Abstract
Significance:
The successful adaptation of microorganisms to ever-changing environments depends, to a great extent, on their ability to maintain redox homeostasis. To effectively maintain the redox balance, cells have developed a variety of strategies mainly coordinated by a battery of transcriptional regulators through diverse mechanisms.
Recent Advances:
This comprehensive review focuses on the main mechanisms used by major redox-responsive regulators in prokaryotes and their relationship with the different redox signals received by the cell. An overview of the corresponding regulons is also provided.
Critical Issues:
Some regulators are difficult to classify since they may contain several sensing domains and respond to more than one signal. We propose a classification of redox-sensing regulators into three major groups. The first group contains one-component or direct regulators, whose sensing and regulatory domains are in the same protein. The second group comprises the classical two-component systems involving a sensor kinase that transduces the redox signal to its DNA-binding partner. The third group encompasses a heterogeneous group of flavin-based photosensors whose mechanisms are not always fully understood and are often involved in more complex regulatory networks.
Future Directions:
Redox-responsive transcriptional regulation is an intricate process as identical signals may be sensed and transduced by different transcription factors, which often interplay with other DNA-binding proteins with or without regulatory activity. Although there is much information about some key regulators, many others remain to be fully characterized due to the instability of their clusters under oxygen. Understanding the mechanisms and the regulatory networks operated by these regulators is essential for the development of future applications in biotechnology and medicine.
I. Introduction
Aerobic metabolism provides significant advantages in energy production, detoxification of xenobiotics, and virulence of bacterial cells. However, reactive intermediates produced in the reduction of oxygen by electron transfer systems can damage all cellular components. In addition to these reactive oxygen species (ROS), other by-products of metabolism including reactive nitrogen species (RNS) and xenobiotics challenge cellular redox homeostasis. Among the main targets of ROS during oxidative stress are iron-containing proteins, which become severely damaged due to the ability of H2O2 and O2 − to oxidize the iron present in exposed iron–sulfur clusters and other cofactors (164, 170). Furthermore, the release of Fe2+ from these centers promotes the Fenton reaction, which produces even more reactive hydroxyl radicals with deleterious consequences for cells (394).
As a response to this scenario, bacteria have developed a set of redox-responsive proteins that trigger the appropriate inducible response according to the level of stress. Redox sensing by the cell is a complex process that integrates diverse stimuli such as O2 tension, nutrient availability, light intensity, RNS, and ROS, among other parameters. Transduction of these redox signals is frequently carried out by transcriptional regulatory proteins through a variety of mechanisms (237, 334, 364). Owing to the tight relationship between iron metabolism and redox homeostasis, the activity of many major regulators relies on iron, either as an ion cofactor assembled in iron–sulfur clusters or as heme-based sensors (80, 121, 134, 274, 288, 291).
Other essential metal ions such as zinc or manganese also play important roles in re-establishing the redox balance (71, 234) and act as cofactors in transcriptional regulation (101, 200, 297, 325). Besides their role in zinc metalloregulation, zinc ions often prevent the oxidation of redox-sensitive cysteines that work as thiol-based redox switches in numerous transcriptional regulators (149, 200).
As will be highlighted in further sections, multiple transcriptional regulatory mechanisms as response to oxidants are widespread strategies in most prokaryotes. Another interesting issue is the diversity of mechanisms developed by the cell to detect and detoxify the same signal. Thus, in
To better understand redox regulation beyond the identification of the antioxidant defensive genes, numerous mechanistic studies of the diverse regulators that convert redox signals into regulatory outputs have been carried out in recent decades. In this comprehensive review, we revisit the main groups of redox-responsive transcriptional regulators with a particular emphasis on recent findings concerning the structural and mechanistic basis of their regulatory functions.
II. Direct Redox Sensing and Regulation
Most of the major direct redox sensors monitorize the redox state of the cell through oxidant-sensitive metal–sulfur clusters or nonmetallated thiol-based switches. Other relevant mechanisms include those used by heme-based sensor proteins. A summary of the direct redox sensors discussed in this section can be found in Tables 1 and 2.
Main Properties of Wbl Paralogs from
Summary of Representative Regulators Involved in Direct Redox Sensing and Regulation
FNR, fumarate nitrate reductase regulator; Fur, ferric uptake regulator; MarR,
A. Regulation involving oxidant-sensitive iron–sulfur clusters
Iron–sulfur clusters function as cofactors of a wide range of transcriptional regulators that exploit the redox and coordination properties of these clusters to act as sensors of environmental conditions.
1. SoxR
SoxR is a conserved regulator in Enterobacteriaceae that belongs to the MerR family of transcriptional regulators. MerR homologues share similar N-terminal winged helix-turn-helix (wHTH) DNA binding regions, whereas C-terminal effector binding regions appear specific to the effector recognized (37). Most members of the family respond to stress signals, such as oxidative stress, heavy metals, or antibiotics. SoxR was initially identified as an O2 − stress sensor (221, 370), although further studies showed the activation of SoxR by nitric oxide and a variety of endogenous and xenobiotic redox-cycling agents (267, 412). In addition, SoxR becomes activated through DNA-mediated oxidation by guanine radicals that are produced in the early stages of oxidative stress (217).
In
The mechanism of action of SoxR in enteric bacteria has been extensively investigated. In solution, SoxR is a homodimer that exhibits one [2Fe–2S] cluster per monomer coordinated by the four cysteines in the conserved sequence (CysX2CysXCysX5Cys) near the carboxyl terminus. In the absence of oxidative stress, SoxR with the reduced [2Fe–2S] cluster may bind to DNA but it is inactive for transcription initiation (Fig. 1A).
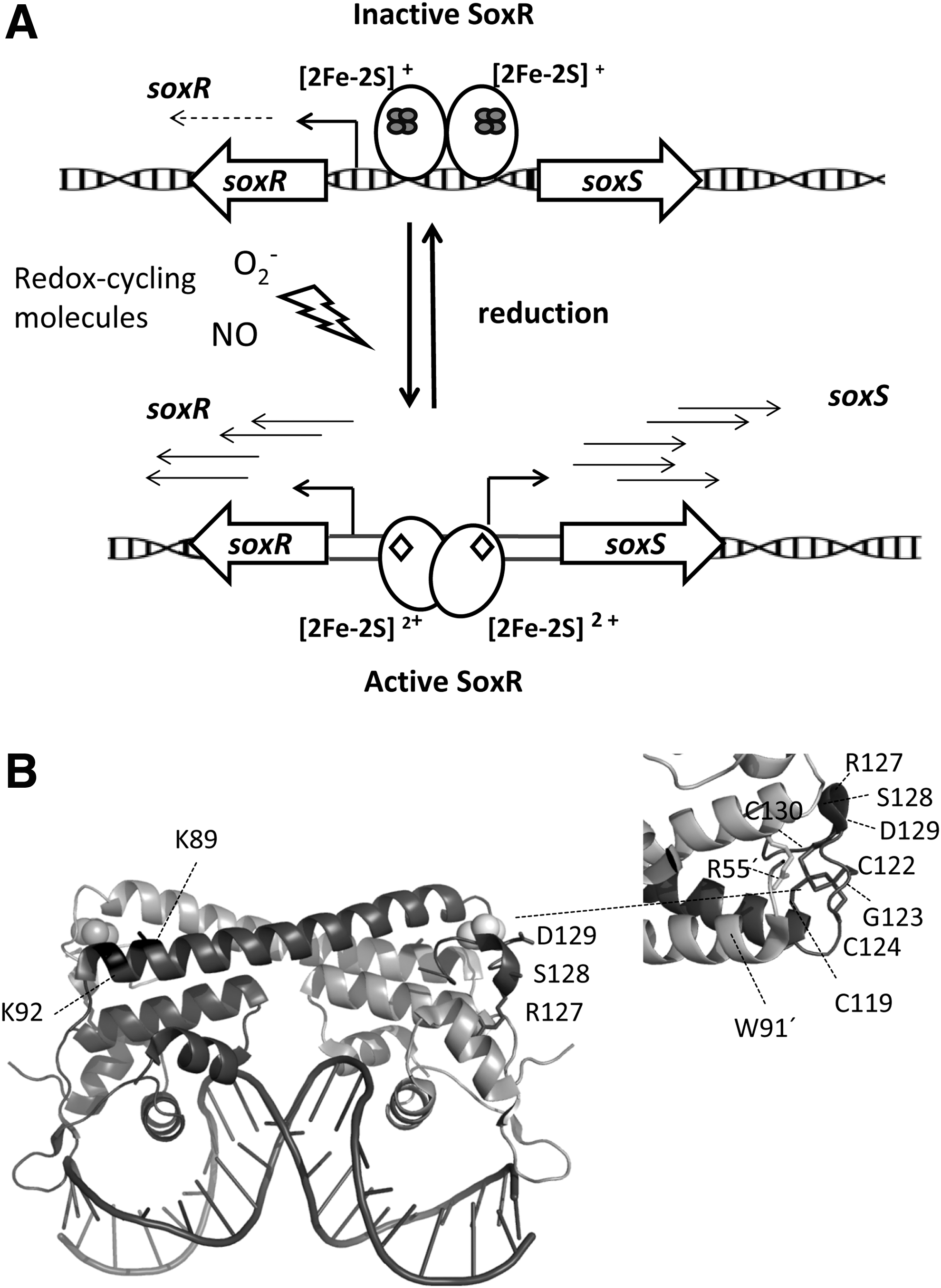
As a MerR-like regulator, SoxR controls the expression of genes whose promoters contain suboptimal 19-bp spacers between their −35 and −10 elements, which are not recognized by the sigma factor of RNA polymerase (RNAP) (37). SoxR activates transcription of its counterpart
Notably, SoxR is the only regulator able to modulate its transcriptional activity undergoing a single change in the cluster redox state (64). The large conformational change of SoxR and the target promoter triggered just by cluster oxidation could be elicited by the remarkable asymmetric environment of the [2Fe–2S] cluster observed in the structural analysis of
Previous electrochemical studies suggested that upon binding to DNA targets, the reduction potential of SoxR [2Fe–2S] has undergone a shift from −285 mV (free regulator) to +200 mV (complexed to DNA) (80). Those results suggested that only strong oxidants would be able to oxidize the cluster to the +2 state and were not in good agreement with the ability of SoxR to react with some redox-cycling drugs and other weak oxidants. However, a recent study using DNA-modified electrodes concluded that DNA binding causes a moderate shift in the reduction potential of SoxR, namely −320 mV of the SoxR bound to DNA
The absence of SoxS in
Another interesting issue is the different selectivity against redox-cycling drugs of SoxR regulators from diverse species. Mutational studies demonstrated that small alterations in the SoxR structure can lead to the evolution of proteins with distinct specificities for redox-active small molecules (335). In addition, a series of physicochemical and mutational studies evidenced the importance of two lysine residues in the vicinity of the [2Fe–2S] cluster, namely Lys89 and Lys92 in
2. IscR
IscR belongs to the Rrf2 family of wHTH transcription factors. Members of the Rrf2 superfamily are widespread in bacteria and respond to different signals, such as nitric oxide (nitric oxide sensitive repressor [NsrR]), iron limitation (RirA), cysteine availability (CymR), or O2 (RsrR) (93, 146, 167, 256). IscR senses the iron–sulfur cluster status in the cell and it is an outstanding example of the integration of redox, sulfur, and iron availability signals.
IscR was first identified in
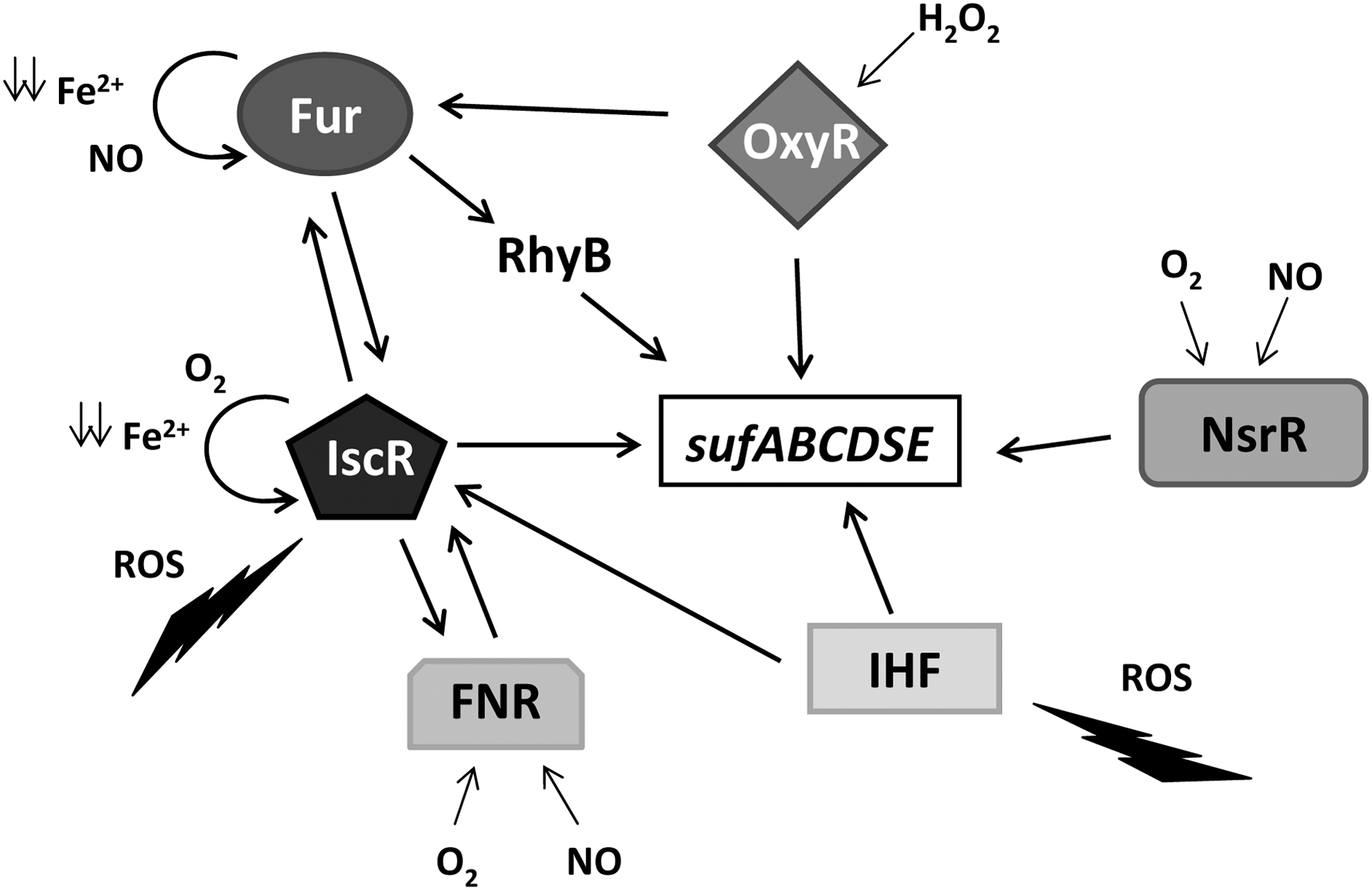
Owing to the prominent role of [Fe–S] clusters in metabolism, IscR is considered a master regulator that controls >40 genes of 20 predicted operons in the
IscR also plays a critical function in
Although some aspects of the molecular mechanism of IscR remain unknown, significant advances in the understanding of IscR–ligand interaction have been made through the characterization of the IscR [2Fe-2S] cluster from
Coordination of the [2Fe–2S] cluster takes place through three conserved cysteine residues at the C-terminus (Cys92, Cys98, and Cys104 in the
The three-dimensional structure of apo-IscR proteins exhibits an overall architecture similar to Rrf2 regulators harboring a wHTH DNA-binding motif and a dimerization domain mainly composed of helix α5 in monomer 1 and helix α6 in the adjacent unit that stabilizes dimer formation mainly by hydrophobic interactions. Analysis of the apo-IscA–DNA interface (Protein Data Bank [PDB] ID: 4CHU), together with sequence alignments of the DNA-binding domains, led to the identification of relevant residues for specific DNA recognition and highlights the role of Glu43 as a selectivity filter in apo-IscR to discriminate against type-1 binding motifs (316). Moreover, the characterization of IscR orthologs from different organisms suggests a high conservation of this unique mechanism of sequence discrimination, unveiling a similar regulation of [2Fe–2S] cluster biogenesis to maintain a perfect balance between favorable and adverse conditions (316).
