Abstract
Introduction
H
miRNAs control posttranscriptional gene silencing (PTGS) primarily via binding to 3′ or 5′ UTR of the target transcripts causing translational repression or mRNA degradation. The first evidence of miRNA binding to the 3′-UTR of target mRNA came with the discovery of lin-4 miRNA in C. elegans. Lin-4 was noted to contain sequences that were complementary to 3′-UTR of lin-14 gene, suggesting an antisense RNA-RNA interaction (44). miRNAs are evolutionary conserved across even distant related species, suggesting probable uniformity in the molecular evolution of cellular miRNA gene activity (57). Studies indicate that posttranscriptional regulation of genes by miRNAs is a prevalent phenomenon. According to bioinformatics-based predictions, almost one third of all protein coding genes are posttranscriptionally regulated by miRNAs (21). Recently, a miRNA reporter library study identified functional “miRNAome” of a cell, which represents about 40% of the total miRNAs with any detectable activity (54). This study raised a significant issue with miRNA profiling, where the number of miRNAs differentially expressed in a cell will be much larger than the actual “functional miRNAome” (54).
Studies during the past two decades have identified and highlighted the significance of miRNAs in a multitude of physiological processes and pathological functions, including development, cell proliferation, hematopoiesis, and cell death. Emerging studies indicate that tissue repair and injury, immune and inflammatory responses are substantially regulated by miRNAs (16, 76, 78). Cells of the monocyte/macrophage lineage are critical for mounting and resolving the inflammatory response during tissue injury and repair (17). In this context, this review focuses on miRNAs that control the development, differentiation, and function of monocytes/macrophages by targeting molecules critical in each of these processes.
miRNAs Regulating Development of Myeloid Cells
Circulating monocytes that originate from adult hematopoietic stem cells (HSCs) are primary precursors of macrophages (88). During homeostasis, HSCs are located in bone marrow (BM) niches. Upon injury, HSCs get mobilized to the spleen and liver (91). Microenvironmental niches in BM dictate the maintenance and differentiation of HSCs (91). miRNAs have been identified as key regulators of HSC renewal and fate (27). Initial evidences pointing toward the significance of miRNAs in HSC regulation came from studies performed on Arsenic resistance protein 2 (Ars2/Asr2) null mice (31). Ars2 is a RNA binding protein that contributes to the delivery of stable miRNA transcripts to the pri-miRNA microprocessor complex, Drosha and DGCR (31). Ars2 null mice suffered from BM failure, suggesting a critical role of miRNAs in HSC renewal and fate (31). Conditional ablation of Dicer, an RNase III enzyme required for miRNA biogenesis, from the HSC compartment resulted in hematopoietic stem/progenitor cell apoptosis (32). A subset of miRNAs, including miR-29a, miR-126, miR-155, and miR-125a/b that were specifically enriched in the HSC compartment compared to other cells in the BM (58). Elaborate activities of miR-126, miR-130a, and miR-196b were observed in the most primitive HSCs; a loss in such activities was noted with differentiation of HSCs to more mature progenies (27). miR-125a alone was sufficient to augment the stem cell pool specifically targeting proapoptotic protein BAK1 (32). miR-126, abundantly expressed in HSCs, controls expansion of stem cells via targeting the PI3K/AKT/GSK3β pathway (43). Forced expression of miR-126 in HSC impaired cycle progression of HSC, while knockdown of miR-126 resulted in increased HSC proliferation without exhaustion (43). Homozygous ablation of the miR-29a/b-1 in mice indicated a critical role of miR-29a/b-1 in HSC function by silencing Dnmt3a (33). These studies recognize miR-29a, miR-126, miR-130a miR-155, and miR-125a/b as key miRNAs that are capable of controlling HSC biology (58) (Fig. 1).
