Abstract
Introduction
T
Bacterial toxins developed sophisticated solutions for the transcellular and intracellular delivery of protein subunits. Viruses are the number one carriers for intracellular nucleic acid delivery. However, only minor changes in their structures can be made, the size and type of cargo is limited, and foreign proteins and viruses trigger immune reactions. Both such protein toxins and viral nanoparticles, on the one hand, present excellent natural examples for how to design carriers for intracellular delivery. On the other hand, synthetic carriers that mimic these transfer processes but are not dependent on naturally available nucleotides and amino acids gain more and more attention.
These synthetic carriers should handle a lot of challenging tasks. They should offer an optimal extracellular stability and an efficient uptake into the cell. Uptake usually results in delivery into endocytic vesicles, which often end in degradative lysosomes. Therefore, the same carriers should facilitate an escape from endosomes after uptake and release their nucleic acid or protein cargo to the cytoplasm, which is optionally followed by transfer to the nucleus (as required in the case of pDNA). In this process, the carriers should act similar to shuttles: Although stably associated with their cargo in a covalent or noncovalent manner in the early extracellular delivery steps, they should not disturb the protein or nucleic acid cargo in functionality inside the cell and should dissociate or disintegrate after delivery.
Not only the demands on delivery differ at the different extracellular and intracellular sites, but also the microenvironments provide unique different properties. Carriers can be designed to sense these environmental differences and to utilize them to undergo changes that are beneficial for the delivery process (116). The molecular programming of such dynamic carriers may include sensors that react to changes in enzyme activities, pH, or the redox microenvironment (Fig. 1). Among the different sensors, bioreducible elements are important tools that are used to distinguish between extra- and intracellular sites, where the redox potential displays big differences. As reviewed in the next sections, the use of disulfide bonds as sensors for reductive environments have a growing impact in the development of oligocation-based macromolecular drug carriers (48, 91, 104).
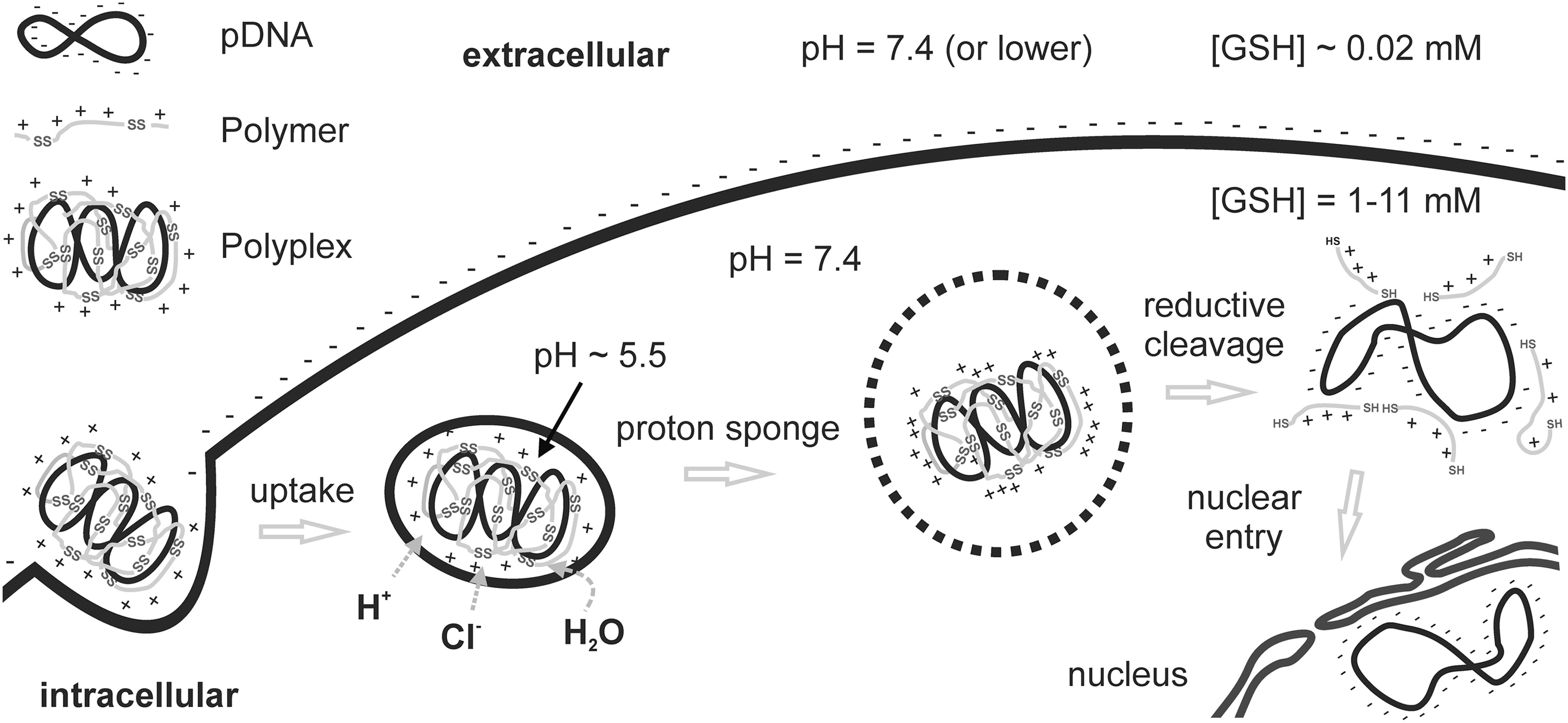
Outside Stabile, Inside Labile—The Delivery Paradox
Polyplexes are nanoscaled, interelectrolyte complexes consisting of cationic polymers and negatively charged nucleic acids (23). The extracellular stability is of highest importance. In the bloodstream, these particles are exposed to nucleases, which can degrade the cargo, and other serum proteins, which can destabilize the complex by electrostatic exchange actions. In addition, nucleic acids are compacted to convenient polyplex sizes of virus-like dimensions (around 100 nm). In case of larger pDNA, this results in favorable compaction and protection. In case of small nucleic acids such as siRNA or oligonucleotides, this results in increased nanoparticle sizes due to packaging as multimers, which prevents the particles from being cleared too rapidly by the kidney (102). High-molecular-weight (HMW) polymeric carriers usually exhibit better transfection efficiency than do low-molecular-weight (LMW) carriers largely because of insufficient polyplex stability of the latter ones (55, 100, 119, 129, 130). LMW carriers, however, have significant advantages with regard to reduced cytotoxicity (124) and better biocompactibility; for example, a reduced complement activation (84) that is relevant for in vivo applications. Moreover, too tight binding by HMW polymers can hinder intracellular vector unpacking (41, 95) and, therefore, reduce transfection efficiency. pDNA should be accessible to the transcriptional machinery in the nucleus. siRNA should be accessible to the RNA-induced silencing complex (44), and the activity of therapeutic cargo proteins should not be affected by the carrier after delivery to the cytosol. Often, the window between sufficiently large polymer size for effectivity and sufficiently small polymer size for high biocompatibility was either small or nonexistent (7). To achieve a better carrier profile with stability on demand, strategies using biodegradable polymers have been explored. The concept often included bioreversible cross-linkage of nontoxic LMW oligomers into HMW carriers via hydrolysable ester, acetal bonds, or reducible disulfide bonds (4, 26, 31, 34, 49, 51, 79, 109). Thus, extracellular polyplex stability and transfection efficiency is partnered with intracellular disassembly and polymer degradation into nontoxic fragments.
