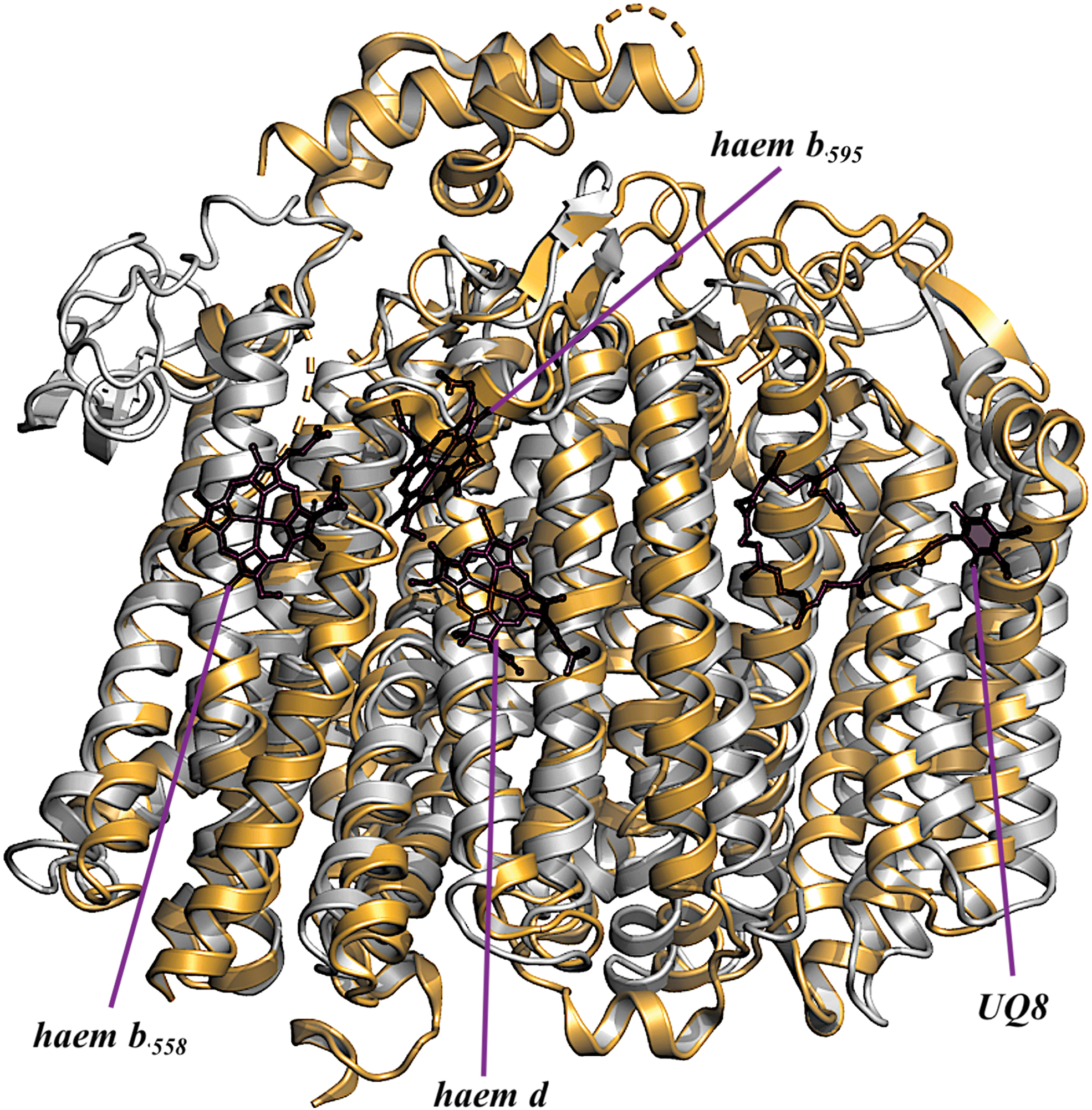
In 2019, using single-particle cryoelectron microscopy (cryo-EM), two structures of the E. coli cytochrome bd-I, in lipid nanodiscs with a bound Fab fragment (280) and in the presence of aurachin C (317), were determined at 2.68 Å (PDB: 6RKO) and 3.3 Å (PDB: 6RX4) resolutions, respectively. The results of the two articles (280, 317) are consistent. It turns out that cytochrome bd-I is a four-subunit enzyme (Fig. 5B). The arrangement and architectures of subunits CydA, CydB, and CydX are similar to the overall structure of the cytochrome bd from G. thermodenitrificans K1041 (Fig. 7). A fourth, previously unknown, subunit, absent in the G. thermodenitrificans oxidase, was named CydH (280) [or CydY (317)]. CydH appears to be a single transmembrane subunit that binds in the cleft between transmembrane helices α1 and α9 of CydA.
Cytochrome bd displays a unique binding domain for the oxidation of quinol called the Q-loop (218, 226). This is the loop connecting the α6 and α7 helices of CydA, facing the outside of the prokaryotic cell (Fig. 8A). The Q-loop is of variable length within the family (239, 282). Based on its size, the bd-type oxidases are divided into two subfamilies: L (long Q-loop) and S (short Q-loop) (7, 48). Cytochromes bd from G. thermodenitrificans and E. coli belong to S and L subfamilies, respectively. The quinone-analogue aurachin D is a powerful and relatively selective inhibitor of the E. coli cytochrome bd-I (221). Duroquinol:O2 oxidoreductase activity of the isolated bd-I oxidase is inhibited by aurachin D with IC 50 of 35 nM (317). In accord with these data, aurachin D inhibits O2 consumption of cytochrome bd in inverted membrane vesicles of M. smegmatis with an IC 50 of ∼400 nM (212).
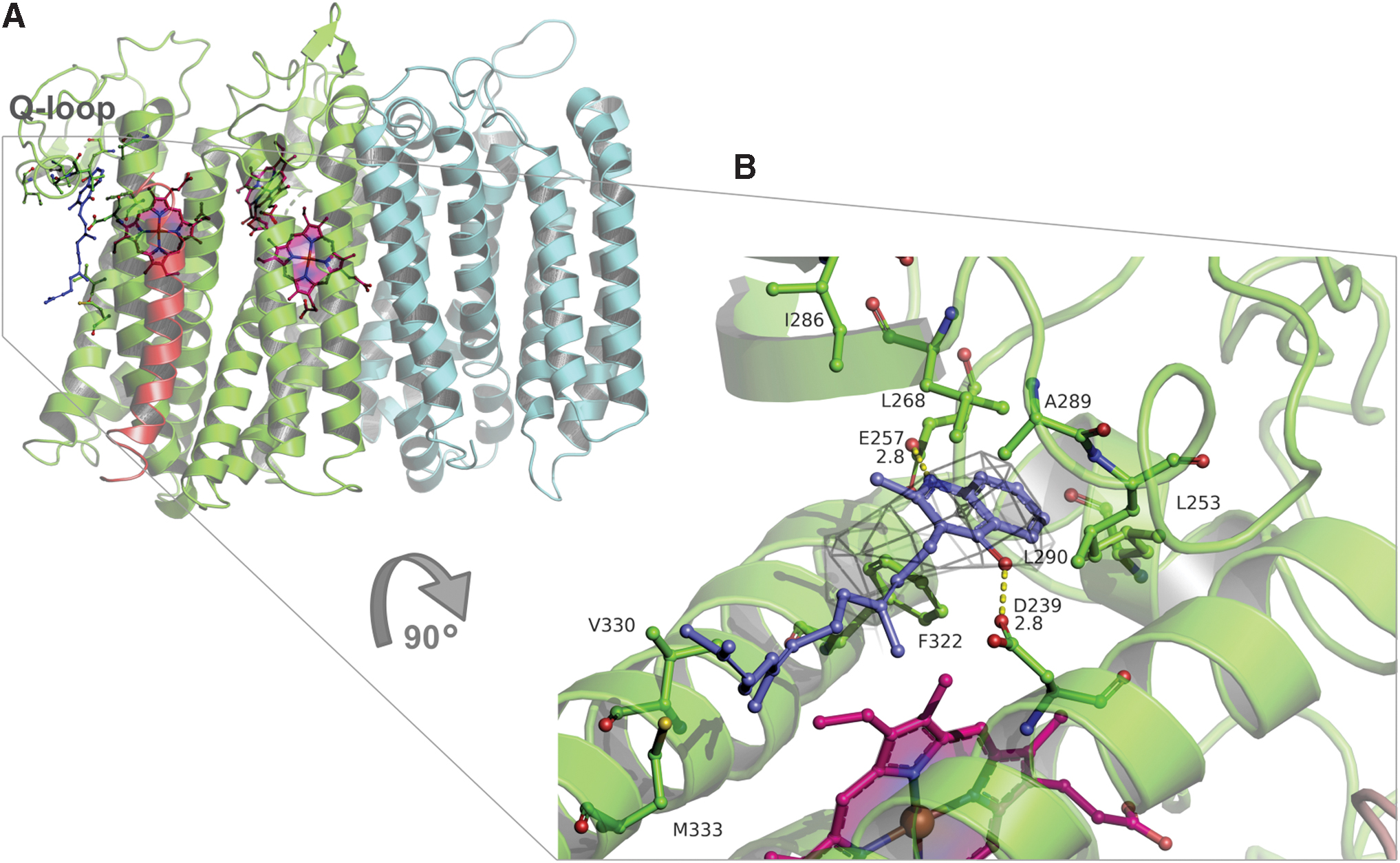
An in silico docking model of the complex between aurachin D and cytochrome bd (Paiardini, unpublished data) suggests that indeed aurachin D binds at the cytochrome bd quinol oxidation site (Fig. 8B), in close proximity to the Q-loop. The docking data (Fig. 8B; Paiardini, unpublished data) are consistent with the fact that, in the structure reported by Theßeling et al. (317), there is a small but significantly unexplained electron density in a pocket comprising E257 and K252, possibly due to a bound aurachin C. The role of the Q-loop in the as-isolated cytochrome bd from E. coli without substrate, in the presence of ubiquinone-1 (substrate analogue), and a quinolone-type inhibitor AD3–11 was investigated (280). The Q loop is divided into two domains: a flexible, disordered QN (N-terminal part) and a rigid, well-ordered QC (C-terminal part). The QC domain that defines the L-subfamily does not appear to be involved in substrate binding.
A feature that emerged from the crystal structure of the cytochrome bd from G. thermodenitrificans was the unexpected triangular arrangement of the three haems located on CydA. This suggests a direct electron transfer from haem b 558 to haem d, followed by equilibration with haem b 595 (Fig. 9A and section V). The evolutionarily conserved W374 could participate in the electron transfer between haem b 558 and haem d (281).
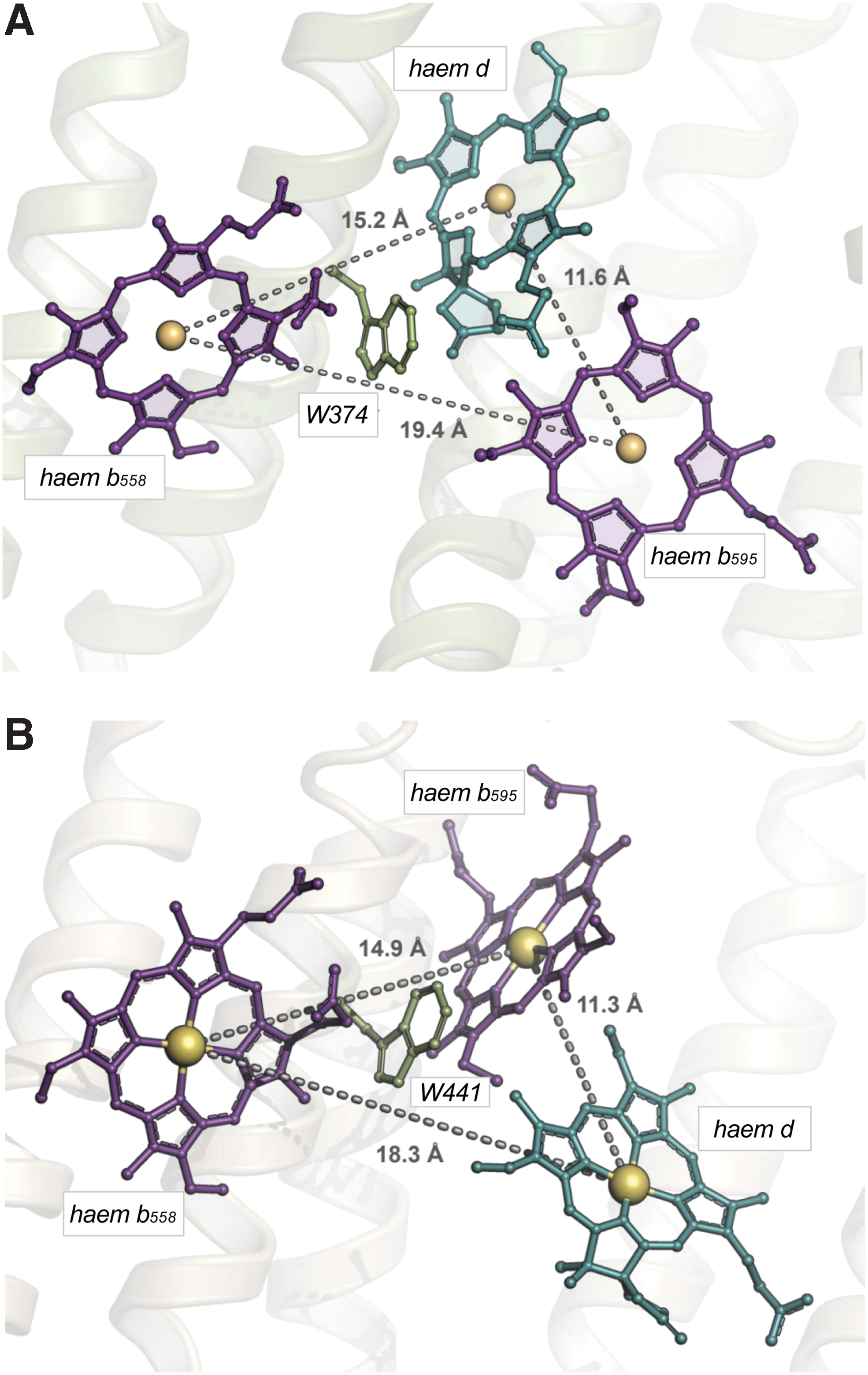
The E. coli cytochrome bd-I preserves triangular organization of the haems (Fig. 9B). The location and coordination of haem b 558 are equivalent in both enzymes. It is coordinated by H186 and M325 (M393 in E. coli). Surprisingly, however, haem b 595 and haem d in the E. coli structure are interchanged with respect to the G. thermodenitrificans enzyme (Figs. 5 and 9). In the latter structure, haem b 595 is hexacoordinate having H21 and E101 (H19 and E99 in E. coli) as axial ligands. It is buried in the protein interior deeper than haem d. Haem d has E378 (E445 in E. coli) as the axial ligand. In the E. coli structure, conversely, haem b 595 is pentacoordinate with E445 as the axial ligand. Furthermore, haem b 595, rather than haem d, is located near the periplasmic surface.
There is a discrepancy in the nature of the axial ligand to haem d between the two E. coli bd-I structures. Safarian et al. (280) claim that this is H19, whereas Theßeling et al. (317) report that the ligand is E99. The difference in the haem d axial ligand observed in the two static structures may indicate flexibility in the haem iron/ligand coordination bond, assuming that the haem d iron may coordinate no more than one protein axial ligand at a time. In line with this suggestion, lability (transient formation/breaking) of the haem d iron bond to a protein ligand in one of the states of the catalytic cycle (the one-electron reduced “mixed-valence” state) was noted earlier in time-resolved experiments (304). This implies the possible functional significance of the intraprotein axial ligand exchange in the haem d coordination sphere during catalysis.
