Abstract
Aims:
Bacillithiol (BSH) is the major low-molecular-weight thiol of the human pathogen Staphylococcus aureus. In this study, we used OxICAT and Voronoi redox treemaps to quantify hypochlorite-sensitive protein thiols in S. aureus USA300 and analyzed the role of BSH in protein S-bacillithiolation.
Results:
The OxICAT analyses enabled the quantification of 228 Cys residues in the redox proteome of S. aureus USA300. Hypochlorite stress resulted in >10% increased oxidation of 58 Cys residues (25.4%) in the thiol redox proteome. Among the highly oxidized sodium hypochlorite (NaOCl)-sensitive proteins are five S-bacillithiolated proteins (Gap, AldA, GuaB, RpmJ, and PpaC). The glyceraldehyde-3-phosphate (G3P) dehydrogenase Gap represents the most abundant S-bacillithiolated protein contributing 4% to the total Cys proteome. The active site Cys151 of Gap was very sensitive to overoxidation and irreversible inactivation by hydrogen peroxide (H2O2) or NaOCl in vitro. Treatment with H2O2 or NaOCl in the presence of BSH resulted in reversible Gap inactivation due to S-bacillithiolation, which could be regenerated by the bacilliredoxin Brx (SAUSA300_1321) in vitro. Molecular docking was used to model the S-bacillithiolated Gap active site, suggesting that formation of the BSH mixed disulfide does not require major structural changes.
Conclusion and Innovation:
Using OxICAT analyses, we identified 58 novel NaOCl-sensitive proteins in the pathogen S. aureus that could play protective roles against the host immune defense and include the glycolytic Gap as major target for S-bacillithiolation. S-bacillithiolation of Gap did not require structural changes, but efficiently functions in redox regulation and protection of the active site against irreversible overoxidation in S. aureus. Antioxid. Redox Signal. 28, 410–430.
Introduction
S
The success of S. aureus as a leading pathogen is caused by high diversity of different virulence factors, such as toxins, proteases, lipases, and superantigens, as well as efficient protection mechanisms against the host immune defense during invasion. During infections, S. aureus has to cope with the oxidative burst of activated macrophages and neutrophils, including reactive oxygen and nitrogen species (ROS, RNS) and the strong oxidant hypochlorous acid (HOCl) (75, 76). HOCl is produced in neutrophils by the enzyme myeloperoxidase (MPO) from hydrogen peroxide (H2O2) and chloride (44). The involvement of HOCl as prime mechanism for oxidative killing of S. aureus by neutrophils has been shown using MPO inhibitors (29). Moreover, killing of many bacteria by isolated neutrophils is strongly inhibited in the absence of MPO (44).
Using quantitative redox proteomics, 58 redox-sensitive protein thiols were identified in the methicillin-resistant Staphylococcus aureus strain USA300 that showed >10% increased oxidation under NaOCl stress. The glyceraldehyde-3-phosphate dehydrogenase Gap was identified as most abundant target for thiol oxidation and represents the major S-bacillithiolated protein in S. aureus cells. Molecular docking of bacillithiol (BSH) into the active site suggests that S-bacillithiolation does not require major structural changes. Finally, our biochemical assays confirm that S-bacillithiolation efficiently protects the Gap active site against overoxidation by H2O2 and NaOCl and inhibits Gap activity, which can be reversed by the bacilliredoxin Brx in vitro.
S. aureus uses several redox-sensing virulence regulators, such as SarA and the MarR/OhrR-type regulators, MgrA and SarZ, for defense against oxidative stress. These control large regulons of virulence factors, antibiotic resistance determinants, and ROS detoxification enzymes (11 –13, 35). MgrA and SarZ are both single Cys MarR/OhrR-type repressors that sense and respond to ROS via thiol-based redox switches and by Cys phosphorylation (63, 70). In addition, S. aureus uses the low-molecular-weight (LMW) thiol bacillithiol (BSH, Cys-GlcNAc-Mal) to maintain the reduced state of the cytoplasm. BSH plays an important role in detoxification of redox-active compounds in S. aureus since bshA mutants displayed increased sensitivities to ROS, hypochlorite, electrophiles, and the antibiotic fosfomycin (52, 64, 65). Moreover, BSH mediates protection under infection-like conditions as shown in phagocytosis assays using human macrophages (64, 65). Apart from BSH, also CoenzymeA (CoASH) and cysteine are found as abundant alternative LMW thiols in S. aureus cells (58).
Under hypochlorite stress, we have shown that BSH is also used for S-thiolation of redox-sensitive Cys residues and forms mixed disulfides with proteins that are termed as S-bacillithiolation. S-bacillithiolation protects protein thiols against overoxidation to sulfonic acids and is an important redox regulatory device in Firmicutes analogous to S-glutathionylation in eukaryotes (15, 16, 47, 52). The presence of CoASH and cysteine as LMW thiols suggests that alternative S-thiolations are also possible in S. aureus, such as S-cysteinylation or CoASH mixed disulfides. In support of this notion, S. aureus encodes a CoASH disulfide reductase (Cdr) that functions in reduction of CoASH mixed protein disulfides (55).
Using shotgun proteomics, we have previously identified 54 S-bacillithiolated proteins in different Bacillus species and Staphylococcus carnosus (16). Among these are eight conserved S-bacillithiolated proteins, such as the methionine synthase MetE, the inorganic pyrophosphatase PpaC, and the inosine-5′-monophosphate (IMP) dehydrogenase GuaB. The glutaredoxin-like YphP protein of the UPF0403 family was also S-bacillithiolated in Bacillus subtilis in vivo at its CGC active site motif (15). YphP and its paralog YqiW were renamed as BrxA and BrxB based on their function as bacilliredoxins in the reduction of S-bacillithiolated OhrR and MetE in vitro (24). Reduction of S-bacillithiolated proteins leads to Brx-SSB formation, which requires BSH and a still unknown BSSB reductase for recycling (24, 25, 31, 52). We have recently fused the YphP homolog (Brx) of S. aureus USA300 (SAUSA300_1321) to roGFP2 to construct a dynamic biosensor to monitor BSH redox potential changes in vivo (51). Brx-roGFP2 was highly specific to recognize BSSB, which confirms the role of Brx as bacilliredoxin also in S. aureus.
The physiological role of S-bacillithiolation in redox regulation has been demonstrated for the redox-sensing OhrR repressor and the methionine synthase MetE under hypochlorite stress in B. subtilis. S-bacillithiolation of MetE in its active site Zn center leads to its inactivation and subsequent methionine auxotrophy (15). The DNA-binding activity of the organic hydroperoxide repressor, OhrR, is inhibited by S-bacillithiolation under sodium hypochlorite (NaOCl) and cumene hydroperoxide stress, which results in the expression of the OhrA peroxiredoxin as ROS protection mechanism (15, 47).
However, the targets for S-bacillithiolation or reversible thiol oxidation under hypochlorite stress are unknown in the major pathogen S. aureus, which could provide leads in drug design to treat MRSA infections. In this study, we have combined the quantitative redox proteomic approach OxICAT (9, 48, 49) and shotgun proteomics to quantify NaOCl-sensitive proteins and to identify S-bacillithiolated proteins in S. aureus USA300. We found that 25% protein thiols showed >10% increased oxidation under NaOCl stress. The glycolytic Gap was identified as the most abundant S-bacillithiolated protein in S. aureus. Our results document that S-bacillithiolation protects the active site against overoxidation and inhibits Gap activity in vitro.
Results
Identification of 58 NaOCl-sensitive proteins using the quantitative redox proteomic approach OxICAT in S. aureus USA300
We were interested to study the role of BSH for S-bacillithiolation and the global thiol oxidation state under hypochlorite stress in the major pathogen S. aureus. Thus, we performed a quantitative thiol redox proteomic approach based on OxICAT (48, 49) and analyzed the percentages of thiol oxidation levels in S. aureus USA300 in response to 150 μM NaOCl stress, as determined previously (51). OxICAT is based on the differential thiol labeling of reduced Cys residues with light isotope-coded affinity tag (12C-ICAT), followed by reduction of reversible thiol oxidation (e.g., protein disulfides and S-thiolation) with Tris (2-carboxyethyl) phosphine (TCEP) and subsequent labeling of previously oxidized thiols with heavy 13C-ICAT reagent (48). Light and heavy ICAT-labeled peptide pairs show a mass difference of 9 Da after separation using mass spectrometry (MS). The quantification of the percentage of thiol oxidation for each Cys peptide is based on the calculation of the intensity of the heavy ICAT-labeled Cys peptide in relation to the total intensity of the light and heavy ICAT-labeled Cys peptides.