Within physiological proteins, the presence of disulfide bonds dominates outside the cell, for secreted proteins, matrix proteins, or proteins at cell surfaces (21). Inside the cell, the combination of high glutathione (GSH) concentrations and enzymes from the thioredoxin family (39) forms a reductive environment of the cytosol. With a concentration of 1–11 mM GSH (94), the intracellular concentration of GSH is approximately 100–1000 times higher than outside the cell. This attribute can be also used by artificial reducible systems to be intracellularly cleaved and change their properties. Although extracellular stability is expected to be significantly improved by disulfides, the situation is not so clear-cut. Disulfide cleavages may also occur in the in vivo situation outside the cell (9). This was already observed in an early in vivo work using immunotoxins (60, 110). The liver as the major organ for GSH production (15, 127) has been considered as responsible for this extracellular degradation of disulfide bonds. Protein disulfide isomerases (PDI), widely known to be localized in the endoplasmatic reticulum (ER), can also occur on the cell surface of hepatocytes and may cleave disulfide bonds (77) Low concentrations of cysteine (∼8 μM) and GSH (∼2 μM) present in human plasma (43) are not considered to significantly contribute to extracellular bioreduction. The extracellular stability of disulfides can be tuned by their structural environment (110, 126); see also last section. Thus, appropriate redox chemistry is considered an excellent approach to solve the conflicting carrier requirements between extracellular stability and intracellular instability.
For example, in an early work, Rice and coworkers (68) introduced defined oligolysine structures containing one to five cysteine residues (
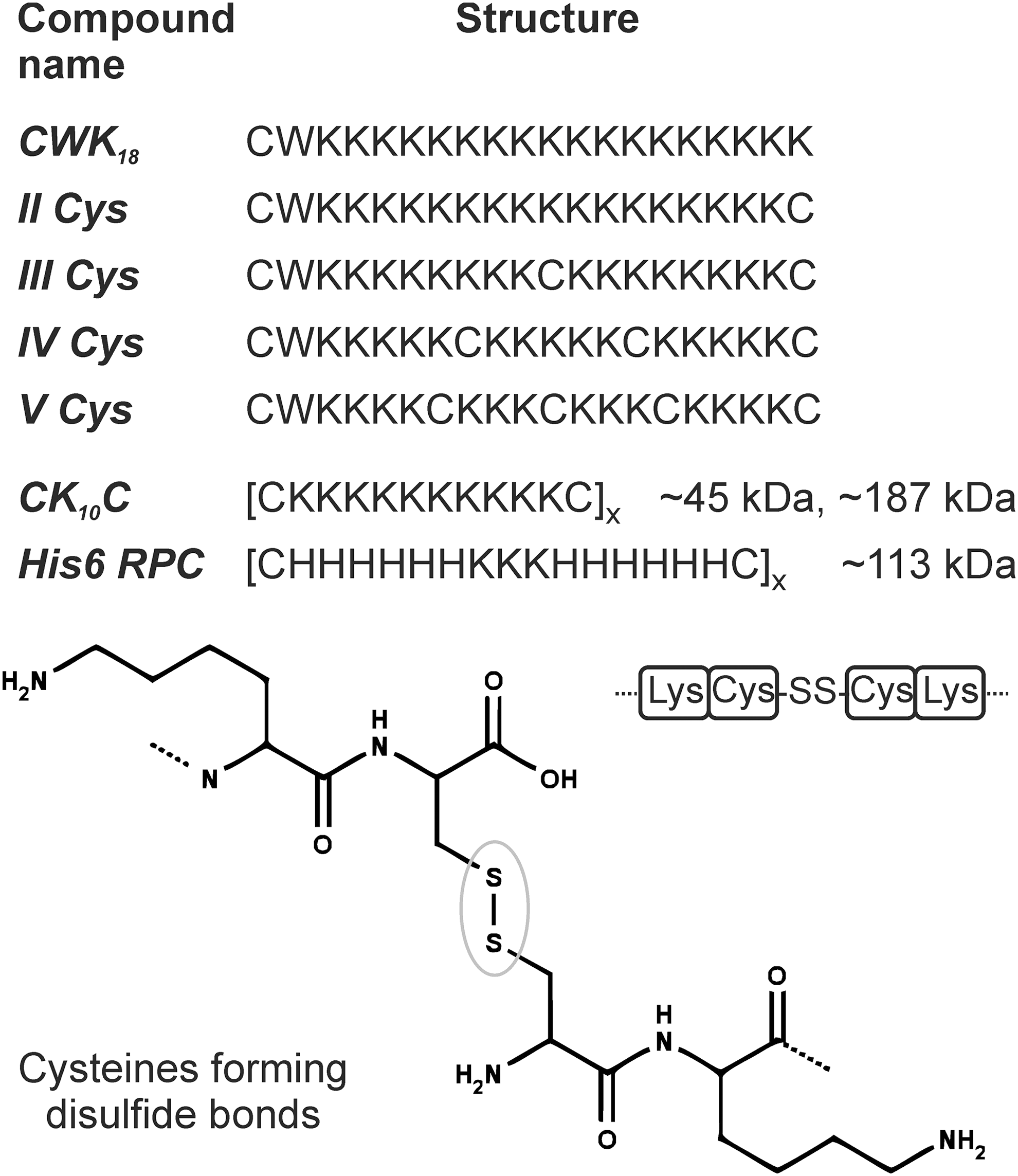
Bioreducible pDNA Carriers with Endosomal Escape Function
For successful gene transfer, several delivery barriers should be overcome (see Fig. 1). Release from endosomes after cellular uptake presents a major bottleneck. Polyethylenimine (PEI), a polymer often used for pDNA transfections, combines optimized DNA binding ability and endosomal buffer capacity within one structure (3, 131). Its diaminoethane motif (108) offers a remarkable pH-reversible protonation and buffer