The short edge-to-edge distance (6.7 Å) between haem b 558 and haem b 595 and the interchanged positions of the high-spin haems suggest a sequential interhaem electron transfer in the E. coli oxidase, b 558 → b 595 → d, as proposed earlier (48, 258) (Fig. 9B and section V). The totally conserved W441 possibly mediates electron transfer between b 558 and b 595. It is worth noting that the plane of haem b 558 is organized more-or-less at right angles to the membrane plane in both the E. coli and G. thermodenitrificans oxidases, whereas either d or b 595 is oriented at about 45o to the membrane plane in the two oxidases; these two high-spin haems are interchanged in the two species (280, 281, 317). Such an arrangement was predicted by electron paramagnetic resonance analyses of membrane multilayers rotated within the instrument cavity in 1980 before any structural data were available (250).
CydB contains a tightly bound ubiquinone-8 (UQ8; Fig. 5B). The molecule is located at a hydrophobic pocket, about 3.5 nm from haem d, in a near-symmetric conformation relative to haems b 558 and b 595. The significant distance from the haems makes its direct participation in electron transfer to/from the haems or oxygen reduction unlikely. The bound UQ8 possibly plays a structural role (280, 317).
Cytochrome bd is not active as a transmembrane pump, but a net proton flux results from quinol oxidation and consequent proton ejection at the periplasmic side of the membrane and uptake of cytoplasmic protons during oxygen reduction. Two alternative proton pathways in the G. thermodenitrificans structure have been indicated, one in CydA and the other in CydB, which converge on haem b 595 (Fig. 10) (281). In the E. coli enzyme, there is a hydrophilic, water-filled H-channel that connects the cytoplasm to haem d by the CydA pathway (Fig. 11). A chain of water molecules runs until D58 of CydB, and then, the conserved S108, E107, and S140 of CydA continue the pathway to haem d. Near D58, the H-channel branches and runs along the WDGNQ motif of CydB forming the CydB pathway branch (not shown). Surprisingly, in the center of CydB at W63, the hydrophilic branch merges with the hydrophobic oxygen-conducting channel (O2 channel, Fig. 11).
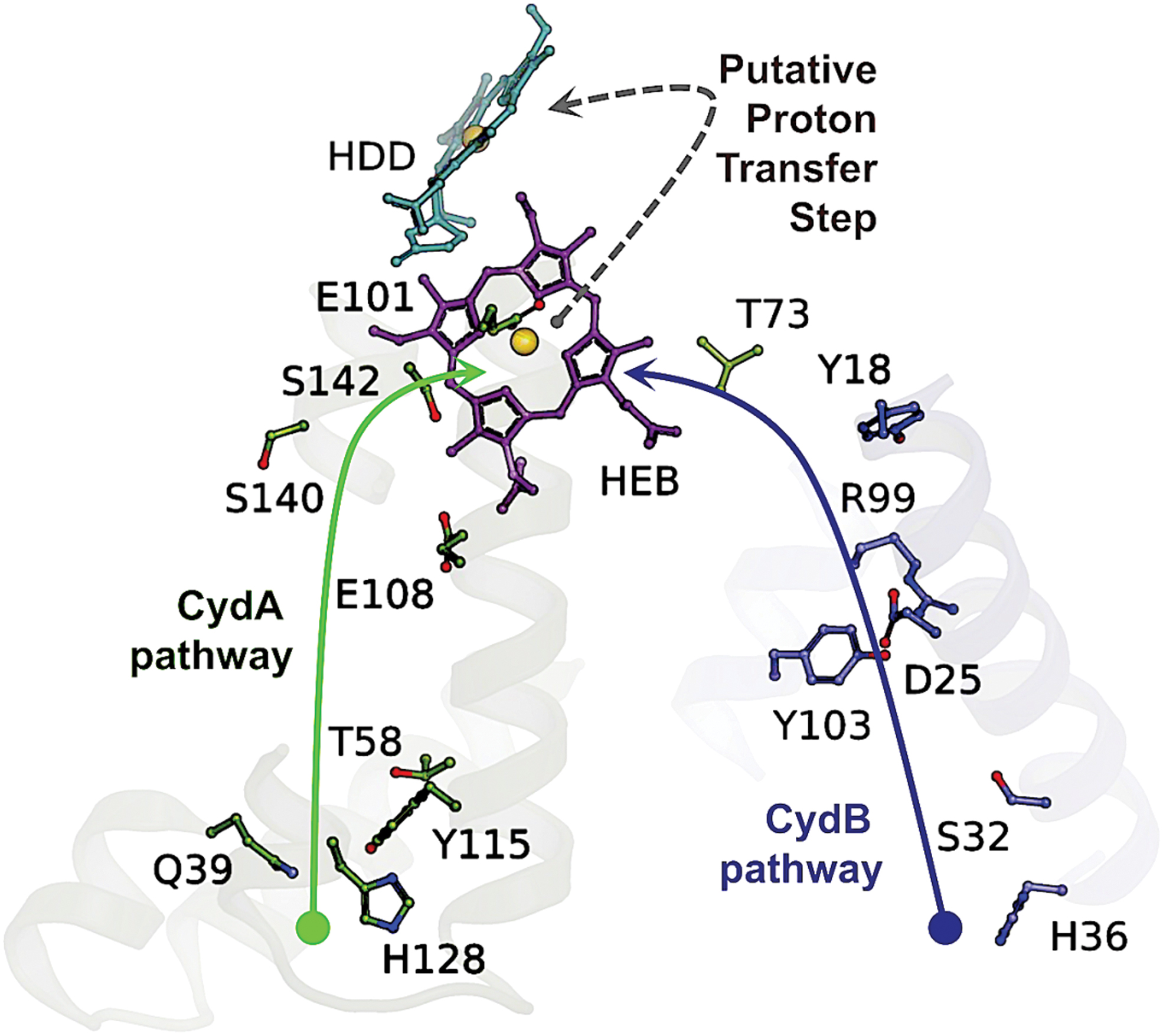
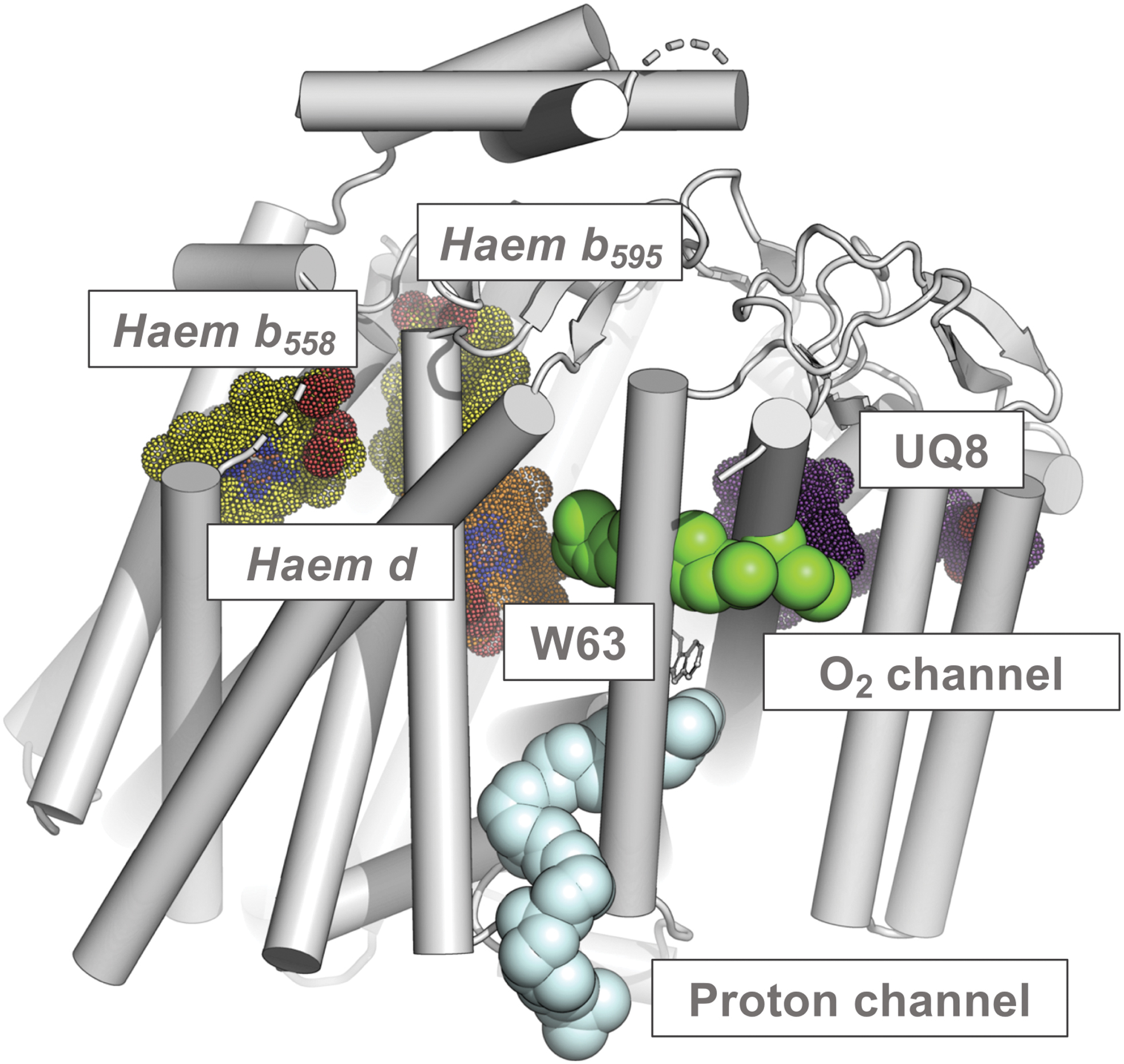
The O2 channel that goes parallel to the membrane plane connects the lipid interface to haem d (280, 317). In the G. thermodenitrificans structure, due to the lack of voluminous pathways there, O2 may gain access to haem d via a potential O2 entry site located laterally at a short distance from haem d. Thus, for the G. thermodenitrificans cytochrome bd, O2 present within the membrane could bind to haem d without the need for any functionally conserved protein cavity (281). The E. coli CydH occupies the O2 entry site in the G. thermodenitrificans enzyme, where CydH is absent, thereby preventing access of O2 to haem b 595 from the hydrophobic lipid bilayer (280, 317).
B. Assembly—the role of CydDC
Understandably, when molecular genetic tools were brought to bear on cytochrome bd, the structural genes (i.e., encoding the oxidase subunits) were the earliest focus. Mutants lacking the oxidase could be isolated and the encoding genes were cloned, sequenced, and analyzed, thus laying the foundations for studies on oxidase organization and function. However, other genes, involved in assembly of the oxidase complex in E. coli, were quickly found and these too have generated new information and hypotheses.
In E. coli, the cydDC genes are distant from the structural genes on the chromosomal map (123), but in certain bacteria they occur in a single operon with cydAB thus being expressed on one continuous transcript, as occurs in Bacillus subtilis (337). Other bacteria have oxidases that resemble cytochrome bd-I, yet have no spectrally detectable high-spin haems (b 595 and the d-type haem) as in Campylobacter jejuni; nevertheless, two genes (cj0081, cj0082) that encode proteins similar to the CydAB proteins in E. coli were identified from genome sequences (241). These issues are discussed in Poole et al. (253).
Neither cydD nor cydC is an essential gene in E. coli, as evidenced by the strategies that identified cydC and cydD mutants. However, Eng et al. (108) describe E. coli cydC as an essential gene and suggest that cytochrome bd-I (i.e., CydABX) and CydDC do not always co-occur: among the 1965 genomes analyzed, 407 species exhibited highly conserved cydC genes, but only 53 showed a cydB homologue. It is puzzling that cydDC genes appear to be found in so many genomes that do not encode cytochrome bd-I; Kranz et al., in the 33 bacterial genomes analyzed, found none (197), although six species encode the oxidase but not CydDC, namely Rickettsia prowazekii, Chlamydia tracomatis, C. jejuni, Porphyromonas ginigivalis, Aquifex aeolicus, and Thermotoga maritima. All were claimed to lack both CydDC and cytochrome bd-I by Eng et al. (108).
Furthermore, neither Geobacillus nor Mycobacterium species were found by Eng et al. to encode cytochrome bd-I, despite the fact that other authors report functional or structural studies on cytochrome bd-I from these bacteria (23, 114, 281). Caution is urged when deducing the apparent absence of genes from a genome, particularly genes such as cydDC that are members of a very large and diverse family—the adenosine triphosphate (ATP)-binding cassette (ABC)-type transporters.
1. The cydDC genes
In addition to the structural genes that encode the cytochrome bd complex, detailed above, it is now clear that, in many bacteria, two additional genes function in proper assembly of the oxidase. In E. coli, where information is most comprehensive, and most bacteria, they are named cydC and cydD. These genes encode a transport system in the ABC class and are widely thought to be the sole two components of a membrane-integrated export system.
By the early 1990s, cydA and cydB encoding the cytochrome bd-type terminal oxidase (bd-I) (61) were mapped to an operon at 16.6 min (188) on the chromosomal map. Remote from the cydAB locus and located at 19.2 min (123), a further gene, implicated in “cytochrome d assembly,” was also identified. Mutants in this gene (named cydC) were devoid of absorbance bands in the visible spectrum that could be attributed to cytochrome bd-I; however, the spectrum was returned to wild-type (WT) characteristics by introducing the cloned cydC + gene on an episome.
Transcription/translation experiments and Western immunoblotting of membranes from a cydC strain showed that the CydA and CydB subunits were present but diminished in membranes from a cydC strain relative to the isogenic cydC + strain (123). Expression of the oxidase subunits in a cydC strain demonstrated that the b 595 and b 558 haems were overproduced, but the haem d component was absent. Thus, a plausible hypothesis was that CydC is involved in biosynthesis of haem d; in the absence of this haem, the oxidase subunits are largely absent and destabilized (123). Note that Siegele et al. independently described the cydC gene but named it surB, because its gene product was required for E. coli cells to exit (i.e., survive) in a stationary phase aerobically (299).
In 1989, we identified a fourth cyd locus by adopting a markedly different strategy (260): survivors of a classical nitrosoguanidine mutagenesis were screened using a hand spectroscope (on samples held at 77 K) for loss of the characteristic absorbance at 630 nm of reduced cytochrome bd. The mutant gene in one such isolate was mapped to 19.3 min (260). Furthermore, a gene implicated in the ability to survive at elevated temperatures, htrD, was shown, after correcting a missing G in the earlier sequence, to be identical to cydD (98).
Cloning of the cydC and cydD genes (255) resulted in a fragment of chromosome originating in the 19-min region of the Kohara map, and thus consistent with earlier P1 mapping data (260). When such plasmids were used as templates for in vivo protein synthesis, two proteins identified as CydD (61 kDa), calculated to be 63 kDa, and CydC (59 vs. 63 kDa) (255) were generated. CydD and CydC were similar: the deduced amino acid sequences revealed 50% similarity and 27% identity. Hydropathy profiles predicted that the CydDC complex comprises two similar membrane proteins, and it was identified as the first documented example of a bacterial heterodimeric ABC transporter, probably an exporter, by comparison with known ABC family members.
2. The CydDC proteins: structure and function
Based on analysis of hydrophobicity of CydD and CydC, it was predicted that both subunits would have six transmembrane helices, the C-terminal portion of each polypeptide being hydrophilic and containing an ATP-binding site (255). A membrane topology model for CydDC predicted both subunits to have six transmembrane regions separated by two major cytoplasmic loops; both ends of each polypeptide chain were predicted to be located in the cytoplasm (86). Subsequent modeling of topography (253) was consistent with those models, and used to highlight the Walker A motif (which binds ATP), the Walker B motif (which interacts with Mg(II)), and conserved amino acids (histidine, glutamate, aspartic acid) that are part of the H-, Q-, and D-loops, respectively.
The periplasm of an E. coli cydDC knockout mutant is “overoxidizing” (126) indicating that CydDC may catalyze cell export of reductant(s). Indeed, loss of CydDC diminishes the level of reduced thiol detected in the extracytoplasmic compartment, while overexpression of CydDC decreases the cytoplasmic reduced thiol pool (146). These observations are consistent with in vitro studies that measured import of 35 S-labeled cysteine into everted membrane vesicles and demonstrated that, in vivo, CydDC mediates energy-linked export of cysteine (247).
