Abstract
Introduction
R
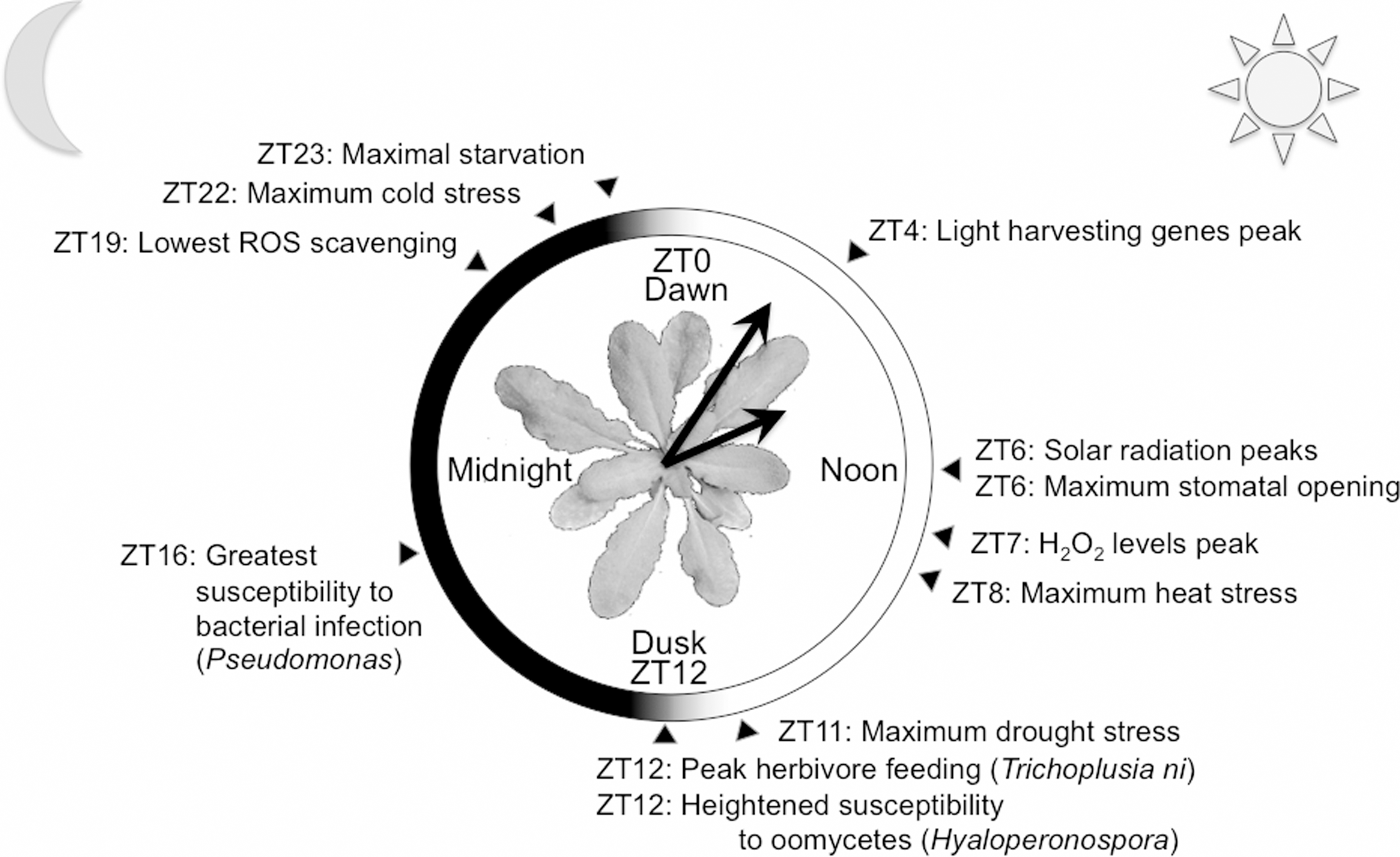
Environmental stress in plants can be roughly divided into abiotic and biotic factors. Abiotic stresses are numerous and include solar radiation, high or low temperature, drought or soaking, lack of nutrients, or osmotic stress. Incidence of abiotic stress is in many cases temporally variable over the circadian or annual cycle, and due to their sessile nature, one could say a plant has to dress for all weathers (Fig. 1). Furthermore, plants are, similar to us, under constant attack from a diverse range of potentially pathogenic organisms. Disease is a relatively rare occurrence, because plants posses an intricate immune system (53). Evidence for the involvement of circadian clocks in timed up-regulation of plant immunity has recently been gathered (9, 40, 140). Redox-based signals are involved in many, if not all, biotic and abiotic stress responses in plants (118, 123). Human populations can be greatly affected by reduced plant productivity as a result of stress. With the world's projected population increase, the global demand for food will drastically rise. Demand of plant-based energy production is also increasing, as emission of greenhouse gasses resulting from the use of fossil fuels will need to be reduced to slow down global warming. In the light of these human challenges, research on plant well-being is crucial to our development as a species (52), and the circadian clock is increasingly found to regulate more and more aspects of plants living healthy lives. The current discovery of rhythms in cellular redox (60, 98), associated with an ancient timekeeping mechanism (28), implies strict interplay between the circadian clock, stress responses, and cellular metabolism. Here, we will review this and other recent links as far as current knowledge allows, and discuss the many ramifications of these novel insights.
Plant Circadian Rhythms
In photosynthetic organisms such as plants, daily changes in light availability constitute a major metabolic change in a cell, and the circadian clock strictly regulates rhythmic photosynthesis (25, 45). In addition, the circadian clock regulates responses to predictable daily environmental stress, as well as plant hormonal signaling (109). In total, roughly a third of Arabidopsis transcripts oscillate with a circadian rhythm (20), compared with about 10% in any human tissue (127). In all taxa, transcriptional circadian changes are driven by clock genes, which themselves oscillate in a circadian manner (14). These clock genes feed back to directly or indirectly regulate their own expression, forming a network of transcriptional/translational feedback loops (TTFLs). Controlled by post-translational processes, the proteins engaging in TTFL oscillations drive more global changes in the cell's transcriptional housekeeping, dependent on circadian time.
Transcriptional/translational feedback loops
The structure and transcriptional targets of this transcriptional oscillator have been studied in great detail in many different organisms. Although the arrangement into TTFLs is shared between higher taxa, the identity of TTFL genes known today is unconditionally different. We will limit ourselves here to elaborating on the plant TTFL structure [excellent reviews exist that detail the clock structures of the mammalian (130), Drosphila (37), fungal (26), and cyanobacterial (50) circadian clocks].
Ongoing experimental efforts as well as mathematical modeling by many labs have created a reasonable understanding of how TTFL loops are wired together to collectively deliver robust ∼24-h oscillations in plants. Although more than 18 years after the identification of the first plant clock mutants in Arabidopsis thaliana (80 –82), new TTFL components and new functions for old components are still identified at a high frequency. The current understanding of central hubs of the plant TTFL network can be summarized as follows (Fig. 2). A morning-expressed heterodimeric transcription factor complex of late elongated hypocotyl (LHY) and circadian clock-associated 1 (CCA1) negatively regulates the expression of the timing of CAB1 expression (TOC1). LHY and CCA1 are partly redundant and still regulate transcriptional activity as homodimers, but the circadian clock runs too fast in the respective single mutants (79). Heterodimers have been shown to bind more strongly to their cognate promoter elements than homodimers in vitro (99). A so-called evening element (EE) is over-represented in the promoters of the many CCA1/LHY regulated genes (45). The EE is appropriately named, as the vast majority of genes downstream of EEs share a peak phase late in the day (45). CCA1/LHY binding at the EE in the TOC1 promoter prevents histone H3 acetylation (101), presumably maintaining a dense chromatin structure. Inhibition of deacetylation changes the circadian expression pattern of TOC1. The associated acetylases and deacetylases remain unknown, but additional regulation is identified by CCA1 paralog REVEILLE8, which promotes H3 acetylation at this site (29).