capacity over a wide pH range, which turns it into an almost perfect “proton sponge” (1, 2, 105, 115). Endosomal protonation of PEI was discovered as an excellent way for triggering lipid membrane disruption and release into the cytosol. In sum, PEI has been established as being a very effective as well as a significantly cytotoxic (32, 74) gene carrier. In order to create a less toxic, bioreducible branched PEI, LMW (<4.6 kDa) linear PEI (
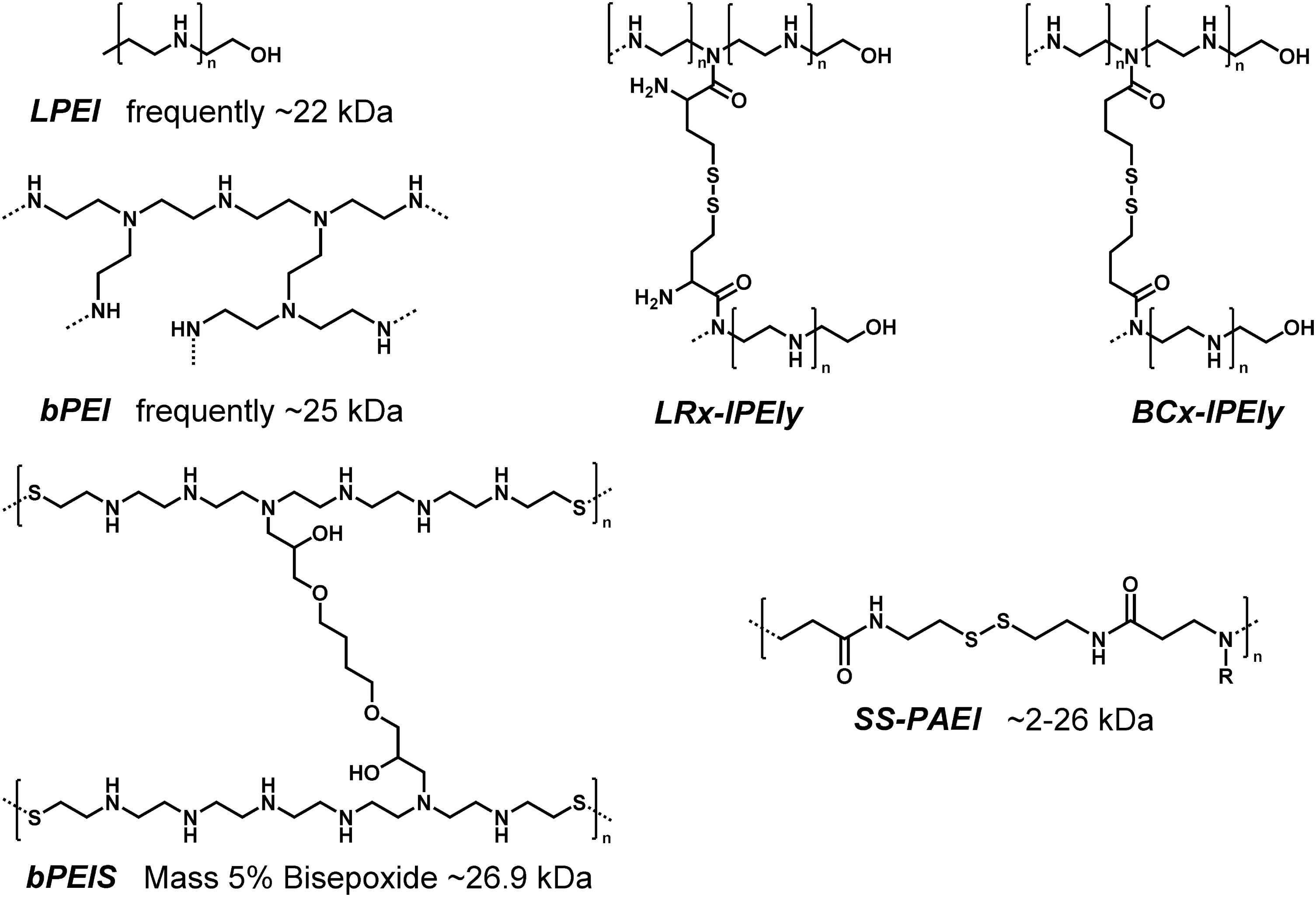
A related effort to produce a biodegradable PEI was taken by Park and coworkers (52). They synthesized a bioreducible linear PEI with a diaminoethane motif that was separated by a disulfide bond after every six protonatable amines (
Disulfide containing poly(amido ethylenimine)s (
A modification that strongly improved the efficacy of polylysine and oligolysine shuttles was the introduction of histidines (73, 81, 88). The second imidazole nitrogen exhibits ideal basicity (pKa 6.0) for an increase of endosomal buffer capacity (lower protonation at physiological than at more acidic endosomal pH), thereby facilitating endosomal release by the “proton sponge effect” that was previously hypothesized for PEI. Thus, it was possible to mediate efficient delivery not only for pDNA but also for mRNA and much smaller siRNA (more information on siRNA delivery in the next section) with one single vector (88). A histidine/lysine ratio of 4:1 in a Cys-His6-Lys3-His6-Cys monomer (
Design of Precise Oligomers for pDNA Transfection
To combine the beneficial transfection properties of bioreducible PEI and related oligoethyleneimine polymers with a precise chemical structure, solid-phase-supported polymer synthesis (35
–38) was introduced. In particular, artificial amino acids containing short defined repeats of the diaminoethane motif were prepared in boc/fmoc protected form, which was appropriate for standard automated or manual peptide synthesis (96). These building blocks, optionally in combination with natural amino acids such as cysteine and other units, were incorporated into peptide-like oligomers. The artificial amino acids provide the oligomers with proton sponge capacity. For example, Stp (succinoyl tetraethylene pentamine) within a polyamide chain provides three protonatable amines, and the building block Sph (succinoyl pentaethylene