The OxICAT analysis enabled the quantification of the percentages of reversible thiol oxidation for 228 Cys peptides in the thiol redox proteome of S. aureus USA300 (Supplementary Table S1; Supplementary Data; Supplementary Data are available online at
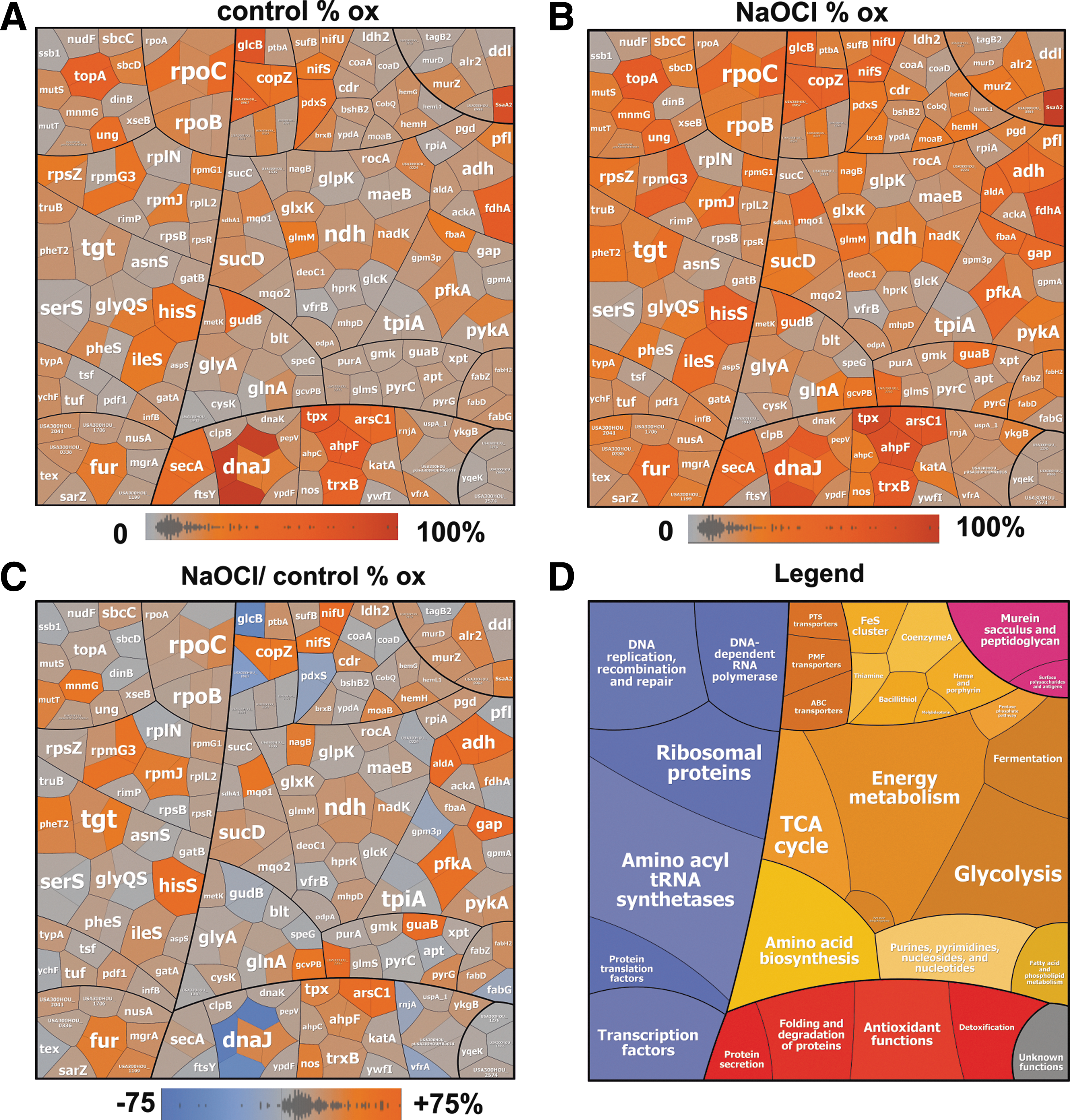
In untreated S. aureus cells, we identified 193 Cys residues (84.6%) with a thiol oxidation level of <25%, including 107 Cys residues (46.9%) with <10% oxidation, indicating that the majority of thiols are in a reduced state (Tables 1 and 2; Supplementary Table S1). Only 35 Cys residues (15.3%) showed basal-level oxidation of >25% in the control. These basal-level oxidized proteins include predicted redox-sensitive proteins (21), such as the thiol peroxidase Tpx, the alkyl hydroperoxide reductase large subunit AhpF, the arsenate reductase ArsC1, and the thioredoxin reductase TrxB1. Tpx and AhpCF were previously found as basal-level oxidized in the redox proteomes of Escherichia coli and Bacillus species (16, 48). Tpx was also S-mycothiolated in Corynebacterium glutamicum at the conserved active site Cys60 (14). In addition, the topoisomerase TopA and the DnaJ chaperone are basal-level oxidized at their Zn-binding Cys residues.
Overview of % thiol oxidation of 228 Cys peptides identified in the redox proteome of the S. aureus USA300 under control and NaOCl stress, as revealed by OxICAT. All reduced Cys peptides have an oxidation degree of <25% that include those with <10% oxidation, shown in parenthesis. Oxidized Cys peptides have an oxidation degree of >25%. The % thiol oxidation increase includes Cys peptides with no significant increased oxidation (<10%) and those with >10% increased oxidation in response to NaOCl stress compared with the control. The % thiol-oxidation and % thiol-oxidation increase and related Cys numbers are shown in bold-faced. The percentage of Cys numbers in relation to all Cys residues is shown in non-bold faced.
NaOCl, sodium hypochlorite.
Quantification of 58 Cys peptides with reversible thiol-oxidations in S. aureus USA300 that showed >10% increased oxidation under NaOCl stress using the OxICAT method. S. aureus USA300 was harvested before (control) and 30 min after exposure to 150 μM NaOCl. Reduced and reversibly oxidized Cys residues were labeled with light and heavy ICAT reagents, respectively, using the OxICAT method. Quantification of % thiol oxidation was performed using the MaxQuant software (
BSH, bacillithiol; CV, coefficient of variation; NADH, nicotinamide adenine dinucleotide; ICAT, isotope-coded affinity tag; RSA, relative surface accessibility; DH, dehydrogenase.
To discover novel NaOCl-sensitive proteins, we analyzed the percentages of thiol oxidation levels under NaOCl stress and its oxidation increase using OxICAT (Fig. 1 and Tables 2; Supplementary Table S1). The OxICAT approach enabled the identification of 58 NaOCl-sensitive Cys residues (25.4%) with >10% increased oxidation, including 19 Cys residues with 20–30% oxidation change under NaOCl stress (Tables 1 and 2 and Supplementary Table S1). Several NaOCl-sensitive proteins have antioxidant functions, such as the AhpCF peroxiredoxins, the thioredoxin reductase TrxB1, and the arsenate reductase ArsC. Furthermore, interesting proteins are the nitric oxide synthase (USA300HOU_1916) and the CoASH disulfide reductase Cdr (USA300HOU_0929), the latter is oxidized at the conserved Cys16. Apart from Cdr, the putative BSH disulfide reductase YpdA (USA300GOU_1417) was oxidized at the same conserved Cys14, but its oxidation is not increased under NaOCl stress (Supplementary Table S1). Moreover, we observed a slightly increased oxidation of the deacetylase BshB2 involved BSH biosynthesis and of the bacilliredoxin YqiW (BrxB) under NaOCl stress. The oxidation of Cdr, YpdA, BshB2, and BrxB could indicate increased S-bacillithiolation and CoASH mixed protein disulfides under NaOCl stress.
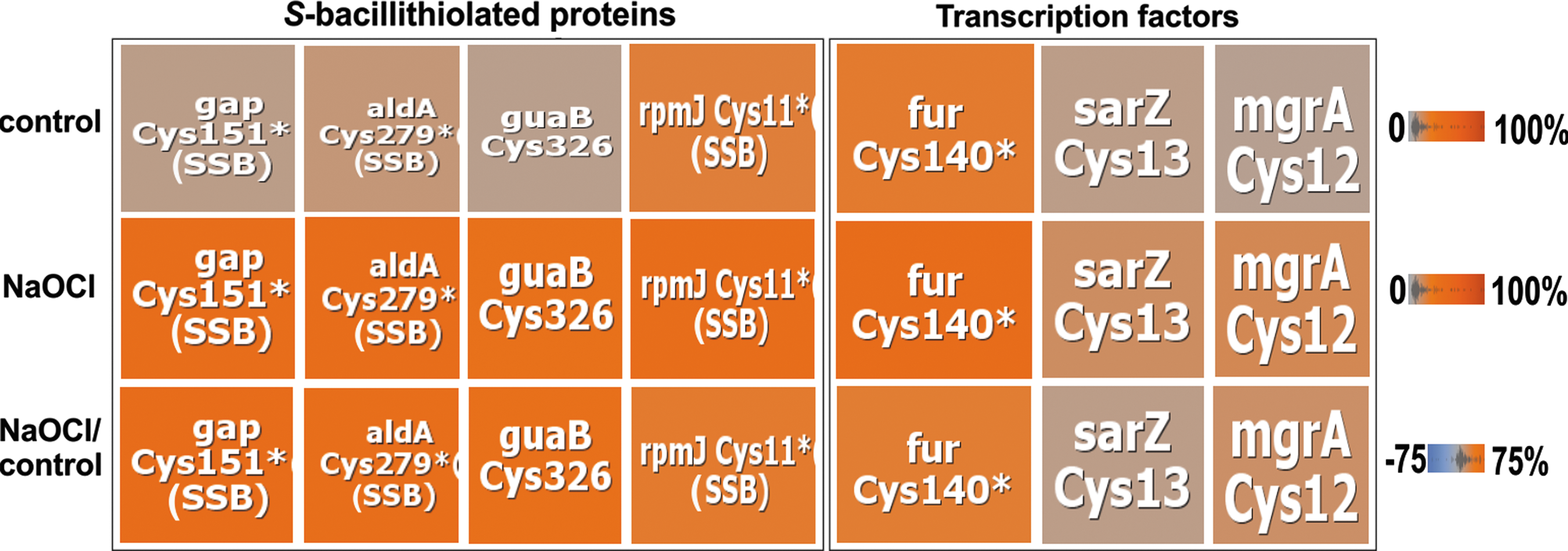
NaOCl-sensitive proteins are often oxidized in CxxC motifs and at conserved Zn-binding sites. Examples for Zn redox switches are the Zn-containing alcohol dehydrogenase Adh (USA300HOU_0610), the ribosomal proteins RpmG3 (USA300HOU_1553), and RpmJ (USA300HOU_2218). Zn-containing ribosomal proteins share three to four Cys residues that are suggested to serve as reservoir for Zn storage (54). As another Zn redox switch, we identified the ferric uptake repressor Fur that showed 16.6% increased oxidation at its Zn-binding site at Cys 140 and Cys143 under NaOCl stress (Tables 2; Supplementary Table S1; Figs. 1–2). Fur contains two CxxC motifs that form a structural Cys4:Zn site and are required for stability. In addition, two regulatory iron-binding sites are present in Fur (32). FurA of Anabaena was described as redox switch under oxidative stress and Cys101 in the CxxC motif is essential for iron-sensing and DNA-binding activity (7).