3. NsrR
NsrR is the master regulator of nitrosative stress response in most β and γ
NsrR is a member of the Rrf2 family that can harbor either a [2Fe–2S] or a [4Fe–4S] cluster, depending on the organism and the purification conditions (167, 371, 407). In
In
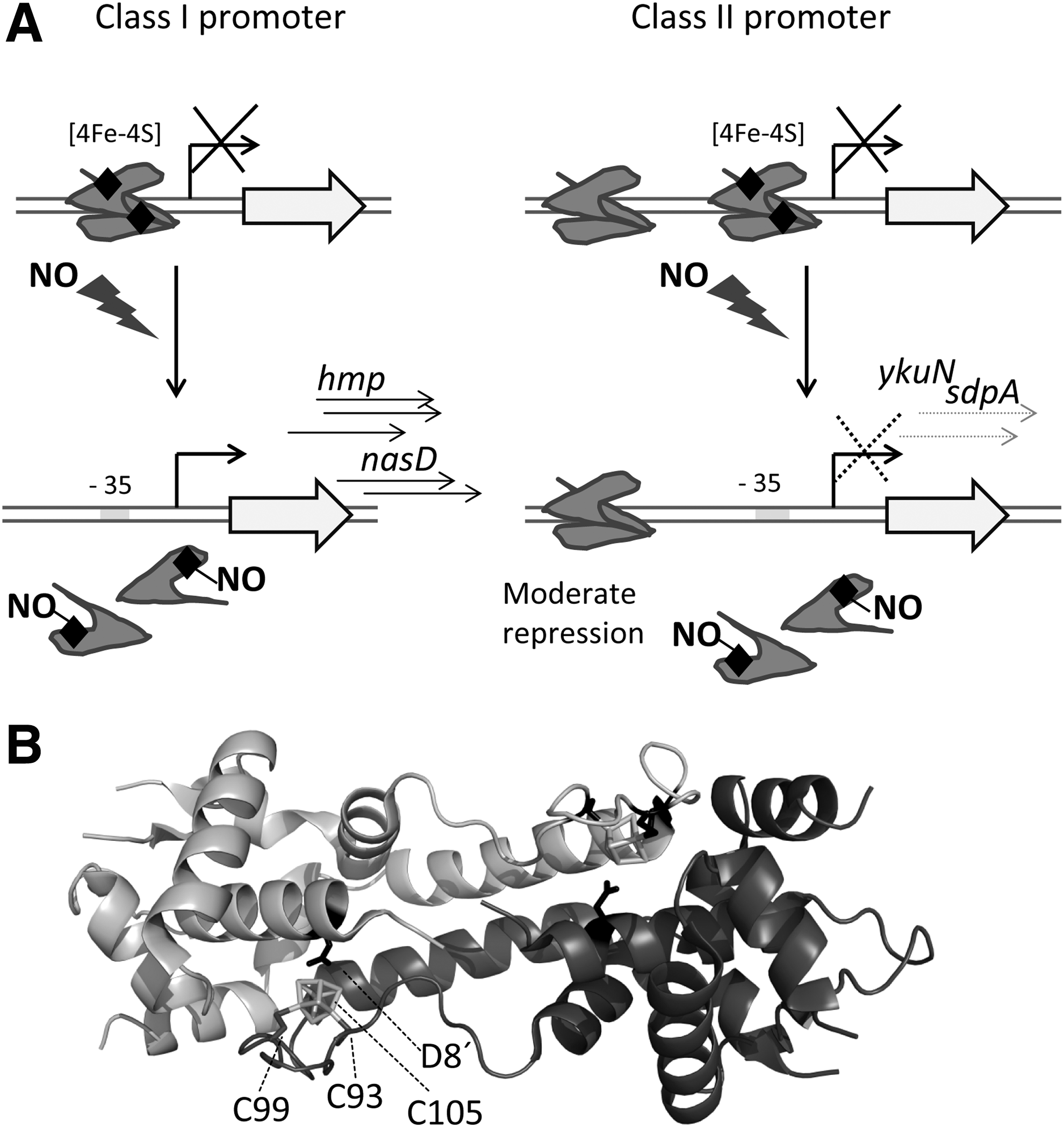
The observation of active NsrR with [2Fe–2S] and [4Fe–4S] clusters has raised some controversy about the physiologically relevant structure of the iron–sulfur cluster in this regulator. Inactivation of aerobically purified [2Fe–2S] NsrR regulators from
Resolution of the crystal structure of [4Fe–4S] NsrR from
4. RsrR
The Rrf2 regulator RsrR (Sven6563) was initially annotated as an NsrR homologue in
RrsR DNA-binding activity is controlled by the status of its [2Fe–2S] cluster that works as a redox switch in a manner similar to SoxR. Under anaerobic conditions, RsrR is a dimer with each monomer containing a reduced [2Fe–2S]+1 cluster that is rapidly oxidized to [2Fe–2S]+2 by O2, increasing
5. FNR
FNR is a widespread sensor of environmental O2 that switches the transition between aerobic and anaerobic respiration. Identified in
In anaerobic conditions, FNR contains one [4Fe–4S]2+ cluster per monomer that in the presence of O2 is rapidly converted into a [2Fe–2S]2+ form through a [3Fe–4S]1+ intermediate, releasing Fe2+ and O2 − with the subsequent DNA damage (63, 64). The conversion of the cubic [4Fe–4S]2+ center to planar [2Fe–2S]2+ requires a series of rearrangements in the proximity of the cluster that, in turn, modifies the dimerization interface, causing dimer dissociation and release from DNA (64). This process may be reverted under low O2 tension. However, if O2 persists in the environment, the [2Fe–2S]2+ clusters are slowly degraded to produce inactive apo-FNR that, depending of the redox status in the cell, will either work as a scaffold for the incorporation of novel [4Fe–4S]2+ clusters or will be degraded by the ClpXP protease (Fig. 4) (104, 245).
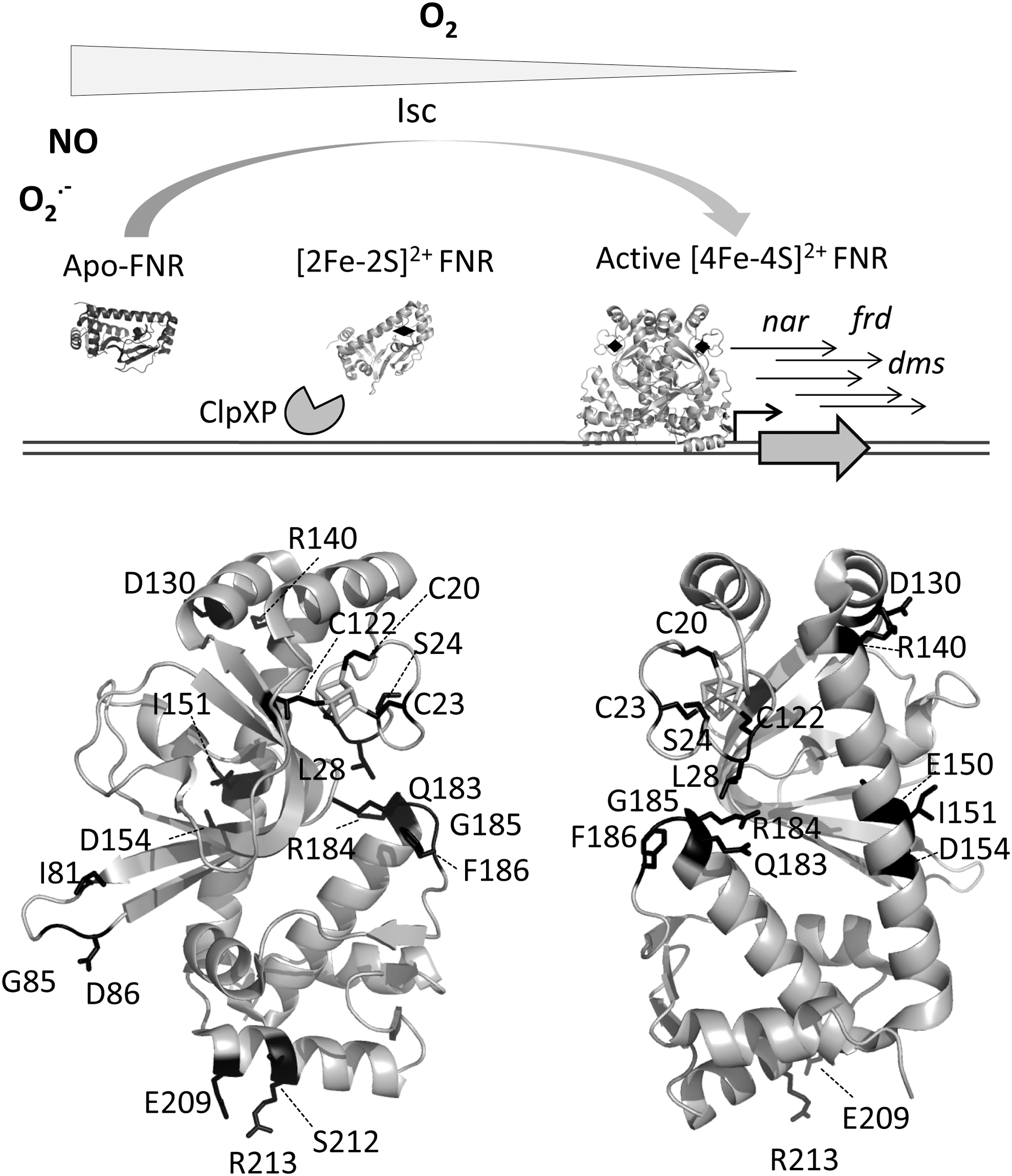
The active dimeric [4Fe–4S]2+–FNR is also sensitive to NO, which generates a mixture of monomeric and dimeric dinitrosyl–iron–cysteine complexes suppressing its ability to bind DNA (69). Therefore, the behavior of FNR differs considerably from that of most CRP family members, which are stable homodimers and bind DNA upon activation by their corresponding effectors. In contrast, a dimer–monomer transition driven by ligation of one [4Fe–4S]2+ cluster per subunit and/or O2 tension is critical for precise FNR activity. Exceptions to this general working model are the FNR regulators from
FNR can work as either a repressor or an activator of a different set of genes. For positive regulation, FNR contains three individual activating regions that mediate contacts with RNAP depending on the promoter architecture (324). Class II promoters, whose FNR binding site is around 41.5-bp upstream of the transcription start site, predominate over class I with the recognition site at −61.5-bp. Different studies have unveiled the complexity of the FNR regulon in
The core of the FNR regulon appears to be conserved across many facultative anaerobes and, typically, contains operons associated with anaerobic respiration (
Before the resolution of the FNR structure from
A detailed comparison between the structural information provided by
The N-terminal region, which contains the four cysteine ligands of the [4Fe–4S]2+ cluster, exhibits high conformational flexibility and is more disordered than the rest of the protein. Cluster assembly seems to organize the FNR N-terminal region eliciting O2 accessibility to the redox center. A network of hydrophobic interactions proximal to the redox cluster that involves residues of the A, B, and C α-helices would serve as a signaling relay between O2 mediated cluster oxidation and dimer dissociation (244, 384). Residue Asp154, which has an inhibitory effect on dimerization, together with Glu150 forms a negatively charged pocket in holo-FNR, proximal to Ile151. In contrast to what was previously proposed, Ile151 does not shelter Asp154 to afford dimerization, but establishes intersubunit van der Waals contacts that are critical for dimer stability. Another important amino acid is Arg140, which enables the O2 sensitivity of the FNR monomer–dimer equilibrium through the formation of a salt bridge with Asp130 belonging to the αB helix of the opposite subunit (244).
Previous studies as well as a comparison with the structure of the FNR-homologue FixK2 from
The variability across species of the residues composing the cluster-binding domain in the vicinity of the four conserved cysteines likely determines cluster sensitivity to O2 in each organism, according to the environment (86, 244, 396). Moreover, the occurrence of multiple FNR proteins in several organisms, such as
6. WhiB/Wbl
WhiB and Wbl (WhiB-like) are a family of multifunctional proteins exclusive to actinomycetes. Wbl proteins play diverse roles in morphogenesis, cell division, metabolism, virulence, and antibiotic production. WhiB was first discovered in
Anaerobically isolated Wbl proteins contain a C-terminal DNA-binding domain and a [4Fe–4S]1+ redox cluster at the N-terminus. The DNA-binding domain contains a Trp/Gly-rich motif, predicted to form a β-turn, followed by two positively charged amino acid motifs with different degrees of similarity with the DNA-binding motif known as AT-hook (11, 300, 314, 345). The [4Fe–4S]1+ cluster is NO sensitive, although in some paralogs it also becomes oxidized upon exposure to O2. The redox cluster is coordinated by four conserved cysteines, two of them in a CysXXCys motif, commonly found in the thioredoxin fold and in oxidoreductases.
Nitrosylation of the [4Fe–4S]1+ cluster is a multistep process that consumes up to eight NO molecules, and if the stress persists, it is followed by complete loss of the cluster (67). Depending on the environment, the coordinating cysteine residues may then remain as −SH, or establish intramolecular disulfide bridges. Unlike other transcriptional regulators containing iron–sulfur clusters, nitrosylated and apo-Wbl strongly bind DNA. The oxidized disulfide-containing apo-protein exhibits the highest DNA-binding affinity. In contrast, DNA interaction with the holo-form is very weak or null (314). Therefore, besides gene regulation mediated by the redox status of the cluster, changes in the redox state of the cysteines provide Wbl with an additional layer of regulation. Moreover, most Wbl proteins present disulfide reductase activity, representing a novel redox system in
The increasing knowledge of this intriguing family of proteins has revealed significant differences among them, possibly related to their functional diversity. The chromosome of
Several important differences may account for the functional divergence among them. First, the variability in cluster environments and exposure to oxidants indicate differences in the redox potentials, consistent with the unequal O2 sensitivity among WhiB paralogs (5). Their similar behavior against NO is supported by mechanistic studies evidencing that nitrosylation takes place through a common mechanism in phylogenetically unrelated regulatory proteins (65). Second, unconserved amino acid residues between the key CysXXCys motifs will certainly result in different redox potentials for each WhiB paralog and, therefore, different disulfide reductase activities. As an exception, the WhiB2 paralog lacks disulfide reductase activity and, instead, has a chaperone-like function (Table 1) (198).
Furthermore, a comparative study of the thermal stability of all seven WhiB proteins from
B. Redox sensing by thiol-based targets
The activity of transcriptional regulators with thiol-based switches builds on the oxidation state of cysteine thiol groups. Thus, −SH groups can be reversibly oxidized upon exposure to redox-active compounds so that a chemical signal is transformed into a biological signal through a conformational change in the regulator that modifies its DNA binding affinity. Several classes of structural changes undergone by the regulator can be observed: major reorganization of the polypeptide backbone in association with disulfide redox activity, order/disorder transitions, changes in the quaternary structure, or disulfide oxidation after the expulsion of metals (95).
A comprehensive review of a large number of bacterial thiol-based redox sensors that specifically sense ROS, reactive electrophile species (RES), and HOCl
1. Sensors involving cysteine–zinc clusters
a. DksA
Prolonged oxidative stress leads to the inactivation of key enzymes of the central metabolism, leading to nutritional starvation. As a response, a transcriptional program known as the stringent response provides bacteria with survival advantages and efficient environmental adaptation (295).