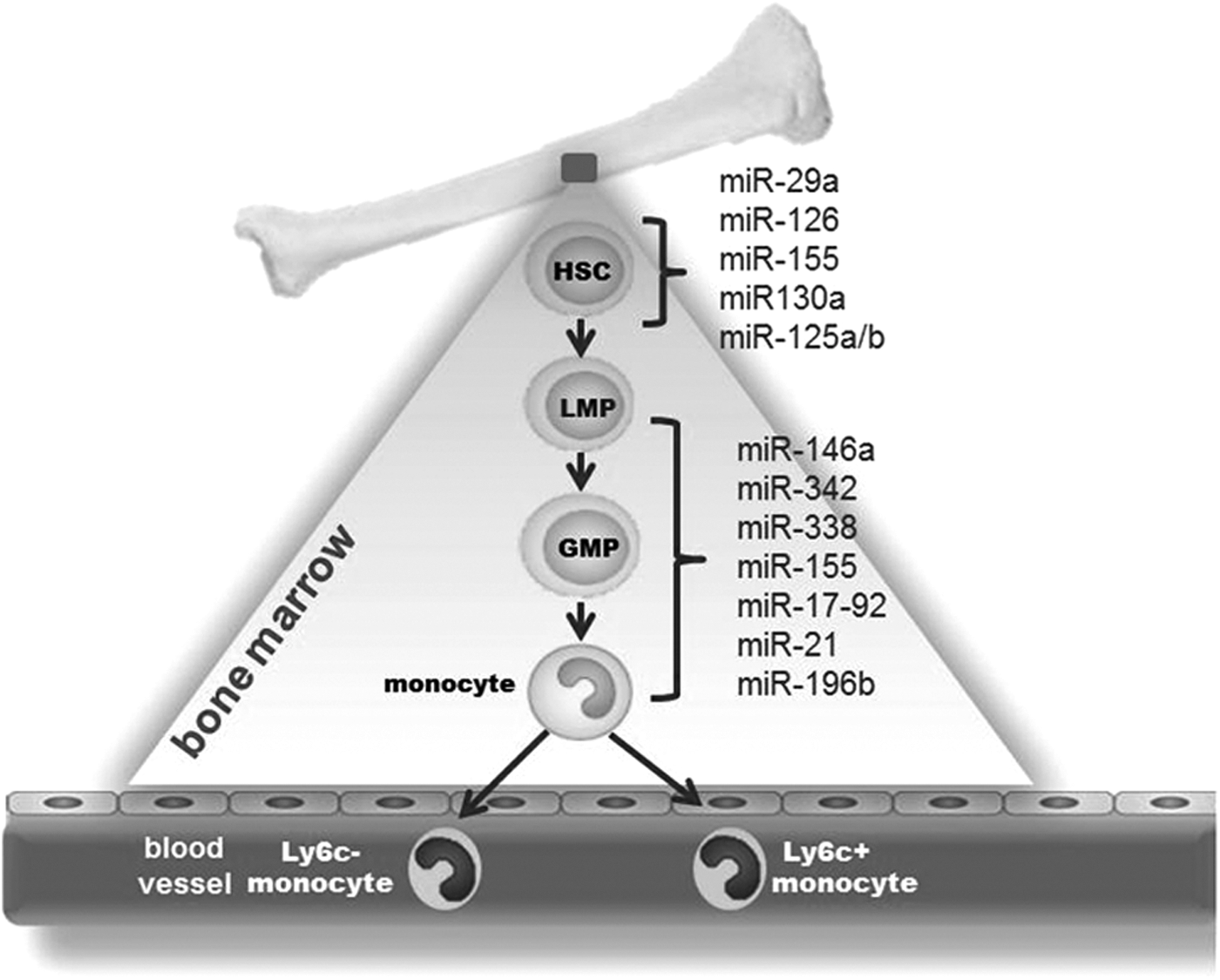
The origin of monocytes from HSC involves differentiation from monoblasts to promonocytes, followed by maturation to monocytes (29). The transcription factor PU.1 facilitates commitment of HSC to lymphoid–myeloid progenitor (LMP) fate via suppression of GATA1 activity (24). The CCAAT/enhancer binding protein (C/EBP) alpha promotes differentiation of LMP to the granulocyte–monocyte progenitor (GMP) stage (24). Increased PU.1 activity favors differentiation of GMP to monocytic lineage (24). miRNAs control multiple steps in the monocyte/macrophage maturation process. PU.1 induces the expression of a subset of four miRs (miR-146a, miR-342, miR-338, and miR-155). This constitutes the initial step in the myeloid cell differentiation/maturation process (28). Forced expression of miR-146a, a negative regulator of innate immune response, was sufficient to drive maturation of stems cells to monocyte/macrophages during adult hematopoiesis (28). In addition to promoting expression of certain miRNA, PU.1 suppresses miR-17p-92 during myeloid differentiation by targeting Egr2 (64). EGR2, in turn, recruits histone demethylase Jarid1b resulting in demethylation of CpG island located at the miR-17–92 promoter (64). The miR17-92 cluster comprises the following six miRNAs: miR-17, miR18a, miR-19a, miR-20a, miR-19b-1, and miR-92a. This cluster is present in high levels in early stem and progenitor cells. Expression of the cluster is downregulated upon inception of differentiation to myeloid lineage (64). PU.1 controls a regulatory circuitry involving transcription activation of miR-424 that leads to monocyte differentiation via translational repression of NFI-A (69). A comprehensive list of miRNAs involved in myeloid cell development has been provided by El Gazzar and McCall (20). The studies described above uphold the extraordinary significance of miRNAs in controlling monocyte/macrophage development (20, 28).
The relative expression levels of PU.1 and C/EBPα dictate the fate of myeloid progenitor cells to monocytic versus granulocytic lineages. Low levels of C/EBPα prevent GMP formation, whereas higher levels promote granulopoiesis over monopoiesis (25). C/EBPα induces miR-223 resulting in degradation of NFI-A mRNA to promote granulopoiesis (25). Both miR-21 and miR-196b promote the formation of monocytes while suppressing granulopoiesis (89). Interaction of colony stimulating factor-1 (CSF-1) with its CSF-1R promotes differentiation and maturation to monocytic lineage. CSF-1R expression is controlled by Runt-related transcription factor-1 (RUNX1, also known as AML1). miRNAs 17-5p, 20a, and 106a target the 3′-UTR of Runx1 and suppress its translation (22). These studies point toward a central role of miRNAs in controlling major stages of monocyte/macrophage development from HSCs. Studies using appropriate conditional knockout/transgenic mice are required to understand specific regulation of these miRNAs in myeloid cell lineage commitment and maturation.