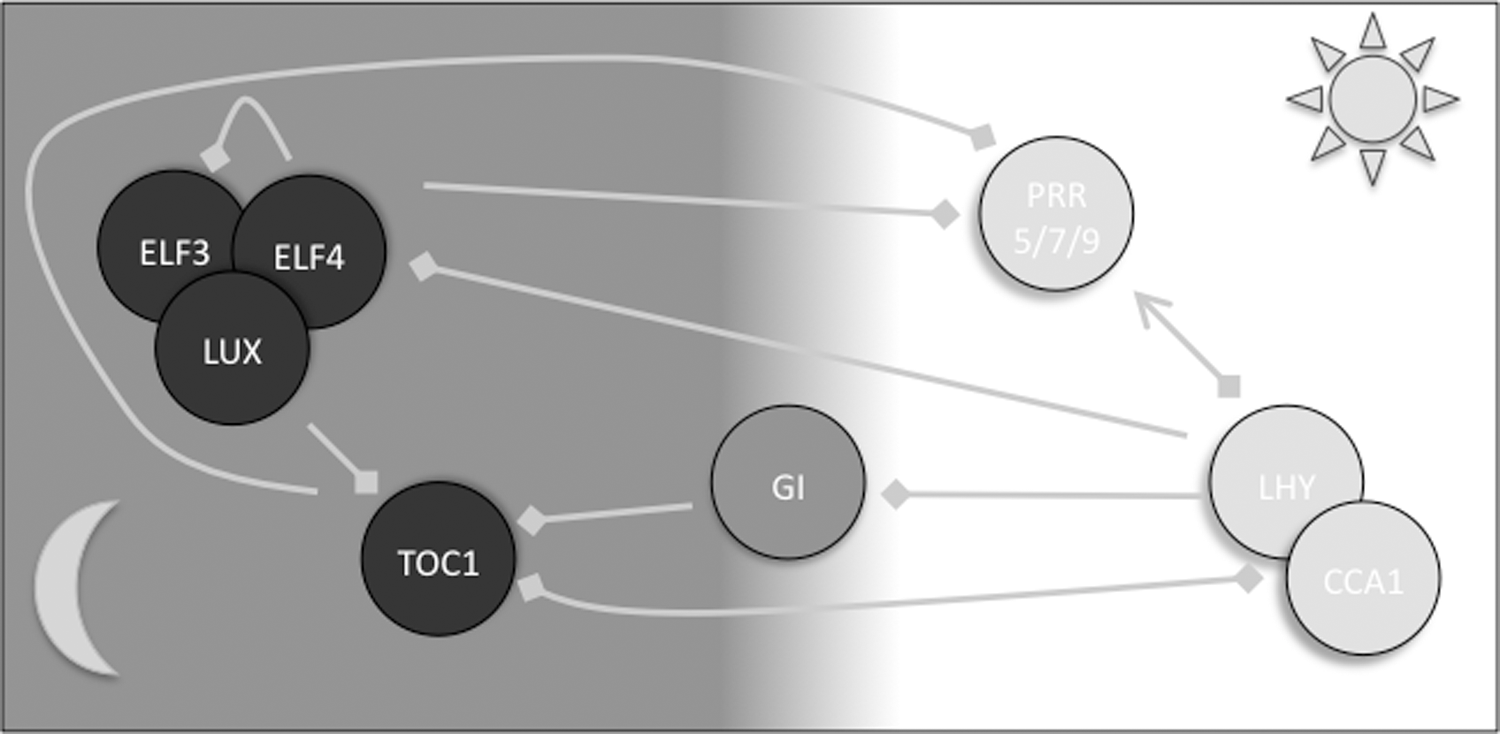
TOC1 feeds back to CCA1 and LHY by directly or indirectly binding their promoters (104). Reminiscent of the reciprocal promoter interaction, TOC1 binding to CCA1/LHY promoters is associated with changes in histone acetylation (93). Until very recently, TOC1 was believed to feed back to CCA1/LHY positively (2, 69), but, in fact, TOC1 is a transcriptional repressor (38, 48, 102). Several peripheral loops feed into this CCA1/LHY-TOC1 “repressilator” structure. A set of morning-phased TOC1-related genes known as pseudo-response regulators (PRRs) is positively regulated by CCA1/LHY (30, 44). PRR5, 7, and 9 repress expression of CCA1/LHY (91, 103). The evening-phased Gigantea protein is negatively regulated by CCA1/LHY, and, in turn, negatively regulates TOC1 (102). The most prominent force in the evening-phased clock appears to be a trimeric protein complex between early flowering (ELF) 3 and 4 and lux arrhythmo (LUX) (96). The members of this so-called evening complex (EC) are strongly repressed by CCA1/LHY (102). The EC itself represses expression of its member proteins ELF3 and LUX (47, 56). Furthermore, ELF3 and ELF4 repress TOC1 (24). The EC binds promoters of clock-regulated downstream genes (24, 47), and night-phased, EC-centered oscillations are thought to explain why sustained rhythmicity can be observed in double-mutant cca1 lhy plants (102).
In spite of the remarkable complexity of TTFL structures, the arrangement of clock genes and proteins in loops is not sufficient to explain all aspects of rhythmic behavior. An intricate system of post-translational regulation acts on TTFL proteins, tuning period length, amplitude, and phase of the transcriptional clock. Across circadian organisms, phosphorylation by evolutionarily ancient kinases such as casein kinase 1 and 2 and glycogen synthase kinase 3 is essential for maintaining healthy endogenous clocks (14, 34, 37, 75, 150). SUMOylation has been shown to be involved in regulating the stability of clock protein transcriptional activity (15, 63). The clock influences output pathways by moderating chromatin arrangements via various redox-dependent histone modifications (90, 101). In addition, ubiquitination is important for timed proteasomal degradation of clock proteins. In plants, ubiquitin modification is the most-studied post-translational modification in TTFL rhythmicity (134), with roles for redox-sensitive F-box proteins FKF1 and LKP2 (6), both oscillating clock components in their own rights. In fact, in the marine unicellular algal species Ostreococcus tauri, targeted protein degradation is important for proper timekeeping at any given phase of the circadian cycle (135). In contrast, transcription and translation are not essential at large phases of the cycle (98), implying that in green lineage timekeeping, when to degrade is more important than when to synthesize.
The hormonal clock
The circadian clock regulates key aspects of plant growth, development, and environmental stress responses by strong and diverse connections to phytohormone signalling (20, 25, 45, 109). Hormone levels themselves oscillate, and the effects of hormone exerts are often circadian gated, meaning that the effect of a hormone stimulus will be more or less pronounced depending on the time of day.