hexamine) provides four protonatable amines (Fig. 4). By solid-phase synthesis technologies, such defined oligomers can be generated with high precision in libraries of various sequences and defined lengths. Initial screenings of such a small library based on the Stp building block were performed for pDNA transfer activity (98). Compared with standard polymers, the oligomers present LMW carriers. Consistent with their incomplete protonation at physiological pH, oligomers did not display significant cytotoxicity and were well biocompatible. Not unexpectedly, their limited size also provides limited polyplex stability. Compound
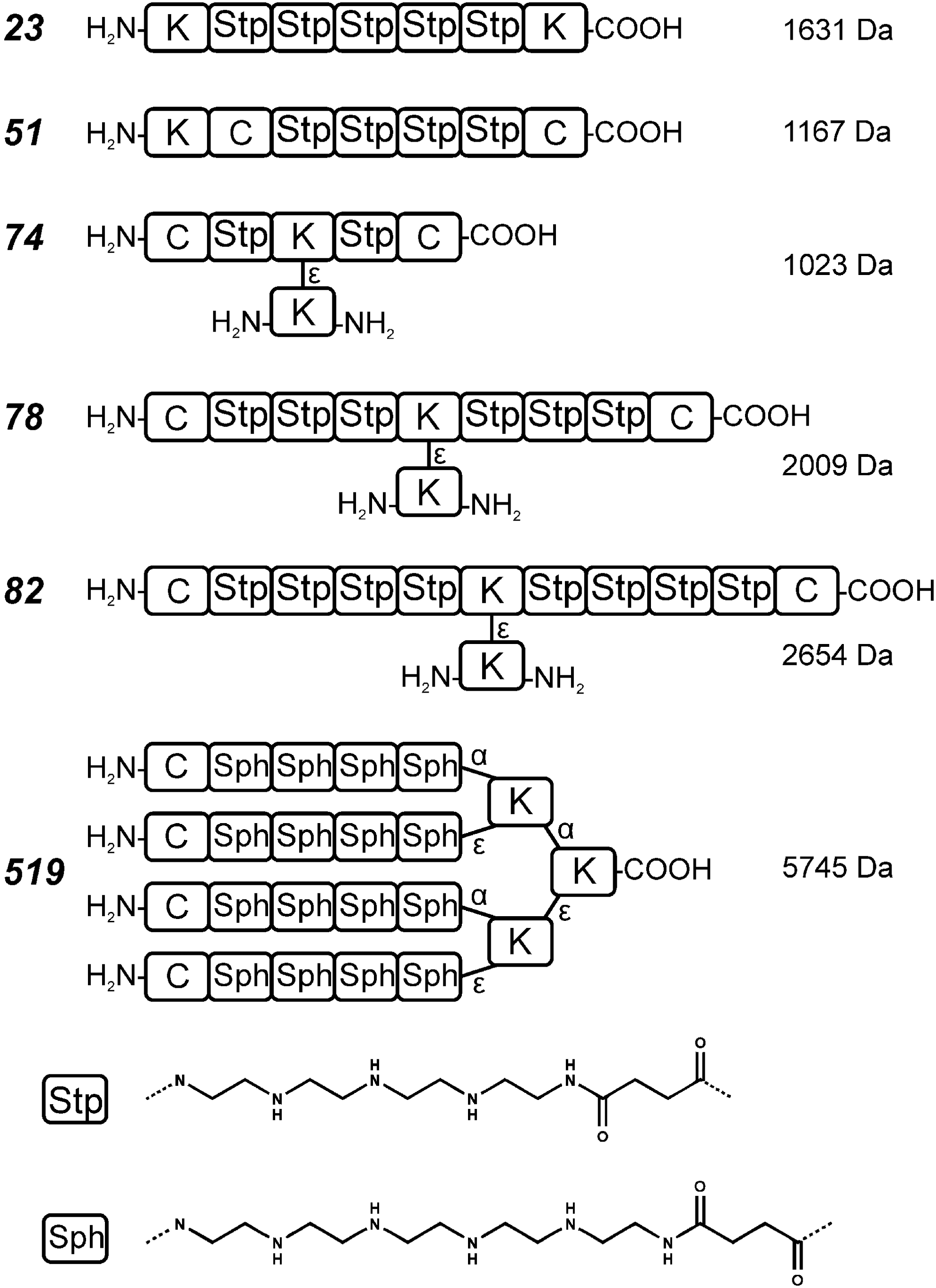
Standard
Screening for pDNA transfection showed the following: The number of nitrogens per building block (Sph containing 6N>Stp with 5N>Gtt with 4N) was more important for pDNA polyplex condensation and transfection than the total number of nitrogens per oligomer. Sph-based four-arm structures with three repeating units per arm (Sph3, 72 ethylenimine nitrogens in total) displayed comparable transfection activity as
Cargo Matters: Reversible Disulfide Bonds Stabilizing siRNA Polyplexes
The delivery of siRNA with cationic systems poses different challenges than pDNA (57, 102, 118). Although both nucleic acids contain a negatively charged backbone (phosphodiester for physiological or phosphorothioate in case of some stabilized nucleid acids), the number of base pairs (bp) per molecule and their topology are different. With a 7 nm rod-shaped size, 21–23 bp, and 42–46 anionic charges per molecule (16, 117), siRNA is comparatively small and cannot offer thousands of negative charges that a long cyclic pDNA provides for binding. This results in a lower electrostatic stabilization, less gain in entropy by complex formation, and, therefore, less stable siRNA polyplexes. Most of the previously described systems for the transport of pDNA, including polylysine (71) and PEI (57), show much less efficiency in siRNA transfection unless appropriate modifications are introduced. One of the encouraging options presents the incorporation of bioreducible stabilizing disulfide bonds, by the cross-linking of siRNA into more “DNA-like” oligomers (58, 75), by linking siRNA with the cationic carrier (19, 70, 80), or by cross-linking the cationic carrier molecules.