Control of Macrophage Polarization and Activation
Macrophages are dynamic cells and have the ability to undergo transition through a continuum of phenotypes and activation states primarily based on the microenvironmental milieu. The outstanding significance of microenvironmental cues to direct macrophage polarization at the tissue injury/repair site has been recognized (17). Although contentious, macrophages are broadly classified as M1 and M2 polarization states (48, 53, 55). M1 macrophages, also known as “classically activated,” are induced by cytokines primarily secreted by Th1 cells, including IFN-γ and TNFα. In addition, granulocyte–macrophage colony-stimulating factor (GM-CSF), lipopolysaccharide (LPS), and other Toll-like receptor (TLR) ligands are also known to induce M1 polarization in macrophages. The M1 polarized macrophages support Th1 responses and produce copious amounts of reactive oxygen species (ROS) to help in killing pathogen (48). Conversely, activation of macrophages with Th2 cell factors results in M2 macrophages or “alternatively activated” macrophages. M2 macrophages influence the processes of infection, tissue repair, vascularization, and tumor promotion/invasion. These macrophages are characterized by their high expression of scavenger receptors in addition to high IL-10, VEGF, and MMPs. (48). The microenvironmental signals elicit a transcriptional response that regulates the phenotype and function of the macrophages (42). The signal transducer and activator of transcription 1 (STAT1) and interferon-regulatory factor 5 (IRF5) play a major role in dictating M1 macrophage phenotype (42), whereas, STAT6, peroxisome proliferator-activated receptor-γ (PPARγ), and IRF4 direct M2 macrophage polarization (42) (Fig. 2). These transcription factors are directly controlled by specific miRNA (52, 86, 92, 94) (Fig. 2). PU.1, on the contrary, irrespective of polarization, serves as the master transcription factor for maintenance of macrophage identity and differentiation by acting on macrophage-specific enhancers (42).
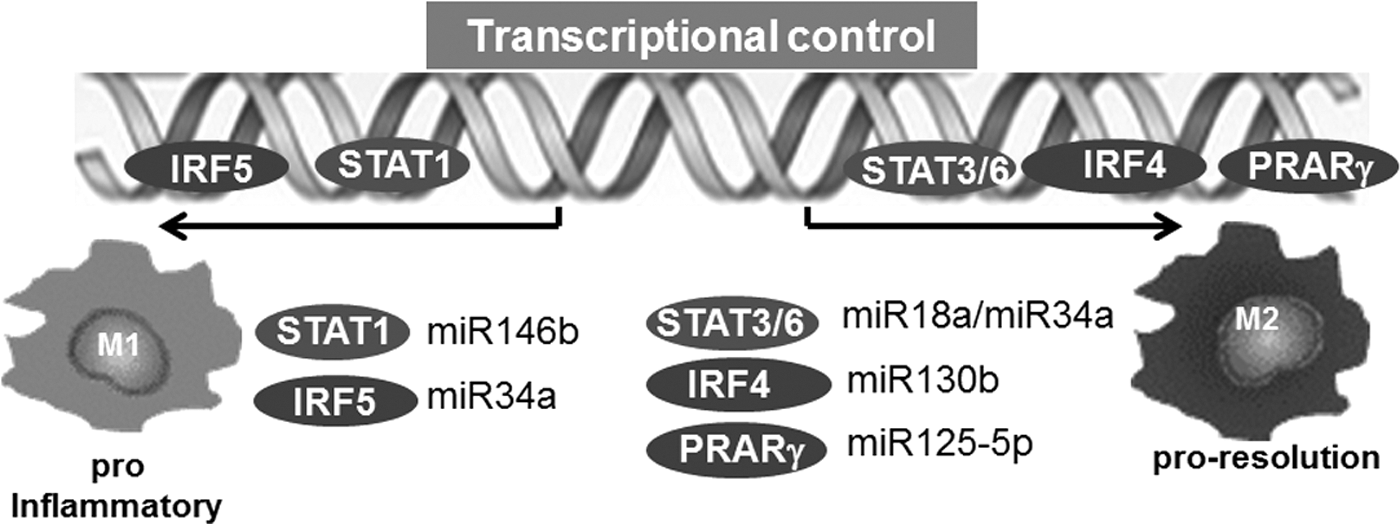