CydDC also transports outward, and thus presumably into the periplasm, the tripeptide glutathione (L-γ-glutamylcysteinylglycine [GSH]), a major regulator of cellular redox poise (248). The transport rate for GSH by CydDC was fivefold higher than for cysteine; therefore, given the abundance of GSH in the bacterial cytoplasm (248), GSH is likely to be a major substrate for CydDC. Addition of GSH and cysteine both stimulated the ATPase activity of purified CydDC (342), supporting an enzymic role in the export of reductant. The complexity of the process is probably not fully understood: Eser et al. (110) studied the effects of mutating three genes (cydD, ggt—encoding periplasmic γ-glutamyl transpeptidase—and mdhA—encoding a multidrug-resistant-like ABC transporter) that might influence periplasmic GSH pools, but none affected the ability of glutaredoxin-3 (GrxCp) to catalyze the formation of disulfide bonds, suggesting the existence of other routes for GSH export in E. coli.
Because CydDC is an exporter and is required for haem assembly into the oxidase, it was hypothesized that CydDC might export haem, a proposal tested experimentally. However, use of everted vesicles and radiolabeled haem in vitro failed to demonstrate this function for CydDC (74). Later work exploited a purified form of CydDC with an absorption peak at 410–412 nm; pyridine haemochrome analyses indicated the presence of a bound b-type haem with a CydDC:haem ratio of 5:1 (342). This bound haem was reducible and oxidizable, and bound CO. Haemin and GSH/cysteine had synergistic and stimulatory effects on the ATPase activity of the complex. This suggests that the haem cofactor has a significant but obscure role in CydDC function.
Certain other reduced thiols (including homocysteine and methionine) also activated CydDC. Control experiments with S-substituted/nonthiol analogues and haem lacking the central iron (protoporphyrin) did not stimulate rates of ATPase activity, and inclusion of nonthiol reductants decreased the ATPase rate. These experiments suggest the need for either thiols or an iron-containing tetrapyrrole in CydDC function. Histidine was an intriguing exception: it gave a twofold increase in ATPase activity, which increased to eightfold on additional inclusion of 1 μM haemin. Since axial ligands to haem include both histidine and reduced thiol compounds, this suggests that the haem-ligating capacity of reduced thiols contributes to the enhancements in ATPase activity observed for GSH and cysteine (342).
3. Structural investigations into the CydDC complex
Two-dimensional crystals have been obtained by incorporating purified CydDC into E. coli lipids, thus permitting cryo-EM (342). The electron densities reveal arrays of dimeric units in “up” and “down” orientations, indicating CydDC heterodimers in the crystal lattice. Although no three-dimensional crystal structure has been reported, Shepherd and colleagues used a structural modeling approach to study the roles of individual residues in catalysis and cofactor binding (253).
The two-point mutations in the earliest cydD1 allele are G319D and G429E (86). Based on homology modeling, the G429 residue is close to a conserved aspartate residue; it is postulated that substitution for a glutamate (also negatively charged) could disrupt the Walker A motif and abolish function. Mutation of G319, which was found to be buried at the bottom of the hydrophobic pocked, may also impact upon conformational changes during the catalytic cycle [see Poole et al. (253)].
4. Physiological impacts of CydDC function
Based on the above data on the effects of CydDC on periplasmic physiology, the pleiotropic phenotype of cydDC strains may result predominantly from the disruption of disulfide folding in that compartment (248). Indeed, sensitivity of the cydDC mutant to benzylpenicillin was attributed to misfolding of the disulfide-containing penicillin-binding protein 4. The observed loss of cell motility in cydDC strains may result, for example, from a defective P-ring motor protein (247), since exogenous cystine (i.e., oxidized cysteine) corrects a motility defect in a dsbB mutant (89). Complementation of cydDC strains with exogenous reductant largely complemented the phenotypes associated with defective disulfide folding (248).
It is important to differentiate between deficiencies of oxidase per se and the additional defects in cydDC mutants, which do not assemble cytochrome bd. This distinction is difficult because both cydDC and cydAB mutants display diverse and sometimes overlapping phenotypes (252). All cydAB mutants (i) appear to lack spectroscopically detectable cytochrome bd, (ii) fail to survive as robustly as WT cells in stationary phase (299), (iii) are sensitive to inhibitors (cyanide, azide, Zn(II) ions) (260), and (iv) intolerant of oxidative and nitrosative stresses (117, 124, 146, 295). cydDC mutants also exhibit low levels of other cytochromes, particularly those haemproteins in the periplasm (254), and have a more oxidized periplasm. The failure of cydDC mutants to export to the periplasm GSH and cysteine results in a more oxidized periplasm. However, we do not understand why bd-type oxidase assembly is inhibited by these defects. Among plausible hypotheses are the following: The haems exported by CydDC to the periplasm (74) may be assembled onto outward-facing domains of the oxidase subunits. The finding that haem stimulates the ATPase activity of purified CydDC is intriguing (Section III.B.2). The haems exported by CydDC to the periplasm CydDC occupy there critical sites such as on chaperones required for oxidase assembly. The processing and assembling processes of the periplasm are intolerant of the oxidized nature of the periplasm in cydDC mutants. In addition to GSH and cysteine, which were identified as substrates for CydDC in transport studies, CydDC may export other thus-far unidentified metabolites.
We discuss the implications of cytochrome bd deficiency (and by extension of cydDC mutations) in Sections VI to VIII and in Poole et al. (253).
5. Conclusions
CydDC is of great interest for its profound impact on oxidase assembly and also because it was the first heterodimeric ABC-type exporter to be described in prokaryotes. The genes may or may not be found in an operon with the structural oxidase genes. The loss of bd-type oxidase assembly and function in cydDC mutants reveals CydDC to be crucial for this respiratory complex. The CydDC system transports to the periplasm reducing molecules, notably GSH and cysteine, which stimulate the ATPase activity of the isolated CydDC complex. We lack information on other potential substrates for CydDC and unambiguous evidence that haem is transported outward to the periplasm where it may be assembled into the oxidase. A direct involvement of haem with CydDC function is, however, suggested by studies in vitro of the ATPase activity of CydDC.
IV. Spectral and Redox Properties
The absorption spectra of the three haems, b 558 and b 595 (which are protohaems IX) and d (which is a cis-haem d hydroxychlorin γ-spirolactone) constitute the overall absorption spectrum of cytochrome bd. To date, the E. coli cytochrome bd-I is the only cytochrome bd for which, by means of spectroelectrochemical redox titrations, the difference (reduced minus oxidized) in spectra of all three haems in the α-, β- (32, 185, 210), and Soret (32) regions was resolved (Fig. 12). The position of the Soret band maximum for haem b 595 is consistent with that unveiled before by femtosecond spectroscopy (50, 328) and by investigations on the E445A mutant (6, 269). The spectral contribution of haem d to the aggregated Soret band is much smaller than those of either b-type haems.
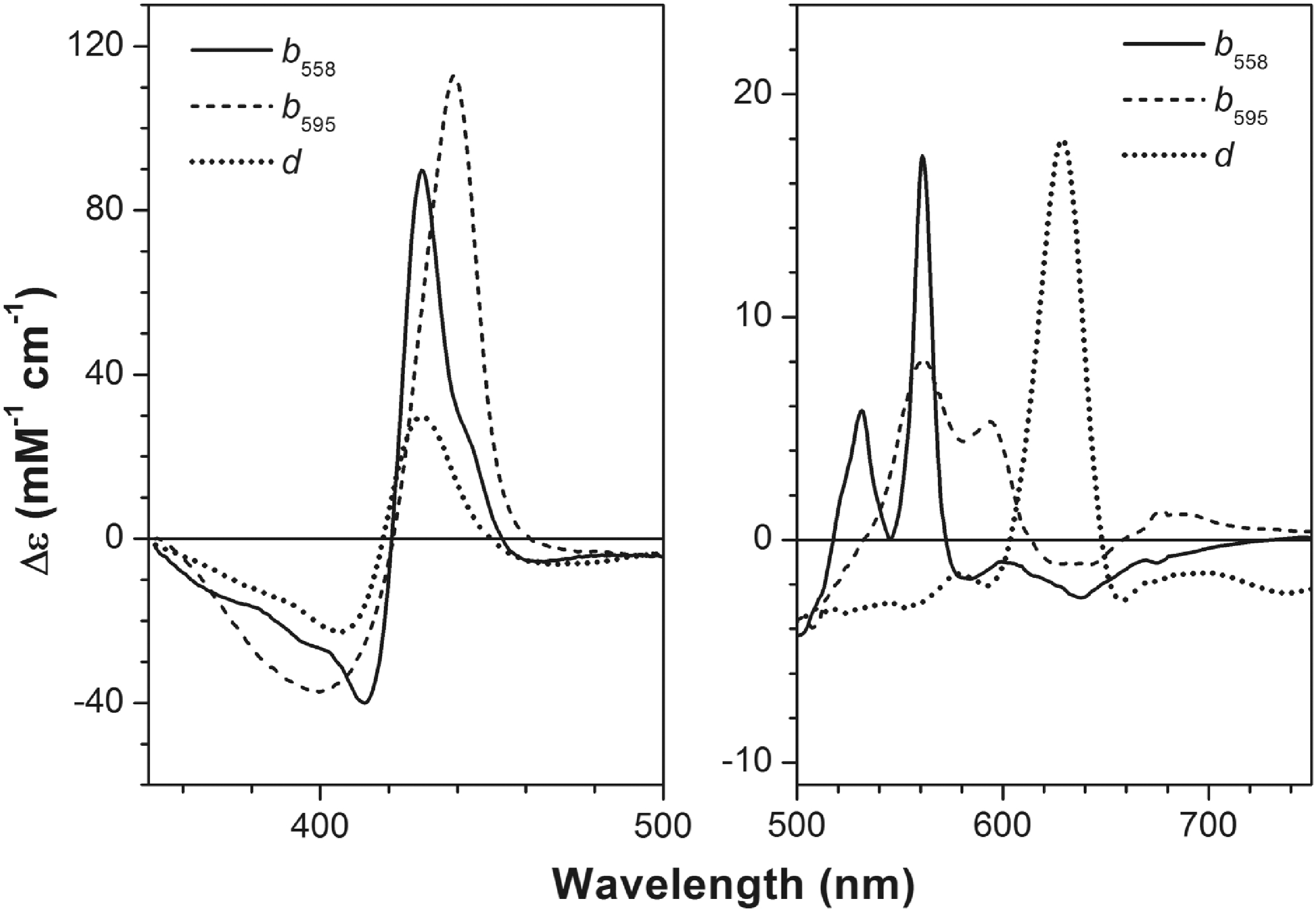
Upper asymptotic redox potentials of haems b 558, b 595, and d in the E. coli cytochrome bd-I in sucrose monolaurate at pH 7.0 are +172, +182, and +256 mV, respectively (32). These values are consistent with those for apparent redox potentials of the haems in bd-type oxidases from E. coli and A. vinelandii reported earlier (17, 169, 185, 211, 220, 264, 277). The detergent's nature affects the apparent redox potentials for haems b, particularly haem b 558, but has virtually no effect on the haem d potential (32, 211). In the absence of haem d (in the E99L mutant), a large interaction potential (about −90 mV) between haem b 558 and haem b 595 is observed (32). The presence of haem d reduces the redox interaction between the two haems b by ca. three times.
In contrast, redox interaction between haem d and either b-type haems is weak. The latter apparently contradicts the fact that the distance between haems b 558 and b 595 is longer than that between haems d and b 595 (281). However, a strong redox interaction between redox-active metal sites does not always mean a short distance between them. Lack of significant electrostatic interaction at short distance may be due to screening of the electron charge by the proton taken up on reduction (32). For instance, in cytochrome c oxidase, the distance between haem a 3 and CuB is much shorter than that between haem a 3 and haem a (154, 322). At the same time, redox interaction between haem a 3 and CuB was not reported, whereas the redox interaction between haems a and a 3 is significant (236). Notably, the redox interaction between haems a and a 3 in cytochrome c oxidase (−115 mV at pH 8.0) (128) is significantly larger than that in cytochrome bd-I.
This may be relevant to the proton pumping mechanism in cytochrome c oxidase (10, 196), although transient states occurring during the enzyme's catalytic cycle may not be accessible in equilibrium redox titrations (300). A possible redox interaction between semiquinone and the haems in cytochrome bd remains to be examined, although the midpoint potential of the bound quinone in cytochrome bd-I was reported (139).
Being high-spin pentacoordinate, haem d can bind not only the O2 enzyme natural substrate but also exogenous ligands, such as CO (7, 34, 37, 38, 44, 50, 52, 54, 142, 143, 165 –167, 169, 210, 232, 269, 302 –304, 328), NO (34, 41 –44, 148, 163, 167, 168), cyanide (34, 167, 177 –179, 181, 198, 249, 262, 263, 277, 312, 313, 321), and H2O2 (35, 43, 49, 116, 155, 172, 209, 259). A marginal ligand reactivity of the haems b was also observed (34, 37, 52, 54, 143, 302). Detailed discussion of the reactions of cytochrome bd with the exogenous ligands is beyond the scope of this review but can be found in Borisov et al. (48) and Junemann (163).
V. Catalytic Cycle
Whereas the superfamily of haem/copper oxidases comprises both quinol and cytochrome c oxidases, the bd-type oxidases characterized thus far are all quinol oxidases. Unlike cytochromes c, quinols are two-electron-donating substrates. Yet, they are oxidized by cytochrome bd through sequential one-electron transfer steps, with O2 acting as the electron acceptor (36, 48, 163). Three quinol types with distinct structural differences can act as physiological substrates for cytochrome bd: ubiquinol (UQH2), menaquinol (MQH2), and plastoquinol.
The nature of the quinol substrate can be species-specific or dependent on growth conditions. For instance, mycobacterial and cyanobacterial bd enzymes utilize MQH2 and plastoquinol, respectively (24, 25, 73, 246), while E. coli cytochrome bd can use UQH2 or MQH2 depending on O2 availability (24, 25, 246, 323). Interestingly, some cytochromes bd were found to copurify with tightly bound quinols (19, 20), and a structural role in dimer stabilization was proposed for the UQ8 molecule detected in the recently solved structure of E. coli cytochrome bd-I (280).
The primary acceptor of electrons donated by the quinol substrate is haem b 558 (Fig. 13). From there, electrons are transferred intramolecularly to the haems b 595 and d, but it is unclear which of the two other haems is reduced first upon oxidation of haem b 558. The structure of the G. thermodenitrificans enzyme (281), with the haems b 558 and d significantly closer to each other than the two b-type haems (edge-to-edge distance of 5.9 vs. 8.5 Å, and Fe-Fe distance of 15.2 vs. 19.4 Å), points to a direct electron transfer from haem b 558 to haem d.