The major regulatory component of the stringent response is hyperphosphorylated guanine [guanosine pentaphosphate, (p)ppGpp] whose accumulation in the bacterial cell modifies the transcriptional profile through the binding to RNAP, inducing a large-scale restructuration of metabolic gene expression (366). Together with the alarmone (p)ppGpp, the global regulator DksA is critical for the stringent response of most Gram-negative bacteria. Without binding to DNA, as a consequence of amino acid limitation, DksA binds to the RNAP secondary channel, destabilizing the open promoter complex and impairing transcriptional initiation. In this way, bacteria quickly reprogram transcription in response to changes in nutrient availability (287).
Beyond its participation in the stringent response and independently of (p)ppGpp, DksA works as a redox-dependent global regulator that has different organism-specific functions (Fig. 5). In
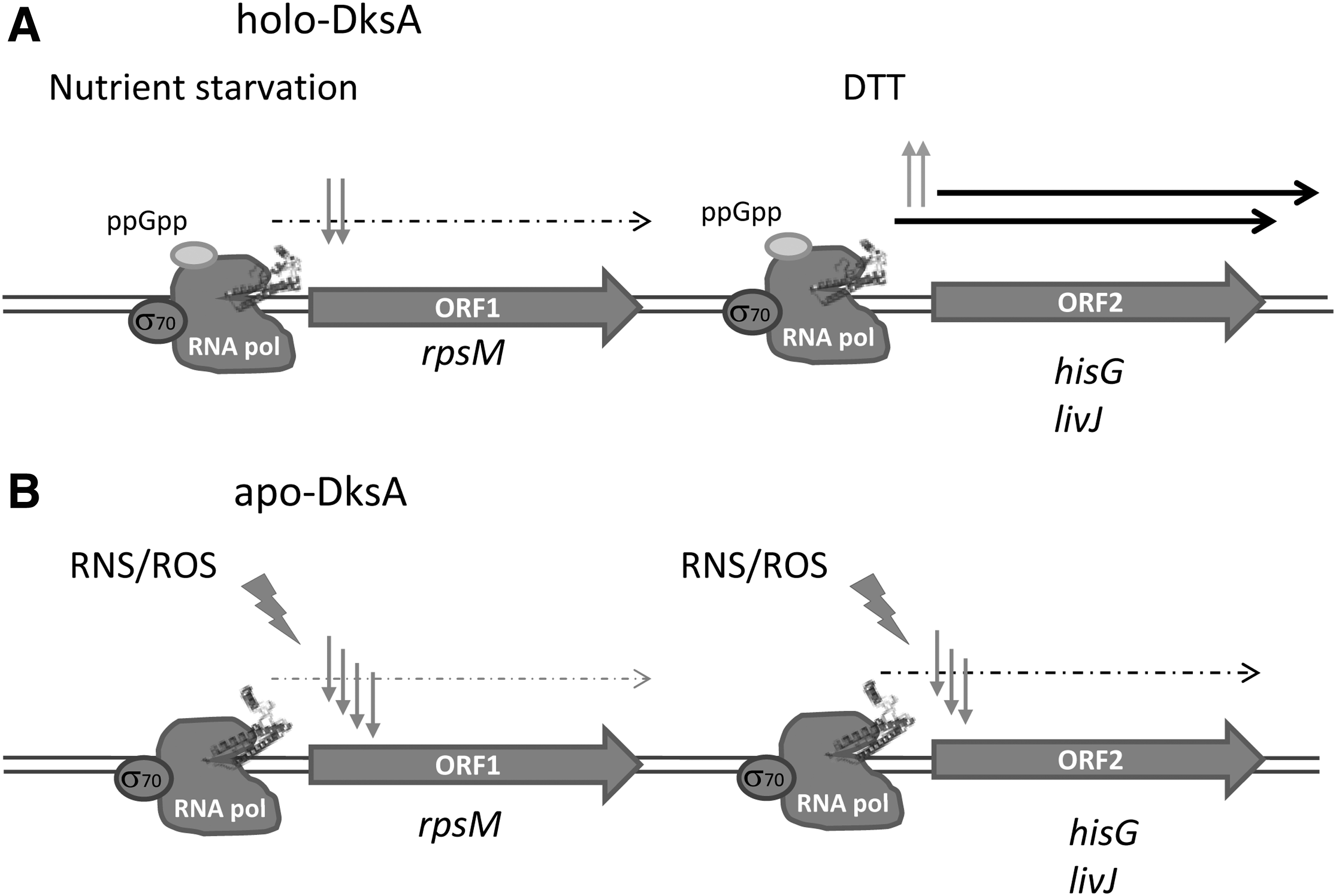
In most Gram-negative bacteria, DksA consists of a coiled-coil domain separated from the C-terminal α-helix by a hinge region containing a 4-cysteine zinc finger motif (290). The coiled-coil domain presents an AspXXAspXAla motif in the loop at its tip that is essential for the protein function (220), whereas the relationship between the presence of Zn2+ and the functionality of the protein seems more complex.
It has been established that thiols in the 4-cysteine zinc finger motif sense oxidative and nitrosative stresses by releasing the zinc ion, independently of the second messenger guanosine tetraphosphate (ppGpp) (Fig. 5) (139). Zn2+ release causes an evident loss in α-helicity of the protein, likely due to disulfide bond formation, since it can be reverted by dithiotreitol (DTT). This oxidation, which has been suggested to occur in the complex DksA–RNAP, converts DksA into a stronger repressor of downregulated genes or, alternatively, fails in the transcriptional activation of DksA-activated promoters, increasing the threshold of the stringent response. These results are consistent with a role for Zn2+-bound thiolates as redox sensors of nitrosative and/or oxidative stress and evidence the ability of 4-cysteine DksA to rapidly integrate nutritional, oxidative, and nitrosative signals into a coordinated transcriptional response (139, 162).
Interestingly, under conditions of zinc limitation, some organisms express DksA paralogs that do not contain zinc, such as DksA2 from
In addition to DksA2 from
b. TraR
TraR is a 73-amino acid protein that exhibits 29% sequence identity with the C-terminal half of DksA (26). TraR is encoded in the
c. RsrA
RsrA functions as a σR-specific zinc-associated antisigma factor that inhibits σR-directed transcription under reducing conditions, regulating the response to thiol oxidative stress in the cytoplasm of
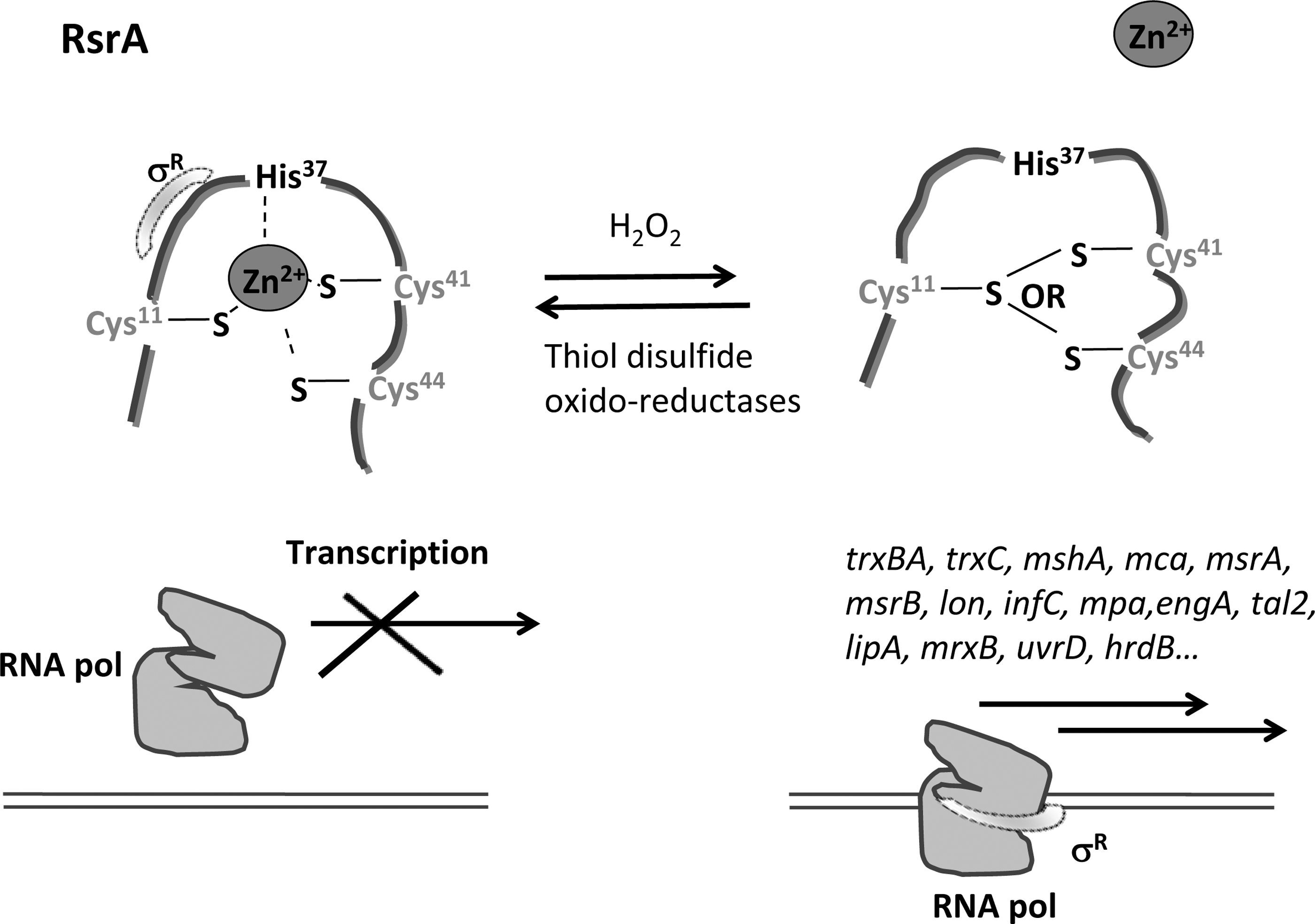
Several σR target genes have been identified. Their products include thioredoxin systems (TrxBA and TrxC), the protein MshA involved in mycothiol synthesis, mycoredoxin-1 (Mrx-1), proteolytic components (Lon, PepN, and ClpX), ultraviolet (UV) resistance components (UvrA system), and proteins involved in cysteine production (CysM), methionine sulfoxide reduction (MsrA and MsrB), guanine synthesis (GuaB), ribosome-associated function (RpmE and RelA), or detoxification of electrophiles (179, 182, 189). Apart from the control of σR by the antisigma factor RsrA, SigR activity is also controlled at the translation level (99).
2. Nonmetallated thiol-based switches
a. OxyR
OxyR was the first transcriptional regulator discovered to have the ability to sense ROS. It regulates the expression of defensive genes against the harmful effect of H2O2 such as those encoding catalases (
OxyR from
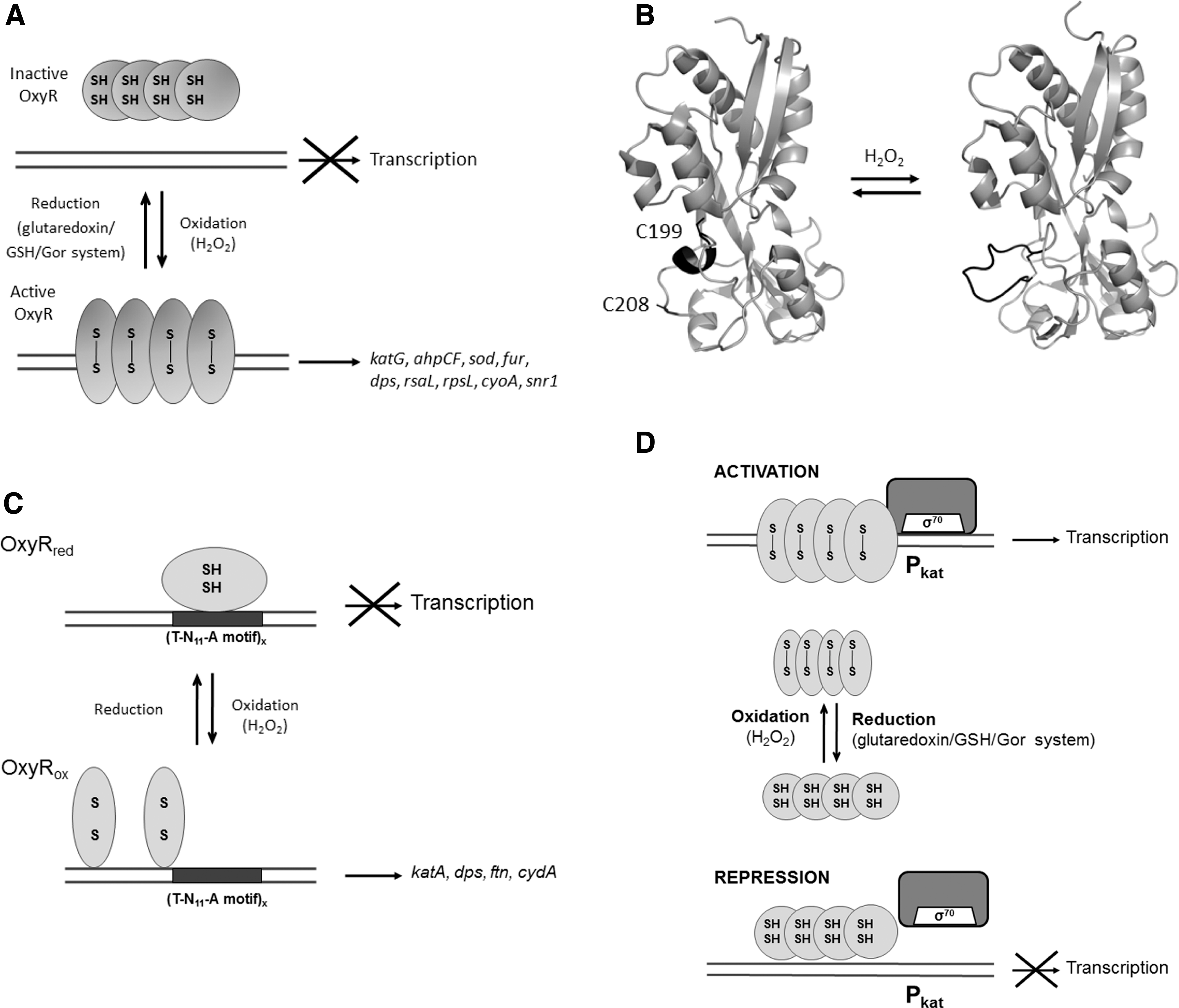
As other members of the LysR family of transcriptional regulators, the OxyR fold consists of two domains: the N-terminal DNA binding domain containing a HTH motif and a C-terminal regulatory domain (319, 363). In reduced OxyR, the regulatory domain consists of two α/β domains that exhibit a similar folding pattern. The redox-active Cys199 resides between two α/β domains, whereas Cys208 is located at the lower part of one of domains separated from Cys199 by ∼17 Å (Fig. 7B). Upon disulfide bond formation between Cys199 and Cys208, the short helix formed by residues 199–203 in the reduced structure uncoils, leading to a significant rearrangement of the secondary structure of the domain that allocates Cys208 (55).
The oxidation of OxyR involves changes in the orientation of monomers in the dimer, compared to the reduced form, that affect the interdimer orientation in the tetramer and eventually the binding to DNA (Fig. 7A). Oxidation of OxyR by H2O2 proceeds in a two-step mechanism that involves selective oxidation of Cys199 to form sulfenic acid (Cys199-SOH) and its subsequent reaction with Cys208, resulting in an intramolecular disulfide bond (211). The structure of a
Moreover, the crystal structure of the full-length
In some pathogenic bacteria,
However, the OxyR thiol–disulfide switch model appears to be more complex since this regulator can be activated by different post-translational thiol modifications. In this way, anaerobic respiration on nitrate of
Moreover, cysteine overoxidation has emerged as a mechanism of regulation of OxyR2 from
b. PpsR/CrtJ
PpsR, directly or indirectly, controls the synthesis of all the different components of the photosystem in purple bacteria. It belongs to the LuxR family of transcriptional regulators and is redox sensitive through the formation of an intramolecular disulfide bond (171). Unlike OxyR, formation of this disulfide bond in PpsR is insensitive to the addition of H2O2 but is sensitive to O2, indicating a different mechanism of disulfide formation from that used by OxyR, where disulfide bond formation is stimulated by trace amounts of hydrogen peroxide even under reducing conditions (Fig. 8) (12).