Based on the knowledge of pDNA delivery systems, new carriers have been invented to accomplish the delivery of siRNA. The approximately 4 kDa pseudodendritic degradable oligomer
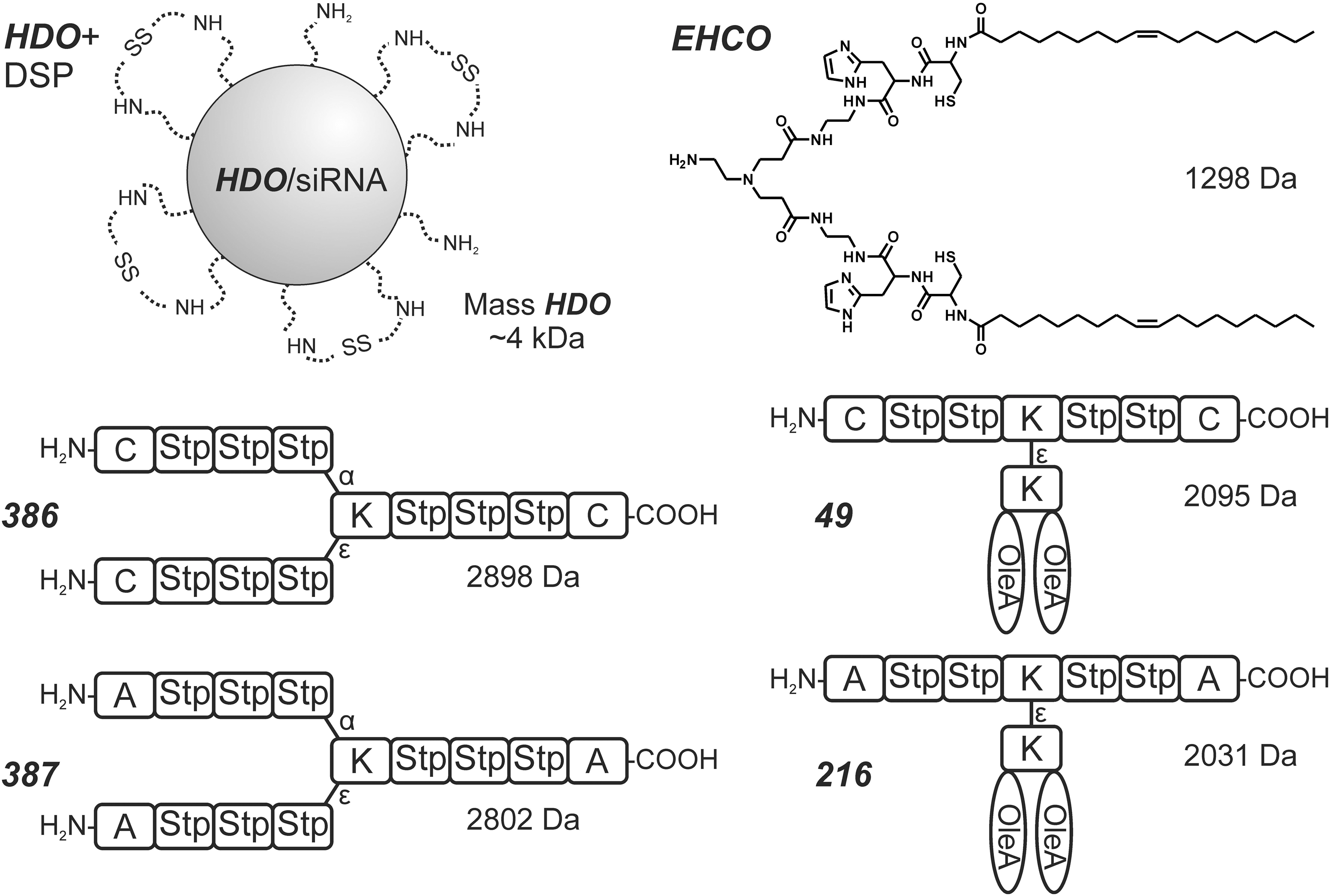
The importance of disulfide-based cross-linking for siRNA can be observed when using precise sequence-defined oligomers that are built by solid-phase synthesis as described earlier. Three-arm structures such as
T-shaped lipooligomers such as
In order to improve polylysine properties, Kataoka and coworkers used a polyethylene glycol (PEG) -polylysine block polymer, which was reacted with iminothiolane (67). Next, a few thiols were incorporated onto lysines without losing cationic charges, because the primary amines are converted into positively charged amidines. With siRNA, so-called polyion complex (PIC) micelles are formed. The presence of the iminothiolane modification results in disulfide-based cross-linked polymers stabilizing the siRNA containing PIC micelles. These micelles were able to achieve 100-fold higher transfection efficiencies than the analogous micelle without cross-linking. Stability tests with increasing concentrations of sodium chloride revealed that the stability of cross-linked polyplexes was significantly increased and could be abolished with reducing agents. Polyplexes with unmodified PEGylated polylysine were not even stable at a physiological ionic strength.
Many multifunctional carriers for siRNA that are based on the same principle but including shielding and targeting domains also rely on disulfide bonds (18). For example, the folate receptor-targeted oligomer
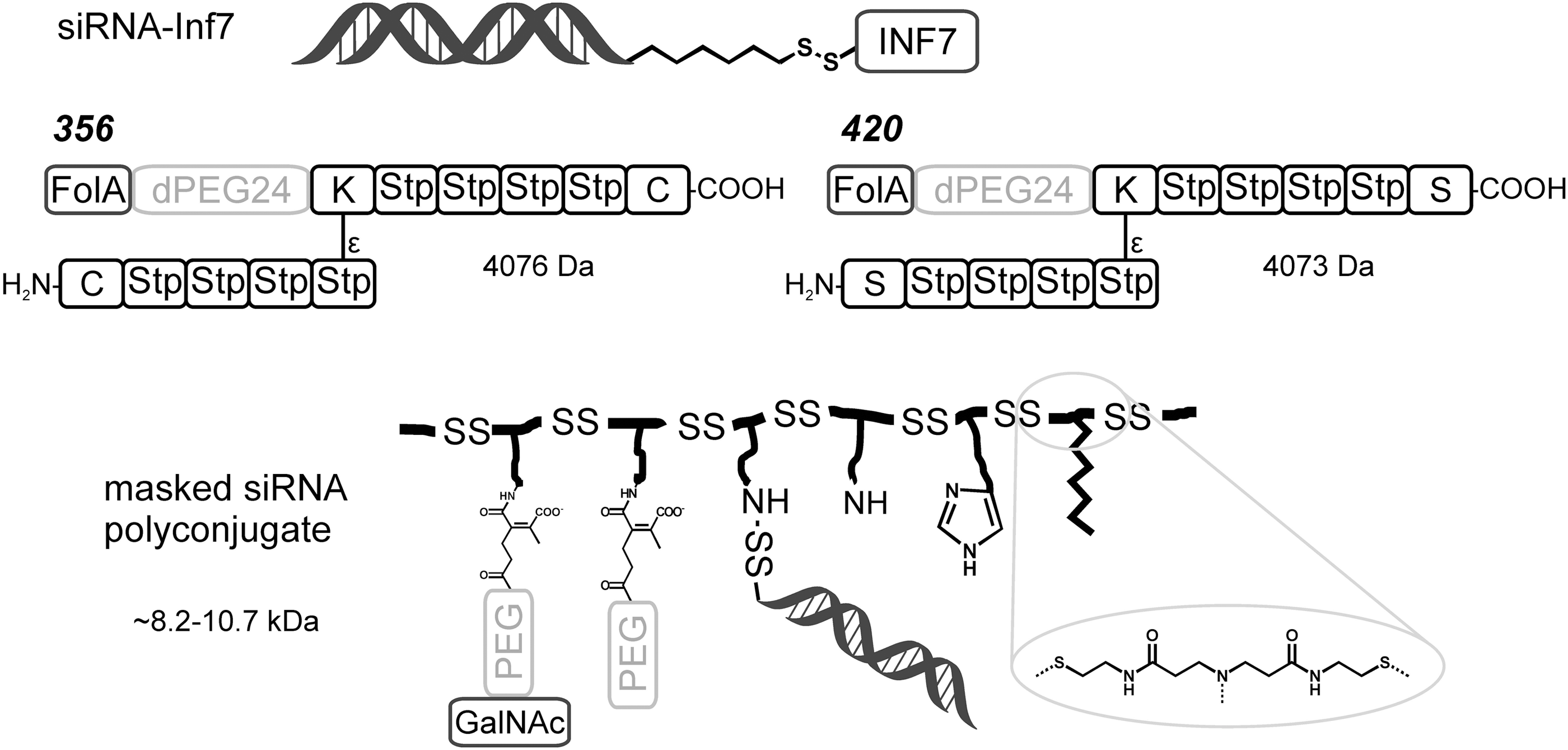
This reversible linking can also be used for attaching siRNA directly to a carrier. Using click chemistry, a PEG folate was covalently attached to siRNA via a disulfide-containing linkage (19). In addition, polycations were attached in this manner. Disulfides have been used to connect siRNA covalently to a pH-responsive masked endosomolytic polylysine carrier (70). Wang and coworkers used it for coupling to a bioreducible poly(amido amine) backbone (Fig. 6, bottom) (80). The side chains of the backbone in this carrier were functionalized with hydrophobic modifications, imidazoles, and pH-reversible targeting ligands. These multifunctional systems with bioreducible properties demonstrated efficient, targeted gene silencing in vivo. The best performing polymer with a mixture of 40/30/30 molar ratio of 2-(2-aminoethoxy)ethyl/2-(1H-imidazol-4-yl)ethyl/dodecyl and a molar weight of ∼8.64 kDa showed 80% mRNA knockdown in mice.