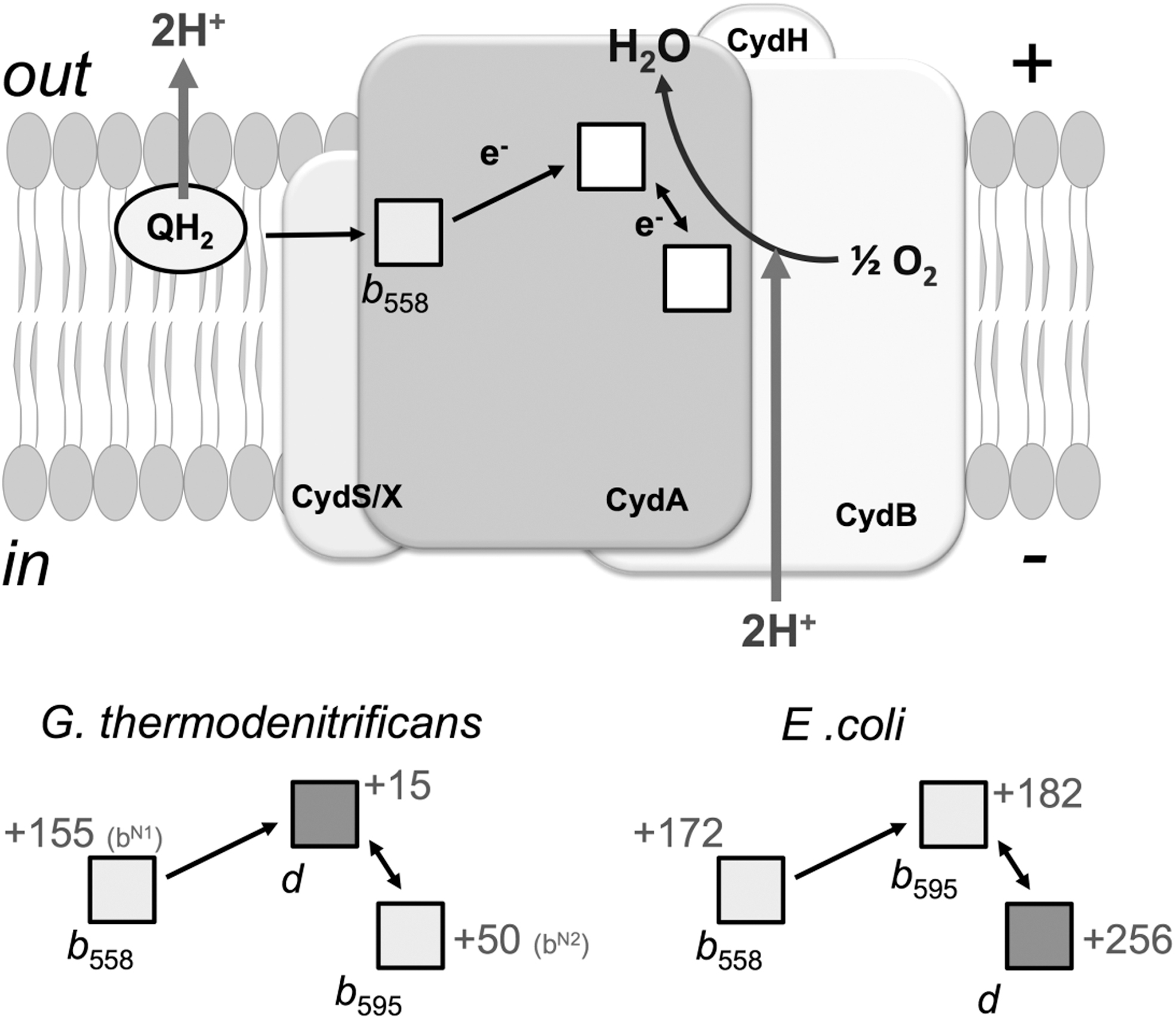
Conversely, the recently solved structure of E. coli bd-I enzyme (280) with the same triangular arrangement of cofactors and similar interhaem distances, but interchanged haems b 595 and d, suggests that electron transfer proceeds along the pathway haem b 558 → haem b 595 → haem d (Fig. 13). Despite this uncertainty, in both structures the haems b 595 and d, without forming a binuclear site with their Fe atoms, are in van der Waals contact (edge-to-edge distances of 3.5 and 3.8 Å in the G. thermodenitrificans and E. coli enzyme, respectively). Therefore, regardless of which haem acts as the primary acceptor for electrons donated by haem b 558, electron equilibration between the haems b 595 and d is expected to be very rapid.
The intramolecular electron transfer processes taking place inside cytochrome bd are not coupled to a proton pumping activity (267). Nevertheless (see section III.A), electron transfer is electrogenic and leads to generation of a transmembrane electric potential; its existence has been demonstrated for the enzymes from E. coli and A. vinelandii with diverse experimental approaches (19, 20, 26, 38, 51, 155, 183, 186, 187, 223, 224, 267). Generation of the transmembrane electric potential occurs as the electrons are transferred from haem b 558 to the haems b 595 and d (19, 20, 38, 155).
The potential arises from uptake and release of protons during electron transfer, but not from the electron transfer itself, in line with the information that the three haems are located at relatively similar depths inside the membrane. Experimental evidence suggests that the transmembrane electric potential results from the combined release into the periplasmic space and uptake from the cytoplasmic space of protons, respectively, associated with quinol oxidation and intramolecular reduction of haems b 595 and d (19, 20, 38, 155) (Fig. 13). Thus, in the absence of a true redox-coupled proton pumping activity, combination of these events accounts for a vectorial proton transfer across the membrane, thereby contributing to the electric potential buildup.
Cytochrome bd in turnover conditions processes O2 to H2O with a very high affinity for O2. For the E. coli bd-I and bd-II oxidases, low K
m(O2) values were determined, respectively, in the 0.003–0.3 and 0.24–2.0 μM range, depending on the experimental approach used (16, 88, 156, 217). An usual feature of cytochrome bd among terminal oxidases is that in the single-electron reduced state (
Interestingly, the single-electron (
Binding of O2 to the fully reduced enzyme leads to the four-electron reduction of O2 to 2H2O via several catalytic intermediates, which are similar in nature to those populated in the O2 reaction with haem/copper oxidases. A scheme of the catalytic cycle is presented in Figure 14. In flow-flash experiments coupled with spectroscopic or electrometric measurements, it was found that haem d in the
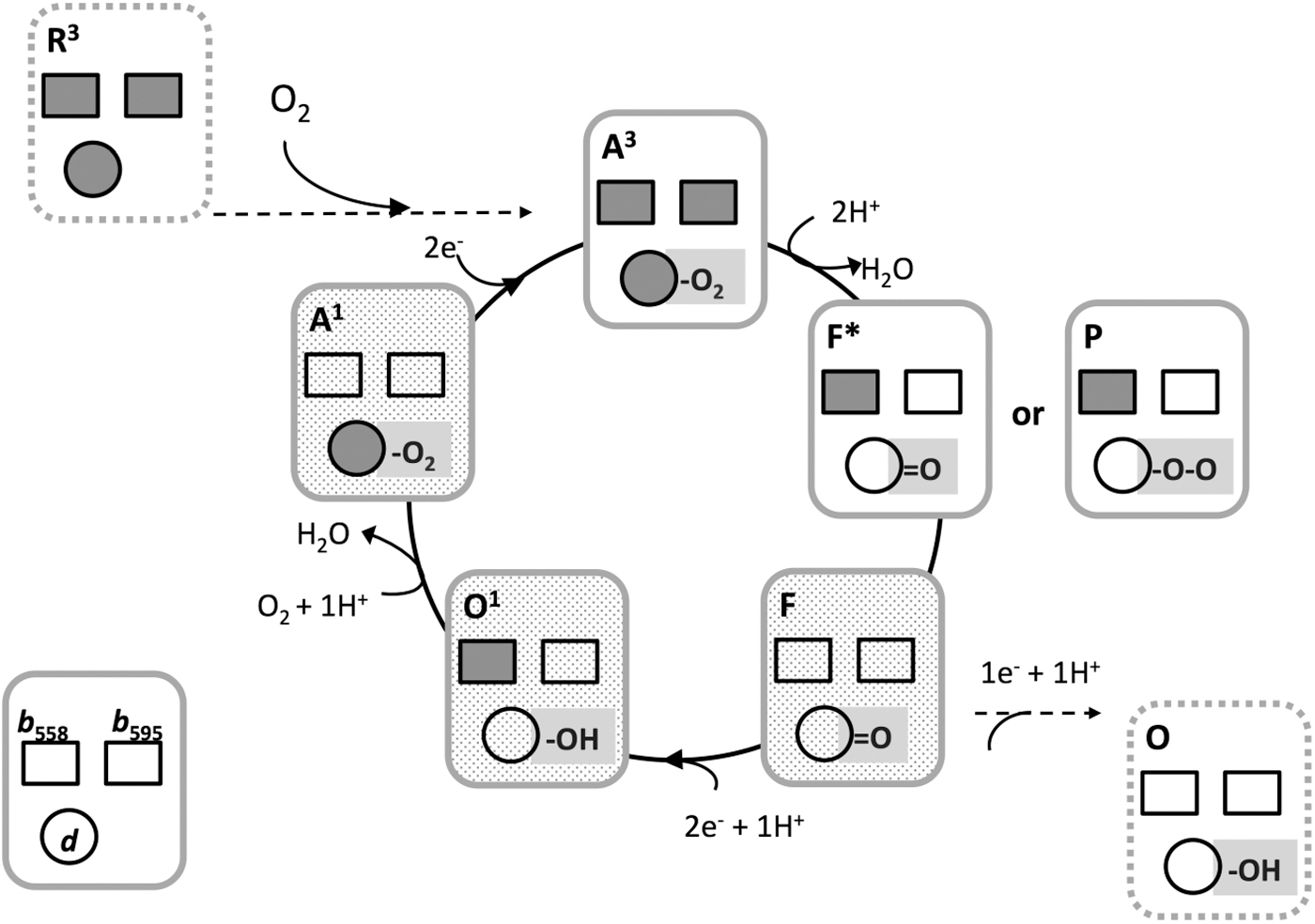
Although the detailed chemical nature of this intermediate remains uncertain, it was noted that the recently solved structure of the E. coli enzyme does not rule out the occurrence of a true
In stopped-flow multiwavelength spectrophotometric experiments, the steady-state level of the catalytic O2 intermediates of cytochrome bd was measured in turnover conditions sustained with excess dithiothreitol and Q1. Under these conditions, at steady state, the mostly populated intermediates proved to be the ferryl (
VI. Physiological Functions
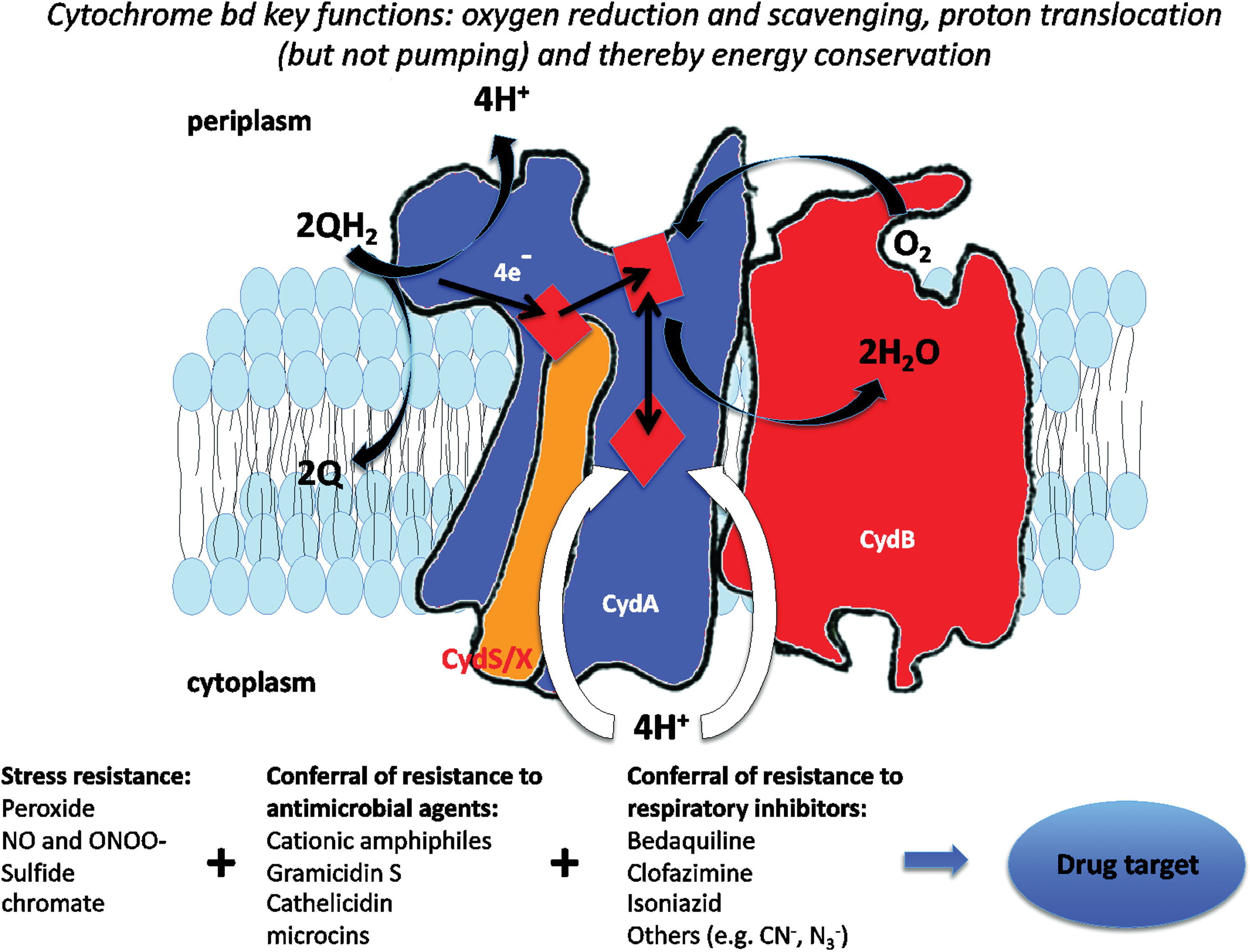
The core functions of terminal oxidases are oxygen reduction, generally to water, reductant consumption (in this case ubiquinol), and proton movements across the membrane, either by proton pumping mechanisms (as in the case of the haem/copper superfamily) or balanced proton extrusion to the outside (periplasm) and uptake from the inside (cytoplasm), as in the case of cytochrome bd (Fig. 15). However, in the case of cytochrome bd, we can discern extra physiological functions or attributes of great interest. These are summarized at the foot of Figure 15 and described in detail in the following sections.
A. Respiratory protection of nitrogenase
In addition to its unique structure, characteristic modes of expression, oxygen kinetics, and resistance to inhibitors (sections C and VII), cytochrome bd is unusual in being implicated in a physiological mechanism termed respiratory protection. This is the maintenance of function of bacterial nitrogenase, an oxygen-labile enzyme, even under aerobic conditions, this being attributed to the phenomenal capacity of cytochrome bd for rapid oxygen consumption.
The free-living obligate aerobe Azotobacter can fix dinitrogen over a wide range of oxygen concentrations, despite the notoriously oxygen-sensitive activity of its nitrogenase. Azotobacter adapts its respiratory rate in response to changing oxygen tensions (101) and, when fixing nitrogen, has one of the highest rates of respiration known (256). Dalton and Postgate proposed that “uncoupled” respiratory activity might prevent oxygen inhibition of nitrogenase activity (90).
The evidence that this involves cytochrome bd is manifold: (i) the level of the oxidase increases when oxygen supply increases (e.g., Moshiri et al. (230), but not in E. coli, where cydAB is expressed maximally at low O2 tensions; see section II.D); (ii) consumption of carbon and energy sources is partially uncoupled from anabolism, specifically via cytochrome bd; and (iii) critically, Cyd− mutants cannot fix nitrogen in air (180). However, a cytochrome bo3 mutant is aerotolerant during nitrogen fixation (203). The Fnr-like transcription factor CydR (338, 339) controls the expression of both cytochrome bd and the uncoupled NADH-ubiquinol dehydrogenase (Ndh) (27), which is thought to supply electrons to cytochrome bd-I and is also essential for aerotolerant nitrogen fixation (28).
The conclusion that cytochrome bd affords respiratory protection is supported by genetic studies of other bacteria. In Azorhizobium caulinodans, both cytochrome bd and cytochrome cbb 3 contribute equally to nitrogen fixation in root nodule symbiosis; the double mutant totally lacked symbiotic N2 fixation (175). In Klebsiella pneumoniae, N2-fixing ability was severely impaired in a cyd mutant even at low oxygen concentrations (170).