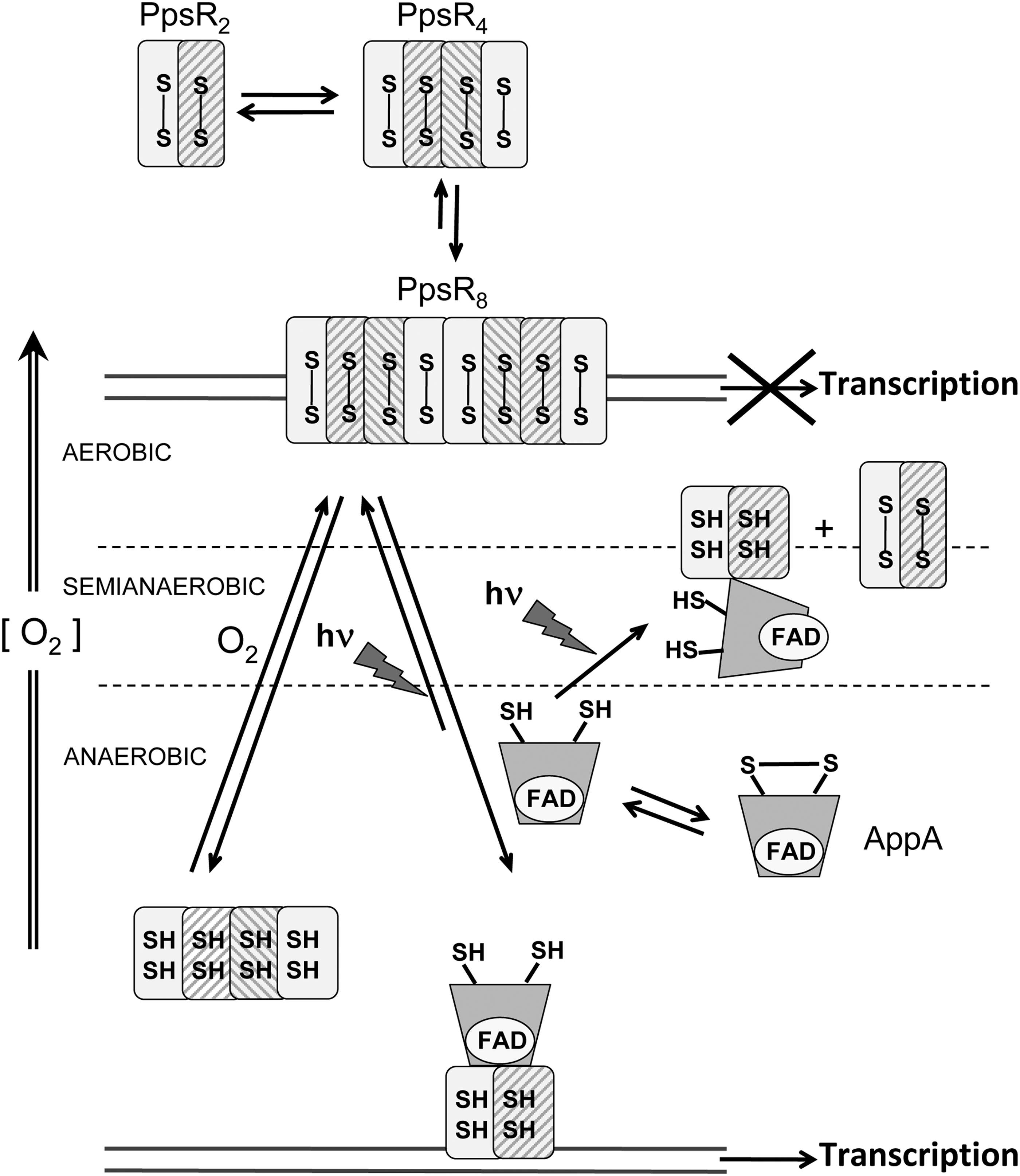
The PpsR proteins have been mainly characterized in two related species
PpsR and CrtJ share 53% identity, and both proteins contain a HTH type DNA binding motif placed at the C-terminal region. In PpsR from
The DNA binding mechanism of the PpsR proteins from other organisms shows variations.
PpsR1 and PpsR2 show fundamental differences with the PpsR/CtrJ family of regulators. In particular, PpsR1 is a redox-sensitive activator through the formation of a disulfide bond that unlike PpsR is intermolecular. Furthermore, oxidation of PpsR1 remains very limited in response to O2. Unlike PpsR1, PpsR2 does not contain cysteine residues and is not redox sensitive. Therefore, the DNA binding affinity of PpsR2 is redox independent (171). Regulation of this type of regulator can undergo further modulation by its association with other proteins. In
c. OhrR
The OhrR family of regulators sense organic hydroperoxides (OHPs) and other ROS by oxidation of a critical and highly conserved cysteine residue. OhrR belongs to the MarR (
Two OhrR subfamilies have been described based on their peroxide sensing mechanism (Fig. 9): the single cysteine class, represented by
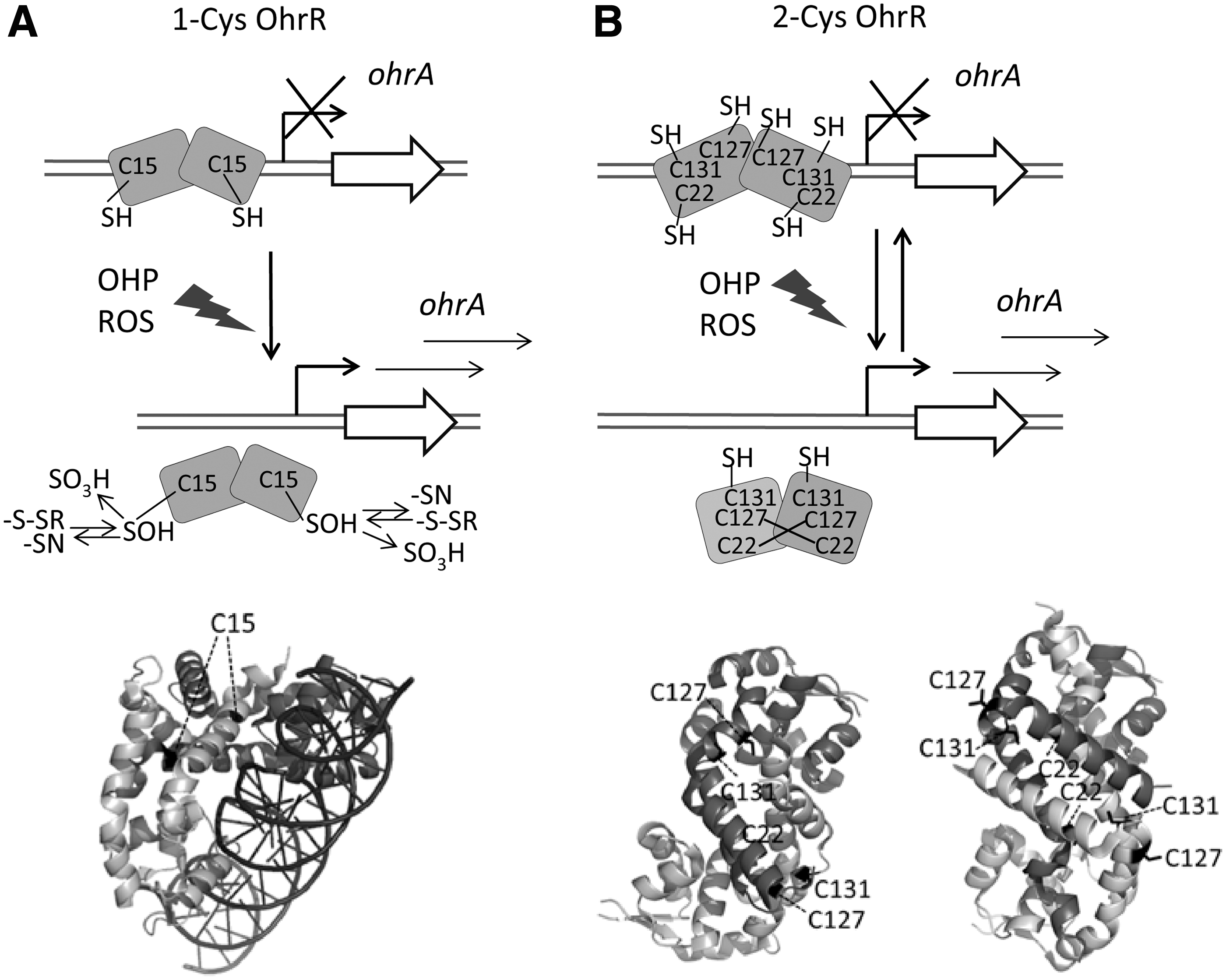
In both 1-Cys and 2-Cys OhrR subfamilies, the initial step leading to transcription derepression mediated by peroxide involves oxidation of a sensing cysteine to sulfenic acid (Cys-SOH) that is not sufficient to derepress transcription. In the first case (Fig. 9A),
OhrR homologues that contain a single cysteine have been characterized in
Therefore, depending on their amino acid content, OhrR proteins isolated from different bacteria can exhibit varied DNA binding properties. Even oligomerization can be important in controlling OhrR activity.
In
According to MgrA homodimer crystal structure, a unique cysteine residue located at the protein dimer interface can be oxidized by H2O2 and OHPs leading to dissociation of MgrA from DNA (47). MgrA activity can also be reversibly regulated by cysteine phosphorylation (355). In the same way, SarZ can be controlled by cysteine phosphorylation (355). SarZ is a global transcriptional regulator that uses a single cysteine (Cys13) to sense peroxide stress and control genes involved in hydroperoxide resistance (
Besides MarR/OhrR family of redox sensors, there are other MarR-type regulators that belong to the MarR/DUF24 subfamily, conserved in Gram-positive bacteria (7). Unlike OhrR, in
YodB contains three cysteine residues (Cys6, Cys101, and Cys108) that are involved in its mechanism of inactivation. The accepted model of functioning indicates that upon treatment with diamide and quinones, YodB is inactivated by formation of a Cys6–Cys101 intersubunit disulfide both
d. NemR
The NemR repressor (formerly named YdhM) belongs to the TetR family of transcriptional regulators and responds to cysteine-modifying electrophiles, alkylation, and reactive chlorine species (RCS) (375). The oxidation of cysteine residues by RCS is a reversible process that leads to a decrease in NemR DNA binding affinity and the consequent derepression of transcription of the NemR-controlled genes
According to crystal structure data of
e. FurA
Fur is the master regulator of iron homeostasis in most heterotrophic bacteria, where it works as a classical repressor. In a simplified model of regulation, Fur binds to its target sequences using Fe2+ as corepressor to block the transcription of an ample regulon (92, 101). Fur belongs to a superfamily including homologues that control processes intimately linked to redox homeostasis, such as PerR (response to peroxide stress) and Zur (control of zinc homeostasis). Remarkably, peroxide transduction by PerR is carried out through a completely different mechanism than that described by Fur (discussed in the next section). Both PerR and Zur can work with Fur to coordinately regulate a set of genes involved in the response to oxidative stress or virulence, among other important processes (152, 325, 368).
Furthermore, anaerobiosis affects the gene expression programs of Fur and the small RNA regulator RyhB in
Usually, Fur proteins contain a structural Zn2+ ion that is absent in the cyanobacterial regulator (142). Thus, the lack of structural Zn2+ in FurA from
The cyanobacterial FurA regulon contains genes that belong to diverse functional categories, including iron homeostasis, photosynthesis and respiration, heterocyst differentiation, oxidative stress defense, and light-dependent signal transduction mechanisms, among others (125, 127 –129).
FurA contains five cysteine residues, four of them arranged into two active CysXXCys redox motifs (Cys101XXCys104 and Cys141XXCys144) located in the C-terminal domain of the protein (dimerization domain). FurA needs not only metal but also reducing conditions to remain fully active
A mutational study of single cysteines introduced in FurA revealed that Cys101 and its particular redox state are critical for the coordination of the metal corepressor that ultimately controls the FurA ability to bind to DNA
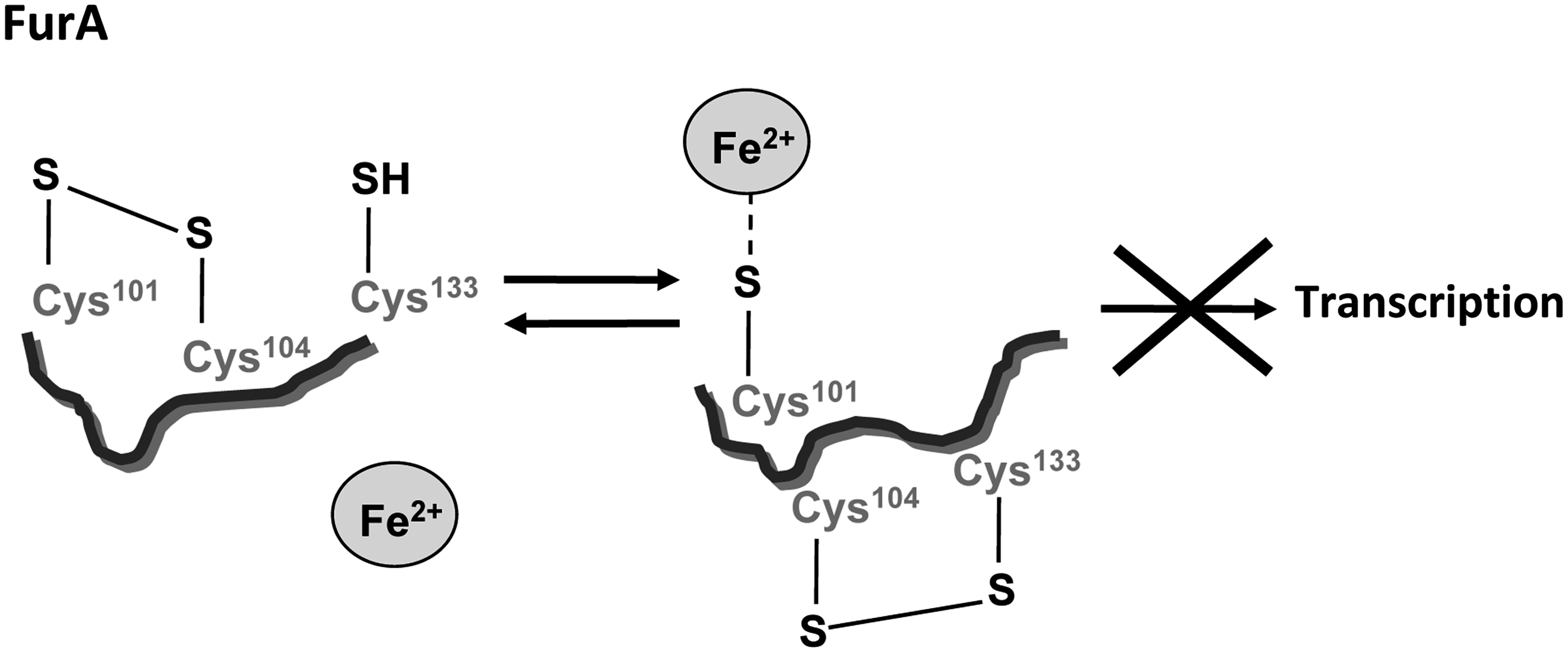
Apparently, this mechanism is specific for cyanobacterial Fur homologues since it relies on Cys133, a residue conserved in cyanobacterial Fur homologues but absent in Fur homologues from heterotrophic bacteria. The FurA redox switch resembles that described for RsrA in the previous section. In both cases, a disulfide bond between both cysteines of a CysXXCys motif controls the redox state of a third cysteine that coordinates the metal ion. However, although in RsrA the coordination of Zn2+ keeps cysteines in a reduced state determining oxidation kinetics of this regulator, in FurA the Fe2+ that binds to the reduced cysteine plays a role as corepressor metal, coordinating in this way iron homeostasis and redox responses.