Highly significant overlap exists between genes responsive to any hormone tested and clock-regulated genes, demonstrating functional crosstalk of clock and hormone outputs (20). Endogenous levels of many hormones, including auxin, jasmonic acid (JA), salicylic acid (SA), abscisic acid (ABA), cytokinin, and ethylene (ET), fluctuate over diurnal light/dark cycles, and for some hormones, even truly circadian oscillations that persist in constant light have been shown (Fig. 3) (19, 20, 40, 54). Interestingly, redox signaling in plants is mediated by hormones as well as the circadian clock (see below), while hormone responses are gated and co-regulated by the circadian clock. For these reasons, links between circadian- and redox signaling in plants cannot be discussed without addressing hormone pathways and links to the clock. One of the processes regulated by hormones and the circadian clock is stomatal opening. The clock allows anticipation of stomatal opening and closing to dawn and dusk, respectively (41). However, during the day, the plant faces a trade-off between opening stomata to allow respiration on the one hand, and closing them to avoid evaporation and drought stress on the other. In the absence of stress, stomatal opening circadially peaks in the middle of the day (41). In addition, ABA stimulates stomatal closing (109). ABA is involved in aspects of development and in responses to biotic and abiotic stress, and more than 40% of ABA-induced genes are clock controlled (20). Water deficiency stress leads to ABA-induced stomatal closing during the day, antagonizing clock regulation. This effect of ABA, however, is gated by the clock, meaning that a weak water deficiency in the leaf will not lead to ABA-induced stomatal closing in the morning, but it will in the middle of the afternoon (18) even though the ABA signal is of the same amplitude. This is an example of clock gating weighing up the advantage of respiration at a phase when temperatures are not yet high in the morning versus the disadvantage of losing too much water at the hottest times of day.
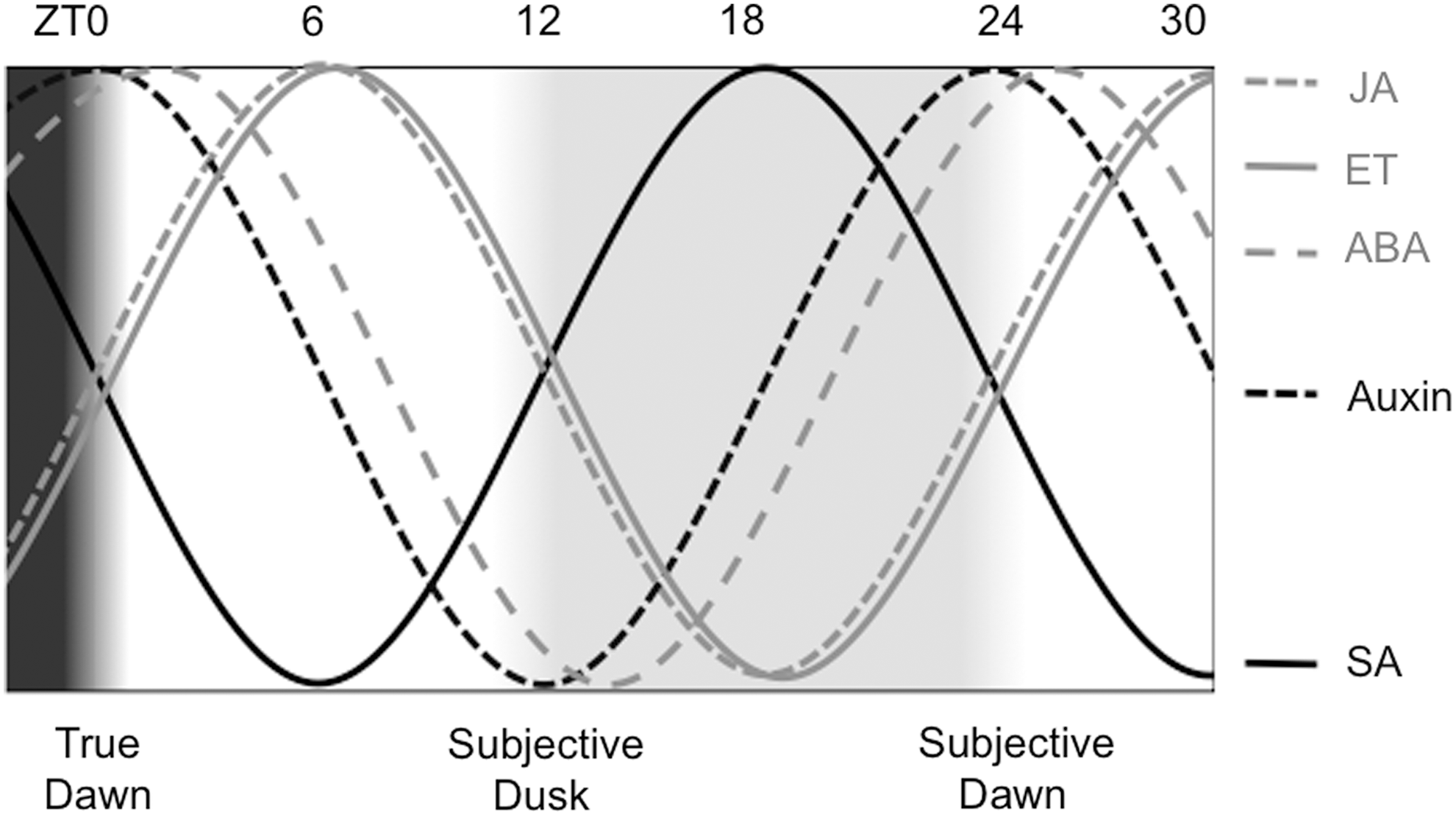
Auxin is the major hormone regulating plant growth and development. Many auxin-responsive and signaling genes turned out to be clock regulated and peak in the middle of the day (19, 20). In fact, auxin levels oscillate in Arabidopsis (19, 54), but circadian-regulated gene expression of auxin-responsive genes does not rely on rhythmic auxin itself. Responses to auxin on transcriptional and developmental level are circadian gated, and are stronger in the subjective night than in the subjective day (19). Auxin also regulates stomata, but unlike ABA, auxin leads to stomatal opening. Clock gating of auxin-mediated stomatal opening results in more effective stomatal opening in response to auxin during subjective day than during subjective night (120). The opposing phase of circadian gating of auxin signals in stomatal opening (gate closed at night) compared with gating in transcriptional and developmental processes (gate closed in daytime) highlights how important clock gating is in translating potentially ambiguous signaling cues to orchestrate the responses that are most appropriate for a specific time of day.
ET is produced in a markedly circadian manner, peaking in the middle of the day (95). This rhythm presumably originates from rhythmic transcription of precursor synthesis genes (132). TTFL function is unaltered by exogenous ET application, and ET-insensitive mutants are not clock compromised, suggesting that there is no feedback from ET back to the clock (109). However, ET has important functions in biotic and abiotic stress responses, which might explain the evolutionary advantage of rhythmic ET emission.
Jasmonates and salicylates are the main hormones involved in activating biotic stress responses in plants; for example, in response to bacterial and fungal attack or herbivore feeding. Basic levels of jasmonates and salicylates are under circadian regulation and peak at opposing phases; jasmonates in the middle of subjective day, and salicylates in the middle of subjective night (39, 40). Feeding of the herbivore Trichoplusia ni is circadian, with the herbivore eating more plant material in the subjective day. In properly timed plants, clock-mediated defenses restrict the damage to plant leaves to small amounts of tissue. However, when defenses are mistimed by differential entrainment of the herbivore and plants, the plants lose about half their leaf material to herbivory. The advantage of proper clock-mediated timing is lost in clock-deficient mutants as well as in jasmonate-deficient mutants (39, 40), solidifying the notion that clock and hormone signaling work together to counteract biotic stress.