The hypothesis of respiratory protection, specifically by cytochrome bd, is widely accepted and the genetic evidence (i.e., that Cyd− mutants cannot fix nitrogen in air) is viewed as the most convincing (238). Nevertheless, objections have been raised. One of these is that total respiratory oxygen consumption is not elevated above 70 μM O2, where protection should be most important. However, A. vinelandii possesses, based on genome interrogation (293), five terminal oxidases: cytochrome c oxidase (Cdt), cytochrome o (Cox), cytochrome bd copy I (CydAB I), cytochrome bd copy II (CydAB II), and cytochrome cbb 3 (Cco).
Note that the mutagenesis approach in Kelly et al. (180), which is taken as the best evidence for respiratory protection, was targeted at cytochrome bd-I. The presence of multiple oxidases allows flux through different branches, only one of which terminates in cytochrome bd-I, to be redistributed during protection of nitrogenase without elevation of total respiratory rates. A second concern is that the efficiency of respiration in protection may be insignificant because the rate of consumption is too low to prevent diffusion of O2 into cells. However, the location of the O2-reducing site deep within the oxidase structure may block O2 access to the cytoplasm.
Clearly though, unanswered questions remain regarding the details of the mechanisms, and other factors (intracellular redox state, ATP provision for nitrogenase, and O2 control of nitrogenase) might be important. Note, however, that “uncoupled” respiration (i.e., respiration that is not coupled to PMF generation) is not an essential requirement of the proposed mechanism, which is only that cytochrome bd-supported oxygen uptake uniquely provides the protection. Other mechanisms probably play roles: thus, the alginate capsule of A. vinelandii is affected by oxygen tension and could create a barrier to the entry of oxygen (279). However, the genetic data do appear to provide the evidence required by Oelze (238) for the hypothesis of respiratory protection.
B. An oxygen-reactive oxidase in anaerobes?
Anaerobic microbes are sometimes defined as those that cannot grow at dissolved oxygen concentrations greater than 5 μM; however, many “anaerobes” survive at such levels. B. fragilis colonizes the colon, even in the absence of facultative anaerobes that could maintain low oxygen concentrations, and grows at oxygen concentrations around 300 nM. B. fragilis encodes a cytochrome bd-type oxidase and consumes oxygen at appreciable rates, but a ΔcydAB mutant was defective in oxygen uptake (13). In this organism, and in many prokaryotes that have been classified as strict anaerobes, cyd genes are widely distributed; these bacteria include Methanosarcina, Archaeoglobus, Moorella, and Geobacter species (13, 92, 205). The term “nanaerobes” has been coined to describe such bacteria that can benefit from, but do not require, oxygen for growth (13).
In the case of facultative anaerobes such as E. coli, survival in low-oxygen environments, such as the mammalian intestine, depends on the organism's respiratory flexibility and in particular the presence of cytochrome bd. Mutants lacking cytochrome bd fail to colonize (159, 160). Expression of genes in E. coli that encode the oxidases and other respiratory chain complexes has been widely studied, and recently in the context of systems biology [for reviews see Bettenbrock et al. (30) and Ederer et al. (104)].
In brief, there occurs a progressive switch to aerobic respiratory metabolism and a remodeling of the cell envelope as oxygen availability increases (276, 319). Maximal levels of cytochrome bd occur at intermediate levels of oxygen supply, that is, at a point approximately midway between anaerobiosis and the onset of aerobic metabolism as inferred from acetate excretion during glucose metabolism (276). In contrast, the cyoABCDE operon was maximally expressed under fully aerobic conditions: changes in abundance of the cydAB and cyoA–E transcripts were reflected cellular contents of cytochrome bd and cytochrome bo3 , respectively, consistent with the previously measured oxygen affinities. Cytochrome bd-II is also operative under limiting oxygen conditions (309).
Oxygen reduction is widespread in sulfur-reducing bacteria including some Desulfovibrio species (91) and supports chemiosmotic energy conservation (100). Although classified as an anaerobe, D. gigas contains a functional membrane-bound respiratory chain, including a canonical cytochrome bd quinol oxidase as its terminal enzyme (202, 215). Lemos et al. (202) demonstrated that membranes from D. gigas reduce oxygen to water and isolated a two-subunit oxidase of the cytochrome bd family, with spectral properties similar to other such oxidases. With NADH or succinate as electron donors, specific oxygen uptake activities were comparable with those of aerobes. Surprisingly, the expression levels of the oxidase were unaffected by 60 μM O2, but modest upregulation was noted in the presence of 150 μM NO (215).
Other established anaerobes such as D. vulgaris respond to 0.1% oxygen exposure at the transcriptomic and proteomic levels (231). The genome sequence (140) indicates the presence of two oxidases: a cytochrome c oxidase and cytochrome bd. Both were confirmed by hybridization experiments with CoxA and CydA probes and further sequence analysis (288). The presence of cydAB genes in D. vulgaris was later confirmed and this oxidase shown to be more highly expressed than the cox operon encoding a cc(o/b)o 3-type oxidase (199). The oxygen affinity of the bd-type oxidase (Km = 600 nM) was measured polarographically (268); however, such measurements probably overestimate the Km value (88).
What are the roles of such oxidases in “anaerobes”? Plausible hypotheses include the following: (i) scavenging of O2 for protection against the damaging effects of oxygen, (ii) ATP gain for survival aerobically in changeable habitats, and (iii) catalyzing a rapid, “uncoupled” electron transfer and burning excess reducing substrates. These functions may have long histories: phylogenetic analyses suggest that the Aquificae phylum is one of the earliest diverging phyla of Eubacteria for which sequence data are available and indicate that cytochrome bd was present in the most ancient Eubacteria (13). This is consistent with the view that sufficient oxygen to support respiration predated the photosynthesis-derived appearance of abundant oxygen on Earth.
C. Environmental stressors and their relationships with cytochrome bd
1. Peroxide
Cytochrome bd may contribute to protection against H2O2-induced stress. Mutant E. coli cells lacking cytochrome bd (126, 206, 329) showed high susceptibility to H2O2, and the same was shown for some pathogenic bacteria (14, 295). Consistently, addition of exogenous H2O2 (206) or endogenous production of ROS increased cytochrome bd expression (67). This protective role of cytochrome bd is not exclusive to E. coli [see Giuffre et al. (124) and references therein]. Hypersensitivity to H2O2 or enhanced ROS production has been described also for other bacteria deficient in this oxidase (106, 340), including bacterial pathogens that inhabit microaerobic environments and are exposed to the ROS produced by the host immune system (107, 201, 212) or as a result of antibiotic treatments (212).
Accordingly, upon exposure to H2O2, an upregulation of cytochrome bd was documented in Staphylococcus aureus (66) and in Mycobacterium tuberculosis, where a catalase-independent hyper-resistance to H2O2 was also observed (305). When anaerobic cultures of an E. coli strain devoid of the antioxidant enzymes KatG, KatE, and Ahp are abruptly aerated, cytochrome bd is able to reduce intracellular H2O2 production (191), suggesting that the oxidase serves as an electron sink by diverting electrons from a fumarate reductase, a major H2O2-generator.
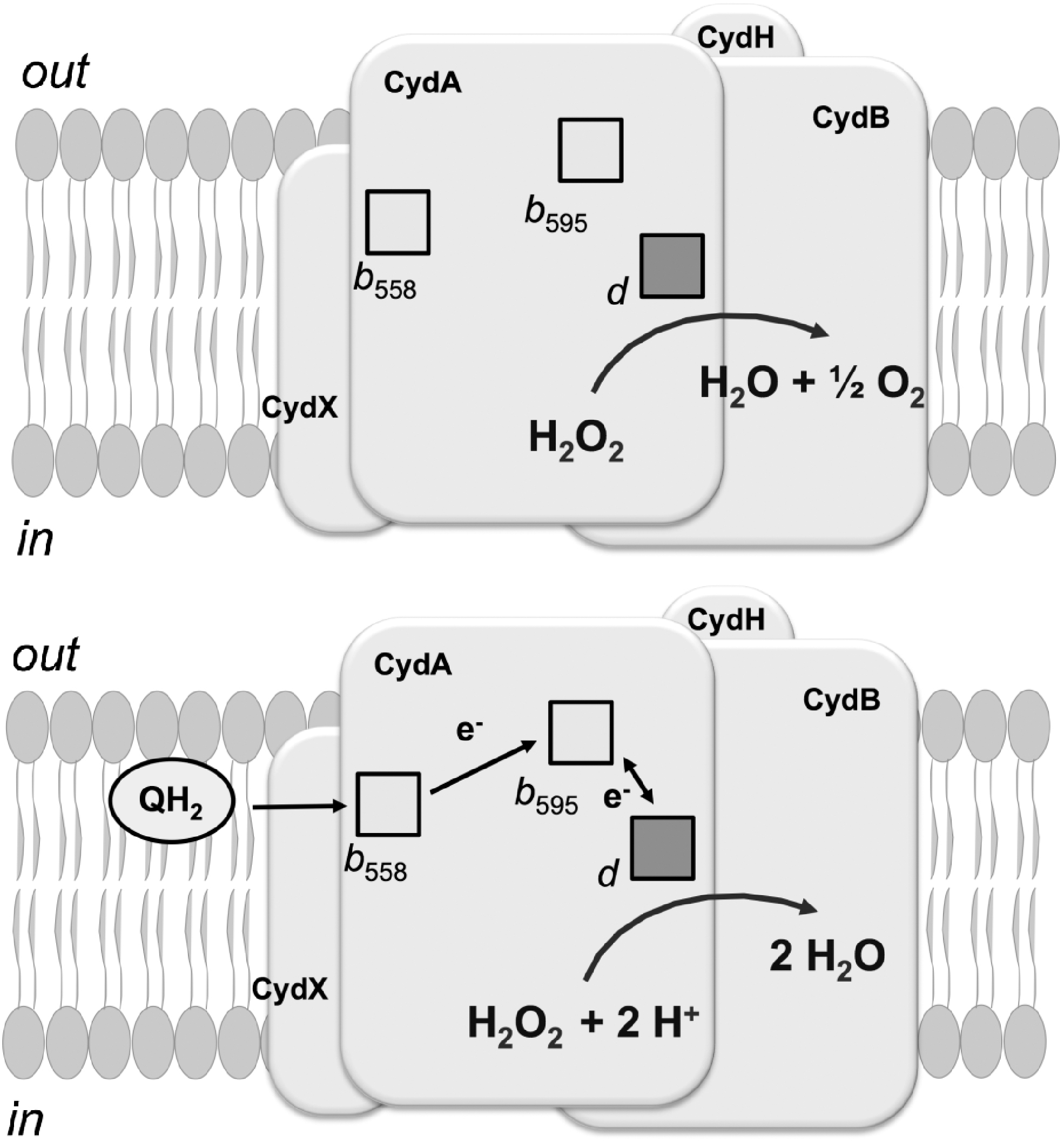
Moreover, E. coli cytochrome bd-I was shown to be capable of detoxifying H2O2 directly (Fig. 16). A high catalase activity was observed in both the isolated untagged bd-I enzyme and in catalase-deficient cells overexpressing cytochrome bd-I (40, 114). Cytochrome bd-I also shows peroxidase activity (1, 39, 182), particularly in the His-tagged enzyme with decyl-ubiquinol as the electron donor (1). The latter preparation, however, displays no catalase activity (1). The issue is discussed in Forte et al. (117). It remains to be established whether the H2O2-metabolizing ability is unique to the E. coli enzyme or is a common property of bd-type oxidases.
2. NO and ONOO−
NO and the product of its reaction with superoxide radical (O2 •−), ONOO−, are produced by the host as part of the immune response to kill invading microbes. Cytochrome bd is involved in protection of bacteria against stress caused by these RNS. Transcriptional upregulation of genes encoding cytochrome bd has been observed in response to NO exposure in E. coli (150, 265) and in other bacteria (215, 229, 273, 296). NO induces greater growth inhibition in cytochrome bd-deficient E. coli strains, compared with cytochrome bo3 -deleted mutants (217). The effect is observed in mutants of both the cytochrome bd-encoding cydAB genes and the cydDC genes (146).
Similar observations were made in cytochrome bd mutants of uropathogenic E. coli (14, 295). Cytochrome bd also protects Salmonella enterica against NO toxicity thereby enhancing its virulence (161). Like cytochrome c oxidase (289, 290), cytochrome bd is potently but reversibly inhibited by NO [IC 50∼0.1 μM NO at 70 μM O2 (42)]. Such inhibition was shown both with E. coli cells (217, 310) and with cytochrome bd isolated from E. coli or A. vinelandii (42). NO reacts with haem d: when haem d is ferrous, oxy-ferrous, or ferric, the end-product is nitrosyl (41, 42, 44), whereas reaction of the ferryl haem d with NO yields nitrite (43) (Fig. 17). Due to an unprecedentedly high off-rate of NO from the ferrous haem d, reversal of cytochrome bd inhibition is much faster than with cytochrome c oxidase (42, 44, 217), explaining why cytochrome bd confers NO resistance to bacteria.
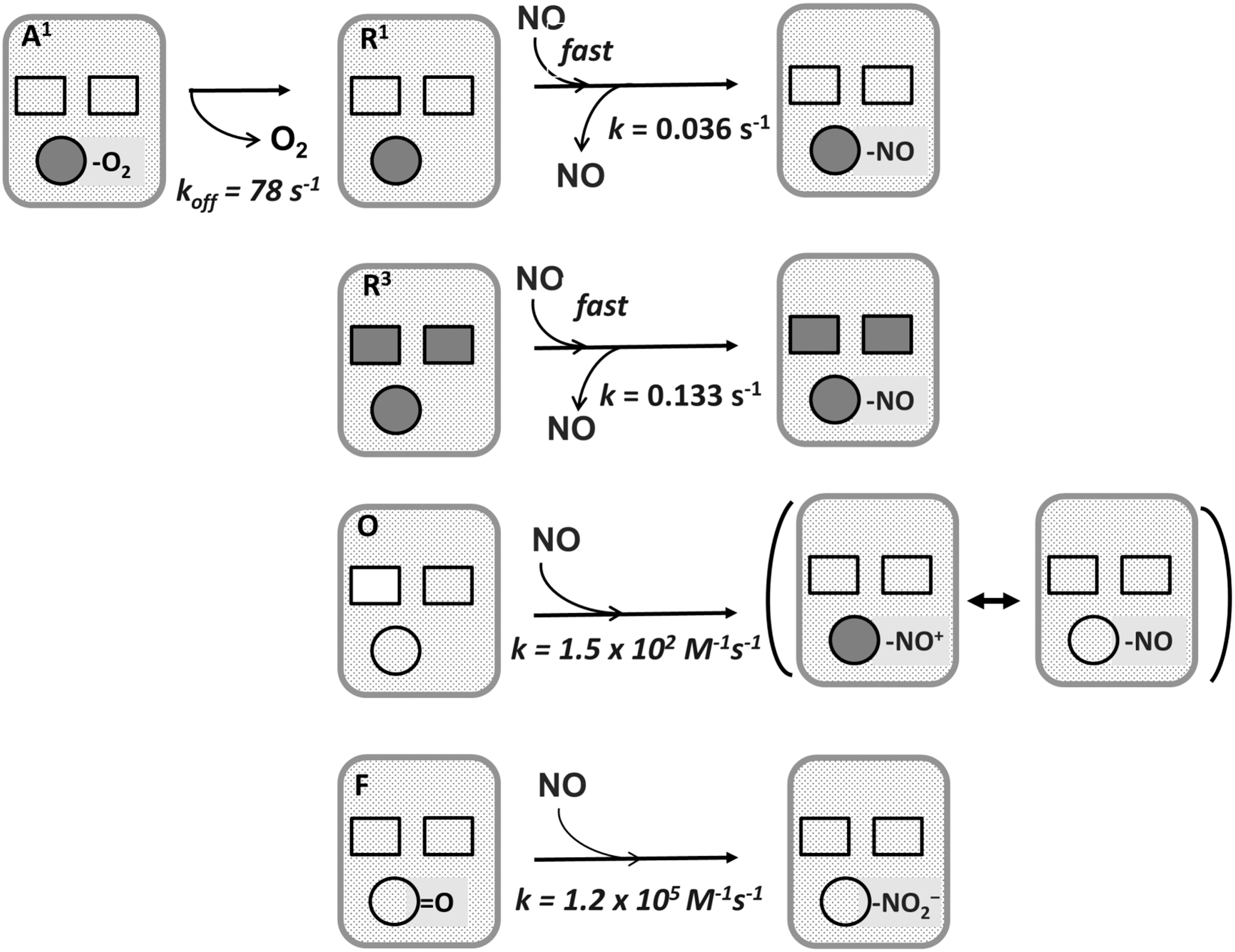
A further reason could be that ferryl cytochrome bd, a highly populated intermediate in turnover (45), converts NO into less toxic nitrite at a rate ∼10 times higher than that of the analogous reaction of the ferryl cytochrome c oxidase (43).