The copper chaperone CopZ was 19.8% oxidized in its CxxC motif that is required for Cu binding (67). The interaction of the B. subtilis CopZ homolog with BSH has been recently studied leading to the formation of S-bacillithiolated apo-CopZ and Cu(i)-bound forms of CopZ (42). In addition, NaOCl-sensitive Cys residues often coordinate FeS clusters or function in FeS cluster biogenesis. The FeS cluster scaffold protein NifU showed 26% increased oxidation at Cys41 that binds the FeS cluster during the assembly. The cysteine desulfurase NifS exhibits 20.6% higher oxidation levels at the catalytic Cys371 that forms the persulfide with the sulfur released during cysteine desulfuration (5). In addition, the FeS cluster assembly protein SufB is oxidized in its FeS cluster binding Cys302. It is interesting to note that the nifS-nifU-sufB genes are cotranscribed in an operon.
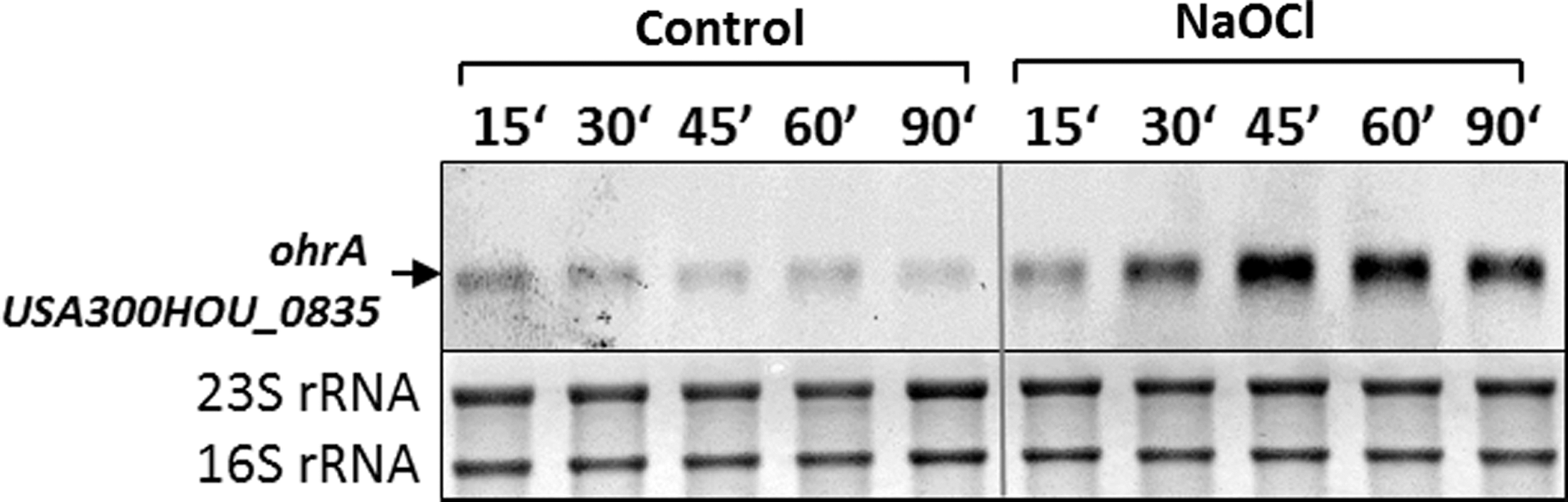
As NaOCl-sensing redox regulators, the MarR/OhrR family repressors, MgrA and SarZ (USA300HOU_0709 and USA300HOU_2368), were identified that showed 10.5% and 6.5% increased oxidation levels under NaOCl stress at their redox-sensing single Cys (Fig. 2). The DNA-binding activity of MgrA and SarZ was inhibited by S-thiolation using a synthetic thiol in vitro (11, 13, 35). In this study, increased oxidation of MgrA and SarZ was found in S. aureus under NaOCl stress, indicating that both could be redox controlled by S-bacillithiolation analogous to OhrR of B. subtilis (47). OhrR and SarZ both control a homologous ohrA peroxiredoxin gene that confers resistance to organic hydroperoxides and NaOCl in B. subtilis (13). Northern blot analyses revealed increased transcription of ohrA under NaOCl stress, indicating that SarZ oxidation leads to its inactivation and derepression of ohrA transcription (Fig. 3). We further noted the 15% increased oxidation of the virulence factor and secretory antigen SsaA2 at its conserved single Cys171 under NaOCl stress. The homologous SceB precursor (Sca_1790) of S. carnosus was previously S-bacillithiolated at the conserved Cys in NaOCl-treated cells (16). Thus, SsaA2 is most likely also S-bacillithiolated in S. aureus.
The NaOCl-sensitive proteins of S. aureus include many metabolic enzymes that function in energy metabolism and in different biosynthesis pathways for amino acids, fatty acids, nucleotides, and cofactors. NaOCl-sensitive enzymes involved in energy metabolism include the glycolytic glyceraldehyde-3-phosphate (G3P) dehydrogenase Gap and phosphofructokinase PfkA (USA300HOU_1685), the alcohol dehydrogenase Adh, the aldehyde dehydrogenase AldA (USA300HOU_2110), the formate dehydrogenase FdhA (USA300HOU_2291), and the malate dehydrogenase Mqo (USA300HOU_2348). Gap and AldA both showed the highest oxidation increase of 29% and 26% under NaOCl stress at their catalytic active sites at Cys151 and Cys279, respectively. Furthermore, the IMP dehydrogenase GuaB and the purine nucleosidase USA300HOU_2265 both displayed 25% increased oxidation under NaOCl stress (Tables 2; Supplementary Table S1).
Among the cell wall biosynthesis enzymes, the alanine racemase Alr2 (USA300HOU_2065) and the UDP-N-acetylglucosamine 1-carboxyvinyltransferase MurZ (USA300HOU_2112) were identified as NaOCl-sensitive proteins. The glucose-inhibited division protein MnmG showed 20.8% increased oxidation under NaOCl stress. Many aminoacyl-tRNA synthetases were strongly oxidized under NaOCl stress. We detected 18–24% higher oxidation levels for the histidine- and phenylalanine tRNA ligases (HisS and PheT2) and for the queuine tRNA ribosyltransferase (Tgt) under NaOCl stress.
Five S-bacillithiolated proteins were identified using shotgun proteomics in S. aureus, including the glycolytic Gap as major target
We used the previously applied shotgun proteomic approach for identification of S-bacillithiolated proteins under nonreducing conditions based on the 396 Da mass increase at Cys residues (16). Five S-bacillithiolated proteins were identified in NaOCl-treated cells of S. aureus USA300, including Gap, AldA, GuaB, RpmJ, and the manganese-dependent inorganic pyrophosphatase PpaC (Table 3; Supplementary Fig. S1
Identification of S-bacillithiolated peptides in the S. aureus USA300 wild type using shotgun LC-MS/MS analysis and their Sequest Xcorrs, ΔCn scores, and mass deviations. The S. aureus USA300 wild type was exposed to 150 μM NaOCl for 30 min and S-bacillithiolated proteins were identified using shotgun LC-MS/MS analysis and the Scaffold proteome software based on the mass increase of 396 Da (+BSH) at Cys peptides. The table lists the Uniprot accession number, protein name, function and molecular weight, the S-bacillithiolated Cys peptide sequence, and the quality control criteria for the peptide-SSB as obtained from the LC-MS/MS analysis and the Scaffold software (Xcorr, ΔCn scores, mass deviations Δppm, and ΔDa, observed and theoretical peptide masses, and charges of the modified peptide).
The S-bacillithiolated Cys is shown in bold-faced.
LC-MS/MS, liquid chromatography tandem mass spectry.