Protein Delivery by Covalently Linked Bioreducible Shuttles
Recombinant proteins present an important and growing class of modern biotherapeutics. Many of them, such as monoclonal antibodies or recombinant growth factors, have targets on extracellular sites (106, 114). Other proteins contained as antigens within vaccines are internalized into phagocytic cells and are degraded into peptides during the immune response. An efficient technology for intracellular cytosolic delivery of intact proteins (“protein transduction”) would tremendously open the applicability of a new class of therapeutic proteins and antibody domains that aim at intracellular molecular targets.
Natural proteins teach us that intracellular protein transduction can be feasible. The human immunodeficiency virus 1 (HIV-1), trans-activator of transcription (TAT) protein (27, 33), or the Antennapedia homeobox protein can cross cytoplasm membranes and internalize into cells under physiological conditions (42). A series of bacterial cytotoxins internalize by endocytosis, followed by subsequent translocation of a cytotoxin subunit across the endosomal membrane. In some cases, the reduction of disulfide bonds facilitates this delivery process. Such reduction-activated cytotoxins have been utilized in drug delivery; for example, listeriolysin-SS-oligocation conjugates for pDNA delivery (11, 90) or numerous immunotoxins (24, 25, 101). Commonly used synthetic protein delivery systems include cationic lipids (13, 14, 62), protein transduction domains (45), such as natural penetratin (85, 93) and HIV-TAT (45, 103), or artificial oligoarginines (93) and cationic polymers (17, 30, 59).
The sequence-defined oligocationic carrier
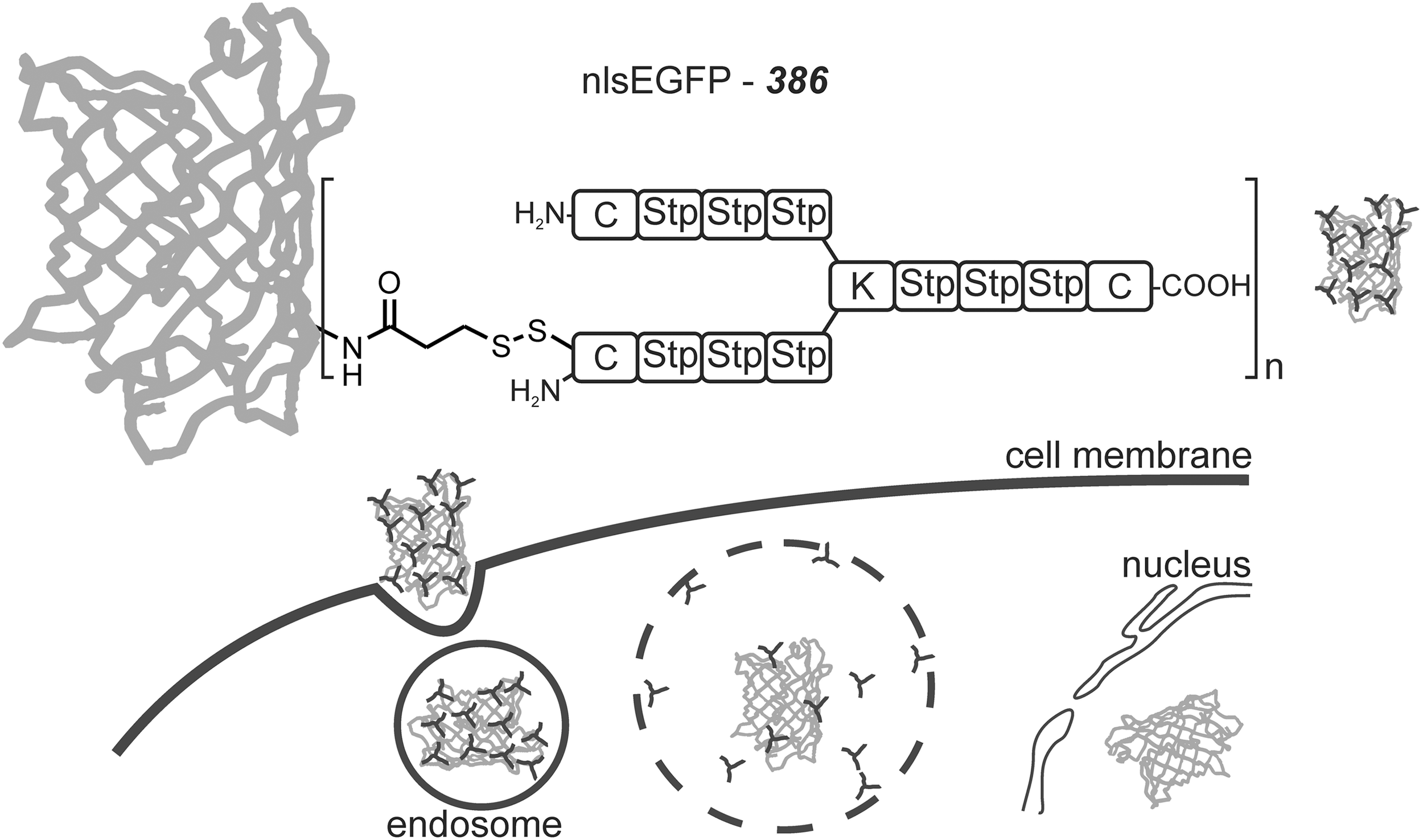
Tuning the Timing and Extent of Bioreduction of Dynamic Carriers
The various approaches for dynamic stabilization of bioconjugates and nanoparticulate carriers reviewed in the earlier sections build on the favorable difference between extracellular/intracellular redox conditions and GSH concentration. However, the frequently claimed notions that a) “disulfide bonds are generally stable outside the cell” or b) that “they are easily cleaved in endosomes” should be questioned as already stated earlier in literature (22), and might be even, in part, misconceptions, or, at least, not generally valid.