ONOO− was reported to irreversibly inhibit cytochrome c oxidase (294), whereas E. coli cytochrome bd-I is highly resistant to inhibition (47). Furthermore, cytochrome bd-I can metabolize this harmful RNS with an apparent turnover rate of ∼10 mol ONOO− (mol enzyme)−1 s−1, thereby playing a protective role against ONOO− damage.
3. Sulfide
Sulfide potently inhibits cytochrome c oxidase, leading to energy depletion and cell death (76, 235, 244, 315). As many bacteria synthesize H2S and inhabit sulfide-rich environments, such as the human colon, they may be endowed with a sulfide-insensitive oxidase, other than cytochrome c oxidase. This hypothesis was tested on E. coli reaching the conclusion that such an oxidase is cytochrome bd (115, 192). Forte et al. found that whereas sulfide is a potent inhibitor of cytochrome bo3 (IC 50 ∼ 1.1 μM), both cytochrome bd-I and cytochrome bd-II of E. coli are insensitive to sulfide up to 58 μM (115).
Furthermore, in E. coli mutants, O2 respiration and growth are impaired by sulfide when respiration is sustained by cytochrome bo3 alone, but unaffected by up to 200 μM sulfide when bd-I or bd-II acts as the only terminal oxidase. Similarly, Korshunov et al. reported that in a cytochrome bd-deficient mutant, both O2 respiration and growth are inhibited by sulfide, exogenously administered or endogenously generated from cysteine (192). The sulfide insensitivity of cytochrome bd is possibly due to the lack of a copper site, since inhibition of cytochrome c oxidase by sulfide is thought to involve a transient binding of H2S to CuB (235).
Based on these data, it was postulated that the sulfide tolerance provided by cytochrome bd can play a role in shaping the composition of human intestinal microbiota, thus impacting human physiology and pathophysiology (62). Interestingly, upregulation of the cytochrome bd-I and bd-II genes was observed in E. coli cells following addition of an H2S donor, alone or in combination with the antibiotic ampicillin, indicating that expression of the sulfide-insensitive oxidases enables bacterial respiration under antibiotic-induced oxidative stress (298). Indeed, E. coli mutant strains lacking bd-type oxidase fail to colonize the mouse intestine, contrary to those lacking bo 3-type oxidase (159).
4. Chromate
Cytochrome bd has been recently shown to contribute to chromate resistance in Alishewanella sp. WH16-1 (340), a facultative anaerobic bacterium isolated from the soil of a copper and iron mine. This strain efficiently reduces sulfate to sulfide and the toxic hexavalent chromate Cr(VI) to the much less toxic Cr(III), thus showing a great potential for chromate bioremediation. Cr(VI), mainly produced by human activities, is considered a severe pollutant and a serious threat to human health, being mutagenic and carcinogenic. The high toxicity of Cr(VI) is linked to its ability to enter the cells and exert a strong oxidizing power. The structural similarity with sulfate allows Cr(VI) to cross the cellular membrane via sulfate transporters (80, 326) and, once in the cell, to generate ROS (306, 326).
Cytochrome bd confers bacterial resistance to chromate by decreasing chromate-induced cellular oxidative stress and allowing sulfide-dependent chromate reduction (340). In Alishewanella sp. WH16 WT and cytochrome bd mutant strains, the addition of millimolar K2CrO4 resulted in a higher H2O2 production in cytochrome bd-deficient cells compared with the WT or the cytochrome bd-complemented strain, suggesting a role for Alishewanella cytochrome bd in H2O2 detoxification, as also shown for the E. coli oxidase (1, 40).
Furthermore, in H2O2 inhibition zone tests, the mutant strain was more sensitive to exogenous H2O2 than was the WT. Since sulfide can be used as a reductant to reduce chromate to less toxic forms, the effect of Na2S on cell growth was also investigated. Whereas growth of the WT cells was unaffected by the presence of 200 μM sulfide in the culture medium, the cytochrome bd-deficient strain was completely inhibited under these conditions. Thus, the cytochromes bd of both Alishewanella Sp. WH16-1 and E. coli (115) contribute to resistance to sulfide and, indirectly, also to chromate. Summing up, cytochrome bd contributes to the remarkably lower minimum inhibition concentration (MIC) for chromate, as well as the ability of WT strains to reduce chromate to less poisonous forms (340).
VII. Antibiotics and Antimicrobial Agents
A. Cationic amphiphilic peptides with antimicrobial activities
Cationic amphiphilic peptides with antimicrobial activities can be divided into two major classes: nonribosomally and ribosomally synthesized peptides (134, 237). Both classes typically further undergo maturation and chemical modifications. Nonribosomally synthesized peptides are produced only in bacteria and fungi. The biosynthesis of these peptides proceeds on multifunctional peptide synthetases in an RNA-independent manner following the multiple carrier thiotemplate mechanism (308). The synthetases have a modular organization, and these modules interact in an ordered manner to produce the peptide product (327). Gene-encoded ribosomally synthesized peptides are produced by all species and can be further subdivided into bacteriocins (produced by bacteria), archaeocins (produced by Archaea), and antimicrobial peptides (produced by eukaryotes) (216, 237).
Bacteria use cationic amphiphilic peptides to kill or limit the growth of competitors that occupy the same ecological niche. In higher organisms, antimicrobial peptides are part of the innate immune response. The peptides are thought to kill bacteria primarily due to their interaction with bacterial membranes or cell walls. Permeabilization of the microbial membrane lipid bilayer is a commonly accepted mechanism of their action, since the peptides usually have a net positive charge and a high ratio of hydrophobic amino acids. These properties enable the peptides to selectively bind to negatively charged bacterial membranes leading to nonenzymatic membrane disruption (347).
However, a growing body of evidence suggests that cationic amphiphilic peptides can have other molecular targets. One such target could be a bd-type terminal oxidase. Indeed, E. coli cytochrome bd-I is targeted by several peptides tested, such as gramicidin S, cathelicidin LL-37, and microcin J25 (MccJ25) (Fig. 18). Thus, cytochrome bd-I would hardly contribute to bacterial defense against these peptides but rather serves as a mediator of their action.
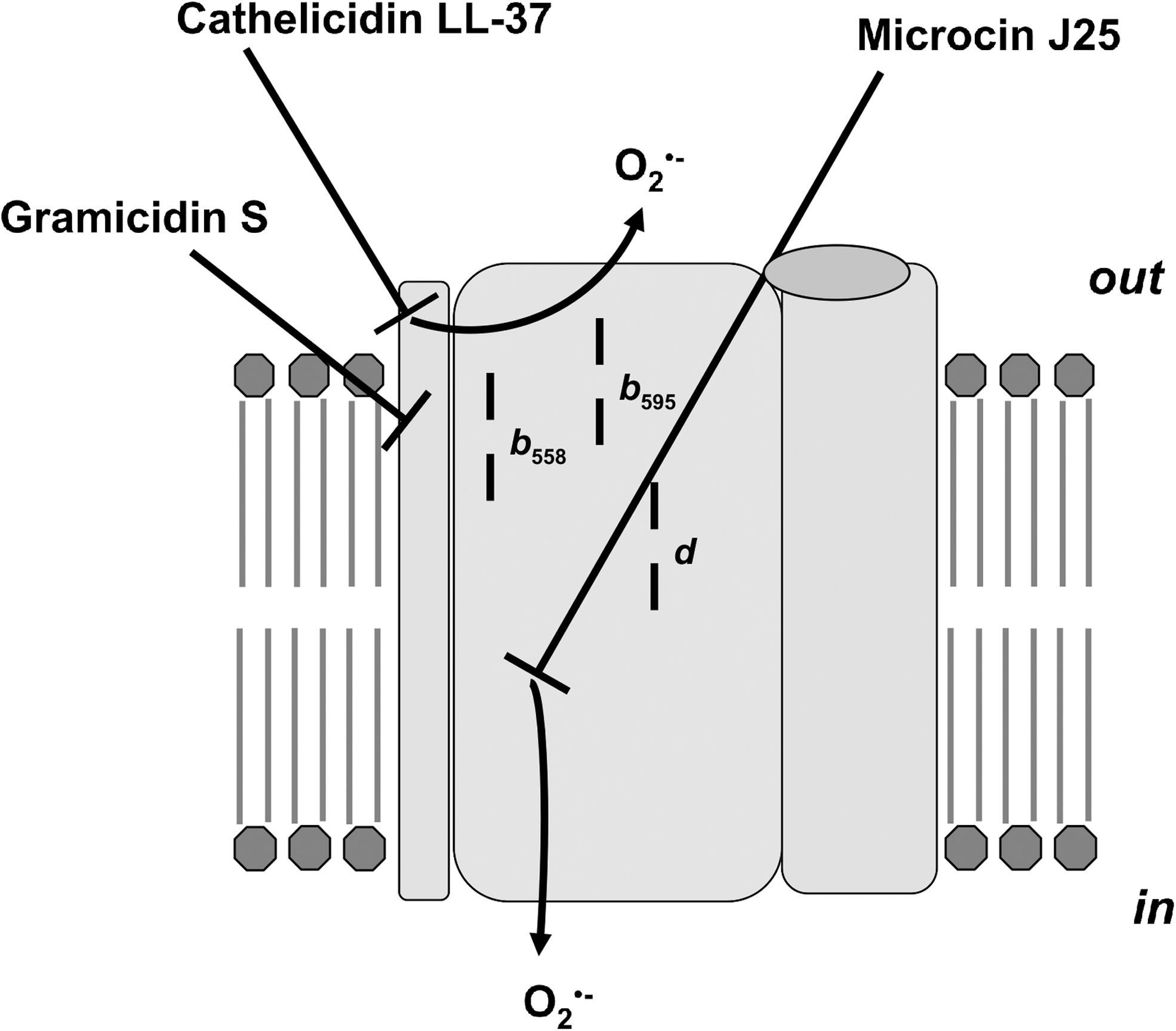
1. Gramicidin S
Gramicidin S (“Soviet”) is nonribosomally produced by the gram-positive soil bacterium Aneurinibacillus migulanus. This is a cationic cyclic decapeptide made up of two identical pentapeptides joined head-to-tail with the primary structure [cyclo-(Val-Orn-Leu-
It was shown recently that gramicidin S is a potent inhibitor of both membrane-bound and purified E. coli cytochrome bd-I (228). On the contrary, the naturally produced mixture of gramicidins A, B, and C, denoted collectively as gramicidin D, which are linear pentadecapeptides forming dimeric cationic channels, does not inhibit cytochrome bd. E. coli cytochrome bo3 and cyanide-insensitive quinol oxidase (CioAB) from Gluconobacter oxydans appeared to be one-order of magnitude less sensitive to gramicidin S. Although submicromolar concentrations of gramicidin S moderately stimulate the ubiquinol-1 oxidase activity of cytochrome bd-I, at higher peptide concentrations, cytochrome bd-I is inhibited. Upon preincubation of cytochrome bd-I with gramicidin S, IC 50 values are in the range of 2.6–5.4 μM.
These values are consistent with the minimum inhibitory concentration (MIC) of E. coli cells (4–16 μM) (190), as well as IC 50 values for 2-heptyl-4-hydroxyquinoline N-oxide (1 μM), antimycin A (5 μM), and piericidin A (10 μM), quinol oxidation site inhibitors (227, 228). This is thought to be mixed-type inhibition, decreasing the apparent V max and the affinity for substrates (228). Gramicidin S does not affect the spectral properties of cytochrome bd-I in the air-oxidized and fully reduced states. The authors conclude that the E. coli cytochrome bd-I is a bacterial membrane target for the peptide, whereas according to current thinking, the principal target of gramicidin S is the lipid bilayer rather than any membrane protein. The underlying mechanism of inhibition may be an alteration of the cytochrome bd-I structure via binding to its hydrophobic surface.
2. Cathelicidin LL-37
Cathelicidins, along with defensins, are the two main families of ribosomally synthesized antimicrobial peptides, effector molecules of the innate immunity of mammals (318). Cathelicidins have a highly conserved N-terminal cathelin domain and a variable C-terminal antimicrobial domain. The latter can be released from the precursor peptide after protease cleavage. LL-37 is an active 37-residue α-helical C-terminal domain of the only human cathelicidin identified to date. The primary structure of LL-37 is LLGDFFRKSKEKIGKEFKRIVQRIKDFLRNLVPRTES (158). This 18-kDa propeptide is denoted as hCAP-18 (human cationic antimicrobial protein).
Recently, it was reported that cathelicidin LL-37 shows a fourfold lower MIC against E. coli in aerobic growth compared with under anaerobic or fermentation conditions (4 vs. 16 μM) (69). Thus, the MIC data suggest that the inhibition of E. coli growth by LL-37 is mediated by oxygen. In aerobic growth conditions, LL-37 induces oxidative stress within seconds of contact with E. coli. To monitor oxidative stress, real-time, single-cell fluorescence imaging with the dyes CellROX Green and Amplex Red was used. ROS formation occurs after entry of LL-37 into the periplasm, but before permeabilization of the cytoplasmic membrane. Entry of the peptide into the periplasm gives it access to the outer leaflet of the cytoplasmic membrane and to external surfaces of cytoplasmic membrane proteins. Strong oxidative signals require a robust transmembrane potential and correlates with the lower MIC in aerobic, compared with anaerobic, growth conditions.
These observations imply that LL-37 affects the aerobic respiratory chain, disrupting electron flow. In aerobic E. coli cultures, the peptide-induced oxidative signals are attenuated in the cytochrome bd-I mutant but not in a cytochrome bo3 mutant (69). This suggests that LL-37 targets cytochrome bd-I, but not cytochrome bo3 , leading to release of ROS, most likely O2 •−, into the periplasmic space. The LL-37 action may be based on its direct interaction with cytochrome bd-I, for instance, due to electrostatic binding. Alternatively, being polycationic (net charge +6), LL-37 may impair cytochrome bd-I function indirectly by perturbation of the lipid environment, for example, via strong interaction with the annular anionic phospholipids such as cardiolipin or phosphatidylglycerol. The authors (69) propose that the host can use the degree of tissue aeration for selective control of the potency of LL-37 and possibly other antimicrobial peptides.
3. Microcin J25
Microcins are ribosomally synthesized bacteriocins produced by E. coli and related enterobacteria. MccJ25 belongs to class I microcins. This is a plasmid-encoded, posttranslationally modified, 21-residue lasso-peptide. The peptide is composed of the N-terminal macrolactam ring and the C-terminal tail. The ring consisting of eight amino acid residues is formed by a lactam bond between the first and eighth amino acids. The ring is threaded and trapped by the C-terminal tail of 13 residues forming a lasso (271). MccJ25 targets essentially foodborne pathogens, such as E. coli, Shigella flexneri, and Salmonella enterica serotypes Newport, Enteritidis, Heidelberg, and Paratyphi B (118).
To cross the outer membrane of a sensitive bacterium, MccJ25 uses the Trojan horse strategy. The peptide is thought to hijack FhuA, an outer-membrane transporter for Fe3+ chelated to the siderophore ferrichrome (285). The process requires a PMF that is transduced to FhuA by the inner membrane-anchored TonB–ExbB–ExbD complex. When in the periplasmic space, MccJ25 is proposed to interact with the inner membrane protein SbmA to cross the inner membrane (286). Once in the cytoplasm, MccJ25 inhibits bacterial transcription via interaction with RNA polymerase (RNAP). The peptide binds to the RNAP secondary channel that directs ribonucleotide precursors toward the active site, thereby blocking its operation in a cork in-the-bottle manner (271).