Plant Redox Rhythms
Rhythmic reactive oxygen species
It was recently shown that peroxide levels oscillate in Arabidopsis plants in light/dark cycles as well as in constant light (60). Direct light effects are clearly manifested by elevated levels of H2O2 in constant light, but the sustained rhythmicity under constant light shows that the circadian clock regulates rhythmic peroxide. In a normal light/dark cycle, recurrent direct light effects will emphasize rhythmic production of reactive oxygen species (ROS). Presumably as an evolutionary strategy to counteract this rhythmic insult, ROS-related genes are also clock regulated (60). More specifically, these genes contain functional CCA1 binding elements in their promoters. Activity of peroxide-scavenging catalase enzymes is also circadian, and this rhythm is a direct result of clock regulation of expression of the three Arabidopsis catalase genes (60). Mutation of CCA1 abrogates rhythmicity of ROS-related genes, of H2O2 levels, and of catalase activity. Furthermore, exogenous application of agents that induce oxidative stress leads to dramatic oxidative damage in CCA1-deficient mutants at concentrations which only moderately affect wild-type plants. Mutation of ELF3, ELF4, PRR5, 7, or 9 also leads to increased susceptibility to oxidative damage, whereas plants that lack TOC1 are more resistant than wild-type plants (60). These results indicate that we lack comprehensive understanding of this phenomenon, but that the clock regulates redox-responsive gene products and redox-related metabolism.
Rhythms in peroxiredoxin oxidation
Recently, redox-based circadian rhythms in post-translational modification of antioxidant peroxiredoxin (PRX) proteins were observed in the marine unicellular green alga O. tauri (98). Post-translational redox-driven S-sulfination (chemistry of which will be detailed later) of 2-Cys PRX peroxidatic cysteines can be monitored using an antibody specifically recognizing PRX in the sulfinic conformation. Using this antibody, rhythms were observed in Ostreococcus cells that were kept in constant darkness (98). This is remarkable, because this obligate phototrophic species shuts down all transcription in constant darkness, meaning that the rhythms in PRX S-sulfination, and therefore assumedly rhythms in H2O2, persist in the absence of CCA1 rhythms or, in fact, any transcription (98). This result indicates that circadian timekeeping is not per se dependent on TTFL loops, and more specifically, that redox rhythms do not depend on CCA1. The limited physiological relevance of studying phototrophic cells in constant darkness arguably restricts drawing broad conclusions from the Ostreococcus result, but due to the extremely high conservation of PRX among higher taxa, the sulfination-specific PRX antibody could be employed to investigate PRX rhythms in other species. Concomitantly with the Ostreococcus experiments, PRX S-sulfination was found to be rhythmic in human red blood cells (97). These cells naturally lack a nucleus and, thus, lack any nuclear transcription or TTFL control. The ex vivo experimental setup also uncouples these cells from any conceivable mobile signals from brain or peripheral circadian locomotors in the bloodstream. When PRX was identified as the first rhythmic molecular output shared between plants and humans, this result was followed up by monitoring PRX states over circadian time series in representative species from every domain of life (28). Astoundingly, PRX S-sulfination was found to be rhythmic in diverse species such as Arabidopsis, mouse, the fungus Neurosopora crassa, the cyanobacterium Synechococcus elongatus, the archaeon Halobacterium salinarum, and the worm Caenorhabditis elegans (28, 100). An Arabidopsis double mutant that lacks functional copies of both 2-Cys PRXs is not arrhythmic, and neither is a PRX-knockout strain of the cyanobateria (28), suggesting that PRX is apparently not a mechanistic component of general timekeeping, unless TTFL rhythmicity is independent of the nontranscriptional processes which drive PRX oscillations. Either way, PRXs are an invaluable (and currently the only) rhythmic marker that traces the so-called nontranscriptional oscillator (NTO).
PRX proteins are evolutionarily ancient and date back as far as 2.5 billion years ago, when the evolutionary invention of photosynthesis by bacteria started a geologically speaking abrupt increase in atmospheric oxygen concentrations (117), in a process known as the great oxidation event. Due to its reactivity, increased ambient oxygen caused substantial difficulties for living cells with mass extinction as a result. However, PRX and other H2O2-scavenging enzymes evolved and provided cells with such large strategic advantages that every current life form on Earth contains these genes (28). Therefore, redox-based rhythms in plant cells are not a priori different from those in other eukaryotic cells, and the existence of a redox-based ancestral proto-clock is not inconceivable (134).
Reversible Redox Modifications
In plants, ubiquitous small-molecule redox couples exist that limit damage from potentially harmful ROS and reactive nitrogen species (RNS) (118). Plants employ ROS-related molecules as bona fide signaling molecules, and induction of stress in plants is commonly associated with oxidative/reductive bursts involving clock-gated hormone signaling. Cellular redox pairs include nicotinamide adenine dinucleotide phosphate (NADP+/NADPH), and reduced/oxidized glutathione and ascorbate. These factors are crucial signaling molecules for plant responses to stress, notably to pathogen attack (123). Clock functions for these redox couples are not known in plants, but well established in mammalian chronobiology. Now that the PRX results have unearthed similarity in rhythmic redox between species (28, 100), this analogy becomes more relevant. Interestingly, plant glutathione pools and the ratio of reduced versus oxidized glutathione is regulated by the circadian hormones JA and SA (123). In addition to small-molecule compounds, ROS-scavenging enzymes are among the most ubiquitous proteins in the plant cell, and as discussed earlier, catalases and PRXs are rhythmically regulated at transcriptional and post-translational levels (60, 134). ROS-induced cellular redox changes lead to oxidation of free thiol side chains on cysteines of regulatory proteins, providing post-translational, redox-sensitive control of these signaling hubs.
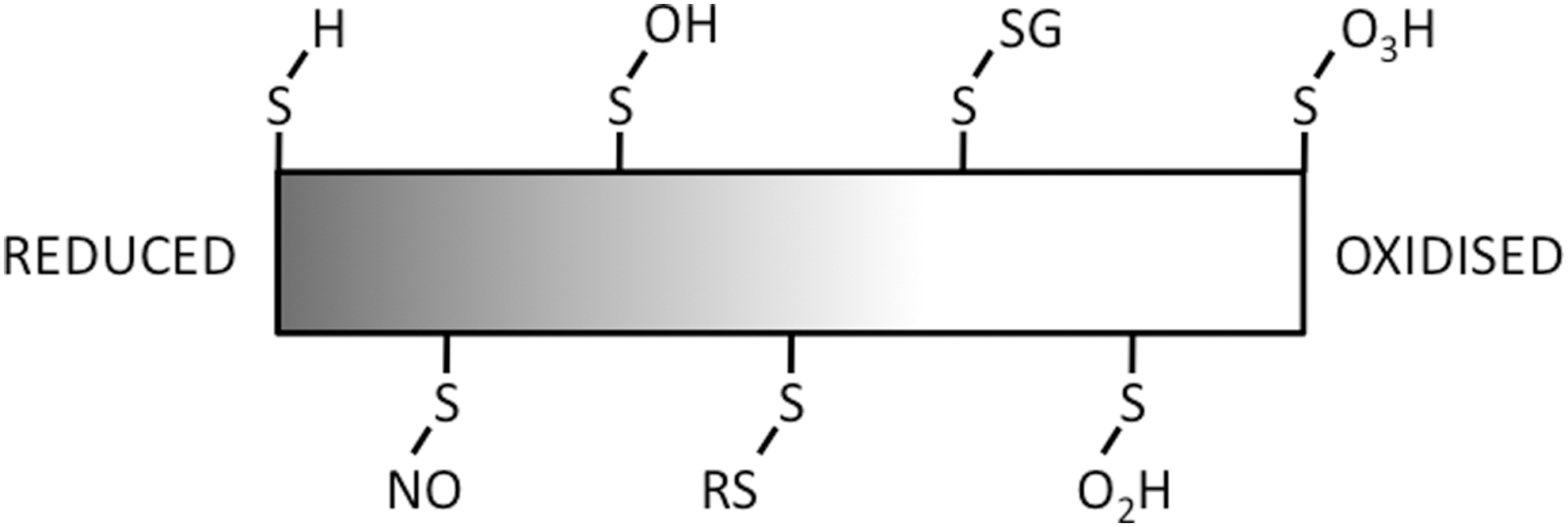
Cysteines are intrinsically nucleophilic, and they can form highly reactive thiolate groups in response to minor redox-state fluctuations. Cysteines are often deeply buried in the protein three-dimensional structure, where availability for redox-active modification depends on the protein conformational state or vice versa, thereby forming molecular switches. Reversible modifications of the cysteine thiol (Fig. 4) include the covalent attachment of nitric oxide (NO) (S-nitrosylation), thiol hydroxylation (S-sulphenation), disulphide bridge formation (S-thiolation), covalent attachment of glutathione (S-glutathionylation), and further oxidation of sulfenic groups to the sulfinic and sulfonic states. As will be discussed later, reversible cysteine modifications that provide these excellent cellular signaling opportunities regulate the function of plant stress responses, providing a redox link between clock-gated hormone signals and downstream activation of stress responses.