The JAK/STAT pathway is under miRNA regulation (96). miR-146b suppress the antiviral function in T cells via directly targeting STAT-1 (90). It is not clear if miR-146a also targets STAT-1 in macrophages to control polarization. In a comprehensive study to decipher the role of miRNAs in macrophage polarization, macrophages were subjected to stimuli inducing four different patterns of macrophage activation/polarization (M1, M2a, M2b, and M2c). The changes in miRNA expression profile were elucidated using TaqMan low-density array human miRNA assays (version 2.0; Applied Biosystems) (30). Specific subsets of miRNAs uniquely expressed under four conditions of polarization were noted. Specifically, miR-125a-3p and miR-26a-2* in M1 macrophages, while miR-193b, miR-27a*, miR-29b-1*, miR-132*, and miR-222* were uniquely expressed in M2b phenotypes (30) (Fig. 3). M2c phenotype did not display any specific change in miRNA expression. Recent studies, including those from our laboratory, demonstrate a critical role of miR-21 in macrophage polarization (8, 16). We reported that miR-21 is central in efferocytosis (engulfment of apoptotic cells)-mediated change in macrophage from a proinflammatory (M1) to proresolving (M2) phenotype (16). In line with this observation, activation of CSF-1R by its ligands CSF-1 and IL-34 plays a major role in macrophage development and differentiation. The molecular signaling elicited by CSF-1R pathway has been shown to regulate macrophage polarization via PI3K→ERK→NFκB and activation miR-21. Inhibition of miR-21 directed macrophage toward the M1 phenotype, suggesting that sufficient miR-21 in macrophages is essential for their M2 polarization (8). A central role of miR-33 in regulation of cellular lipid metabolism by repressing genes involved in cholesterol efflux, and fatty acid oxidation, has been recognized (39). A shift of macrophage metabolism toward glycolysis promotes M1 or proinflammatory phenotype (26). miR-33 was recently reported to promote macrophages toward M2 phenotype via targeting AMP-activated protein kinase, a key player in intracellular energy equilibrium (61).
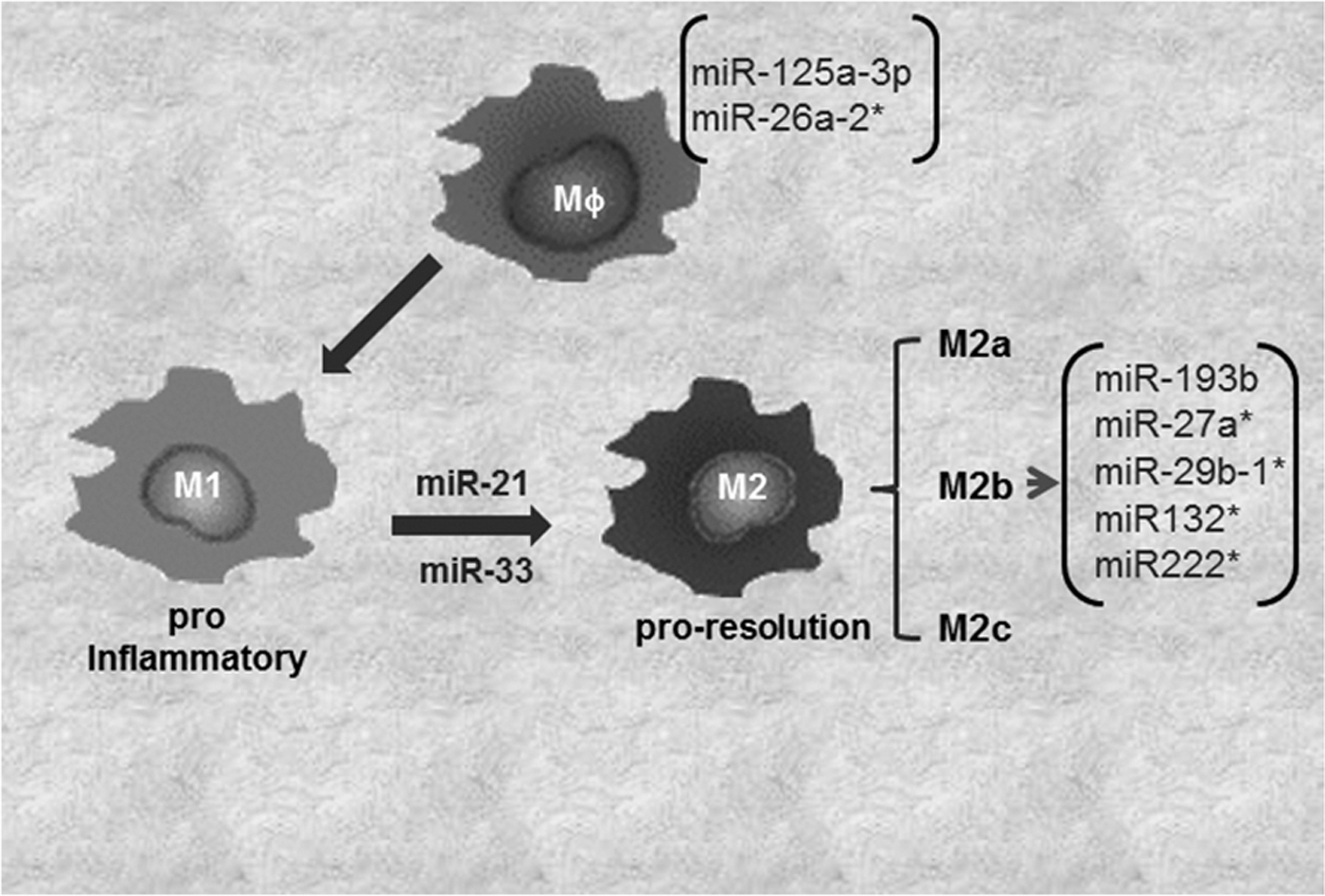
Regulation of Monocyte/Macrophage Function
Macrophages/monocytes are implicated in regulation of a multitude of biological processes, including inflammation and phagocytosis (53, 55). To maintain focus, we limit discussion to the roles of miRNAs in regulation of key functions of monocyte/macrophages related to inflammation and tissue repair (Fig. 4).