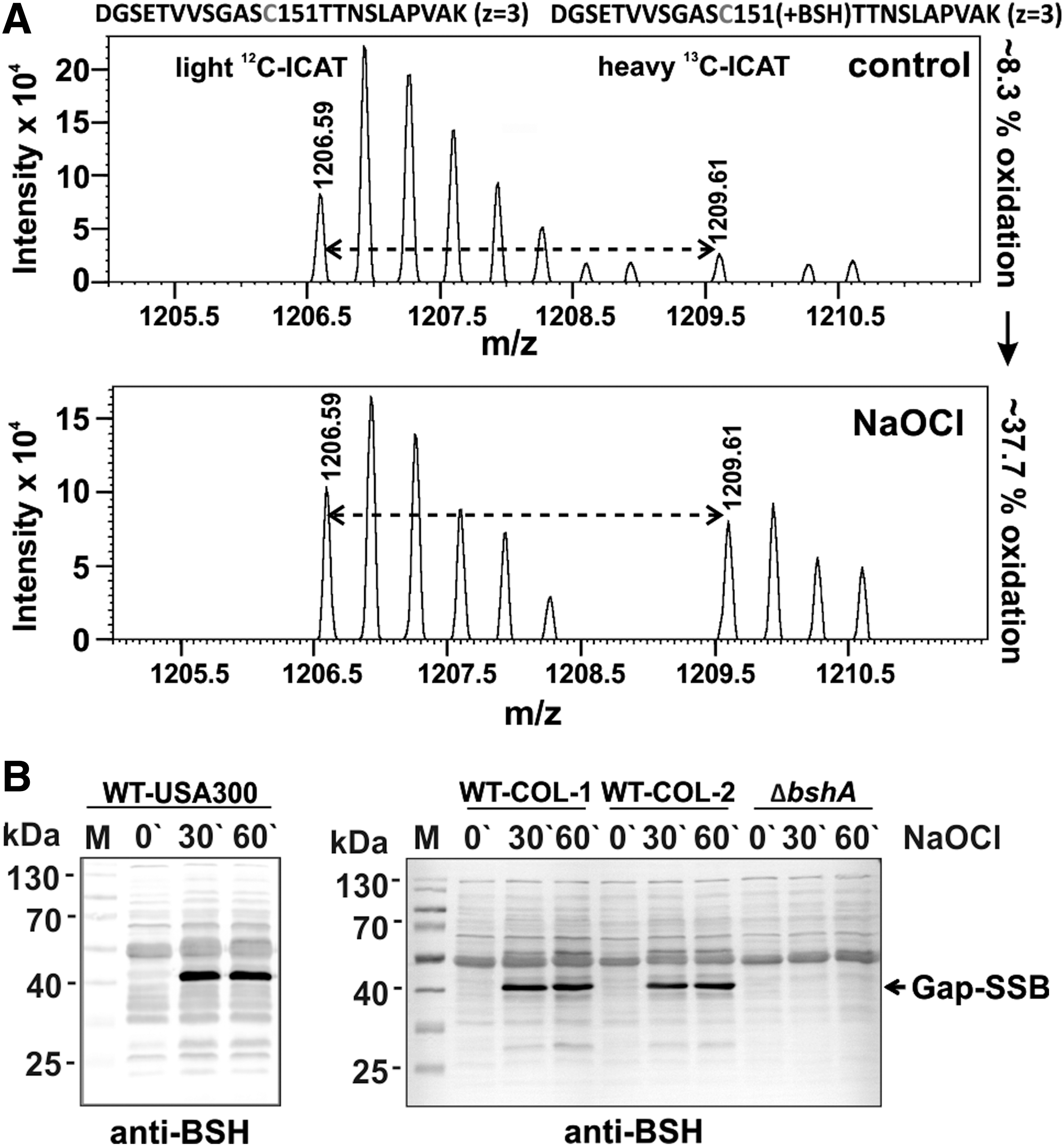
Gap and AldA were S-bacillithiolated at their catalytic active sites at Cys151 and Cys279, respectively (Fig. 2 and Table 3; Supplementary Fig. S1). The AldA homolog of S. carnosus was previously found S-bacillithiolated at Cys279 (16). The active site Cys of Gap is a conserved target for S-glutathionylation in eukaryotic Gap homologs. Cys151 of Gap showed 29.5% oxidation increase under NaOCl stress in the OxICAT analysis, which is reflected also by the mass spectra of the ICAT-labeled Cys151-peptides (Fig. 4A
Gap contributes as most abundant Cys protein with 4% to the total Cys proteome
We were further interested in the contribution of Gap and other S-bacillithiolated proteins to the total Cys proteome of S. aureus. S. aureus USA300 encodes for 2694 proteins. These include 1864 proteins with 4935 Cys residues, indicating that the Cys content is 0.64% in the theoretical proteome (Supplementary Fig. S2A, B
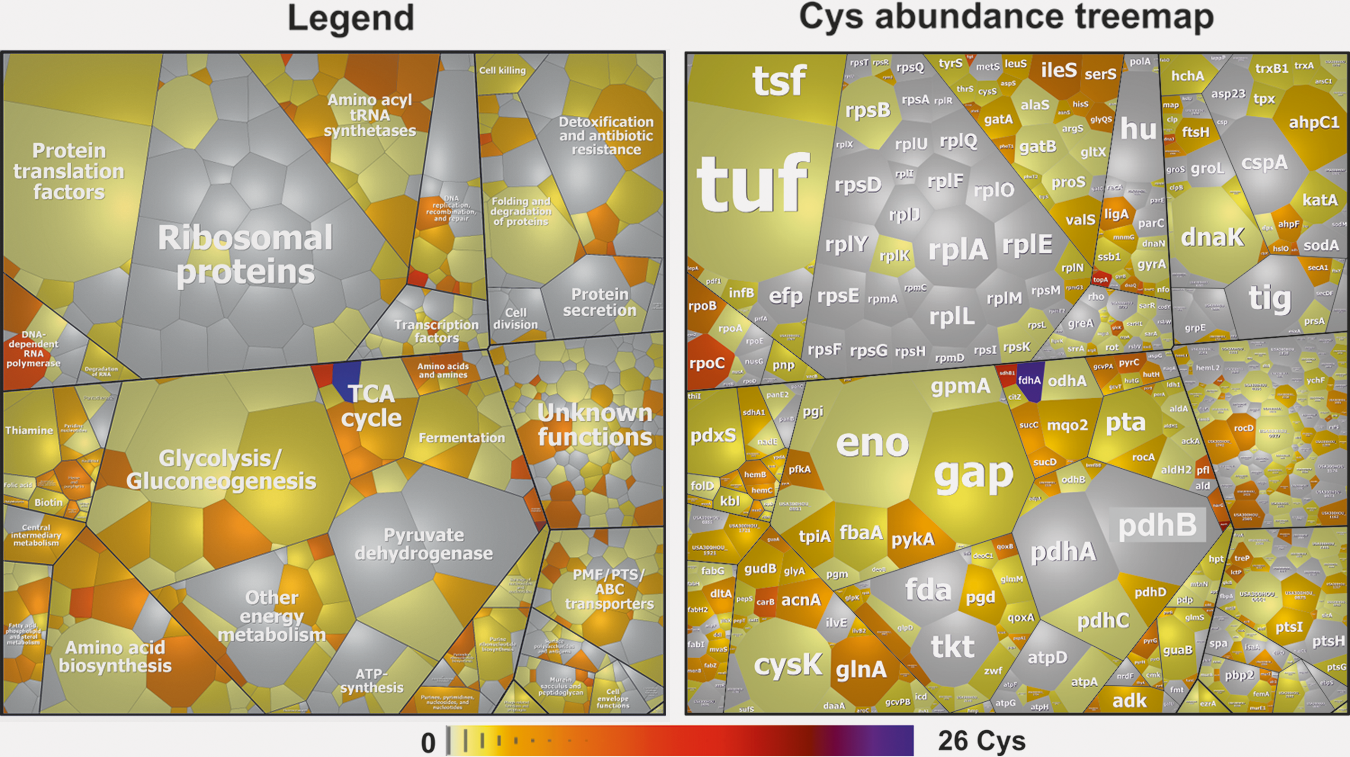
The S-bacillithiolated Gap was identified among the most abundant Cys-containing proteins and contributes with 4% of the total Cys proteome (Supplementary Fig. S2C). This indicates that Gap makes the major contribution to the S-bacillithiolome of S. aureus as visualized also by the BSH Western blots. The other S-bacillithiolated proteins, AldA, RpmJ, GuaB, and PpaC, are less abundant and make with 0.1–0.7% of Cys abundance only a minor contribution to the total Cys proteome (Supplementary Table S2).
H2O2 and NaOCl-induced inactivation pathways of Gap in S. aureus due to overoxidation and S-bacillithiolation in vitro
The active site of Gap is usually present in a highly conserved CTTNC motif in different organisms (Supplementary Fig. S3
Gap of S. aureus was purified as His-tagged protein from E. coli. The inhibition of Gap activity by increasing H2O2 concentrations was monitored spectrophotometrically with G3P as substrate in the presence of NAD+. The remaining Gap activity was determined by nicotinamide adenine dinucleotide (NADH) generation as absorbance change at 340 nm during the slope in the reaction, as described previously (61). Treatment of Gap with 100 μM H2O2 leads to a 50% decrease in Gap activity, while exposure to 1–10 mM H2O2 resulted in complete enzyme inactivation (Fig. 6A). Inactivation of Gap with 1–10 mM H2O2 alone was irreversible due to overoxidation since Gap activity could be not restored with 10 mM dithiothreitol (DTT) (Fig. 6C). To investigate whether S-bacillithiolation can protect the enzyme against irreversible overoxidation, Gap was pretreated with 10-fold molar excess of BSH before H2O2 exposure. Gap activity was already 90% inhibited after oxidation with 100 μM H2O2 in the presence of BSH, while treatment with 100 μM H2O2 alone only led to 50% decreased activity (Fig. 6A, B). Gap inactivation with H2O2 and BSH was caused by reversible S-bacillithiolation since DTT reduction resulted in recovery of Gap activity (Fig. 6C–E; Supplementary Fig. S9). These results support that the Gap active site is highly sensitive to overoxidation, which can be prevented by S-bacillithiolation in the presence of H2O2 and BSH.