C. Regulation by metal-catalyzed oxidation: PerR
The metal-catalyzed oxidation of histidine to 2-oxo-histidine is an important marker of oxidative stress commonly associated with the regulation of enzyme activity (322). Unlike other Fur paralogs that regulate their target genes in response to the availability of different metals, PerR activity is based on metal-catalyzed oxidation of a histidine residue located in the conserved HisHisHisXHisX2CysX2Cys motif positioned at the hinge between the metal-sensing C-terminus and the DNA-binding domain of Fur proteins (173). Oxidation of PerR leads to loss of the iron cofactor and its dissociation from DNA to derepress transcription of genes involved in the antioxidant response. The mechanistic differences between Fur and PerR from heterotrophic bacteria have recently been reviewed (291). PerR may function as both activator and repressor of gene expression. Targets include antioxidant enzymes, virulence genes, and other regulators (36, 306, 377).
D. Methionine oxidation-based transcriptional regulation: HypT
The
HypT has different multimeric forms. On binding to DNA, the dodecameric ring-like structure of HypT dissociates into an active tetrameric form that acts as a transcriptional activator. The current model proposes that the oxidation of methionine residues promotes the transition of the inactive dodecameric form of HypT to the active tetrameric form (83). MsrA and MsrB are required to reverse the oxidation state of the oxidized methionine residues, thus inactivating the activity of HypT1 (83).
E. Redox sensing by heme-based sensor proteins
Bacterial heme-based sensor proteins exploit the redox chemistry of heme to sense environmental gases (
1. CooA
CooA (bacterial CO oxidation transcriptional activator) is a heme-binding protein that controls the expression of a regulon allowing anaerobic growth of
CooA is a homodimer and each monomer contains a
Apparently, this protein exists in the cell in three general heme states (Fig. 11). Under oxidizing conditions, low-spin Fe3+ heme is axially coordinated by Cys75 and Pro2, a residue located in the N-terminus of each protein monomer. In this situation, the protein is unable to associate with CO and consequently to bind to specific DNA sequences efficiently.
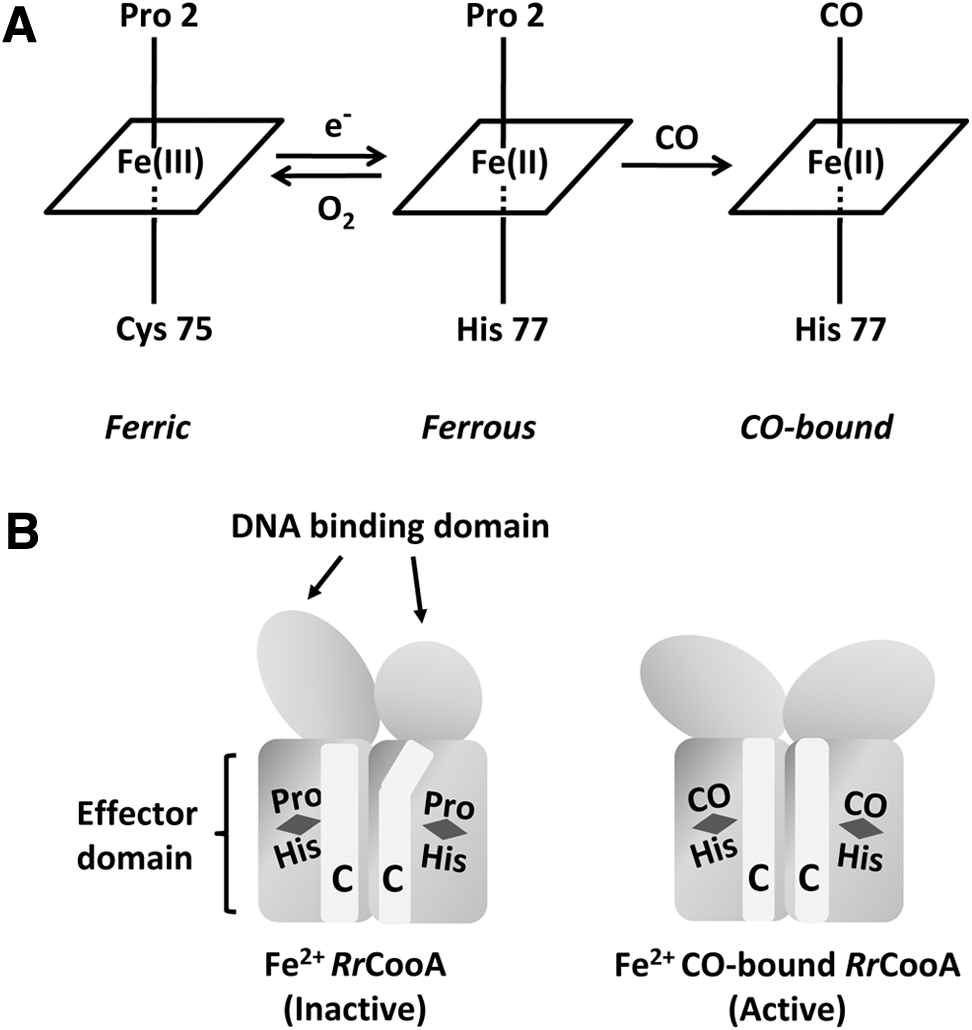
In reducing conditions, Fe2+ heme is obtained and Cys75 is replaced by His77 as an axial ligand of ferrous iron. Therefore, a redox-dependent axial ligand exchange between Cys75 (ferric form) and His77 (ferrous form) occurs on reduction of the heme iron (332). After exposure of RrCooA to CO under anaerobic conditions, CO binds to Fe2+ heme and displaces the Pro2 iron ligand (Fig. 11A) (9). Displacement of Pro2 entails a conformational change leading to a reposition of the heme, exposing the CO-bound heme to the long α-helices (C-helices) that extend along the homodimer interface (Fig. 11B) (188, 406). This interaction stabilizes an alternative conformation of the domain containing these C-helices that alters the hinge region separating the DNA- and CO-binding domains. When the geometry of the hinge region alters, the inactive form of CooA destabilizes and the active form stabilizes (405). Reorientation of CooA DNA-binding domains is necessary to produce a transcriptionally active state (157).
A semi-apo state of the CooA dimer, with a heme-bound monomer in a CO-bound form of
CooA modulates the expression of the
2. RcoM
RcoM (regulator of CO metabolism) is a CO-sensing transcription factor that undergoes a redox-mediated ligand switch and may utilize redox-active heme to sense the redox state of the cell. It couples an N-terminal Per-Arnt-Sim (PAS) fold (like the mammalian NPAS2 CO sensor) to a C-terminal DNA-binding LytTR domain, but its particular heme ligation characteristics, DNA binding modules, and organization of domains are different from those reported for mammalian sensors (187). In different organisms, it appears to regulate
RcoM1 and RcoM2 from
In the aforementioned heme proteins, heme exists as a stable and essential prosthetic group. However, recent studies reveal that the function of some proteins is acutely modulated by the reversible binding of heme that acts as a cellular signaling messenger (257). The association/dissociation of the heme iron complex to/from the protein regulates(s) its functions, including catalytic reactions (kinase and proteolysis) or DNA binding. In contrast to heme-based gas sensors wherein gas molecules (O2 and CO) bind only to the heme Fe2+ complex, proteins that are regulated by the reversible binding of heme associate/dissociate to the heme Fe3+ complex. Under reducing conditions, a heme-responsive heme sensor can be converted into a heme-based gas sensor (337).
3. Heme sensing by thiol-based switch sensors
Some of the previously described regulators whose mechanism of action relies on a thiol-based switch have also shown the ability to bind heme through redox-sensing cysteines affecting their DNA-binding activity. In these cases, the reversible binding of heme plays a pivotal role in up- and downregulation of transcription factors. Heme-responsive proteins of this type display fast easy heme association and dissociation. They contain a HRM characterized by the presence of a CysPro sequence. The cysteine of the CysPro motif is an axial ligand of Fe3+ heme, but upon reduction, a redox-dependent ligand switch occurs and the cysteine of the CysPro sequence is no longer a ligand of Fe+2-heme (160).
It has been demonstrated for both PpsR and FurA that the binding of heme alters their DNA-binding pattern and inhibits their ability to form higher order complexes with DNA. Cyanobacterial FurA binds heme in the micromolar concentration range and this interaction negatively affects its
In the case of PpsR, the Cys424 present in its DNA binding domain is critical for heme interaction. The binding of heme changes PpsR–DNA binding pattern, inhibiting the formation of higher order PpsR–DNA complexes and inducing increased transcription of several PpsR regulated genes. This interaction seems to provide a mechanism for bacteria to react to the unbound tetrapyrrole products since in this way excess heme can quickly change the state of photosynthetic gene expression from inhibition to activation (404). Oxidation of Cys424 in the absence of heme stimulates the DNA binding of PpsR. This redox regulation is also observed while heme is present. It appears that Cys424 could be a versatile target for different types of modification such as forming/breaking disulfide bridges (239), coordinating heme, and potentially being oxidized into different oxidation states.
One notable difference with FurA is that an Ile residue follows Cys424 instead of Pro, as would be expected for a typical heme-sensing protein. However, PpsR utilizes cysteine as the critical axial residue as is the case for FurA and other HRM-containing heme sensors. Thus it is feasible that Cys coordinated to Fe2+/Fe3+ in heme is a conserved strategy for heme sensing (404).
F. Sensors of the NAD+/NADH balance: Rex
Beyond working as a cofactor for oxidoreductase enzymes, NAD+/NADH serves as a substrate for a wide range of proteins and provides a direct link between the cellular redox status and the control of signaling and transcriptional events. The ratio of NAD+ to NADH inside the cell is mainly affected by substrate availability and O2 tension, driving a set of responses aimed at maintaining redox homeostasis. Modification of this ratio is often used in industrial processes to improve the productivity of certain metabolites (225, 333, 410).
The main actor that converts the redox signal provided by the NAD+/NADH balance into a regulatory input is the transcriptional regulator Rex, whose ability to bind DNA is modulated specifically by the NAD+/NADH ratio (Fig. 12) (133). Rex has been identified and characterized in archaea and eubacteria regardless of their O2 requirements (154). Overall, Rex is a transcriptional repressor that remains bound to its DNA targets when the NAD+/NADH ratio is sufficiently high.
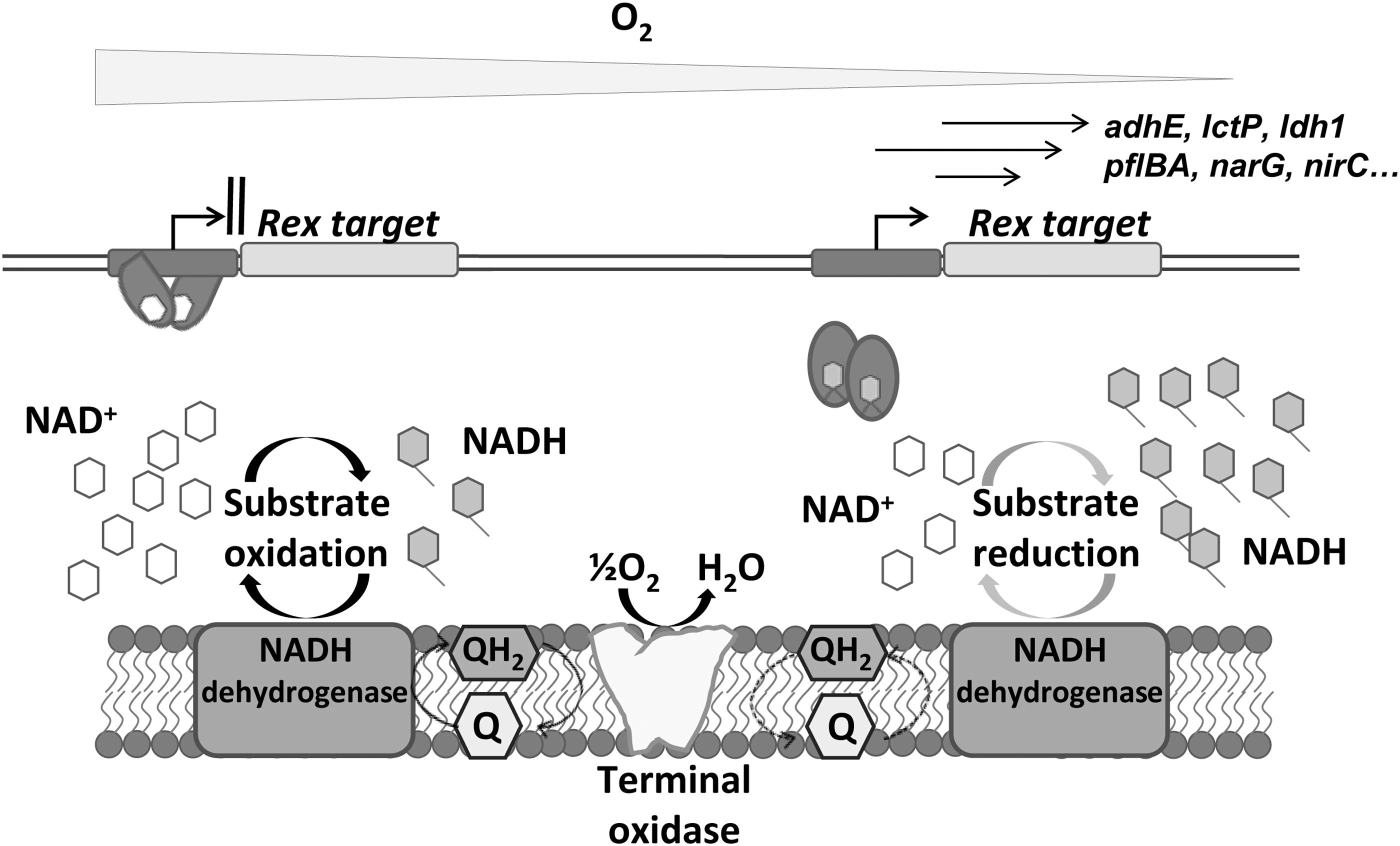
Under microaerobic or anoxic conditions, NADH competitively binds to the Rex C-terminal domain, causing a conformational change of the Rex homodimer and subsequent release from its recognition sites on DNA, allowing transcription of downstream genes. An estimated value of this ratio for transcription derepression has been calculated in
The available crystal structures of Rex in complex with DNA and/or NADH (PDB code 2VT3) indicate that Rex comprises an N-terminal wHTH-fold domain interacting with DNA and a C-terminal Rossmann-fold domain binding NADH and mediating subunit dimerization (241, 339, 388).
Key domains involved in DNA binding and NAD sensing are broadly conserved in Rex orthologs identified in the phyla
In anaerobic bacteria, a different Rex regulation mechanism has been proposed (414), which could be the result of subtle variations in the NAD+/NADH binding motifs of the Rex family, such as the substitution of Tyr98 by histidine in the regulator characterized from anaerobes. The Tyr98 residue, highly conserved in aerobic bacteria, has been proposed to play a key role in the switching mechanism between open and closed dimers through the interaction with a conserved Asp residue that is essential for binding DNA. Furthermore, the DNA-binding motifs recognized by anaerobic Rex-family members exhibit consistent deviations from the consensus established for aerobic bacteria (302, 414). These substitutions in the operator motifs seem essential for optimal protein–DNA interaction and a settling factor for the different structural bases for NAD+/NADH sensing between aerobic and anaerobic Rex proteins.
Rex regulons have been identified in several bacteria, highlighting the importance of this protein in the control of redox homeostasis, central metabolism, or hydrogen production, among other processes (207, 302). In
III. Two-Component Systems: Redox Control of Sensor Kinase Regulation
One of the main mechanisms that allow bacteria to overcome changes in the physicochemical parameters of natural environments is the regulation mediated by two-component systems (199). These systems are usually composed of a sensory protein called sensor kinase or HK and a response regulator (RReg) protein. The sensor kinase is able to sense the signal and as a consequence the protein performs autophosphorylation of a conserved histidine residue located in the HK domain. The phosphate is then transferred to an aspartate residue of the RReg through a process called transphosphorylation. The phosphorylation of the RReg causes its activation and then the RReg binds to promoters of target genes modulating their transcription. However, sometimes the RReg promotes other cellular responses in the cell different to transcriptional modulation. For example, some RRegs contain catalytic domains that once activated by phosphorylation are able to carry out enzymatic activities (111).