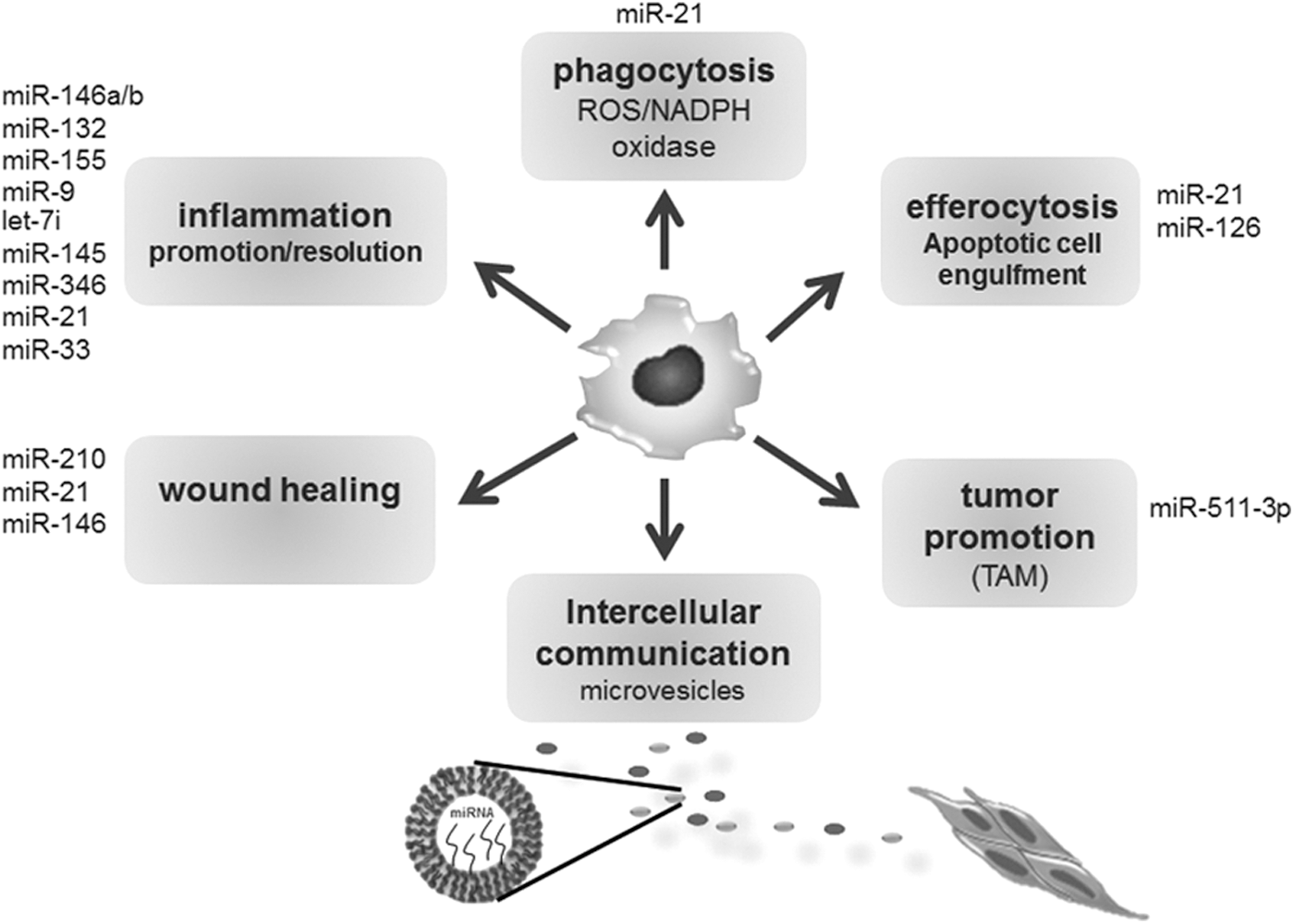
Inflammation
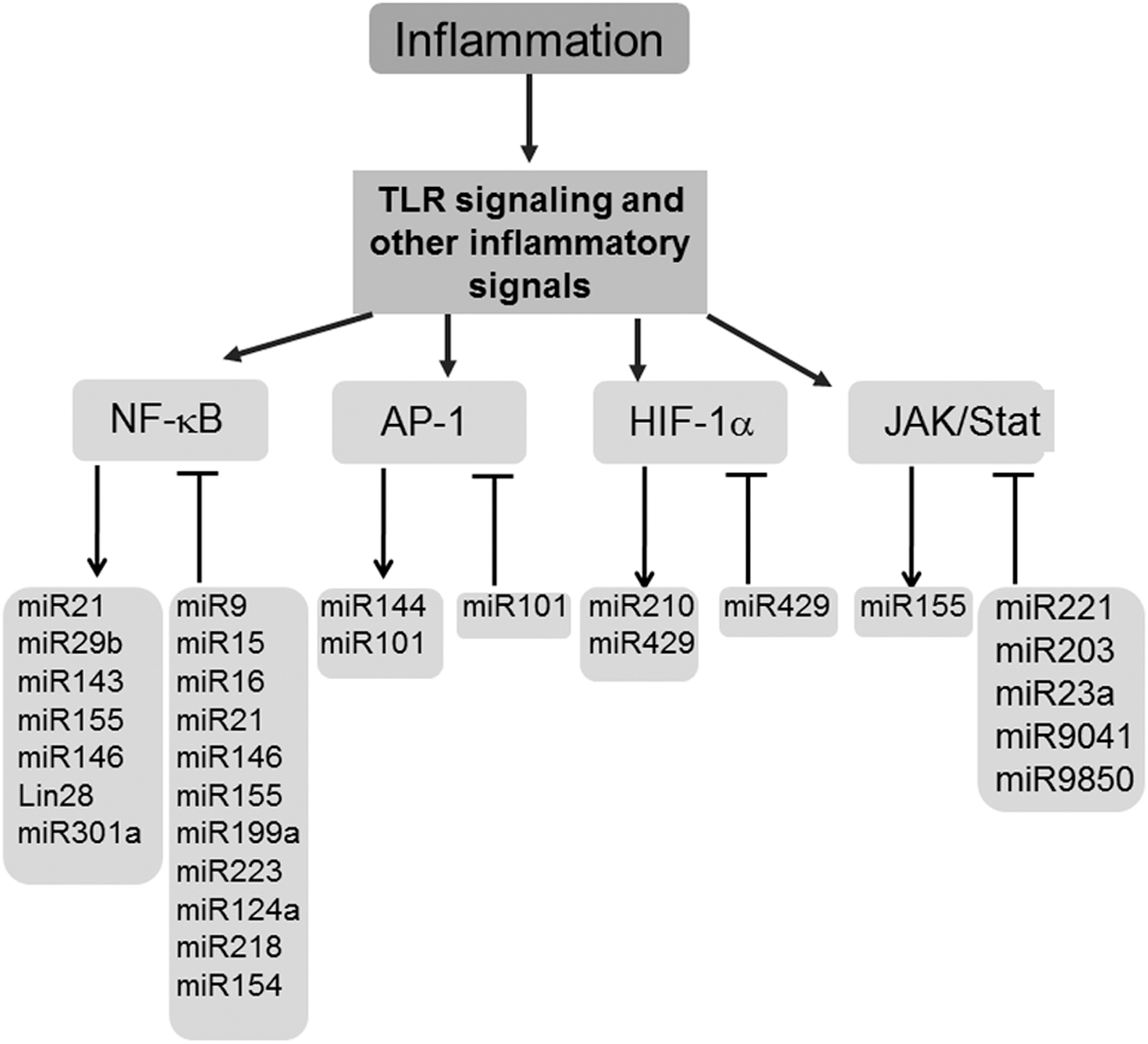
Inflammation is a dynamic process that is elicited following injury/infection and other external stimuli with the objective to repair and restore homeostasis (68). Macrophages are key cells of the innate immune system, an evolutionarily conserved form of host defense against infection (36). miRNAs control multiple regulatory checkpoints to fine-tune macrophage inflammatory response (71, 72). TLR and related downstream signaling cascade pathways are important detection mechanisms that enable the initial steps of the inflammatory cascade. The first evidence recognizing inflammation-sensitive miRNAs in macrophages was provided using miRNA expression profiling in endotoxin-treated murine macrophages (84). Of the 200 miRNAs investigated, miR-146a/b, miR-132, and miR-155 were identified as LPS-responsive miRNAs (84). Promoter analysis studies recognized miR-146a as a NF-κB-inducible miRNA (84). In addition, the study also identified binding sites for miR-146a/b at the 3′-UTRs of the TNF receptor-associated factor 6 (TRAF6) and IL-1 receptor-associated kinase 1 (IRAK1) genes, suggesting these targets may be subjected to PTGS by miR146a/b (84). A key function of miR-146 controlling TLR and cytokine signaling via a negative feedback regulation loop was proposed (84). This study was instrumental in highlighting the significance of inflammation-sensitive miRNAs in macrophages. An induction of miR-155 in mouse macrophages exposed to inflammatory mediators polyriboinosinic:polyribocytidylic acid or the cytokine IFN-β was identified (59). miR-155 tightly controlled the expression of SHIP1 and SOCS1, both critical regulators of the inflammatory response in macrophages (59). Bala et al. reported that increased miR-155 levels contribute to alcohol-induced elevation in TNF-α production (4). miRNAs let-7i, miR-145, and miR-346 were shown to regulate TLR signaling. Of these, only let-7i inversely correlated with TLR signaling and immune response (60, 74). NFκB is a master transcription factor that regulates the expression of proinflammatory cytokines such as