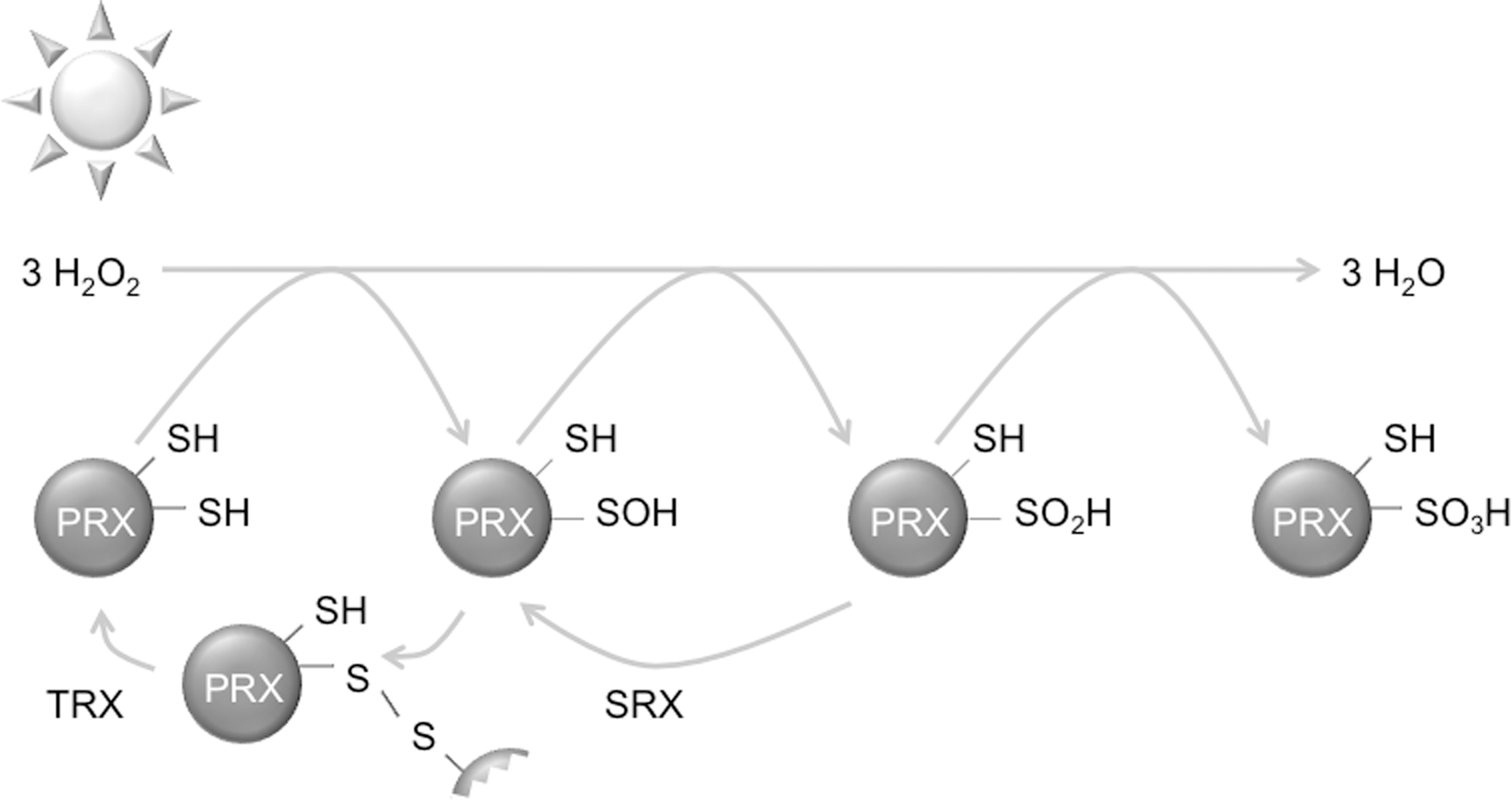
Uniquely, many of these cysteine oxidations can be found in PRX proteins (Fig. 5). Driven by H2O2, PRX undergo iterations of an oxidative/reductive cycle, where an active-site cysteine is oxidized to the sulfenic form, neutralizing one peroxide molecule. PRX is reduced back to the initial state by first forming a disulphide bridge with a second active site cysteine on another PRX molecule. This disulphide bond is reduced enzymatically by thioredoxin (TRX), returning both PRX molecules to the reduced form ready to scavenge another H2O2 molecule (42, 134). Proteins of the 2-Cys PRX class can scavenge further peroxide molecules by over- and hyperoxidation of the peroxidatic cysteine from the sulfenic to the sulfinic and sulfonic form. Dimers of these overoxidized PRX proteins can form higher-order multimers that exhibit chaperoning functions and signaling functions. Overoxidized PRX can be rescued back to the sulfenic state by sulfiredoxins (SRX). In another unicellular alga, Chlamydomonas reinhardtii, protein disulphide isomerase (PDI)2 was found to interact with 2-Cys PRX exclusively during the night phase (32). Potentially, PDI2 could be involved in the reduction of oxidized states of PRX to the ground state during phases of low H2O2. Consistently, PRX S-sulfination peaks during the subjective day phase in Ostreococcus and Arabidopsis (28, 98), coinciding with peak H2O2 levels (60).
The SA-responsive transcriptional coactivator nonexpressor of pathogenesis-related genes 1 (NPR1) provides an example in which redox modifications reversibly influence hormone-responsive transcription after stress. Sets of cysteines in multiple NPR1 molecules form disulphide brides to build homo-oligomeric complexes in the plant cytosol (88). Disulphide formation is facilitated by previous S-nitrosylation on a specific cysteine, presumably bridging the energetic gap between reduced and disulphide-bonded cysteines or between NPR1 conformational states. Formation of NPR1 multimers is essential for maintaining a large pool of NPR1 that is available for acute responses to biotic stress: pathogen attack leads to SA-regulated cycles of cellular reduction and oxidation, which, respectively, reduce or promote formation of disulphide bonds between NPR1 molecules. This transiently releases NPR1 monomers, which are translocated to the nucleus where they signal transcriptional activation in a process involving proteasome-mediated turnover of NPR1 (125, 126, 129). Nuclear NPR1 exhibits its transcriptional role in concert with heterodimers of thymine–guanine–adenine (TGA) transcription factors (12, 23, 55, 66). Two of these, TGA1 and TGA4, are thought to contain an intramolecular disulphide bridge in their “resting” state, preventing interactions with NPR1. Cellular reduction may break this bond, allowing physical association with NPR1; while subsequent S-nitrosylation and S-glutathionylation of TGA cysteines result in enhanced DNA binding and transcriptional activity of the complex (23, 66). In this process, redox-based modification of cysteines on NPR1 and TGA factors, thus, regulates multimerization, stability, subcellular localization, intermolecular interactions, and transcriptional activity in plant innate immune responses.