Canonical two-component systems perform transphosporylation in one step, whereas phosphorelays work in multistage processes to allow a fine-tune regulation of the system. Phosphorelays are found in nonorthodox two-component systems and in hybrid sensor kinases (199).
Sensor kinases may sense a wide variety of environmental signals including single molecules, such as nitrate or citrate, and more complex signals such as light or gas molecules (O2, CO, or NO) (34, 121, 184, 210). In this review, we focus on sensor kinases that perceive redox signals and respond to these signals modulating the transcriptional regulation of the cell. A summary of the two-component systems reviewed in this work can be found in Table 3.
Summary of Two-Component Systems Involved in Redox Sensing
FAD, flavin adenine dinucleotide; GAF, cGMP-specific phosphodiesterases, adenylyl cyclases, and FhlA; PAS, Per-Arnt-Sim; sMMO, soluble methane monooxygenase; TCA, tricarboxylic acids.
A. Indirect redox sensing based on PAS–GAF domains
The widespread PAS domain functions as a transduction module, acting as a sensor of environmental stimuli such as light, redox state, respiration, O2, and overall energy level of the cells, among others (364). PAS domains are present in both one-component and two-component systems, as well as in many other proteins that may contain an ample range of different domains simultaneously.
The PAS domains have a highly conserved three-dimensional structure, although they exhibit low sequence homology. They consist of ∼300 amino acid regions with several imperfect repeats, sometimes associated with a PAS-associated C-terminal motif. These motifs contain a conserved sequence of ∼40 amino acids at the C-terminal of PAS domains, contributing to the correct structure and folding of the PAS (140).
The highly versatile and multipurpose PAS scaffold can bind a broad range of redox ligands, including heme, flavins, and metal ions (140). PAS domains may also determine the specificity of transcriptional factors in modulating the expression of target genes. Some proteins, such as cGMP-specific phosphodiesterases, adenylyl cyclases, and FhlA (GAF), contain domains with a very similar fold to PAS, known as GAF domains (150).
Classification of the PAS–GAF proteins is difficult because tandem and multiple PAS domains are common in individual proteins, and often many other domains are also present; about one-third of PAS proteins contain two or more PAS domains (140). PAS proteins may include simultaneously PAS domains that bind heme, flavin mononucleotide (FMN), flavin adenine dinucleotide (FAD), 4-hydroxycinnamic acid, C3–C4 carboxylic acids (malonate, malate, and succinate), C6 carboxylic acids (citrate), and divalent metal cations. It has also been suggested that fatty acids may play a role as a PAS ligand (192). In any case, PAS domains have evolved as multifunction protein modules and they have very diverse functions, including redox sensing.
1. Redox control of sensor kinase regulation involving metal clusters
Sensor kinases are able to sense the presence of signal molecules by using sensor domains (
a. PAS domain heme: FixL–FixJ two-component system
The FixL–FixJ two-component system is involved in the regulation of nitrogen fixation genes that are tightly controlled by O2 availability. This system has been widely studied in symbiotic bacteria
Once the activation of FixL sensor kinase occurs in
FixL in
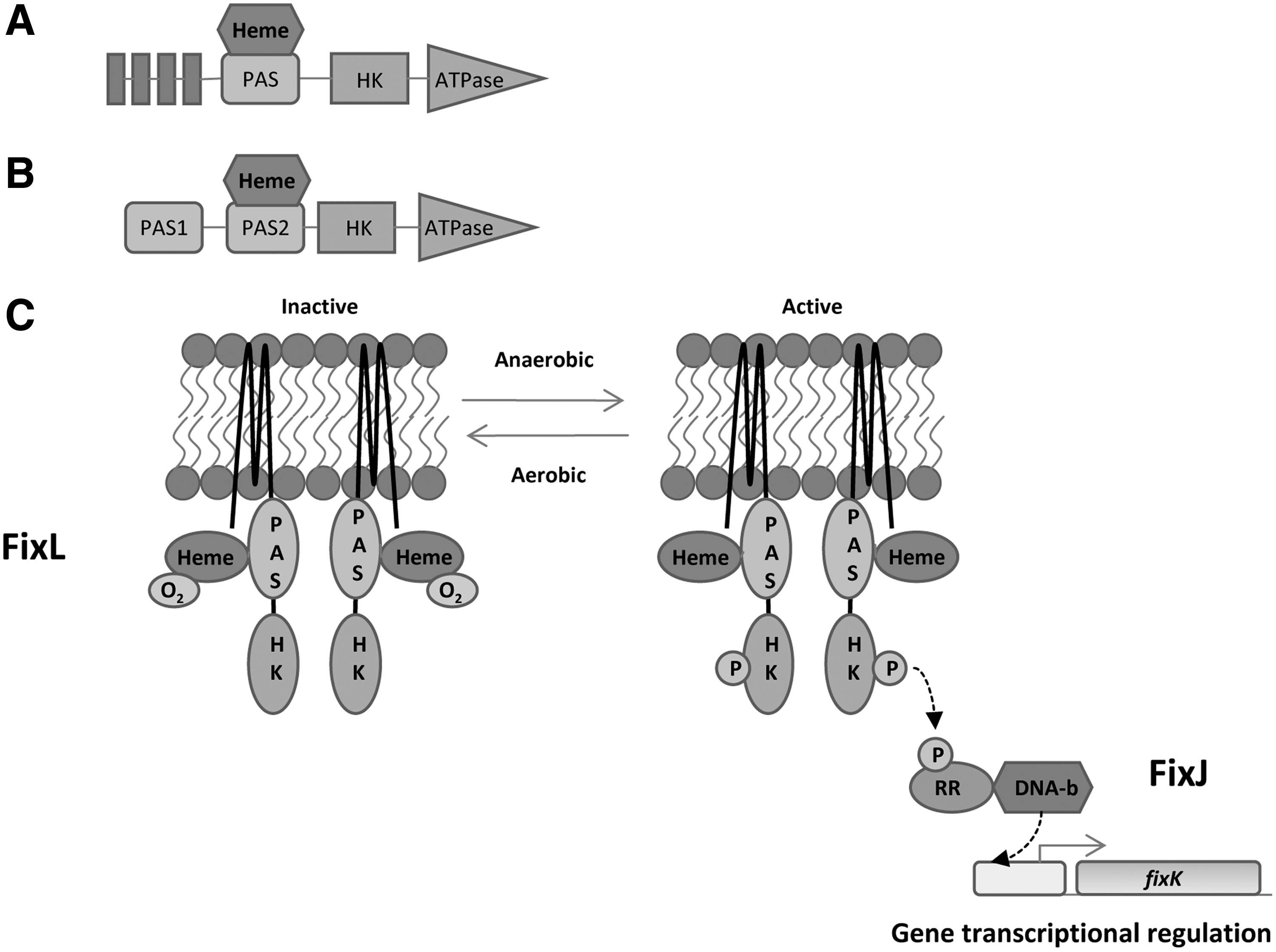
Biochemical studies revealed that changes in the spin state of Fe inside the heme moiety can modulate the regulatory effect mediated by the sensor domain (123). The authors proposed that under oxygen-limiting conditions, no O2 was bound to the sensory domain and the heme assumed a high-spin state. In this configuration, the autophosphorylation of FixL was allowed, after which FixL phosphorylated FixJ that, in turn, activated the transcription of
b. GAF domain heme: DosS–DosR two-component system
DosS–DosR is a two-component system involved in the dormancy process of
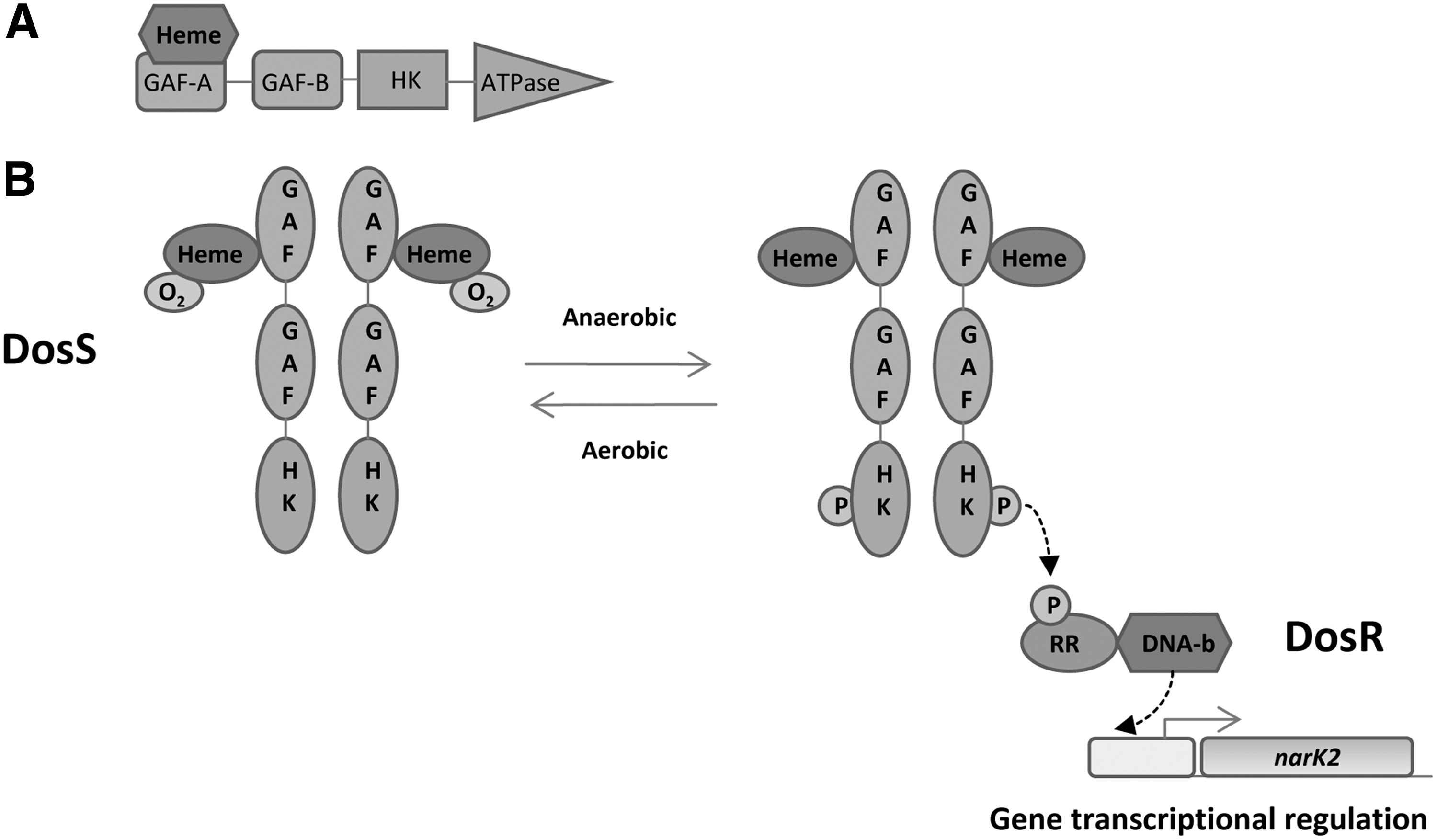
It has been reported that the hydrogen-bonding network is a key factor in gas molecule recognition. Two amino acids, Tyr171 and Glu87, seem to play an important role in gas discrimination (19). DosS shows kinase activity in Fe2+deoxy, Fe2+–CO, and Fe3+–NO forms and has little activity in the Fe3+ and Fe2+–O2 forms (Fig. 14B) (165, 166). The formation of Fe2+–O2 complex has been described, but other groups have described the oxidation of Fe2+ to Fe3+ when it is exposed to O2 (54, 165, 203, 285, 350). In view of these contradictory reports, the detailed mechanism of O2 sensing remains unknown. In contrast, it has been reported that DosS can act as a bifunctional enzyme, showing both kinase and phosphatase activities (185). In response to the availability of O2, NO, CO, and ascorbic acid, the DosS–DosR two-component system induces the expression of ∼50 genes (202, 282, 336, 362). The role of DosR in the regulation of
c. PAS domain Fe–S cluster: NreB–NreC two-component system
Some sensor kinases use FeS-containing PAS domains to sense changes in O2 tension. NreB sensor kinase is the cognate partner of the NreC RReg, both present in
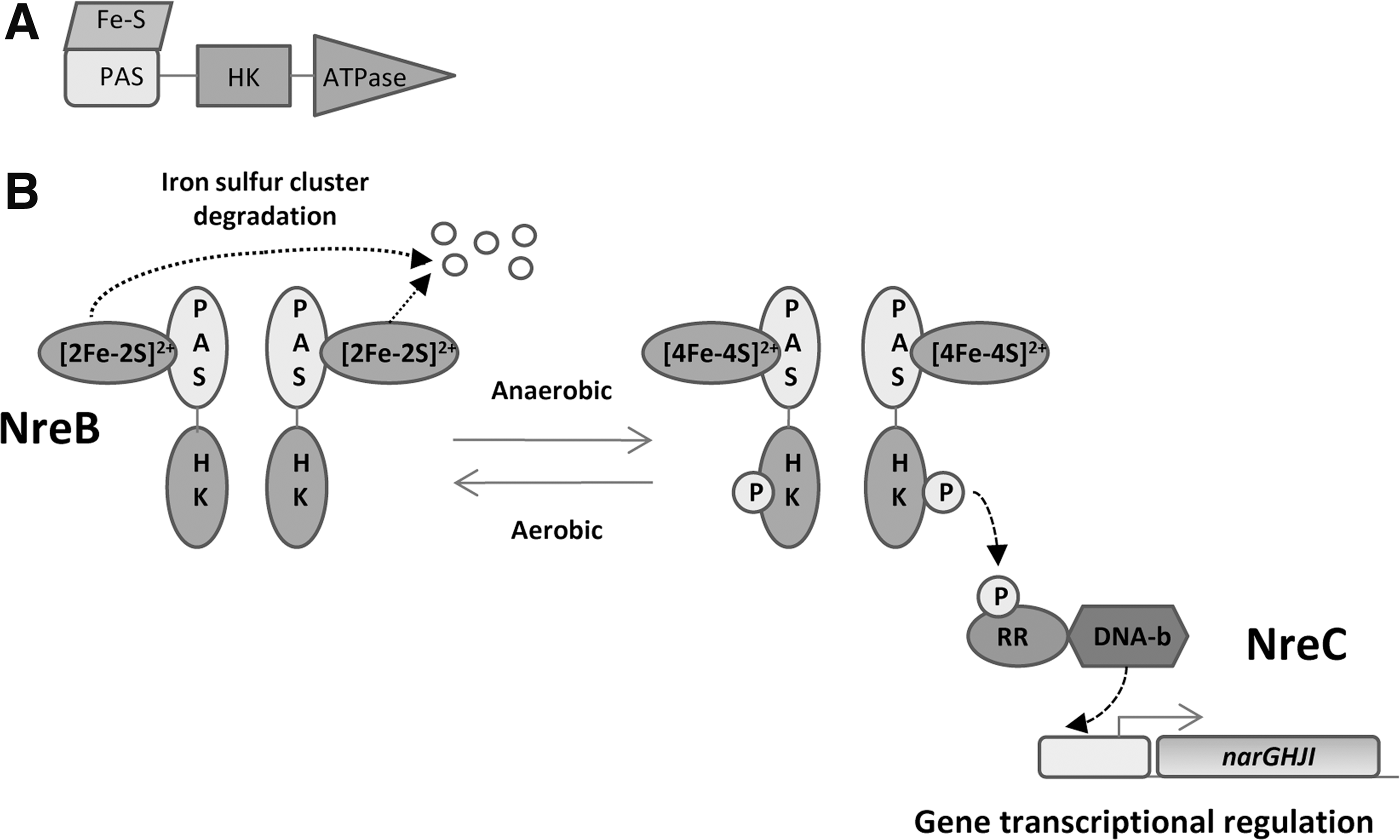
Under anoxic conditions, the PAS domain holds an [4Fe–4S]2+ iron–sulfur cluster coordinated by four conserved cysteines (180, 255). In this configuration, it performs autophosphorylation and transphosphorylation of its RReg NreC. Once activated, this RReg achieves transcriptional regulation (Fig. 15B). In the presence of O2, the [4Fe–4S]2+ cluster is converted into [2Fe–2S]2+, which is unstable and becomes degraded (Fig. 15B). Recently, it has been reported that a third protein called NreA interacts with NreB and is involved in the regulation of the NreB phosphorylation level. The NreA crystal structure shows that the protein binds one molecule of nitrate at its GAF domain. The authors suggested a nitrate/O2 cosensing by NreA–NreB system as part of what is called the NreABC system (265, 266).