TNF-α, IL-1β, IL-6, and IL-8 in macrophages (37, 38). miR-146, one of the first miRNA identified to be involved in the NFκB activation pathway, constitutes a negative feedback loop where NFκB activates miR-146, which in turn suppresses NFκB signaling by directly targeting TRAF6 and IRAK1 (84). Conversely, miR-181b indirectly regulated NFκB in a positive feedback loop that included NFκB → IL-6 → STAT3 → miR-181b → CYLD → NFκB (46). NFκB-dependent reporter assay screening of several hundred miRNA genes identified miR-301a as one of the most effective activators of NFκB by repressing NFκB repressing factor (46) (Fig. 5). miR-9 has been linked with innate immune response; the expression of this miRNA contributes to a negative feedback loop in the NFκB pathway, where miR-9 is activated by TLR4→NFκB in human PMNs and monocytes target NFκB subunit p50 (6). The studies discussed above clearly indicate that miRNAs fine-tune the inflammatory response by controlling major pathways such as TLR and NFκB. miRNA control of other major transcription factors implicated with inflammatory response has been highlighted (5, 10, 14, 34, 45, 56, 62) (Fig. 5). Such control by miRNA is likely to prevent an exaggerated inflammatory response.
ROS production and function
In response to inflammation, phagocytosis, or environmental stimuli, macrophages produce copious amounts of ROS by a process known as “respiratory burst” (2, 63). NADPH oxidases are primarily responsible for the “respiratory burst” action in cells of myeloid origin (3). NAPDH oxidase is a multisubunit complex that comprises membrane-bound cytochrome component, that is, gp91phox (phagocyte oxidase) and cytoplasmic components, including p47phox, p67phox, and p40phox, and rac (23, 63, 75). Upon activation, the cytosolic subunits translocate to the membrane-bound gp91. In 2008, our laboratory reported first evidence demonstrating that the rate-limiting p47phox subunit of NADPH oxidase is a target for miRNA (80). Global lowering of cellular miRNAs by knocking down Dicer, a key enzyme of miRNA biogenesis, resulted in decreased production of inducible ROS (80). Recently, a cross talk between ROS and