Given the ubiquitous nature of rhythms associated with redox and hormone signaling, diverse redox-based post-translational modifications discussed here are likely to be instrumental in regulating cellular transcriptional rhythmicity. Following this reasoning, the enzymes associated with regulating reductive states of proteins are highly relevant to plant chronobiology, and will be discussed next.
Enzymatic Reductions and Cysteine Modifications
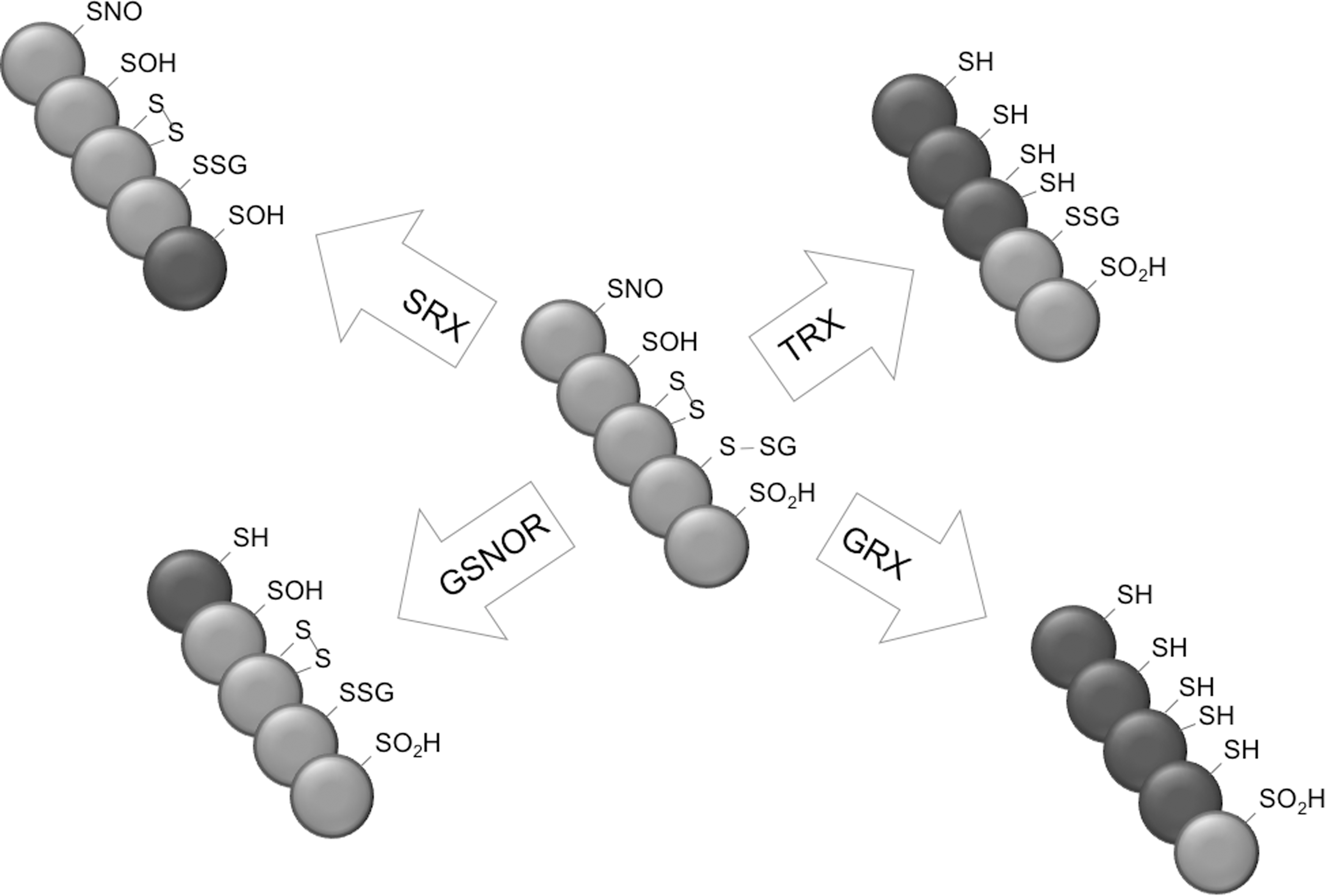
The variety of cysteine modifications provides significant complexity to the proteome. Nonetheless, proteomic profiling of hyper-reactive cysteines in eukaryotic cells indicates that these residues are a relatively rare feature (141), in line with cellular signaling functions. Accordingly, it has now emerged that each of the varying degrees of oxidative cysteine modifications may regulate the function, localization, activity, and stability of the proteins that harbor them (121), as discussed earlier. Cellular utilization of cysteine modifications as signaling cues for the regulation of proteins is complicated by the fact that most occur spontaneously on exposure to different small redox molecules (i.e., cellular redox buffers, ROS and RNS). To circumvent this problem, it appears that cells have evolved a large array of redox enzymes which are capable of reducing cysteine modifications (Fig. 6), thereby providing a redox-dependent signaling switch.
Disulphide reduction
Enzymatic reduction of the disulphide bond has been widely studied and, consequently, is well understood. Disulphide reduction is most prominently catalyzed by the TRX system. TRX is a small antioxidant enzyme with a conserved -Cys-–Gly-–Pro-–Cys-– active site sequence. The active site is a part of a characteristic structural fold common to all members of the TRX superfamily. The first cysteine of the active site resides at the N-terminal end of an α helix, providing it with a permanent dipole that lowers the pK
a of this cysteine (59). The low pK
a of this cysteine maintains it in a reduced state with a sufficiently high reduction potential and nucleophilic character, allowing it to attack disulphide bonds in target proteins. Interestingly, disulphide attack results in the formation of a mixed disulphide bond between the first active site cysteine of TRX and the cysteine of the target protein. This mixed disulphide is subsequently resolved by the second active site cysteine, yielding the TRX oxidized and the target protein reduced (Eq. 1). Mutation of the second active site cysteine prevents it from resolving the mixed disulphide, thereby trapping the target protein covalently onto TRX. The strong nature of this bond has been extensively exploited by molecular biologists to identify TRX target proteins. Plant biologists were among the first to apply this method to identify TRX targets that are specific to different plant cell organelles, bringing the total to more than 500 different targets (3, 4, 86, 87).
In contrast to those in animals, plant TRXs belong to two distinct redox systems that recycle their activity using different reducing agents. In the chloroplast, the powerhouse of the plant cell where oxidative stress and modifications are prevalent, TRX activity is recycled by ferredoxin–thioredoxin reductase (FTR), which uses the reducing power of ferredoxin (FDX) to reduce the active site of chloroplastic TRX proteins (Eq. 2). Since FDX is reduced in the light, the TRX/FTR system is in a more reduced state during the day; while at night, molecular oxygen oxidizes the system. In the second redox system, prevalent in other organelles and the cytosol, TRX activity is recycled with the reducing power of NADPH by NADPH-dependent thioredoxin reductase (NTR) (Eq. 3). Three partially redundant isoforms of NTR have been described in plants. Isoforms NTRA and NTRB reside in the cytosol and mitochondria (107), while NTRC is localized to the chloroplast (116).