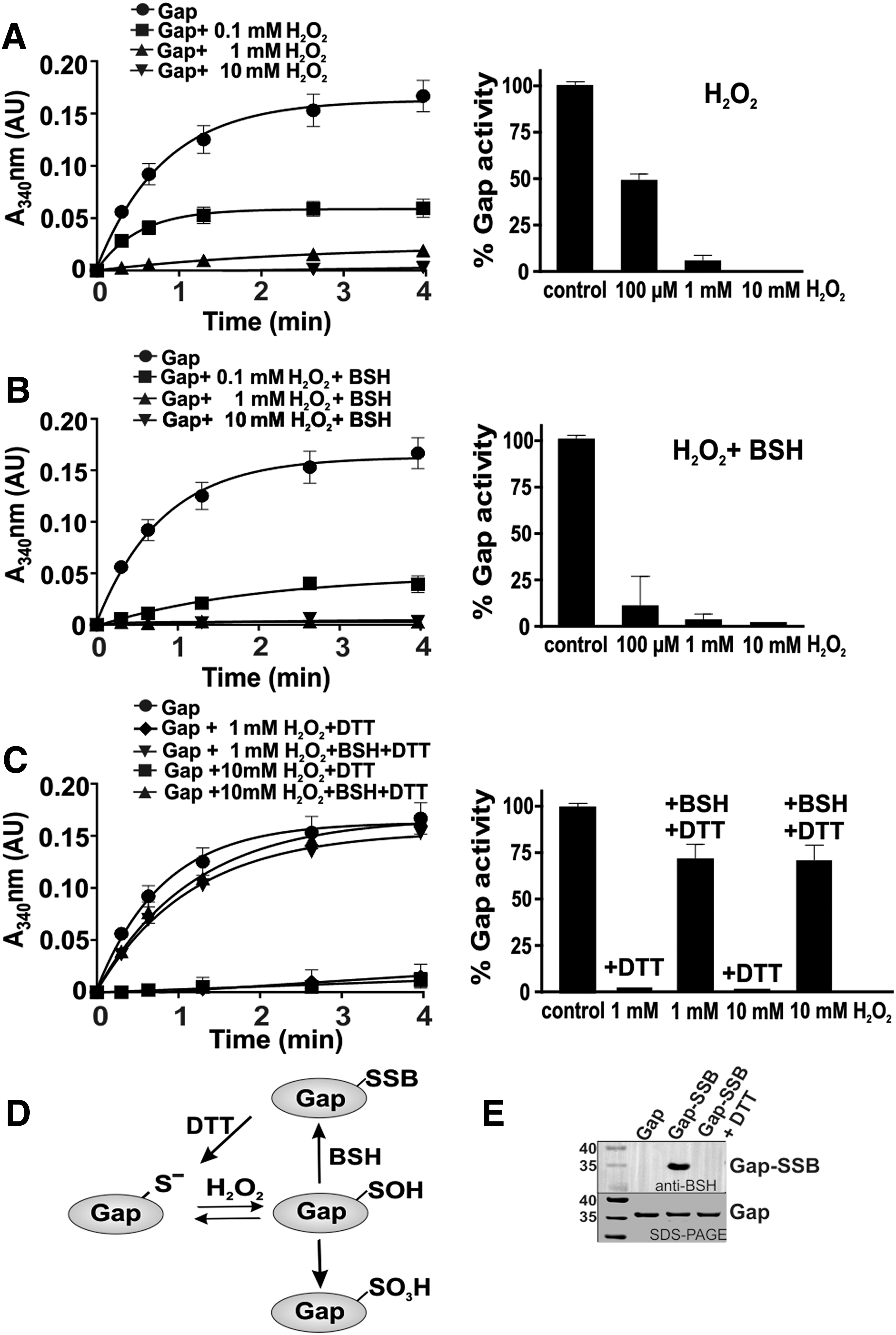
Next, we determined the time-dependent Gap inactivation by both H2O2-dependent oxidation pathways (Supplementary Fig. S4). Gap was treated with 1 mM H2O2 on ice with or without BSH and the remaining Gap activity was determined after different times of H2O2-dependent overoxidation and S-bacillithiolation. The Gap activity assays revealed that both S-bacillithiolation and overoxidation lead to 80% enzyme inhibition after 7.5 min of H2O2 treatment (Supplementary Fig. S4A). In addition, we analyzed the time course for the detection of Gap-SSB or the overoxidized Cys151 under H2O2 treatment with or without BSH using BSH-specific Western blots or MS, respectively. The MS results identified the overoxidized Cys151 sulfonic acid (Cys151-SO3H) after 1 min of H2O2 treatment (Supplementary Fig. S5). The S-bacillithiolated Gap could be also detected after 1 min of treatment with BSH and H2O2 (Supplementary Figs. S4B and S9). These results suggest that overoxidation and S-bacillithiolation occur at similar rates under H2O2 treatment in vitro. However, the Gap activity assays after treatment with different H2O2 concentrations indicate that Gap inhibition is faster with 100 μM H2O2 in the presence of BSH compared with 100 μM H2O2 alone, which only leads to 50% enzyme inhibition (Fig. 6A, B). Thus, S-bacillithiolation of Cys151 by H2O2 in the presence of BSH serves to protect the active site against overoxidation.
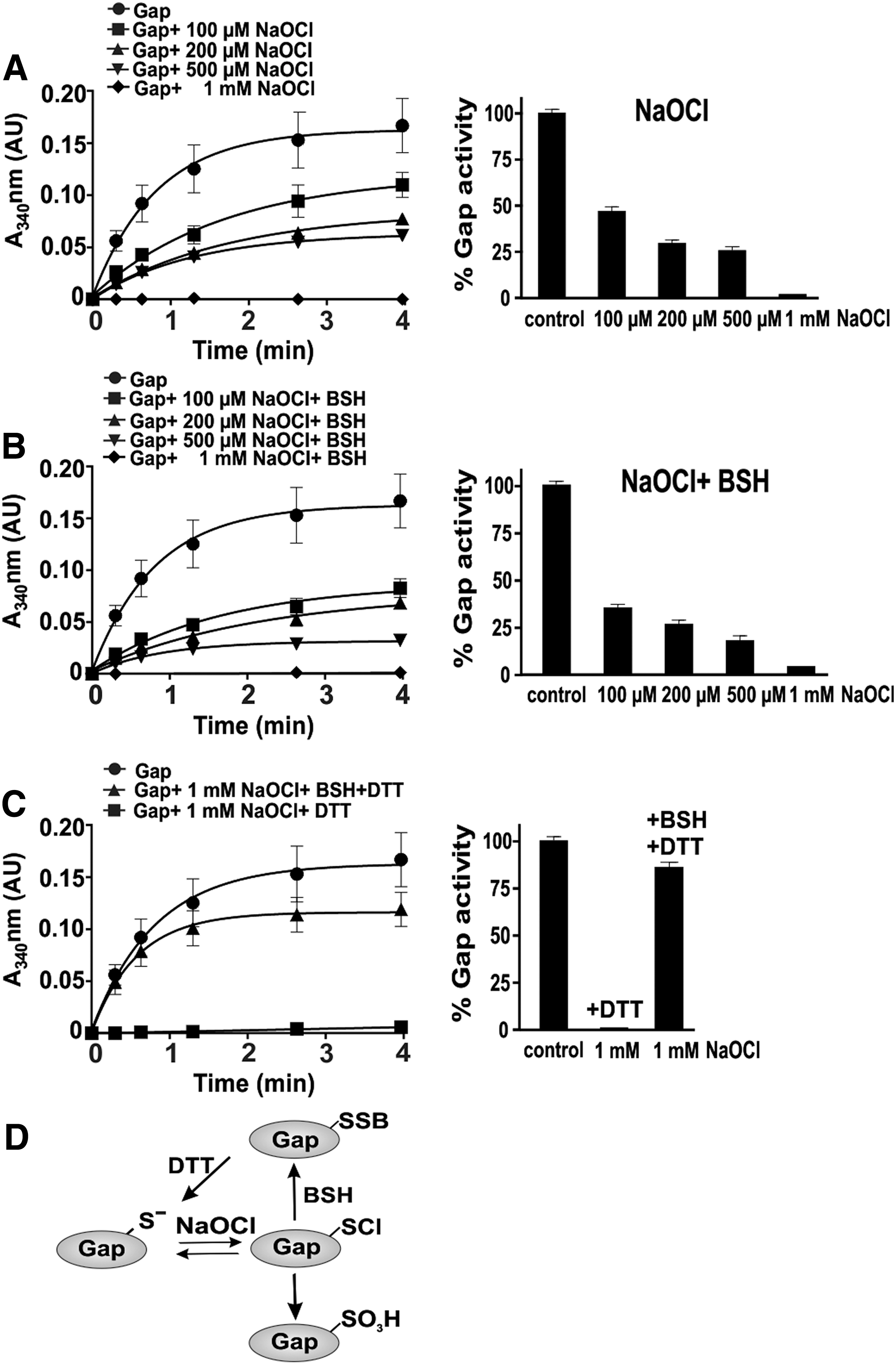
Since S-bacillithiolation of Gap was observed under NaOCl stress in vivo, we studied the dose-dependent Gap inactivation by NaOCl with or without prior exposure to BSH (Fig. 7). Treatment of Gap with 100–500 μM NaOCl led to 50–75% inhibition of Gap activity. Pretreatment of Gap with BSH before exposure to 100 μM NaOCl resulted in 70% activity decrease. Gap was fully inactivated with 1 mM NaOCl in the absence or presence of BSH. Treatment of Gap with 1 mM NaOCl alone resulted in irreversible inactivation due to overoxidation since Gap activity could be not restored using DTT. In the presence of BSH, Gap inactivation by NaOCl was caused by reversible S-bacillithiolation since 85% Gap activity could be restored by DTT reduction (Fig. 7C, D). Next, we studied the time course for NaOCl-induced overoxidation and S-bacillithiolation pathways in the presence of 1 mM NaOCl. The Gap activity assays with or without BSH showed that Gap inhibition is faster with BSH and NaOCl compared with NaOCl alone (Supplementary Fig. S6). These results indicate that S-bacillithiolation can efficiently prevent overoxidation of the Gap active site under NaOCl in vitro, supporting our in vivo finding.