The other targets of MccJ25 are respiratory chains of sensitive bacteria. Although the inhibition mechanism is not yet completely clear, it was reported that the peptide can inhibit the activity of respiratory chain complexes and enhance O2 •− production (21).
Recently, Galvan et al. examined the effect of MccJ25 on the properties of the E. coli terminal oxidases using bacterial cells, membranes (119), and the isolated enzymes (118). It was shown that E. coli mutant strains lacking cytochrome bd-I or cytochrome bo3 , in which the entry of MccJ25 is facilitated by overexpressing FhuA, are less sensitive to MccJ25 than the WT (119). The MIC of MccJ25-GA (a variant of MccJ25 in which the C-terminal carboxyl group is blocked with an L-glycine methyl ester that leads to the loss of its ability to interact with RNAP) for an E. coli cytochrome bd-I mutant increases 16-fold compared with WT. In the presence of MccJ25, membranes of the E. coli mutant expressing cytochrome bd-I as the only respiratory oxidase show significant ROS overproduction. The ROS overproduction observed with the membranes, however, is not seen with whole cells, possibly because detoxifying enzymes in the cytosol rapidly quench the ROS produced.
Membranes of the E. coli mutant expressing cytochrome bo3 as the only respiratory oxidase show no ROS overproduction in the presence of MccJ25. Galvan et al. concluded that MccJ25 affects E. coli cytochrome bd-I making it a major site of O2 •− production. Interestingly, while the Δbo 3 and Δbd-I mutants become less sensitive to the peptide compared with the WT, in the Δbo 3/Δbd-II and Δbd-I/Δbd-II double mutants, recovery of the original, WT-like sensitivity to MccJ25 and MccJ25-GA is observed. This observation allowed the authors to suggest that E. coli cytochrome bd-II may protect the cell from the deleterious effects of MccJ25 and MccJ25-GA (119). The protection could be achieved by ROS detoxification, but further investigation is required.
The inhibitory action of MccJ25 on enzymatic oxygen consumption was studied in detail with the isolated E. coli quinol oxidases, bd-I and bo 3 (118). MccJ25 was found to inhibit the ubiquinol-1 oxidase activity of cytochrome bd-I with an IC 50 of 76 μM. This is accompanied by a decrease in V max, but K m for ubiquinol-1 does not change significantly. Based on these effects of the peptide on the kinetic parameters, the authors proposed that MccJ25 inhibits cytochrome bd-I in a noncompetitive way (118). The influence of MccJ25-GA on the cytochrome bd-I activity appears to be similar to that of MccJ25. In contrast, the cytochrome bo 3 activity is not affected by MccJ25, and V max and K m for ubiquinol-1 with and without peptide are not significantly different. When the isolated cytochrome bd-I is treated with MccJ25 or MccJ25-GA, ROS generation initiated by the addition of ubiquinol-1 increases by 39% or 31%, respectively.
Since superoxide dismutase suppresses ROS production, it was concluded that the generated species is O2 •−. Neither MccJ25 nor MccJ25-GA increases ROS production by the isolated cytochrome bo 3. The addition of MccJ25 to the air-oxidized cytochrome bd-I in the presence of 6 mM KCN induces absorption changes corresponding to the reduction of haem b 558 and the displacement of bound O2 from haem d 2+. Treatment of cytochrome bo 3 with MccJ25 and cyanide causes no spectral change. Thus, the peptide can reduce cytochrome bd-I. This is consistent with the fact that MccJ25 is a redox-active peptide capable of forming a long-lived tyrosyl (Tyr9) radical (64).
However, it is still not clear whether the peptide-induced change in the oxidation state of cytochrome bd-I is related to its inhibitory action. It is possible that the binding of MccJ25 to the bd-I oxidase slows the rate of intraprotein electron transfer, for instance, by affecting quinol oxidation via conformational change in the protein. This could lead to the release of O2 •−, fast enough to compete with electron transfer to the oxygen-reducing site to form H2O. It is clear, however, that to have a significant inhibitory effect on both RNAP and the respiratory chain, the peptide should accumulate to high levels in the sensitive bacterial cell.
B. Other antimicrobial compounds
Cytochrome bd appears to be involved in the defense of mycobacteria against antibiotic-induced stress. Targeting the respiratory chain enzymes is now considered a new promising strategy for fighting multidrug-resistant, extensively drug-resistant, and totally drug-resistant strains of Mycobacterium tuberculosis (11, 72, 136). As for most bacteria, M. tuberculosis has a branched respiratory chain (71, 73). Electrons are transferred from the type II NDH or other dehydrogenases to the MQ pool, and then to O2 through either the bcc-aa 3 supercomplex, formed by cytochrome bcc (a variant of the bc 1 complex) and cytochrome aa 3, or cytochrome bd (11, 72, 136, 152). The PMF thus generated powers of ATP synthesis by the ATP synthase.
1. Bedaquiline
Bedaquiline (BDQ; Sirturo™), a diarylquinoline compound (Fig. 19), was the first drug approved by the FDA and EMA that targets M. tuberculosis energy metabolism. It selectively inhibits the M. tuberculosis ATP synthase by binding to the c-ring rotor of its Fo subcomplex (4, 193, 261). As a result, ATP synthesis is inhibited and intracellular ATP levels decrease significantly (193, 194). The inhibition mechanism is possibly based on the ability of BDQ, upon its localization at the ATP synthase, to create an uncoupled microenvironment by acting as an H+/K+ ionophore that causes dissipation of transmembrane pH and potassium gradients (137). The activity of BDQ appears to be specific for mycobacteria and is not observed with nonmycobacterial strains (4).
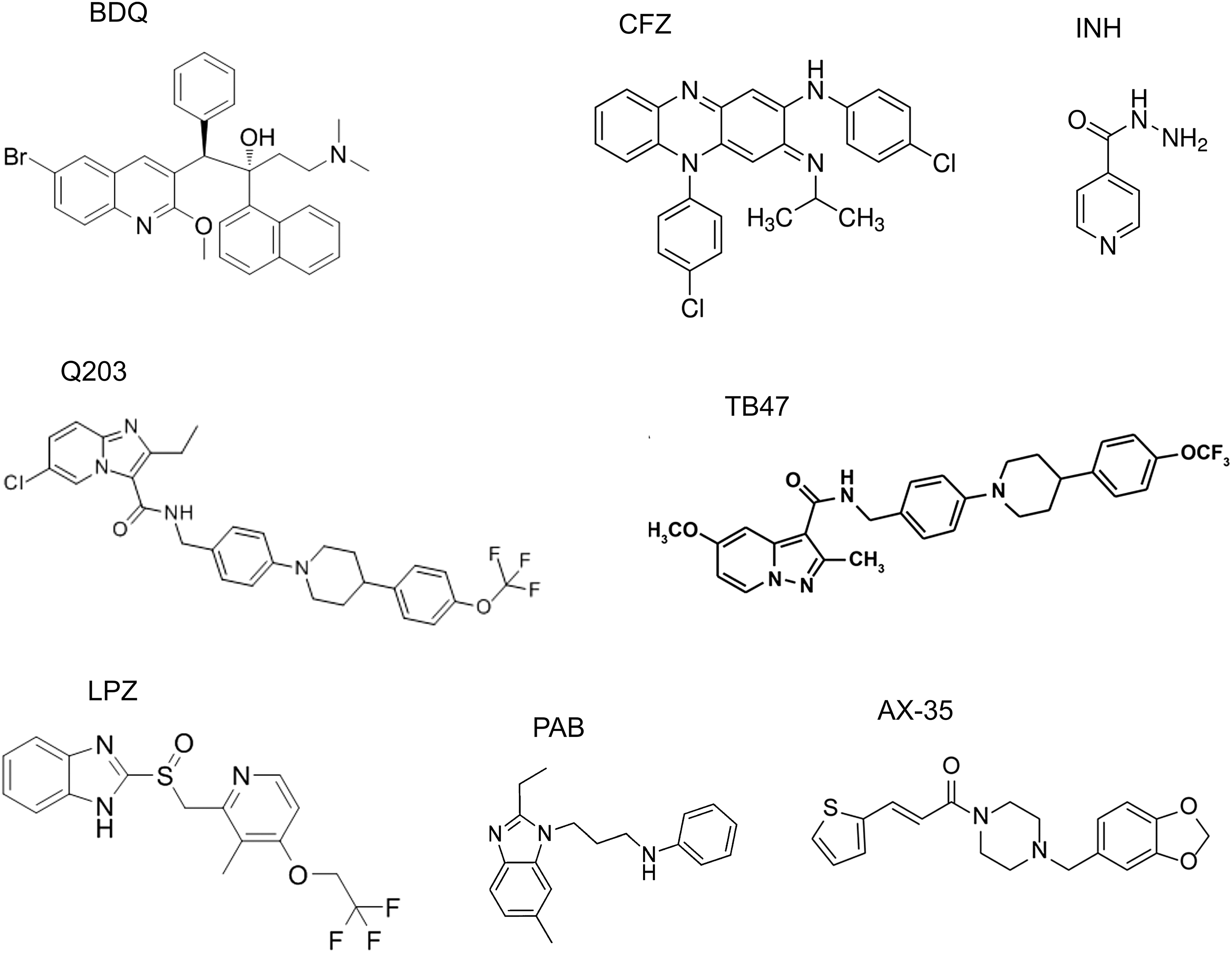
In mammalian cells, the compound affects neither the mitochondrial ATP synthase (133) nor the membrane potential (112). Killing of M. tuberculosis by BDQ is not concentration- but time-dependent, its bactericidal activity showing a delayed onset (195). The rate of M. tuberculosis H37Rv killing, however, is increased by 55% if cytochrome bd is mutated by replacing the cydA gene with a hygromycin cassette via specialized transduction (23).
While BDQ is bactericidal against M. tuberculosis, it is bacteriostatic against the nonpathogenic M. smegmatis (4). If the cydA gene is disrupted, susceptibility of M. smegmatis to BDQ increases (138) and the drug becomes bactericidal (212). Accordingly, both M. tuberculosis and M. smegmatis WT cells treated with BDQ show a marked increase in cytochrome bd expression levels (138, 195).
2. Clofazimine
Figure 19 shows the riminophenazine derivative clofazimine (CFZ) (12), a well-known antileprosy drug. The drug can also undergo reduction with NADH catalyzed by NDH-2. Reduced CFZ is unstable and spontaneously oxidized by O2, producing ROS (344). As NADH is continually produced in the Krebs cycle and fatty acid β-oxidation, the redox cycling of CFZ will continually run. The viability of the M. smegmatis cydA mutant is strongly reduced in response to CFZ compared with WT strains (212).
3. Isoniazid
Isoniazid (INH) (Fig. 19) is a first-line bactericidal antituberculosis drug that inhibits the biosynthesis of mycolic acids, the core components of the mycobacterial cell wall (270). Its activity, however, is reduced significantly under stress conditions, such as low aeration (335) and nutrient starvation (31). This indicates that the bactericidal effects of INH may not be due only to inhibition of the known target. INH rapidly increases ATP levels and oxygen consumption, and dissipates membrane potential in Mycobacterium bovis BCG. The fact that Q203 and BDQ compromise INH-mediated ATP increase and bactericidal activity suggests that a killing mechanism of INH involves perturbation of the mycobacterial respiratory chain (346). Indeed, M. tuberculosis mutants defective in cydC essential for cytochrome bd assembly appeared to be more susceptible to INH in mice (99).
Accordingly, inhibition of cytochrome bd by aurachin D significantly reduces cell recovery in a culture settling model (346). This points to a contribution of cytochrome bd in protection against INH stresses such as hypoxia (346). Interestingly, the rate of ROS production increases in INH-treated mycobacterial cell extracts (297), and disruption of superoxide dismutase A increases INH sensitivity (103). If INH induces ROS by interacting with the respiratory chain, cytochrome bd could protect mycobacterial cells against the drug via ROS detoxification.
Thus, protection against CFZ, BDQ, and INH provided by cytochrome bd may be related to the ability of the oxidase to scavenge and/or prevent generation of ROS [e.g., see Al-Attar et al. (1), Borisov et al. (39), Borisov et al. (40), Forte et al. (114)] induced by the drugs (Fig. 20).
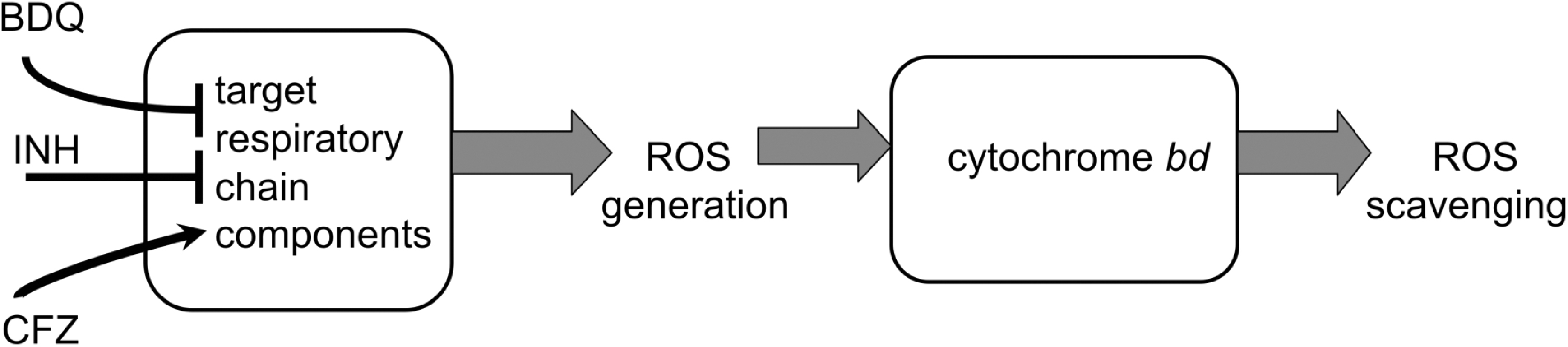
4. Cytochrome bd protection of mycobacteria from compounds targeting cytochrome bcc
Another drug targeting the M. tuberculosis respiratory chain is Q203 (Fig. 19), now designated telacebec by its developer, Qurient Co., Ltd. This imidazopyridine amide (IPA) compound and a clinical-stage drug candidate inhibit cytochrome bcc by binding to the quinol oxidation site (Qp) in the cytochrome b (QcrB) subunit (245). However, Q203 is only bacteriostatic and does not kill the drug-tolerant persisters because, following bcc inhibition, electron flow rerouting to cytochrome bd is sufficient to support sufficiently high oxidative phosphorylation and prevent Q203-induced death (173). Accordingly, while the growth of most M. tuberculosis clinical strains is fully inhibited by IPAs, the laboratory-adapted strains H37Rv, CDC1551, and Erdman can overcome this growth inhibition by upregulating the cydA gene, whose deletion makes the H37Rv strain highly susceptible to IPAs (5). The potency of Q203 is modulated by carbon catabolism and the composition of the culture broth medium (174). It is alleviated by glycerol supplementation that correlates with the overexpression of the cydABDC operon. The cydAB deletion annuls the detrimental effect of glycerol on the efficacy of Q203.
A study in a mouse model of tuberculosis reveals a powerful synthetic lethal interaction between cytochrome bcc and cytochrome bd (173). A synthetic lethal interaction refers to a type of genetic interaction where the single inactivation of two genes affects cell viability only slightly, whereas their simultaneous inactivation causes lethality (349). This also includes the case when the combination of a mutation and treatment with a chemical compound results in cell death, while the mutation or compound treatment alone is nonlethal. Simultaneous inactivation of both mycobacterial respiratory oxidase complexes (via genetic deletion of the cydAB genes and chemical inhibition of cytochrome bcc by Q203) results in complete inhibition of respiration, killing of phenotypic drug-tolerant persisters, and rapid eradication of M. tuberculosis infection in vivo (173). Thus, at least in the mouse lung microenvironment, the ability of M. tuberculosis to multiply and persist is dependent on oxygen respiration.