miR-451 was identified using BM-derived macrophages from p47phox null mice (65). Furthermore, miR-21 via targeting PDCD4-AP-1 pathway was observed to regulate H2O2-dependent responses in smooth muscle vascular cell injury model. Using a global profiling approach, miRNA signature in response to oxidative stress such as ionizing radiation and H2O2 was determined (81). The pathways underlying oxidant-induced miRNA expression remain to be characterized.
Efferocytosis
Macrophages are responsible for cleaning up host tissue of invading pathogens as well as dead/dying cell debris. The process of engulfment of dead/dying cells by macrophages is referred to as “efferocytosis” (19). Efferocytosis is well known to play a central role in the resolution of inflammation (70). Despite the resemblance between the processes of efferocytosis and phagocytosis, the two processes are clearly distinct with respect to receptors and ligands involved (67, 87). Efferocytosis initiates signaling events involving two main complexes, (i) CrkII/ELMO/Dock180 or (ii) ABCA1/GULP (40). Activation of either complex leads to activation of Rac1, which initiates cytoskeletal rearrangement and subsequent engulfment of apoptotic cells (40). miRNAs have been implicated in the signaling cascade involving efferocytosis-induced resolution of inflammation, more specifically the change in macrophage phenotype following efferocytosis (16). Engulfment of apoptotic cells by macrophages dampened proinflammatory response and promoted an anti-inflammatory milieu by secretion of cytokines such as IL-10. Studies from our laboratory identified that in macrophages miR-21 can be induced by efferocytosis. Elevated macrophage miR-21 supported efferocytosis and silenced target genes PTEN and PDCD4, which in turn upregulated IL-10 and suppressed TNFα production, respectively, accounting for a net anti-inflammatory M2-like phenotype observed in postefferocytotic macrophages (16). CrkII, a protein of the efferocytosis engulfment complex, has also been proposed to be a direct target of miR-126, indicating a potential regulation of miRNA at an upstream efferocytosis-mediated signaling pathway (15).