Regeneration of S-bacillithiolated Gap using the bacilliredoxin Brx (SAUSA300_1321) in vitro
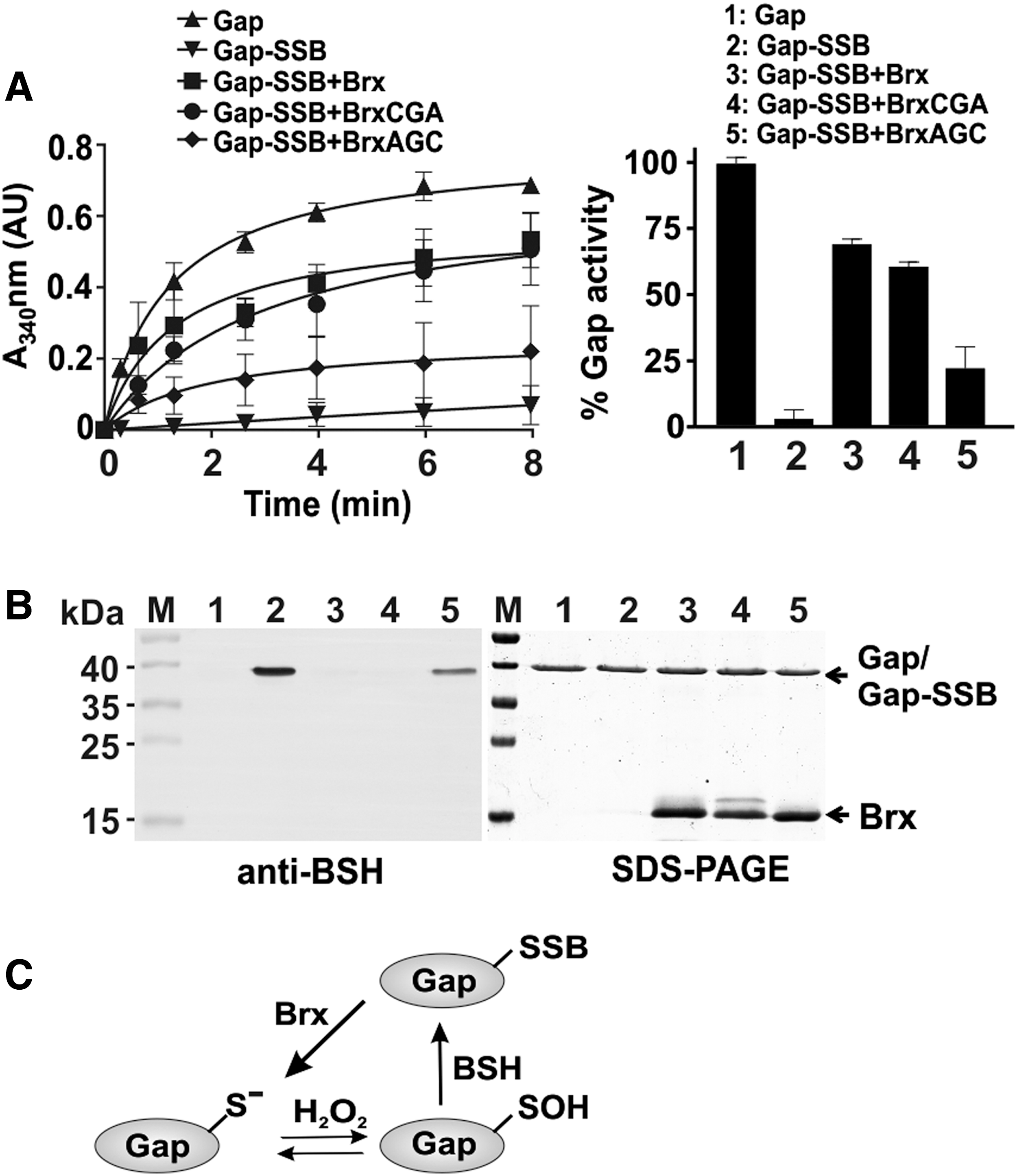
The reversal of S-bacillithiolation was shown to require the glutaredoxin-like bacilliredoxins, YphP (BrxA) and YqiW (BrxB), in B. subtilis (24). Using a Brx-roGFP2 biosensor, we demonstrated recently that the YphP homolog of S. aureus (SAUSA300_1321 or Brx) is highly specific as bacilliredoxin to recognize BSSB (51). Thus, Gap activity was measured after debacillithiolation of Gap-SSB with Brx and Brx Cys mutant proteins (BrxCGA, BrxAGC) and G3P oxidation was followed by NADH production as absorbance change at 340 nm (Fig. 8A). Gap activity could be restored to 70% and 60% during debacillithiolation with Brx and the BrxCGA resolving Cys mutant in vitro, respectively. However, Gap activity was only 25% recovered with the BrxAGC active site mutant protein supporting the specificity of the Brx active site for the attack of BSH mixed disulfide. Debacillithiolation of Gap-SSB by Brx and the BrxCGA mutant was verified in BSH-specific Western blots (Fig. 8B; Supplementary Fig. S9). These results indicate that S-bacillithiolation of Gap functions in protection and redox regulation of the active site Cys and can be reversed by the bacilliredoxin Brx in vitro (Fig. 8C).
Structural features of the Gap active site during overoxidation and S-bacillithiolation
We were interested in structural changes of Gap after overoxidation and S-bacillithiolation. The crystals of H2O2-treated overoxidized Gap diffracted X-rays to 2.6 Å resolution and belonged to the P212121 space group. Previously, several crystal structures of the Gap holo- and apoenzyme have been reported with the protein always crystallized in the P21 space group (57). The structure of the overoxidized Gap contains four monomers in an asymmetric unit, each consisting of the NAD+-binding domain (residues 1–150) and the catalytic domain (residues 151–336) (57) (Supplementary Fig. S7A
According to our MS results and previous publications (73), the Gap sulfonic acid was identified by MS as overoxidized form. In the structure of overoxidized Gap, the sulfonic acid form could be modeled into the electron density of the active site Cys151 in each monomer (Supplementary Fig. S7B, C). Overoxidation of Cys151 results in enzyme inhibition as supported by our activity assays. During catalysis, the sulfhydryl group of Cys151 attacks the nucleophilic carbon of the G3P substrate to form a covalent intermediate, thiohemiacetal (72). In the active enzyme, His178 forms an ion pair with Cys151, which increases the acidity and nucleophilicity of the thiol group. During G3P oxidation, His178 hydrogen bonds with the acyl carbonyl of the substrate and stabilizes the hemithioacetal intermediate (57). Apart from interfering with the function of Cys151, the sulfonyl moiety of the hyperoxidized Cys151 also interacts with the main chain carbonyl of Asn316 and the imidazole ring of His178 (Fig. 9A; Supplementary Fig. S7D). Thus, hyperoxidation of Cys151 affects the function of two key catalytic residues of Gap, Cys151, and His178, leading to irreversible inactivation of the enzyme.
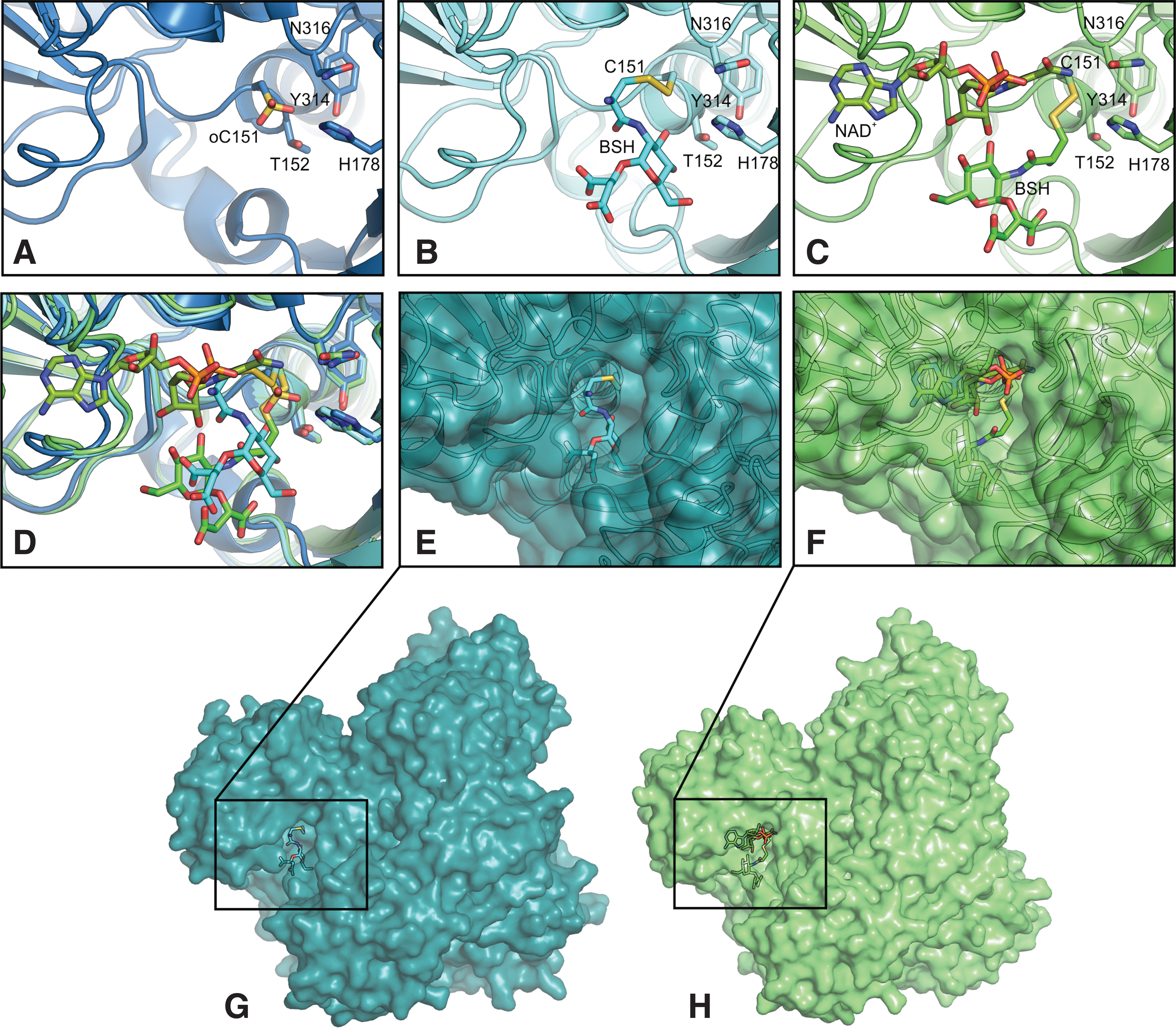
To obtain insights into the structural changes upon S-bacillithiolation, BSH was modeled into the active site of the apo- and holoenzyme structures using molecular docking (Fig. 9B, C). We used a covalent docking algorithm that takes into account the possibility of bond formation between ligand and receptor. Docking of BSH into the apo- or holoenzyme structure resulted in a set of covalent complexes (10 best poses), in which the disulfide bond can readily form and which are structurally very similar at least in the vicinity of the disulfide bond, suggesting a high confidence in the docking pose (Supplementary Fig. S8). In the holoenzyme structure, the NAD+ cofactor partially occludes the binding pocket and narrows the space available for BSH binding. As a result, in the holo-Gap active site, where NAD+ is present, BSH takes up conformations, which differ significantly less compared with the ones in the apoenzyme. When superimposing the two best binding poses, BSH in the apoenzyme structure partially occupies the part of the pocket where NAD+ would be present (Fig. 9D). However, in both cases, S-bacillithiolation of the active site does not require major conformational changes of the protein (Fig. 9E–H). In addition, previous molecular dynamic simulations of human GAPDH (61) suggested little fluctuations of the protein. Taken together, we suggest that BSH can undergo disulfide formation with the active site at little energetic or entropic costs. This may further explain why Gap as the most abundant redox-sensitive protein in the proteome of S. aureus is also the most abundant S-bacillithiolated protein under NaOCl stress.