d. GAF domain Fe–S cluster: AirS–AirR two-component system
The AirS–AirR two-component system (formerly called YhcS–YhcR) contains an [Fe–S] cluster inside a GAF domain. This system has been described in
AirS contains an N-terminal domain that holds the GAF domain followed by the HK domain and the ATPase domain (Fig. 16A). The GAF domain holds a Fe–S cluster-binding motif with four conserved cysteines (Cys-X7-CysXCys-X17-Cys). Experiments performed by Sun
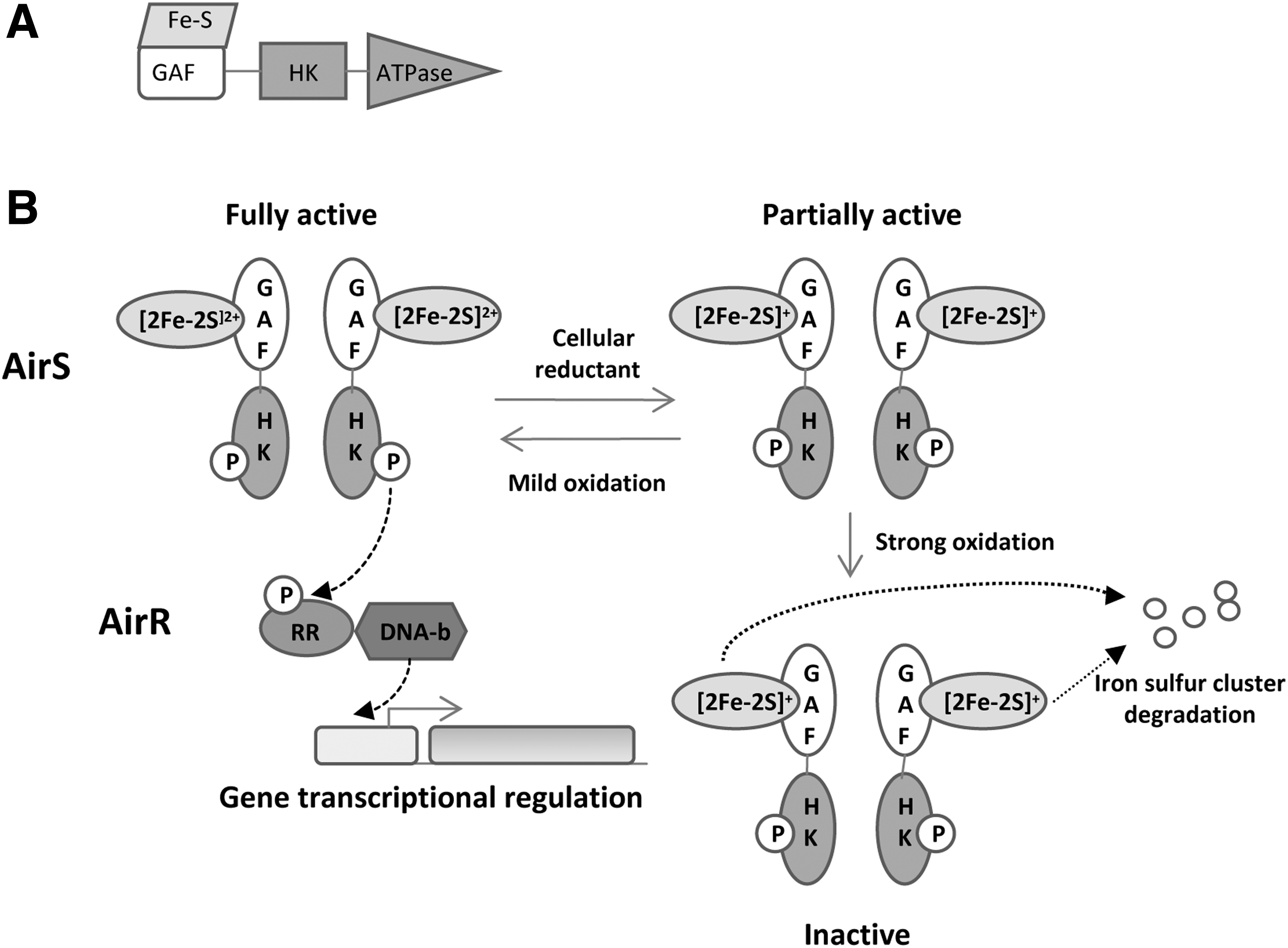
2. Sensing by NAD-binding PAS domains: KinA–KinE–Spo0A system
The KinA–KinE–Spo0A system has been described in Gram-positive bacteria such as
KinA is a cytoplasmic sensor kinase containing three PAS domains called PAS-A, PAS-B, and PAS-C. It has been shown that PAS-A was able to bind NAD+. This binding allowed KinA to sense the intracellular NADH/NAD+ ratio (195). The authors proposed that KinA is inhibited by NAD+ when respiration levels are high. Conversely, when the NAD+/NADH ratio decreases (low respiration levels), the kinase is activated, stimulating the Spo0B phosphorylation (195).
3. Flavin-binding PAS-based HK sensors: MmoS–MmoQ two-component system
Flavin-binding PAS domains are abundant in signaling proteins. The redox state of FAD and FMN cofactors senses the redox state of cytoplasm, the electron transfer chains or the visible light perception, and allows a response to readapt the metabolism to new conditions. In the past few years, several FAD/FMN-containing primary redox sensors have been described. These sensors transmit the redox signal to a secondary downstream effector domain or protein. The MmoS–MmoQ system constitutes a suitable model for the study of sensor kinases containing an FAD-binding PAS domain.
MmoS is a sensor kinase that regulates the expression of a soluble methane monooxygenase (sMMO) in a process that depends on copper availability. sMMO catalyzes the oxidation of methane to methanol. This enzyme is found in methanotrophs such as
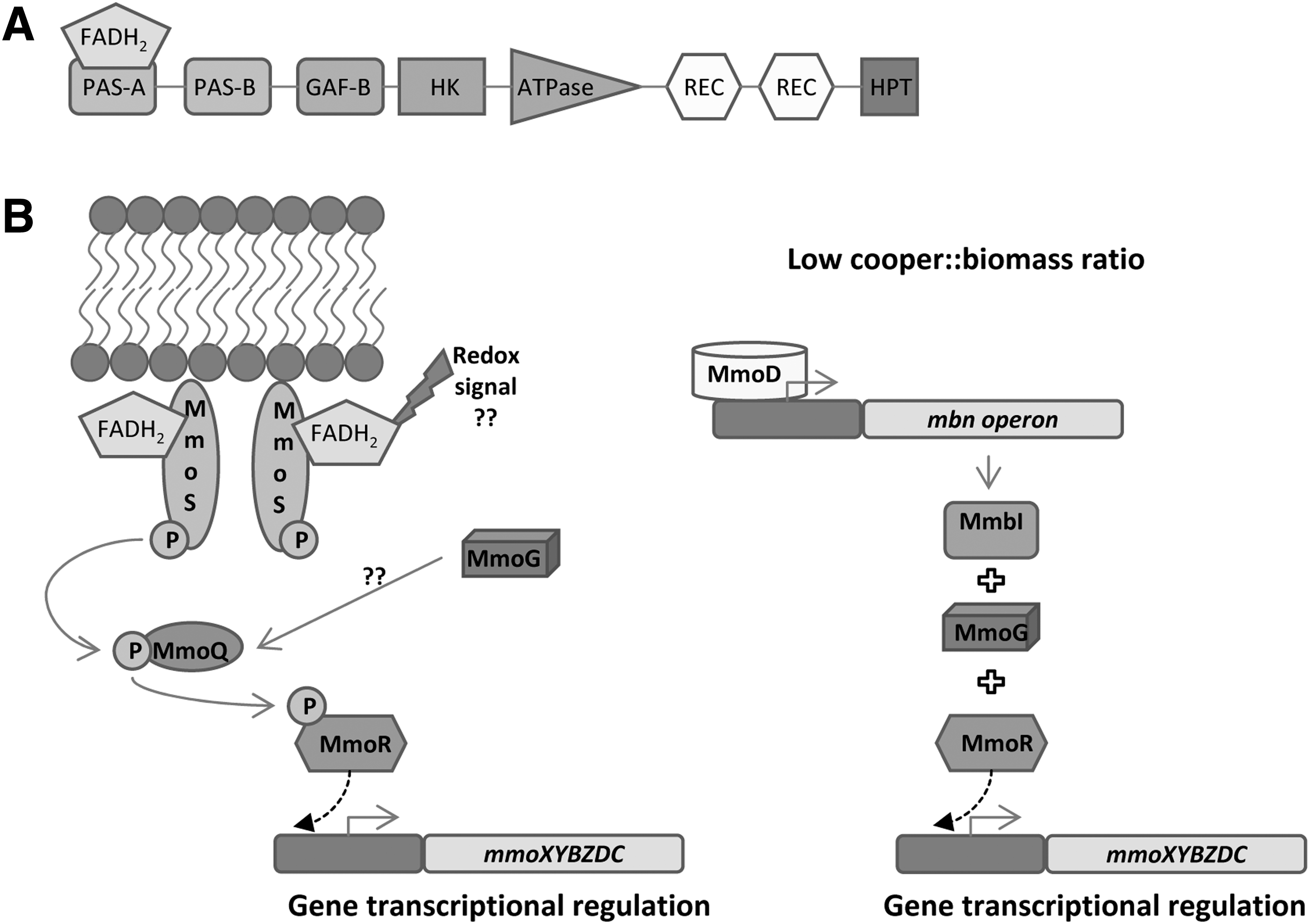
MmoS is a nonorthodox sensor kinase that contains two PAS domains (PAS-A and PAS-B) and a GAF domain followed by a HK domain, two receiver domains and a histidine phosphotransfer domain (Fig. 17A). MmoR is activated by MmoS at low copper levels, activating, in turn, the expression of
The crystal structure of MmoS was resolved by Ukaegbu and Rosenzweig in 2009 (374). A single FAD molecule was found in the PAS-A domain, which is in agreement with the model proposed by the same authors in 2006. Nowadays, the redox signal sensed by MmoS remains unknown. Several hypotheses have been proposed such as MmoS direct sensing of copper reduction or indirect sensing
4. Signal modulation by disulfide bond formation: ArcB–ArcA two-component system
Sometimes sensor kinases sense changes in O2 availability indirectly. In these cases, they are able to sense the redox status of elements that make up part of the electron transport chain such as quinones or cytochrome oxidases and, in turn, activate or repress the transcription. These redox molecules/proteins are able to oxidize or reduce some cysteines in sensor kinases, generating the redox switch. An increasing number of sensor kinases that perform this type of indirect redox sensing are being described in the literature. The best characterized examples are ArcB and RegB sensor kinases.
ArcB senses the redox state of the quinone pool by using a complex and delicate mechanism, whereas RegA kinase seems to be able to sense the redox state of the cell by using two different mechanisms. ArcB is addressed in detail hereunder since its redox-sensing mechanism involves PAS domains. However, RegB sensor kinase is included in Section III.B entitled “Non-PAS domain redox sensing based on disulfide bond formation” because although RegB uses a cysteine-based switch, these cysteines are not held in a PAS domain. Indeed, neither RegB nor its ortholog PrrB includes PAS domains in their architecture.
ArcB is a nonorthodox sensor kinase since it contains a histidine phosphotransfer domain after the receiver domain (Fig. 18A). ArcB also holds a PAS domain in the N-terminal region containing two cysteines, Cys180 and Cys241, which are responsible for autophosphorylation. ArcB has a crucial role in the adaptation of
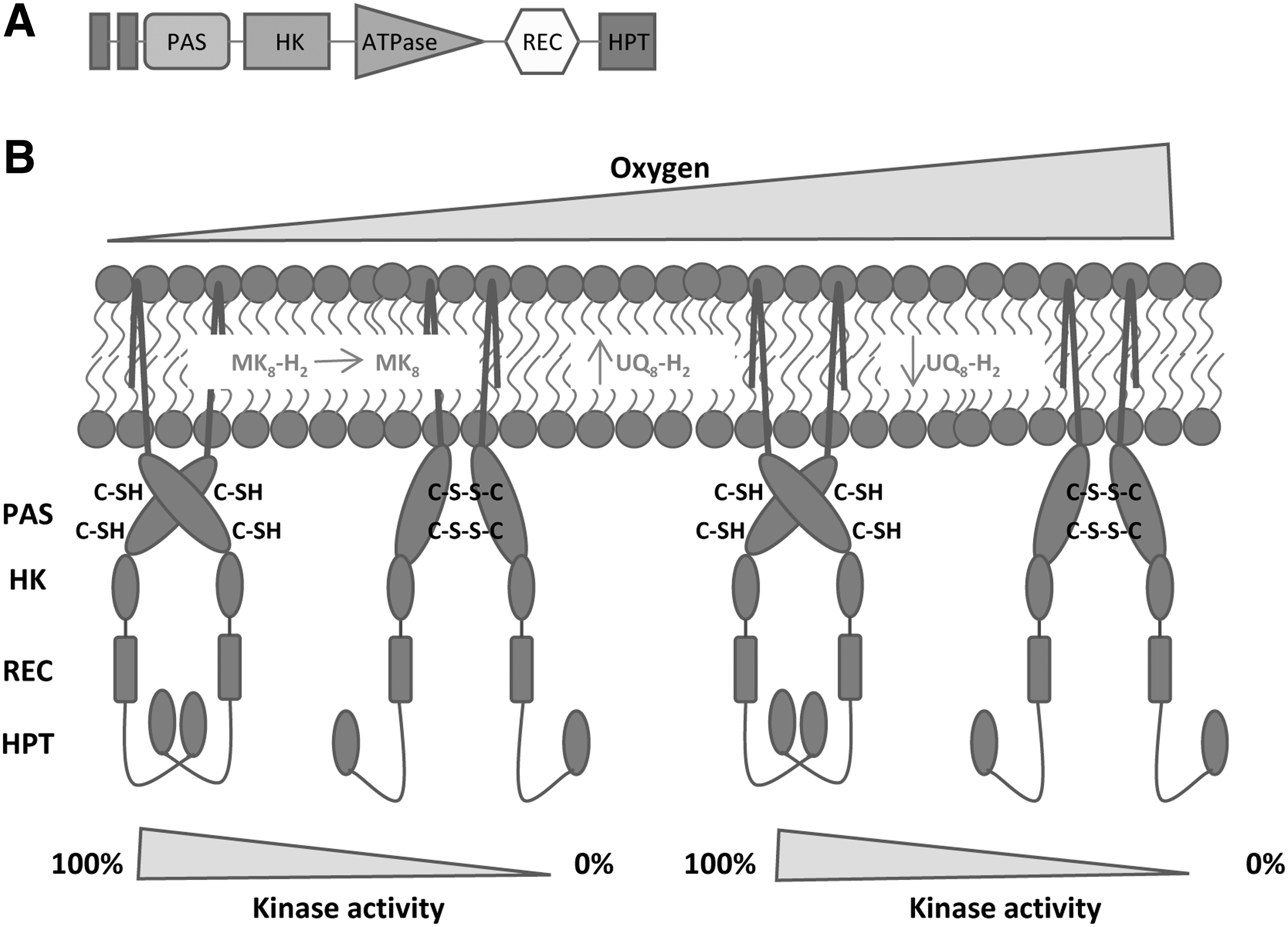
Autophosphorylation of ArcB is inhibited by oxidized ubiquinone-0 and menadione (117). These molecules are soluble analogues of ubiquinone-8 and menaquinone-8. The authors proposed that under anaerobic conditions when the quinone pool was mainly formed by oxidized ubiquinone and menaquinone, the activity of ArcB was silenced. In contrast, when O2 became limited, the quinone pool was transformed into ubiquinol and menaquinol and the autophosphorylation was triggered (117, 233).