Wound healing
Wound healing is a well-orchestrated reparative event that occurs in response to injury and its microenvironment (49). Macrophages infiltrating the wound site postinjury are often termed as wound macrophages (17). Wound macrophages are essential for repair (17, 50, 51). Work from our laboratory has focused on developing the significance of miRNAs in controlling molecular mechanisms that regulates wound healing (7, 11 –13, 76, 77, 79, 80). Here, we limited discussion to those miRNAs that directly regulate macrophage function in wound healing. miR-21 was identified by us recently as a major negative regulator of LPS-induced inflammatory response in wound macrophages (16). miR-146a, an inflammation-sensitive miRNA, was significantly low in diabetic wounds (93). Such decreased miR-146a levels caused upregulation of proinflammatory target genes (93). The study did not, however, identify a role of wound macrophages in miR-146 dysfunction in diabetic ulcers. The studies to explore the significance of miRNAs in wound macrophage activation and function have just begun. Further studies investigating significance of miRNAs in wound macrophage activation and function are required to understand the mechanisms underlying the critical role of macrophages in wound healing.
Tumor-associated macrophages
Macrophages located in tumor stroma are referred to as tumor-associated macrophages (TAMs) (47). These macrophages primarily display M2 phenotype and modify inflammation, promote angiogenesis, matrix remodeling, and tumor cell migration/invasion, thereby suppressing antitumor immunity (47, 82). The presence of a higher number of TAMs is associated with tumor progression and poor patient prognosis (47). Cancer-related studies suggest the involvement of miRNAs in modulating response to TAM by directly acting on these macrophages and the precursors of these cells, including HSCs (82). In mouse models, a specific subset of TAMs that displays high expression of the mannose receptor (MRC1/CD206) and attenuated CD11c expression has been identified with high proangiogenic and tumorigenic activities (83). miR-511-3p, an intronic miRNA, is encoded by both mouse and human MRC1 genes (83). Using reporter vector bioassay, miR-511-3p was noted to show high expression and activity in CD206+ TAMs (83). By targeting CD206 (MRC1), miRNA-511-3p plays a critical role as gatekeepers of the protumorigenic activity of TAMs. Detailed studies investigating relevance of miRNAs in TAM subset phenotype and their influence on the activity of TAM are warranted.
Macrophage Intercellular Communications Via Extravesicular Release
Macrophages are secretory cells that control specific cellular processes via intercellular communication. Extracellular membrane vesicles (EVs) function as shuttles between cells for delivery of cargo such as proteins; mRNAs and miRNAs have emerged as major mediators of intercellular communication (66). Exosomes, microvesicle (MVs), and apoptotic bodies are three broad categories of EVs that are released by a number of cells, including macrophages (95). Exosomes are vesicles of endosomal origin that originate by inward budding and are between 40 and 100 nm in diameter (85). However, the MVs are released from plasma membranes as a result of outward budding and fission (85, 95). Macrophages secrete both MVs and exosomes that carry cargo, including proteins; mRNA and miRNA are for intercellular communication (41). In peripheral blood, the majority of circulating microvesicles originate from platelets, while monocytes/macrophages contribute the second most abundant population of circulating MVs in blood (35). Macrophage-derived MVs have been demonstrated to carry functionally active miR-223 that subsequently induces macrophage differentiation and is an important contributor to the innate immune response (35). Another example of transferring functionally active miRNAs was provided by an observation where human macrophages, via gap junctions, transferred miR-142 and miR-223 to hepatocarcinoma cells (HCCs) (1). Such transfer of miRNAs resulted in PTGS of target proteins, that is, stathmin-1 and insulin-like growth factor-1 receptor in HCC, suggesting that such transfer is not just a passive process, rather it actively regulates functions in the acceptor cell. Another example of active transfer has been recently reported where monocytes exposed to alcohol communicated with naive monocyte via miR-27a cargo transferred from EVs (73). Transfer of miRNA resulted in naive macrophages to polarize to M2 macrophages (73). Taken together, macrophage-derived exosome and MV-mediated transfer of miRNAs add a novel dimension to how macrophages modify inflammatory responses and other cellular activities.
Conclusion
During the last decade, we have witnessed an unprecedented growth in understanding of miRNAs in health and disease. In most cases, miRNAs have been observed to fine-tune cellular responses and, therefore, act as a rheostat rather than an on/off switch. A critical role of miRNAs in the regulation of innate immune response and macrophage biology, including development, differentiation, and activation, has emerged. miRNA expression and activity are evolutionarily conserved and highly cell specific. Increased macrophage-specific knockout/transgenic studies will clarify the significance of these tiny molecular rheostats on development, differentiation, and activity of these cells in regulation of biological responses. Such information will be critical in developing miRNA-directed therapies. In summary, miRNAs fine-tune numerous facets of macrophage development and function. Such fine-tuning is critical in preventing exaggerated macrophage response to endogenous or exogenous stimuli.
Footnotes
Acknowledgments
Wound healing research in the author's laboratory is funded by NIDDK R01 DK076566, NIGMS RO1 GM108014, GM069589, GM 077185, NINR NR013898, & NR015676.