Several disulfide-bonded protein conjugates and immunotoxins have been found to be cleaved in vivo after intravenous application (9, 60). The questions about the exact site of bioreduction might be less important for some nanocarrier designs but critical for others. The majority of the previously mentioned strategies build on cytosolic cleavage of multiple disulfide bonds of oligocationic carriers that had caged their cargo during the extracellular delivery. In those cases, it might be less important whether bioreduction of some fewer bonds starts at the cell surface, significantly takes place in the endosomal vesicle system, or largely happens in the cytosol. When cleavages of large numbers (approximately hundreds) of disulfide bonds per nanoparticles are required, the majority of cleavage processes can be assumed for the cytosolic location because of its larger reduction capacity.
Bioreducible disulfide linkages, however, have also been considered dynamic elements in earlier steps of the delivery process: For example, in the process of nanocarrier deshielding, which might be beneficial to occur at the cell surface or within endosomes, or for activation of endosomolytic domains within endosomal vesicles. The so-called “PEG dilemma” comprises the experiences that surface shielding of nanocarriers with PEG is very beneficial for extended blood circulation and (passive or active) tissue targeting, however, negatively affects intracellular uptake across endosomal membranes. Endosomal pH-sensitive cleavable PEGs are able to overcome this dilemma (46, 50, 72, 76, 121). Thus, bioreductive cleavage of disulfide-bound PEG at the cell surface would be a very useful process (8, 54, 82, 107). In our own research, productive deshielding of PEGylated pDNA/PEI polyplexes was far more successful in utilizing pH-labile bonded hydrazone (121) instead of disulfide-bonded (56) PEG chains.
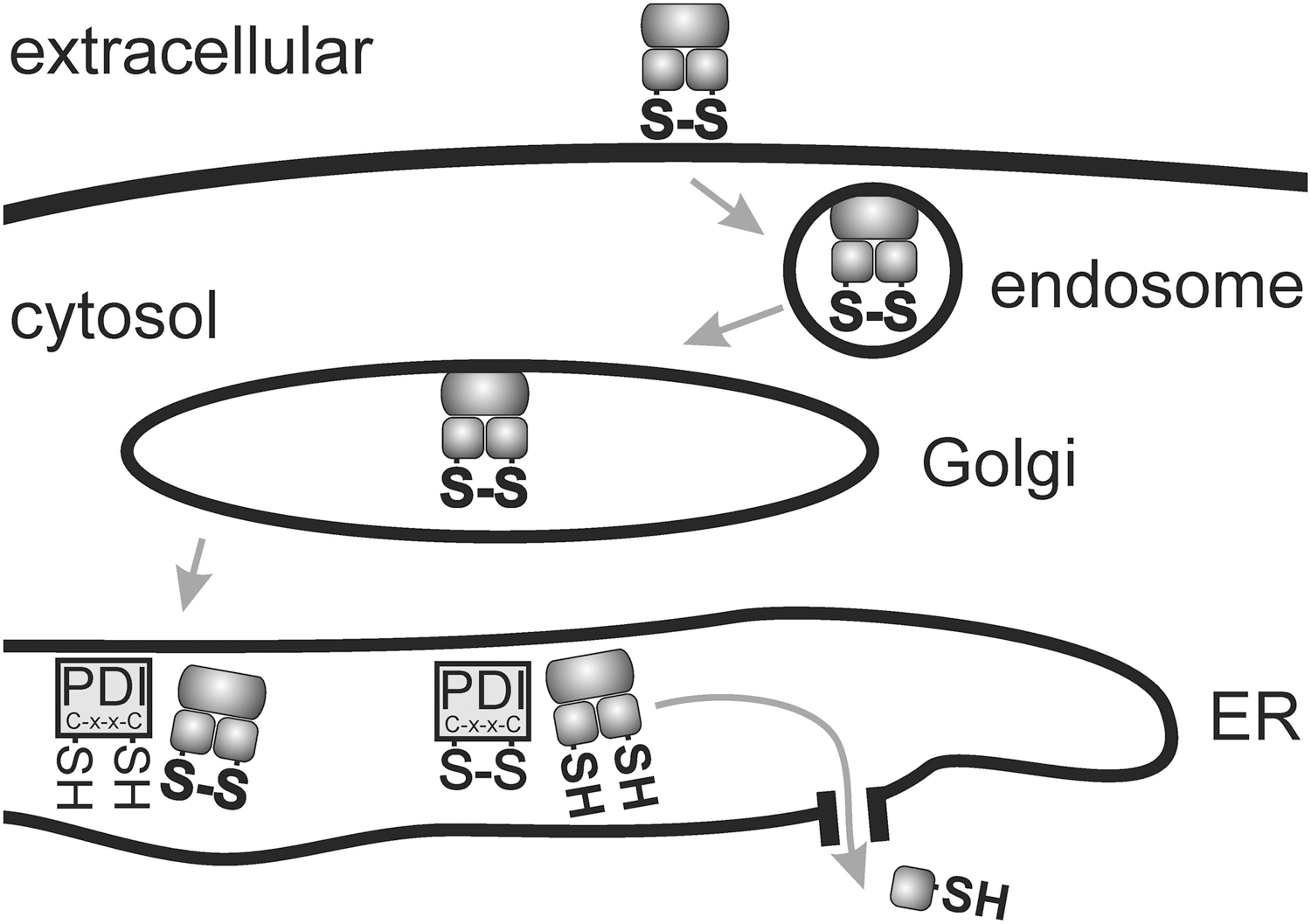
Some approaches such as those reported by Lee and coworkers (90) successfully build on reductive liberation of endosomolytic agents within the endosome for subsequent cytosolic transport of the cargo. Such strategies are rationally supported by natural examples, including toxins and viral translocation that include a disulfide cleavage as a part of the translocation process (69, 78, 83, 86, 99, 112, 120, 123). For example, several toxins of the AB5 family (A toxic subunit, B receptor-binding carrier units), including cholera toxin, shiga toxin, or pertussis toxin, enter cells by endocytosis. This is followed by retrograde transport into the ER lumen, where repair chaperone PDIs recognize single disulfide bonds as “misfolded protein domains.” Then, a physiological repair process named ER-associated degradation (ERAD) retro-translocates the A unit through a pore to the cytosol for proteasome-mediated degradation (Fig. 8) (120). AB-type toxins such as ricin or Pseudomonas exotoxin, or some viruses such as SV40 (99), also contain disulfide linkages, which require cleavage for cytosolic translocation, presumably mediated by PDIs. For translocation, diphtheria toxin requires an analogous cleavage by PDI in the endosomal compartment (86). It, however, should be kept in mind that bioreductive potential in standard acidifying endosomes might be more limited (22) and applicable for special cases only.