Afterward, a more sophisticated mechanism of regulation was proposed, in which ArcB is able to respond to the redox state of the ubiquinone/ubiquinol pool and the menaquinone/menaquinol pool depending on O2 availability (23). The authors suggested that after a transition from anaerobic (0% O2) to low aerobiosis conditions (20% O2), the menaquinone pool is oxidized resulting in the inactivation of ArcB. Upon a shift from low aerobiosis conditions (20% O2) to high aerobiosis conditions (80% O2), the total ubiquinone pool increases and, therefore, ubiquinol reduces disulfide bonds and activates ArcB (Fig. 18B). In aerobic conditions, the quinone pool decreases, the oxidation of cysteines occurs, and the inactivation of ArcB takes place (Fig. 18B) (23). Recently, it has been reported that a third type of quinone (demethyl menaquinone) is involved in ArcB phosphorylation modulation. It seems that demethyl menaquinone is also able to oxydize ArcB and that demethyl menaquinol is able to reduce ArcB (376).
In recent years, several sensor kinases such as EvgS, TodS, and HskA have been found that also respond to the redox state of quinone pool although the detailed mechanisms are not as well understood as for ArcB (27, 330, 340).
5. Atypical signal transduction PAS–GAF-based mechanisms: NifL–NifA system
The NifL protein in nitrogen-fixing organisms senses both the redox and fixed nitrogen status to regulate nitrogen fixation by controlling the activity of the transcriptional activator NifA (82, 148).
In
The NifA protein from
B. Non-PAS domain redox sensing based on disulfide bond formation
1. RegB–RegA two-component system
RegB–RegA in
RegB protein is a classical sensor kinase that contains five transmembrane regions, a HK domain and an ATPase domain (Fig. 19A). RegB sensor kinase is able to autophosphorylate in the presence of specific redox signals and then phosphorylates its cognate RReg RegA that binds its target promoters activating or inhibiting the transcription. RegB senses redox signals by using the complex mechanism shown in Figure 19B and C.
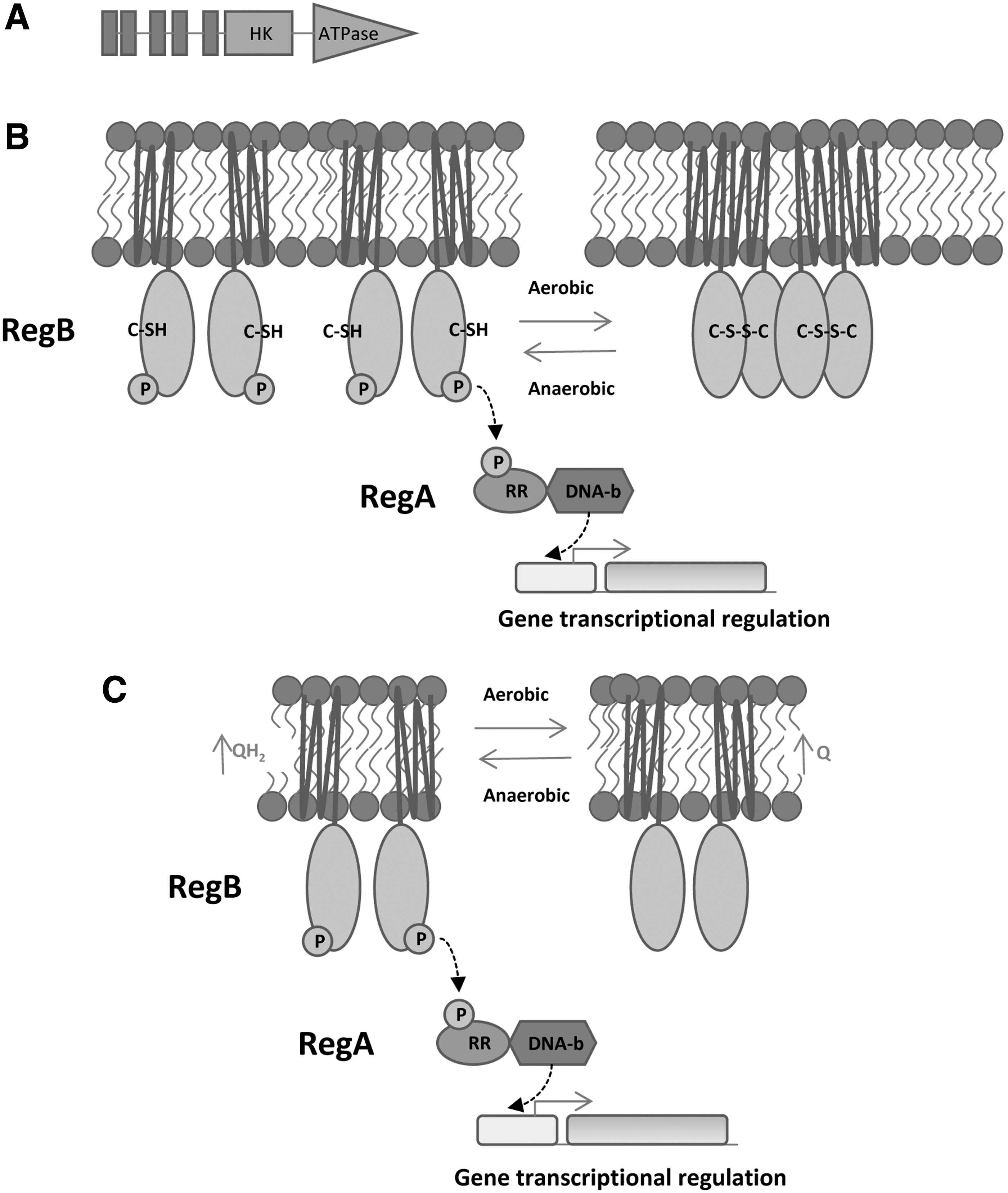
The RegB HK domain holds a highly conserved quinone binding site GlyGlyXXAsnProPhe and a conserved cysteine in position 265. At high O2 conditions, the Cys265 forms an intermolecular disulfide bond that inactivates the autophosphorylation activity of RegB. This molecular bond converts the RegB dimers into inactive tetramers (Fig. 19B). In addition, ubiquinone molecules are able to bind to the quinone-binding site of RegB, inhibiting RegB activity
2. PrrB–PrrA two-component system
A homologue system of RegB–RegA in
PrrB is a redox sensor whose model of action is based on the Cbb3-1 terminal oxidase redox state, although the underlying mechanism that controls HK activity is unknown (191). The model suggests that Cbb3-1 oxidase generates an inhibitory signal on PrrB sensor kinase under aerobic conditions (268, 269). This inhibition of PrrB triggers a silencing of genes related to photosynthesis. Other HKs among the family of RegB and PrrB sensor kinases are ActS in
IV. Redox Photosensors Based on Flavins
There are three main classes of flavin-based photosensors attending to domain receptor type, namely light-oxygen-voltage-sensing (LOV) domains, blue light sensors using FAD (BLUF) domains, and cryptochromes. They regulate many physiological responses to blue light. Changes induced by the light promote conversion between different redox and protonation states of the flavin, which are then coupled to conformational or other changes that signal physiological responses (61). For both the photolyase/cryptochrome and LOV domain photosensors, photoexcitation leads to changes to the flavin that are common in flavin-dependent enzyme reactions: an alteration in the oxidation state of the flavin or the formation of a covalent adduct (119). However, it is not yet clear whether there is a mechanism linking flavin excitation to photoreceptor activation in the BLUF protein family. The formation of a flavin adduct is not involved in BLUF domain activation; therefore, the role of electron transfer and accompanying changes in the flavin redox state remains a subject of controversy (119). A summary of the redox photosensors reviewed in this work can be found in Table 4.
Summary of Redox Photosensors
BLUF, blue light sensors using FAD; DASH,
A. LOV domains, a special class of PAS domains
LOV domains are ubiquitous regulators of phototropic responses, described as a class of PAS domains that binds FMN or FAD noncovalently (57). LOV domains are present in both chemotrophic and phototrophic bacterial species. They have been found in regulatory domains of sensor HKs, diguanylate cyclases-phosphodiesterases, DNA-binding domains, and regulators of RNAP sigma factors (145). LOV proteins exhibit the typical PAS fold, with a globular α/β-fold (LOV core) flanked by variable and often helical N- and C-terminal extensions (140). This kind of domain may be found as a single or multiple domain or associated with additional sensor domains such as GAF, cyclases, and HKs-associated sensory extracellular domains, or other PAS domains (145).
As a consequence of the light signal and changes in the flavin state, a conserved cysteine residue in the LOV domain forms a flavin adduct. Details of the photocycle of this kind of photosensors are extensively discussed in an excellent recent review (231). The LOV domain signaling involves the generation of conformational changes triggered by the conserved photochemistry of the LOV core (61). The mechanism depends on the oligomeric state, structural changes, and the type of effector domain (61). In these processes, blue light induces the unfolding of flanking helices, dimerization, and rotation of the LOV modules (61).
Two major groups of LOV proteins have been described (145). The first group, LOV–HKs, corresponds to ∼50% of bacterial LOV proteins. The second group comprises the so-called LOV–GGDEF–EAL proteins that contain conserved GlyGlyAsp/GluAsp/GluPhe and GluAlaLeu motifs. These domains could participate in metal binding and might form the phosphodiesterase active site. LOV–GGDEF–EAL proteins are predicted to regulate the synthesis and hydrolysis of cyclic di-GMP and constitute ∼20% of bacterial LOV proteins. Other less common LOV signaling proteins include LOV STAS (sulfate transporter anti-σ antagonist) proteins (∼10%), LOV HTH proteins (∼3.5%), and the LOV SpoIIE (sporulation stage II protein E) proteins (∼2%). A small number of LOV proteins with a globin domain, a CheB or CheR chemotaxis domain, or a cyclase 4 domain have also been reported (230).
Several LOV proteins have a specific DNA binding domain, activated by changes in the flavin state. In bacteria, a certain number of LOV proteins with a HTH effector domain have been identified in recent years. However, no bacterial LOV proteins have been described with a zinc finger DNA binding motif similar to the
1. LOV/helix-turn-helix DNA-binding proteins
Among the light-regulated transcriptional regulators bearing the LOV domain, there is an interesting class in which the LOV domain is fused to a HTH DNA-binding domain. Although these proteins have been well described in plants and animals, the information for bacteria is scarce. They have been described in the alphaproteobacterium
In the light-activated EL222 transcription factor from
Light-driven allosteric changes are interesting tools to control gene expression or biochemical activities. LOV/HTH proteins have also been proposed as good candidates for the design of light-controlled systems (415).
2. Short-LOV proteins
Several LOV proteins have been identified in bacteria and fungi as so-called “short” LOV proteins composed of a conserved LOV core and N- and/or C-terminal helical extensions. Owing to the absence of fused effector domain(s), the next step in signal propagation in short LOV proteins is expected to involve partner proteins, which remain unidentified (310). Interestingly, the genes encoding PpSB1–LOV (Q88E39) and PPSB2–LOV (Q88JB0) from
3. Phototropins: YtvA (PfyP) and stress response
Phototropins are membrane-associated LOV proteins that usually possess two N-terminal photoactive LOV domains (LOV1 and LOV2) and a C-terminal serine/threonine kinase domain (229). Blue light photoexcitation of the LOV domains results in the receptor autophosphorylation and initiation of the phototropin signaling (56). Phototropins are ubiquitous and they are especially abundant in plants.
YtvA from
B. Non-PAS domain photosensors
1. BLUF domains
BLUF domains are light-triggered switches that control enzyme activity or gene expression in response to blue light, remaining activated for seconds or even minutes after stimulation (283). BLUF was initially described in purple bacteria for its role in photosynthetic gene expression (238). Well-studied BLUF domains are present in proteins such as AppA, PAC-a/PAC-b, BlsA, BlrB, BlrP (YegF), and PixD (61).
The secondary structure of the BLUF domain is dissimilar to those of the PAS domains or DNA photolyases. Furthermore, no significant similarity was found between the fold of its FAD-binding region and those from other FAD-binding protein families (124). For this reason, the BLUF domain was considered a novel FAD-binding domain involved in blue light- or redox-dependent sensory transduction, with a novel FAD-binding fold (124).
BLUF proteins are unique in being the only family of photoreceptors known to show photo-induced proton-coupled electron transfer (283). The photoresponse of BLUF sensing depends on interactions of the flavin with several conserved residues of the domain. Using ultrafast time-resolved infrared spectroscopy to investigate the primary photophysics of the BLUF domain of the light-activated antirepressor AppA, Laptenok
AppA, a light- and O2-sensor antirepressor from
Crystal structures and hydrogen–deuterium exchange of AppA complexed with PpsR suggested that blue light dissociated multimeric AppA–PpsR complex from DNA but did not appreciably alter the affinity of the two protein components. Figure 8 shows the current model for the mechanism of the AppA–PpsR system (393).
BLUF domains are abundant in cyanobacteria, and several AppA homologues are annotated in the cyanobase. The PixD–PixE system is the best studied, as for instance PixD (
2. Cryptochromes
Cryptochromes are blue light/UVA photoreceptors involved in regulatory processes. They are closely related to photolyases (involved in DNA repair functions, they use light to repair UV-damaged DNA) and form the cryptochrome–photolyase family. Cryptochromes and photolyases bear a conserved N-terminal α/β domain and an α-helical domain that noncovalently binds an FAD in their catalytic center. The catalytic activity of photolyases requires the FAD to be in its two-electron reduced active state as FADH− (386). The N-terminus might bind other antenna chromophores. Cryptochromes are a relatively heterogeneous group found in different types of organisms, and they are not as well characterized as the photolyases.
Cryptochromes act as photoreceptors and transcriptional regulators, depending on the type of organisms. The classical cryptochromes show high sequence similarity to photolyases, but they lack DNA repair activity and only act as signaling molecules, regulating the circadian clock, growth, or development. However, a distinct group of the classical cryptochromes has been identified. Its homologues were found in diverse organisms (
Very few cryptochromes from bacteria have been characterized, and most of them also show photolyase activity. In addition to the Syn-CRY, CryB of
Based on structural data from RsCryB, Geisselbrecht
V. Concluding Remarks
The regulation of redox homeostasis is of paramount importance for the survival of free-living bacteria and species infecting a host. Thus, their ecological success is strongly dependent on the correct performance of a range of transcription factors that trigger the appropriate genetic program in response to different redox signals. Understanding the mechanisms of the different redox-responsive regulators has been hindered by the instability of the redox centers and prosthetic groups that are essential for their activities. It is likely that some of them still remain to be discovered. Moreover, cross talk among several regulators and the diversity of responses displayed against the same signal complicate the identification of their direct gene targets.
Fortunately, in recent decades, the development of novel biophysical tools together with the resolution of the crystal structures of several redox regulators has provided a wealth of knowledge about their response mechanisms. Furthermore, high-throughput transcriptomic analyses have allowed researchers to complete the crossroads of regulatory networks in numerous bacteria. All these data furnish researchers with valuable information that may allow the development of novel drugs and other applications in microbial biotechnology.
Footnotes
Acknowledgments
The authors acknowledge the support from the Spanish Ministry of Economy, Industry, and Competitiveness (BFU2016-77671-P) and the Government of Aragón-FEDER (B18).