In addition to TRXs, the superfamily of TRX enzymes includes glutaredoxins (GRXs). Similar to TRXs, many GRX enzymes contain two active site cysteines that function to reduce disulphide bonds in target proteins. Despite the similar structural fold of the active sites of these oxidoreductases, GRX shows far greater affinity toward protein-S-S-glutathione mixed disulfides. Consequently, GRX is most likely a major player in controlling signaling through protein S-glutathionylation. GRX enzymes with a single active site cysteine are also abundant and utilize a monothiol mechanism for reduction. Both the dithiol and monothiol mechanisms for disulphide reduction or protein deglutathionylation depend on the nonenzymatic recycling of GRX activity by the reducing power of glutathione (Eqs. 4 and 5) (78). Oxidized glutathione (GSSG) is, in turn, reduced by the activity of NADPH-dependent glutathione reductase (GR).
Recent data have shown a surprising connection between the TRX and GRX systems in plant cells. Arabidopsis mutant gr1 plants lack cytosolic GR activity and were shown to accumulate high levels of GSSG (71). Although these mutants have reduced buffering capacity against ROS, gr1 mutants do not exhibit strong phenotypic defects and reduction of GSSG on exposure to oxidative stress was still possible, albeit delayed. These data suggest that an alternative cytosolic mechanism for GSSG reduction exists in plant cells. Genetic crosses of gr1 to ntra ntrb double mutants and biochemical analysis of GSSG reduction by TRX indicated that the TRX/NTR system functions as a cytosolic backup for GSSG regeneration (71) and may also blur the lines between disulphide and deglutathionylation activities of TRX and GRX enzymes.
Protein denitrosylation
Protein S-nitrosylation has recently emerged as an important post-translational signal in a variety of cellular processes. Protein denitrosylases have now been identified that provide protection against S-nitrosylation or allow it to be utilized as a signaling switch. A variety of NO donors may cause protein S-nitrosylation, but of particular biological relevance is S-nitrosoglutathione (GSNO), the S-nitrosylated form of glutathione. GSNO is thought to function as a mobile reservoir of NO bioactivity and is a potent NO donor, causing extensive protein S-nitrosylation (67). GSNO levels are controlled by the activity of GSNO reductase (GSNOR), which shows high specificity for GSNO and can be found from bacteria to yeast, mammals, and plants (31, 67, 112). Genetic knockout of GSNOR results not only in elevated levels of GSNO, but also of S-nitrosothiol (SNO)-proteins, indicating that GSNOR functions as an indirect protein denitrosylase (31, 67). GSNOR denitrosylation activity plays critical roles in human pathology and in plants, it influences both biotic and abiotic stress responses (7, 63, 123).
Comprehensive purification of denitrosylase activity from mammalian cells identified the TRX/NTR system as a potent denitrosylase with direct activity toward SNO-proteins (7). Subsequent experiments have shown that TRX may catalyze the reduction of many different SNO proteins which are involved in a variety of cellular processes [reviewed in (114)]. Thus, TRX enzymes simultaneously harbor disulphide reduction and denitrosylation activities. Denitrosylation of SNO proteins by TRX releases either NO radicals or HNO, but the molecular mechanism remains unclear (Eq. 6). Denitrosylation has been proposed to occur through both mixed disulphide formation and through trans-nitrosylation, a process in which the TRX active site becomes transiently S-nitrosylated (7, 94, 128). Notably, other members of the TRX superfamily, such as GRX and PDI, may also function as potential protein denitrosylases (49, 119).
Reduction of hyper-oxidized cysteines
Higher-order oxidized states of cysteines, including cysteine sulphinic and sulphonic acids, were long considered irreversible and have been associated with loss of protein activity in several disease states and pathologies. This notion was changed for cysteine sulphinic acids with the finding that 2-Cys PRX sulphination is reversible (143, 144). It was subsequently shown that the sulphinic acid reductase SRX was responsible for this reduction (Fig. 5) (10). SRX utilizes an unusual ATP-dependent mechanism in which the target cysteine sulfinic acid is first phosphorylated, creating a leaving group that facilitates the formation of a mixed disulphide S-monoxide between SRX and PRX, which is readily resolved by TRX or GSH (51, 70). A. thaliana SRX (AtSRX) localizes to the chloroplast where it targets 2-Cys-PRXs. While genetic analysis of AtSrx mutants indicated that these plants are sensitive to exogenously applied ROS, surprisingly, they also exhibit increased tolerance to photo-oxidative stress, suggesting that AtSRX has complex functions depending on the type of oxidative stress encountered (68, 108). It should be noted that SRX enzymes are specific to 2-Cys PRXs, so the existence of further reductases that target sulfination awaits discovery.
Circadian Redox and Plant Immunity
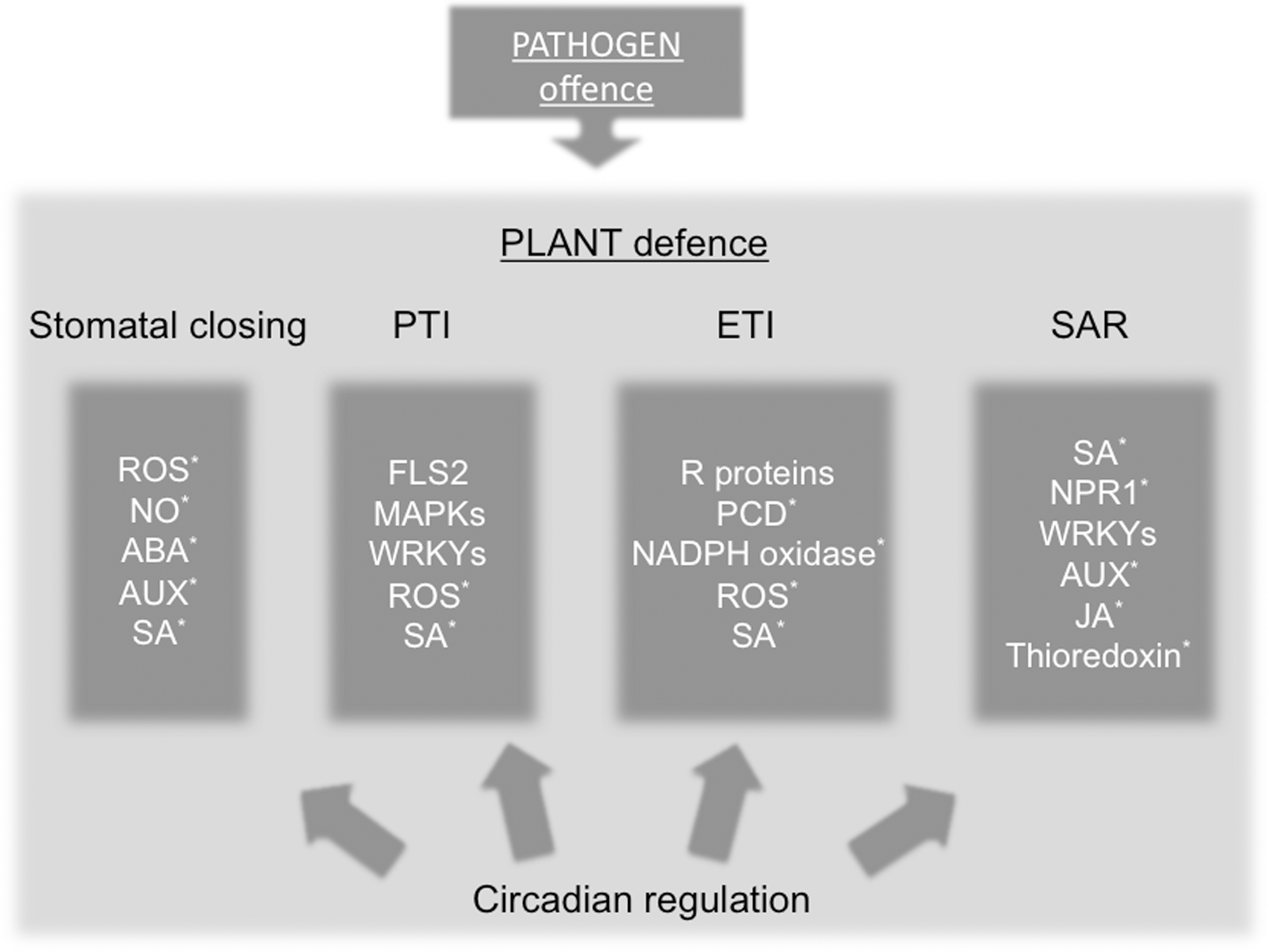
As discussed earlier, many plant hormones and cellular redox molecules are circadian. Importantly, interplay between plant hormones and cellular redox molecules is of vital importance to the regulation of the plant immune system. Combined with recent reports showing circadian regulation of the plant immune system, this suggests that circadian redox rhythms may play an important role in plant-pathogen interactions (Fig. 7).