Mycobacterium ulcerans is known to be the causative agent of Buruli ulcer, a chronic, necrotizing disease that affects the skin and sometimes bone leading to permanent disfigurement and long-term disability. Due to a nonsense mutation in the cydA gene, cytochrome bd is inactive in all classical M. ulcerans strains. This makes cytochrome bcc the only functional terminal electron acceptor in their respiratory chain. As a result, Q203 becomes bactericidal at low dose against these strains both in vitro and in a mouse model. The drug could thus simplify and shorten Buruli ulcer treatment (292).
Figure 19 also shows other compounds targeting QcrB of the M. tuberculosis cytochrome bcc. These are pyrazolo[1,5-a]pyridine-3-carboxamide (TB47) (213), lansoprazole (278), phenoxyalkylbenzimidazoles (29, 65), and the arylvinylpiperazine amide (AX) series AX-35 and four related analogues (AX-36 to AX-39) (113). The compounds are bacteriostatic rather than bactericidal in their action. This is probably due to switching mycobacterial respiration to cytochrome bd that can be upregulated upon compound treatment (113, 278). Accordingly, when cytochrome bd is disrupted, the compounds become bactericidal (29, 113, 213).
Thus, cytochrome bd seems to protect mycobacteria against cytochrome bcc inhibitors by providing an efficient alternative respiratory route (Fig. 21).
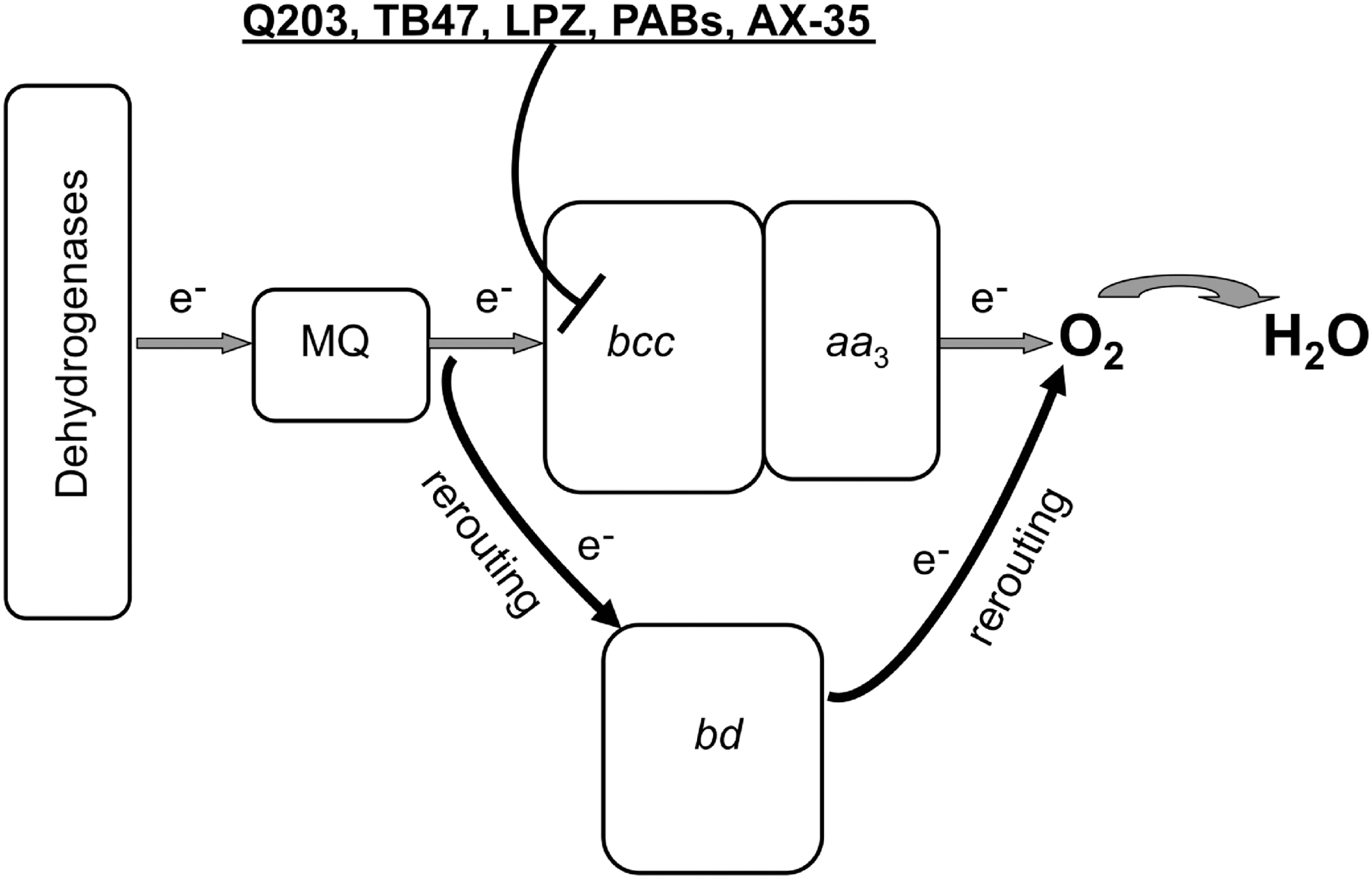
VIII. Cytochrome bd As a Potential Therapeutic Target
During host colonization, pathogenic bacteria face hostile environments, often characterized by harsh conditions, including severe O2 limitation and abundance of harmful species, such as ROS and RNS produced by the host immune system, or high levels of H2S produced by coexisting bacteria. Under these conditions, expression of cytochrome bd is potentially beneficial for pathogens, owing to the ability of this enzyme to function even under O2-limiting conditions, due to its high O2 affinity and resistance to cytotoxic species, such as H2O2, ONOO−, NO, and H2S. Therefore, it is not surprising that, as summarized below, cytochrome bd promotes virulence in several pathogenic bacteria.
In the case of Shigella sp., a well-known foodborne enteric pathogen, a markedly attenuated virulence was observed in mice models when CydAB expression was impaired by mutation of the cydC gene in the same operon. On the contrary, overexpression of CydAB in response to impaired synthesis of ubiquinone or riboflavin resulted in significantly higher bacterial virulence (334).
The role of cytochrome bd in bacterial virulence was also investigated in Group B Streptococcus, a common constituent of the vaginal microflora whose transmission to newborns during delivery can result in life-threatening sepsis. This pathogen, commonly assumed to have a strictly fermentative metabolism, was shown to adopt a quinone- and haem-dependent respiration-based metabolism requiring cytochrome bd (341). Inactivation of cydA in this pathogen was reported to result in reduced growth in human blood and, importantly, attenuated virulence in a neonatal rat sepsis model (341).
Another interesting case is represented by S. Typhimurium, a well-known foodborne pathogen responsible for a broad disease spectrum, ranging from self-limiting gastroenteritis to acute systemic infection in susceptible patients. As in E. coli, this enteric pathogen encodes two bd-type oxidases: cytochrome bd-I and cytochrome bd-II, respectively, encoded by cydAB and cyxAB. Whereas cytochrome bd-II was found to be essential for intestinal colonization following antibiotic treatments depleting the gut of Clostridia species (275), cytochrome bd-I appears indispensable for sustaining tissue infection in mice models (85). Interestingly, the role of cytochrome bd-I in S. Typhimurium virulence was related to the ability of this oxidase to sustain bacterial energy metabolism under O2-limiting conditions and to play a key role in the defence against nitrosative stress together with the NO-detoxifier flavohemoglobin (Hmp) (161).
The importance of cytochrome bd in affording NO-resistant respiration during the infection process was also demonstrated by Shepherd et al. (295) for uropathogenic E. coli (UPEC), a major cause of urinary tract infections, which are the most common nosocomial infections in developed countries. In this study, the relative contribution of the NO-detoxifying systems (Hmp, NorVW, NrfA), the iron/sulfur cluster repair protein YtfE, and the NO-tolerant cytochrome bd-I to UPEC growth and survival during infection was investigated. Cytochrome bd-I was shown not only to confer the highest resistance (together with Hmp) to an NO releaser but also to contribute to resistance to neutrophil-mediated killing and enable survival within macrophages. In addition, unlike the other defence systems (NorVW, Hmp, and YtfE), cytochrome bd-I was found to enhance UPEC survival in a mouse model after two days. On this basis, it was concluded that in UPEC, cytochrome bd-I is the main contributor to NO tolerance and host colonization under microaerobic conditions.
Cytochrome bd was also shown to play a role in the virulence of Listeria monocytogenes (77), another foodborne pathogen responsible for a number of life-threatening infections. This pathogen requires a special ability to adapt to varying O2 levels. It encodes two respiratory terminal oxidases, a cytochrome bd-type (CydAB) and a cytochrome aa 3-type MQH2 (QoxAB) oxidase. Both are used for respiration at different O2 tensions and, importantly, have a role during infection, as mutation of either oxidase results in attenuated infection in mice. However, only the cytochrome bd-type oxidase is essential for aerobic respiration and bacterial replication in HeLa cells.
Less clear is the role played by cytochrome bd during the infection of Brucella spp., the pathogens responsible for brucellosis, a disease characterized by protracted recurrent fever in humans and abortion in ruminants. As a strategy to evade the host defence mechanisms, the Brucella spp. undergoes replication inside host macrophages, facing O2-limiting and hyperoxidative conditions. In an early study on B. abortus, cydB mutants were found to display higher sensitivity to killing in murine macrophages and attenuated virulence in a mouse model, reverted by genes encoding superoxide dismutase or catalase (107). In a later report on Brucella suis, cytochrome bd was found to be the preferentially used terminal oxidase inside macrophages and to contribute to intracellular replication (208). However, in a subsequent study by the same group, lack of cytochrome bd was found to lead to bacterial hypervirulence (not attenuation) in a mouse model (157).
The role of cytochrome bd in microbial virulence was thoroughly investigated also in mycobacteria. M. tuberculosis can survive, but not proliferate, under O2-limiting conditions. As reviewed in Cook et al. (73), this pathogen has a versatile respiratory metabolism enabling bacterial survival at the significantly varying O2 tensions encountered during host infection. The respiratory electron transfer chain is branched and terminates with two terminal oxidases: a bd-type oxidase (CydAB) and an aa 3-type cytochrome c oxidase that form a supercomplex with bc 1 (denoted as bc 1 -aa 3). As revealed by investigating both M. tuberculosis (23) and the closely related nonpathogenic M. smegmatis (176), in the absence of stress factors, cytochrome bd is dispensable for growth in a rich medium. However, it appears to have a role in mycobacterial virulence, particularly at the transition between acute and chronic infection, when cytochrome bd and a nitrate transporter were shown to be transiently upregulated in a transcriptomic study on a murine infection model (296).
Mutation of cydC resulted in a lower bacterial burden in this transition phase, but not in the acute phase of infection, as observed in the same animal model (296). Although this observation was not reproduced in a later study (173), it is conceivable that cytochrome bd has a role during infection, when M. tuberculosis is expected to encounter several stresses, including limiting O2 tensions, NO, and ROS/RNS produced by the host immune system. In line with this view, NO and hypoxic conditions were reported to enhance cydAB expression in M. tuberculosis and other mycobacteria (22, 33, 79, 127, 176, 296), and knockout of cytochrome bd was found to impair mycobacterial growth at low O2 tensions (176, 296) and in the presence of H2O2 (212). Conversely, compared with the WT, a higher resistance to H2O2 was observed in an M. tuberculosis mutant strain with enhanced expression of cytochrome bd (305).
Altogether, the evidence reviewed above points to cytochrome bd as a prospective target for the development of next-generation antibiotics. However, a thorny path may await us. The flexibility of the Q-loop poses a problem to structure-driven design of quinone substrate-like inhibitors (280). For bd-type oxidases structurally similar to the E. coli bd-I, one could try to design hydrophobic compounds that would block the oxygen channel functioning. One more option is to develop quinol-like inhibitors that would compete with ubiquinol-8 for the CydB binding site, thereby preventing correct assembly of a CydAB core dimer (280).
IX. Possible Biotechnological Applications
Oxidases of the bd type have a broad potential application for biotechnology. This may relate to (i) the monitoring of parameters in biotechnological production processes, (ii) the creation of microbial fuel cells, (iii) the selection of industrial strains of microorganisms resistant to toxins and copper deprivation, (iv) various issues of bioremediation and wastewater treatment, and (v) the creation of conductive systems based on immobilized biofilms on the electrode. Some real and potential applications of cytochromes bd in industrial and medical biotechnology are illustrated by the following examples.
Forced aeration is one of the major energy consumption factors of bioleaching processes. The ability of cytochrome bd to be expressed under low oxygen conditions has been used for the identification and validation of genetic markers associated with oxygen availability in low-grade copper bioleaching systems (78).
Strains of C. glutamicum are used in the biotechnology industry for production of million tons of amino acids, mainly
Extracellular electron transfer (EET), that is, electron movements outside the envelope of microbial cells, is highly relevant to the design of electrochemical systems, construction of fuel cells and biosensors, and understanding the physiology of the gut microbiota. Cytochrome bd activity was shown to attenuate EET in the gram-positive lactic acid bacterium Enterococcus faecalis through its effects on the redox status of the MQ pool. Wiring respiratory electron transfer from a bacterium such as E. faecalis to an electrode could be used as a sensor in screens for antimicrobial compounds acting by inhibition of cytochrome bd (240).
Mining, jewelry, and metal-processing industries use cyanide for extracting gold and other valuable metals, generating large amounts of highly toxic wastewater. There are microorganisms (e.g., Pseudomonas pseudoalcaligenes CECT5344) that can grow under alkaline conditions using cyanide, cyanate, or different nitriles as the sole nitrogen source, and are able to remove cyanide from a jewelry industry wastewater. These bacteria will enable the development and optimization of improved strategies for biodegradation and bioremediation of industrial cyanide- and metal-containing effluents. As cyanide generally inhibits cellular aerobic respiration, bacteria possessing an alternative oxidase insensitive to cyanide, such as the quinol oxidase encoded by the cioAB genes, which belongs to the cytochrome bd family (60).
The study of bd-type oxidases also contributes to the development of means to combat biofilms in food and medical biotechnology. Cytochrome bd, a high-affinity quinol oxidase required for aerobic respiration under hypoxic conditions, is the most abundantly expressed respiratory complex in the biofilm community. It was shown recently that the cytochrome bd-expressing subpopulation is critical for biofilm development (14).
X. Conclusions
Cytochrome bd is found only in bacteria but is widely distributed among taxa and physiological types. However, its current importance and high profile were slow to be recognized: since its discovery by Keilin, Warburg and pioneers in respiratory metabolism in the 1930s (when it was named cytochrome a 2), publications on this oxidase accrued very slowly. Based on the Web of Science Core Collection, the number of articles has grown rapidly: in 1990, there was one article (searching for “cytochrome AND bd”), but now that number is ∼1500 per annum. The explosion of interest is a result of the recognition that cytochrome bd represents a novel class of oxidase and that its unique structure, catalytic capabilities, tolerance of stress, and importance to pathogens are worthy of detailed study.
Despite impressive advances, significant challenges remain. From the point of view of fundamental understanding of redox and antioxidant biology, the most pressing are a full description of electron flux through the three haems and the mechanisms of oxygen reduction. Since cytochrome bd is not a proton pump, although contributing to creation of a protonmotive force, accompanying proton movements are less problematic. However, the novel features of cytochrome bd pose additional challenges, particularly its ability to confer upon bacterial resistance to oxidative stress, NO and a number of important antimicrobial compounds. Its unique structure and function make it a profitable area of study for developing novel therapeutics, thereby contributing to overcoming the global scourge of resistance to established, and overused, antibiotics.
Footnotes
Funding Information
This work was supported by the Russian Science Foundation (project 19-14-00063).