Discussion
Dynamic thiol redox switches are the hallmarks of oxidative stress response and the major principle of redox signaling mechanism in prokaryotes and eukaryotes (33). Quantitative redox proteomic methods such as OxICAT allow to determine the redox state of proteins thiols and to dissect redox-sensitive thiols at high resolution (9, 48). To date, global thiol redox proteomics in the major pathogen S. aureus identified only few targets for oxidation under H2O2 stress (19, 77). However, S. aureus shows remarkable resistance to 100 mM H2O2, which is attributed to the constitutive expression of the catalase KatA indicating that S. aureus is well adapted to peroxide stress during infections (36).
In this study, we used OxICAT to monitor the redox state of 228 Cys residues in S. aureus USA300 under more severe infection-like conditions as provoked by hypochlorite. In untreated cells, the majority of thiols (84.6%) are reduced with an oxidation degree <25%, which is in agreement with previous studies in E. coli and yeast cells (9, 48). Under NaOCl stress, 58 NaOCl-sensitive proteins showed >10% increased oxidation, indicating that 25% of all identified protein thiols are redox sensitive in S. aureus. To analyze whether these NaOCl-sensitive Cys residues are buried or surface exposed, we calculated their relative surface accessibilities (RSAs) using NetSurfP (
Among the NaOCl-sensitive proteins with the highest oxidation increase of 20–30%, we identified Gap, AldA, and GuaB as S-bacillithiolated at their conserved substrate-binding active sites. Only a few S-bacillithiolated proteins were found by the shotgun proteomic approach due to the instability and low abundance of the BSH-modified peptides. Thus, we assume that many more NaOCl-sensitive proteins of our OxICAT dataset are modified by S-bacillithiolation, but failed to be identified using the shotgun method.
Apart from S-bacillithiolation, also alternative S-thiolations are possible in S. aureus under NaOCl stress such as S-cysteinylation or CoASH mixed disulfides since cysteine and CoASH are also present in the thiol metabolome of S. aureus (58, 64). Moreover, the Cdr displayed an increased oxidation level under NaOCl stress at its conserved Cys16 in our OxICAT analyses and was previously shown to function in reduction of CoASH disulfides in S. aureus (18, 55). However, S-cysteinylation and CoASH mixed disulfides were not detected by MS due to their low abundance or instability.
Many Zn-containing NaOCl-sensitive proteins were identified, such as the Fur repressor, which is oxidized in its Zn-binding site. Zn-binding sites are common redox switch motifs (37). The best-studied example is the oxidation-sensitive Hsp33 chaperone that responds to hypochlorite by a redox switch in its Zn motif and protects E. coli proteins against oxidative aggregation (26, 38, 46). NaOCl-sensitive Zn-containing proteins include the alcohol dehydrogenase Adh and the ribosomal proteins, RpmG3 and RpmJ. Several previously predicted redox-sensitive Cys residues are found in CxxC motifs, such as the copper chaperone, CopZ, and the antioxidant proteins, AhpC, TrxB, and ArsC (21). These results are in agreement with previous redox proteomic results in E. coli under NaOCl stress (48). Increased oxidation levels were detected for both MarR/OhrR family regulators MgrA and SarZ that are oxidized at their single Cys residues. Using Northern blot analyses, we confirmed that SarZ oxidation leads to derepression of transcription of the ohrA gene (USA300HOU_0835) in S. aureus. Thus, the OxICAT approach identified increased oxidation of both major thiol redox regulators under NaOCl stress in S. aureus.
The most abundant S-bacillithiolated protein was the glycolytic Gap enzyme in S. aureus under NaOCl stress, which was S-bacillithiolated at the active site Cys151. Gap is the most abundant Cys-containing protein contributing 4% to the total Cys proteome. The active site Cys is used for the nucleophilic attack at the aldehyde group of the G3P substrate to catalyze the substrate-level phosphorylation of G3P to 1,3-bisphosphoglycerate with production of NADH (34). Gap homologs are common targets for oxidation in eukaryotes and prokaryotes and subject of different post-translational thiol modifications, including S-sulfenylation, S-glutathionylation, S-nitrosylation, and S-sulfhydration, resulting in reversible enzyme inactivation (10, 34). Inactivation of Gap has been shown to redirect the glycolytic flux into the pentose phosphate pathway to supply nicotinamide adenine dinucleotide phosphate (NADPH) as reducing power under oxidative stress (10, 66).
Gap of S. aureus was previously identified as oxidation-sensitive target for reversible thiol modification (19) and was also found to be overoxidized at its active site Cys151 under H2O2 stress (73). Using biochemistry, MS, and X-ray crystallography, we confirmed previous findings that the glycolytic Gap enzyme from S. aureus is highly sensitive to overoxidation to Cys sulfonic acid in vitro in the presence of H2O2 alone. In this work, we found that Gap is the most abundant S-bacillithiolated protein under NaOCl stress in vivo. We further demonstrate that S-bacillithiolation functions in reversible inhibition of Gap activity under H2O2 and NaOCl treatment in vitro and protects the active site Cys against overoxidation to ensure fast regeneration of this essential glycolytic enzyme during recovery of cells. Our Gap activity assays suggest that both pathways, the overoxidation and S-bacillithiolation, operate at similar kinetics under H2O2 treatment, while inactivation due to S-bacillithiolation was faster under NaOCl stress. Together, our results confirm the preference for formation of S-thiolation in the presence of LMW thiols as observed in many eukaryotic Gap homologs (34, 43, 78).
The reactivity of the active site cysteine toward H2O2 and the substrate G3P was recently shown to depend on two different mechanisms (34, 61). The catalytic Cys is in close proximity with His178 in the structure of Gap of S. aureus that attracts the thiol proton, leading to deprotonation and increased acidity of the catalytic Cys. The acidic nature explains the reactivity of catalytic Cys toward the substrate G3P that covalently reacts with the nucleophilic thiolate during the catalytic cycle (34, 62). However, the increased acidity of Cys151 does not explain its strong reactivity toward H2O2. Instead, the reactivity of the active site thiolate depends on a specific H2O2-binding pocket, transition state stabilization, and a proton relay mechanism promoting leaving group departure (34, 61).
This proton relay mechanism also determines the preferred modification by S-bacillithiolation of Gap in S. aureus under H2O2 in vitro, which requires the initial formation of a sulfenic acid, followed by reaction to the BSH mixed disulfide. HOCl shows very fast reaction rates with thiols (3 × 107 M −1s −1) that are several orders of magnitude higher compared with H2O2 (17, 27, 30, 60, 74). HOCl first leads to chlorination of thiols resulting in an unstable sulfenylchloride intermediate that reacts further to form disulfides in the presence of another thiol. In the absence of proximal thiols, the sulfenylchloride quickly leads to irreversible oxidation stages (17, 30, 52). We have shown in S. aureus that S-bacillithiolation functions in protection and redox regulation of the Gap active site against overoxidation under NaOCl stress in vitro and in vivo. Molecular docking of BSH into the active site of the Gap apo- and holoenzyme was used to model the S-bacillithiolated active site at high confidence. The model of Gap-SSB structure suggests that BSH can undergo disulfide formation with Cys151 without major conformational changes. This may explain why the most abundant Cys protein Gap is the preferred and dominant target for S-bacillithiolation inside S. aureus cells.
S-glutathionylation of the active site Cys of Gap was found in many eukaryotic organisms, such as Arabidopsis thaliana, the malaria parasite Plasmodium falciparum, or in human endothelial cells, leading to reversible inhibition of Gap activity (3, 43, 69). Reactivation of Gap was catalyzed by glutaredoxins and thioredoxins in plants and malaria parasites (3, 43). In B. subtilis, the bacilliredoxins, BrxA and BrxB, were shown to catalyze the reduction of S-bacillithiolated OhrR and MetE in vitro (24). In this study, we showed that S-bacillithiolated Gap is also a substrate for the bacilliredoxin Brx (SAUSA300_1321) in S. aureus, which requires the active site Cys for debacillithiolation in vitro. Thus, the bacilliredoxin pathway is also involved in regeneration of Gap activity in S. aureus.
Materials and Methods
Bacterial strains and growth conditions
Bacterial strains used were S. aureus COL and USA300 and its isogenic bshA mutants as described previously (64). For cloning and genetic manipulation, E. coli DH5a and BL21 (DE3) plysS were cultivated in Luria Bertani (LB) medium. For NaOCl stress experiments, S. aureus USA300 and COL strains were cultivated in LB medium until an optical density at 540 nm (OD540) of 2.0, transferred to Belitsky minimal medium, and treated with 150 μM NaOCl stress as described (51). NaOCl, diamide, DTT, N-ethylmaleimide (NEM), and H2O2 (35% w/v) were purchased from Sigma-Aldrich.
MS-based thiol redox proteomics using the OxICAT approach
S. aureus USA300 was harvested before and after exposure to 150 μM NaOCl for 30 min, respectively. The OxICAT method was performed according to the protocol of Lindemann and Leichert (49) with the modification that cells were disrupted using a ribolyzer. The ICAT-labeled peptides were dissolved in 0.1% (v/v) acetic acid and loaded onto self-packed LC columns with 10 μl of buffer A (0.1% (v/v) acetic acid) at a constant pressure of 220 bar without trapping. Peptides were eluted using a nonlinear 85-min gradient from 1% to 99% buffer B (0.1% (v/v) acetic acid in acetonitrile) with a constant flow rate of 300 nl/min and measured using Orbitrap MS as described (6). The S. aureus USA300 sequence database was extracted from Uniprot and used by the search engine Andromeda and the MaxQuant software (version 1.5.1.2) to quantify the ICAT-labeled Cys peptides. Two miscleavages were allowed, the parent ion mass tolerance was 10 ppm and the fragment ion mass tolerance was 1.00 Da. The average percentage of oxidation of each Cys peptide and the percentage change under NaOCl stress were calculated from 2 to 3 biological replicates using the intensity values provided by MaxQuant. Voronoi treemaps were generated using the Paver software to visualize the percentage oxidation of all identified ICAT-labeled peptide pairs. The OxICAT proteomic data have been deposited to the ProteomeXchange Consortium via the PRIDE partner repository with the dataset identifier PXD004918.