In sum, many bioreductive approaches are built merely on educated guesses that are supported by facts from nature and empirical findings and imagination (hopefully not too far from reality). A more quantitative understanding of the bioreductive behavior of cellular processes, however, would provide enormous help for improved designs and, thus, present important opportunities for the future.
In this direction, Leroux and coworkers (5) developed a well-characterized, defined dendrimeric oligomer probe for analysis of bioreductive disulfide cleavage in cellular systems. The probe consists of generation 3 dendrimer PAMAM-G3 with disulfide-linked dye (BODIPY)2, which provides the ability to distinguish between reduction of internal (dendrimer core) and surface disulfide bonds. Applying this system in four cell lines with different reductive potency (HeLa, A549>PC3>Caco-2 cells), both interesting expected findings and surprising findings were made. For example, data based on pulsed thiol depletion and endocytosis inhibitor studies revealed the rather surprising finding that probe disulfide cleavage occurred exclusively at the extracellular cell surface, possibly triggered by cell surface oxidoreductases such as PDIs or chaperones (also belonging to the PDI family), with secreted thiols such as GSH acting as cofactors. Incubation with a cell membrane-impermeable oxidant completely suppressed bioreduction.
The same study evaluated bioreduction of the PAMAM probe after pDNA complex formation. Incorporation into such DNA polyplexes reduced the cell surface bioreduction, which is consistent with sterically restricted access of the probe to cell surface oxidoreductases. For the most reducing HeLa cell line, the logical distinction between cleavable disulfides in surface and core states became apparent. Moreover, the expected subsequent bioreduction after delivery into the acidic endosomal environment was incomplete. Intentionally, the authors had selected G3 PAMAM, which in contrast to higher-generation PAMAM or PEI does not have “proton sponge” based endosomal escape properties. The addition of chloroquine as an endosomolytic agent strongly enhanced bioreduction, which was consistent with endosomal escape and efficient reduction in the cytosol. As the authors note, “efficient bioreduction in the endosomes, while sometimes reported, is not a universal phenomenon and should be verified for each new system and its corresponding target cell line” (5).
How may we utilize an increased knowledge about reductive characteristics of different biological microenvironments? Not only the biological surrounding influences the fate of disulfide bonds, but also the molecular chemical environment of disulfide bonds within a dynamic carrier can be tuned. For example, by stabilizing disulfide bonds by bulky groups providing sterical hindrance (47, 110), different exposure (core or surface) within a nanoparticle system (as illustrated in the PAMAM polymer example discussed earlier), the electrostatic environment facilitating or restricting electrostatic interaction with a reducing agent (5, 125), or the number and positioning of multiple disulfide bonds (126).
The influence of bulky groups on disulfide stability was shown in vitro with structures with methyl, benzene, and cationic residues localized next to disulfide bonds (110). When treated with 0.03 mM DTT at pH 7.4, the stability was as follows: The most stable derivate was the one with methyl and benzene residue at the α-C next to the disulfide bond, followed by the same derivate without methyl group, followed by a structure with an alkyl chain next to the disulfide group. The presence of a cationic charge in δ-C position next to the disulfide resulted in a faster reduction than with all uncharged linkers mentioned earlier. Based on this information, the most stable and the most labile structure were incorporated in a linker to connect an antibody (OX7) with a Ricin A toxin subunit to form immunotoxins. These immunotoxins were tested in mice for stability in vivo. From the more stable disulfide-linked immunotoxin with methyl and benzene residue, 50% in active form and 50% in free antibody form were present in blood 48 h after an intravenous injection. In contrast, the bioreducible linkage with the cationic element in its environment reached the same cleavage stage (50/50 ratio immunotoxin/free antibody) after only 8 h (110). More recently (47), a systematic study of disulfide-linked antibody-maytansinoid immunotoxins was reported while evaluating the disulfide-linker stability and antitumoral activity in vivo in mice. Sterical hindrance (by methyl group substitutions flanking the disulfide bonds) enhanced stability to reductive cleavage by DTT and plasma stability in mice. In in vivo efficacy testing, a conjugate with intermediate disulfide bond stability (having two methyl groups on the maytansinoid side of the disulfide linkage but no methyl substitution on the antibody linker side) displayed the highest antitumoral efficacy.
Not only the sterical and electrostatic environment of disulfides, but also the reducing agents and the pH have a crucial influence on the stability of a disulfide bond. It has been shown that positively charged amino acids next to a cysteine in a peptide destabilized the disulfide bond between oxidized dimers in a 10 mM GSH solution at pH 4.9 (mimics late endosomal/lysosomal environment) (125) The influence was higher for amino acids with higher pKa values (highest for arginine with pKa 12.1) and if more positively charged amino acids were incorporated. The distance (α- or β- amino acid next to cysteine) played a minor role. Analogous negatively charged amino acids with low pKa values (glutamic acid with pKa 4.3 has the strongest influence) stabilized the disulfide bond. Consequently, the half life of structures with α- and β-arginine was only 0.17 h compared with 8.7 h for structures with α- and β-glutamic acid. If GSH, which is also present in its deprotonated GS
Conclusions
The bioreducible structure of disulfide bonds offers some most beneficial options to oligocationic carriers for gene and protein delivery. Importantly, it provides a straightforward option for the design of dynamic carriers and shuttles discriminating between intracellular and extracellular cytosolic locations. The stability of the cargo polyplex that is needed during the harsh extracellular conditions and during cellular uptake can be provided by cross-links in a two- or three-dimensional manner via covalent conjugation or caging of the cargo. While pDNA polyplexes can be built on purely electrostatic interactions with commonly HMW carriers, the smaller siRNAs and cargo proteins may critically depend on such disulfide stabilizations. Intracellularly, disulfide bonds are efficiently cleaved by the reductive environment, releasing the cargo in a functional form undisturbed from the delivery shuttle. As an additional benefit, bioreduction disassembles oligocationic carriers into smaller, usually nontoxic fragments. Chemical tuning of bioreductive elements for site-/time-specific cleavage presents an encouraging future option for activating delivery functions beyond simple assembly/disassembly, for example, triggered deshielding or unmasking of endosomolytic, cytosolic transport and cargo release elements.
Footnotes
Acknowledgments
The authors thank Olga Brück (LMU) for skillful assistance in preparing this article. This work was supported by the German Research Foundation (DFG) special research focus project grant SFB1032 B4 and the Cluster of Excellence Nanosystems Initiative Munich (NIM).