Identification of S-bacillithiolated and overoxidized Cys peptides using LTQ-Orbitrap MS
For identification of S-bacillithiolated peptides, NEM-alkylated protein extracts were prepared from S. aureus USA300 cells after exposure to 150 μM NaOCl for 30 min as described (15). The protein extracts were separated by 15% nonreducing sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE), followed by tryptic in-gel digestion and LTQ-Orbitrap-Velos MS, as described (15). Post-translational thiol modifications of proteins were identified by searching all tandem mass spectrometry (MS/MS) spectra in dta format against the S. aureus USA300 target–decoy protein sequence database extracted from UniprotKB release 12.7 (UniProt Consortium, Nucleic acids research 2007, 35, D193-197) using Sorcerer™-SEQUEST® (Sequest v. 2.7 rev. 11, Thermo Electron, including Scaffold 4.0; Proteome Software, Inc., Portland, OR). The SEQUEST search parameters and thiol modifications were used as described (15). The scores and mass deviations of the S-bacillithiolated peptides are shown in detail in Supplementary Figure S1, including their fragmentation spectrum and ion tables.
MS of the H2O2-treated overoxidized Gap was performed after in-gel tryptic digestion using nLC-MS/MS by Orbitrap fusion, as described previously (45). For Cys151-SO3H peptide identification and quantification, MS1 data were filtered to the precursor target masses applying an m/z window of 3 ppm. Isotopic distribution and fragmentation spectra were inspected manually in different charge states in successive MS2 scans in different overoxidized Gap samples.
Cloning, expression, and purification of the S. aureus Gap, Brx, and Brx Cys-Ala mutant proteins in E. coli
The previously constructed plasmids, pET11b-Brx-roGFP2, pET11b-BrxAGC-roGFP2, and pET11b-BrxCGA-roGFP2 (51), were used as template to amplify S. aureus brx (SAUSA300_1321), brxAGC, and brxCGA by PCR using primer pairs 1321-roGFP2-For-NheI (5′-CTAGCTAGCATGAATGCATATGATGCTTATATGAAAG-3′) and roGFP2-1321-Rev-BamHI (5′-CGCGGATCCTTAGTGATGGTGATGGTGATGTTTACAATTT TCGTCAAAGGC-3′). The reverse primer also encodes the C-terminal His6-tag. The PCR products were digested with NheI and BamHI and inserted into plasmid pET11b (Novagen) that was digested using the same restriction enzymes to generate plasmids pET11b-brx, pET11b-brxAGC, and pET11b-brxCGA. The primer pairs gap-For-NdeI (5′-GGAATTCCATATGGCAGTAAAAGTAGCAATTAATG-3′) and gap-Rev-BamHI (5′-CGCGGATCCTTAGTGATGGTGATGGTGATGTTTAGAAAGTTCAGCTAAGTATGC-3′) were used to amplify the S. aureus gap gene (SAUSA300_0756) by PCR. Chromosomal DNA of S. aureus USA300 was used as template. The PCR products were digested with the restriction enzymes, NdeI and BamHI, and inserted into plasmid pET11b that was digested with the same enzymes to generate plasmids pET11b-gap. The correct sequences of the cloned genes were confirmed by sequencing. The plasmids were transformed into E. coli BL21 (DE3) plysS (Novagen).
For protein expression, E. coli BL21(DE3) plysS strains with the plasmids, pET11b-gap, pET11-brx, pET11b-brxAGC, and pET11b-brxCGA, were grown in 1 liter LB medium and 1 mM isopropyl-β-D-thiogalactopyranoside (IPTG) was added at the exponential phase (OD600 of 0.8) for 3 h at 37°C. His-tagged proteins were purified using His Trap™ HP Ni-NTA columns and the ÄKTA purifier liquid chromatography system (Amersham Bioscience). The proteins were further concentrated to 2–6 mg/ml using Amicon Ultra concentrators (Millipore). Before the activity assays, Gap and Brx proteins were reduced with 10 mM DTT for 30 min, followed by DTT removal using Micro Biospin 6 columns (Biorad).
Gap activity assay
Gap activity was monitored spectrophotometrically at 340 nm and 25°C by the production of NADH. The oxidation of G3P to 1,3-bisphosphoglycerate (1,3-BPG) was measured in an assay mixture containing 1.25 mM NAD+ and 0.25 μM Gap in argon-flushed 20 mM Tris-HCl, pH 8.7, with 1.25 mM ethylenediaminetetraacetic acid and 15 mM sodium arsenate. After preincubation, the reaction was started by addition of 0.25 mM D,L-G3P. Sodium arsenate was used as a cosubstrate to form unstable 1-arseno-3-phosphoglycerate, as described previously (61). Degradation of the product allows a favorable equilibrium for measuring the rate of Gap activity in the glycolytic forward reaction. Initial rates were determined by calculation of the slope in the linear part of the curve during the first 80 seconds at the beginning of the reaction (linear regression function, GraphPad) as described previously (61). Percentage of Gap activity was calculated as (Rateinactivated/Rateuntreated x 100%). The results are presented as mean ± SEM from at least three separate experiments.
S-bacillithiolation of Gap in vitro and reduction by the bacilliredoxin Brx
About 25 μM of purified Gap was S-bacillithiolated with 250 μM BSH in the presence of 2.5 mM H2O2 for 5 min. Excess of BSH and H2O2 was removed with Micro Biospin 6 columns (Biorad). For the Brx debacillithiolation assay, Gap-SSB was incubated with Brx, BrxCGA, and BrxAGC at 37°C for 30 min, followed by Gap activity assays and nonreducing BSH-specific Western blot analysis, as described (16).
Western blot analysis
The S-bacillithiolated proteins were harvested from S. aureus USA300 wild-type and bshA mutant cells after exposure to 150 μM NaOCl, separated by nonreducing SDS-PAGE, and subjected to BSH-specific Western blot analysis using the polyclonal rabbit anti-BSH antiserum, as described previously (16).
Northern blot experiments
Northern blot analyses were performed as described before (15) using RNA isolated from S. aureus USA300 wild type under control conditions and after treatment with 150 μM NaOCl. Hybridization specific for ohrA (USA300HOU_0835) was performed with the digoxigenin-labeled RNA probe synthesized in vitro using T7 RNA polymerase from T7 promoter containing internal PCR products using the primer pairs ohrA-for, 5′ TGGCAATACATTATGAAACTAAAGC 3′, and ohrA-T7-rev, 5′ CTAATACGACTCACTATAGGGAGATTTAAATCGACATTAATATTTCCTTGA 3′.
Crystallographic procedures
Before crystallization, H2O2-treated overoxidized Gap was concentrated to 11 mg/ml. Crystals of overoxidized Gap were grown at 18°C using the hanging drop vapor diffusion technique and 30% (w/v) PEG 3350, 0.1 M Tris, pH 8.5, as the reservoir solution. Crystals were cryoprotected by transfer into mother liquor mixed with 50% (v/v) PEG 400 in a 1:1 ratio and flash-cooled in liquid nitrogen. X-ray diffraction data were collected from a single crystal at 100 K on beamline 14.1 of the BESSY II storage ring (Berlin, Germany) (56) equipped with a PILATUS 6M detector (Company-REF), with a 0.1 ° oscillation and exposure time of 0.3 s per frame. Diffraction images were processed using XDS (41). Crystal parameters and data collection statistics are given in Supplementary Table S3. The Gap-SO3H structure was solved by molecular replacement with Molrep (71) using the structure of the Gap apoenzyme (PDB entry 3LC7; [57]) as a model. The final model of the Gap-SO3H was generated by iterative rounds of manual model building using Coot (20) and automated refinement using the phenix.refine package in PHENIX (1) with the inclusion of TLS parameters generated by the TLSMD server (59). Coordinates and structure factor amplitudes have been deposited in the Protein Data Bank (4) under the accession code 5T73 and will be released upon publication.
Molecular docking of BSH into the Gap active site
To model a covalent complex between BSH and the S. aureus Gap active site Cys151, docking experiments were performed with the holo form containing NAD [PDB code: 3LVF chain R, (57)] as well as the apo form [PDB code: 3LC7 chain O, (57)] of the enzyme. Before molecular docking, both protein structures were prepared using the protein preparation wizard (68) in the Schrodinger software (Release 2016–1) graphical user interface Maestro. Hydrogen was added according to the protonation states at pH of 7.0 as predicted by PROPKA, bond orders were assigned, and disulfide bonds were allocated. Water with less than three hydrogen bonds to nonwater residues was removed and minimization of heavy atoms was performed using OPLS3. The BSH structure was obtained from Pubchem (ID: CID 42614123) and processed with the ligand preparation wizard. The ligand was protonated at pH of 7.0 ± 2.0 using Epik (28). Covalent molecular docking was performed using CovDock (79), which combines the two programs Glide (23) for docking and Prime (39, 40) for minimization. Cysteine 151 was set as reactive residue, and the reaction type was disulfide formation. All atom positions were fixed, except for the targeted residue and the ligand. Covalent docking was performed with default options and the poses were ranked according to the Prime energy.
Footnotes
Acknowledgments
This work was supported by a grant from the Deutsche Forschungsgemeinschaft (AN746/4-1) within the SPP1710 on Thiol-based Redox switches, by the DFG grants AN746/3-1 and project C1 of the Research Training Group GRK1947, and by the ERC Consolidator Grant (GA 615585) MYCOTHIOLOME to H.A. Protein crystal structure analysis was supported by an Alexander von Humboldt postdoc fellowship to A.J.P.-B. Molecular docking was supported by the Klaus Tschira Foundation to L.T, K.K., and F.G. The authors would like to thank Sandra Maaß and Dörte Becher (University of Greifswald) for MS of the ICAT-labeled peptides. MS of the overoxidized Gap was performed at the Centre for Chemical Microscopy (ProVIS) at the Helmholtz Centre for Environmental Research, which is supported by European regional development funds (EFRE-Europe Funds Saxony) and the Helmholtz Association. The authors are grateful to Ambrose Cheung for the kind gift of S. aureus USA300 bshA mutant.
Author Disclosure Statement
No competing financial interests exist.
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