Stomatal defenses and pattern-triggered immunity
Resistance against the biotrophic bacterial pathogen Pseudomonas syringae pv. tomato (Pst) DC3000 was recently shown to be controlled by the circadian clock. Plants are least susceptible to infection in the subjective morning, and this rhythmicity was lost by CCA1 overexpression or elf3 mutation (9). Resistance against Pst DC3000 is conferred by distinct overlapping layers of the plant immune system (122). To infect a plant, Pst DC3000 should first invade the leaf lamina through stomata. Subsequent recognition of pathogen-associated molecular patterns (PAMPs) by pattern recognition receptors at the plant cell membrane constitutes the first layer of defense, known as pattern-triggered immunity (PTI). Activation of PTI induces rapid stomatal closure to prevent further access of pathogenic bacteria into the leaf intercellular space. PTI-induced stomatal closure requires components from the SA and ABA signaling pathways, both of which have been shown to exhibit circadian regulation (see above), in conjunction with RNS and ROS. PAMPs induce the accumulation of NO in stomatal guard cells, and pharmacological scavenging of NO prevented PTI-induced stomatal closure (76). Moreover, mutation of the NADPH oxidase respiratory burst oxidase homolog D (RBOHD) abolished PTI-induced ROS production, resulting in a failure to close stomata and prevent Pst DC3000 invasion (77). Interestingly, stomatal aperture and conductance show circadian rhythmicity (16, 41). Recalling the fact that many ROS appear to be clock regulated, this raises the intriguing possibility of interplay between the control of stomatal aperture by circadian mechanisms and by ROS/RNS-mediated immune mechanisms.
After the PTI-induced ROS burst, mitogen-activated protein (MAP) kinases are activated that induce a subset of immune genes. Bhardwaj et al. (9) analyzed publically available microarray datasets for the presence of diurnal and circadian rhythmicity in immune-associated genes. Strikingly, nearly all of the more than 100 genes analyzed showed diurnal regulation, whereas slightly more than half also displayed a circadian pattern. Among the latter were genes with crucial roles in establishing PTI, including a signaling sector consisting of the pattern recognition receptor FLS2 that recognizes bacterial flagellin, downstream MAP(K)s, and the PTI-associated transcription activator Trp-Arg-Lys-Tyr (WRKY)22. Accordingly, PTI-induced cell wall reinforcement against pathogen invasion showed time-of-day dependency, which was largely abolished in arrhythmic CCA1 overexpressing plants (9). Clearly, circadian regulation adds a novel layer to PTI against Pst DC3000 in Arabidopsis.
Effector-triggered immunity
Adapted pathogens have evolved effector proteins that suppress the onset of PTI and promote virulence. Co-evolution between plants and pathogens resulted in the development of plant intracellular immune receptors known as resistance (R) proteins that keep the cell under surveillance for the presence of pathogen effectors (122, 136). Activation of R proteins initiates a second layer of plant defences known as effector-triggered immunity (ETI). A major hallmark of R-protein-activated ETI is the rapid onset of the hypersensitive response, a form of programmed cell death that isolates and ultimately kills invading biotrophic pathogens (122). Strikingly, the expression of many R proteins is under circadian control (9, 140). The functional consequences of this were recently investigated in ETI against downy mildew infection. Resistance against this biotrophic pathogen was shown to largely depend on a cluster of genes that initiate defence and programme cell death. This set of immune genes, including the R gene RPP4, which confers full immunity to downy mildew strains, was controlled by the circadian regulator CCA1, exhibiting peak expression at night and early morning, coinciding with downy mildew sporulation and dispersion of spores. Accordingly, artificial infection at dusk rather than dawn resulted in a CCA1-dependent increase of susceptibility to downy mildew infection (140). It, thus, appears that plants time the expression of defence genes and R genes involved in programmed cell death to coincide with the time of day at which pathogen threat is imminent.
ROS/RNS are involved in programmed cell death on activation of ETI (21, 22, 133), and circadian control of their production, scavenging, and gene targets was recently shown (60). The production of ROS during ETI is, in part, due to the activity of NADPH oxidases, some of which show circadian expression (9) and are related to mammalian oxidases that are involved in the pathogen-activated respiratory burst in phagocytes (133). Importantly, ETI-induced activity of the Arabidopsis NADPH oxidase AtRBOHD is itself regulated by RNS. RNS species produced on infection specifically S-nitrosylate AtRBOHD at Cys890, impeding binding of the cofactor flavin adenine nucleotide (FAD) and impairing its ability to produce ROS. In accordance, mutation of Cys890 rendered RBOHD insensitive to RNS, resulting in enhanced pathogen-induced ROS production and programmed cell death (147). Moreover, pathogen-induced S-nitrosylation of AtRBOHD was shown to be governed by the denitrosylase GSNOR1. Mutant gsnor1 plants accumulate high levels of SNO proteins, exhibit low NADPH oxidase activity, and are highly susceptible to pathogen infection (31, 147). Nevertheless, these mutants exhibit accelerated, albeit ineffective programmed cell death on pathogen infection. These findings suggest that GSNOR1-governed SNOs facilitate programmed cell death independent of ROS, while concomitantly suppressing NADPH oxidase activity to fine-tune the production of ROS on activation of ETI. A role for SNOs in regulating ROS levels was further substantiated by the finding that PRX IIE activity is inhibited by S-nitrosylation (111). This antioxidant enzyme detoxifies peroxynitrite (ONOO−), a product formed on reaction of NO with superoxide, which interferes with cellular signalling through protein tyrosine nitration. Thus, SNO regulation of PRX IIE modulates the cellular level of ONOO−.
Systemic acquired resistance
A third layer of defence against pathogen infection is the activation of an immune memory known as systemic acquired resistance (SAR). SAR is activated on primary infection and establishes long-lasting, broad-spectrum resistance against subsequent pathogen attack throughout the entire plant (122). Activation of SAR is dependent on accumulation of the immune hormone SA, as failure to accumulate this hormone results in extreme pathogen susceptibility (36, 142). As discussed earlier, the expression of several SA synthesis genes and the cellular level of SA are also under circadian control (9, 40), implying that the clock influences the efficiency of SAR. SA and its functional analogues transiently alter the cellular redox environment (58, 72, 88, 123, 137). Reactive cysteine residues on signaling proteins sense these redox changes. In plant immunity, the transcriptional coactivator NPR1 senses SA-induced changes in cellular redox by changing its conformation as discussed earlier. Nuclear NPR1 monomers regulate the expression of over 2500 genes, including those encoding circadian-regulated immune WRKY transcription factors (9, 88). Cellular reduction by itself, however, is not the only mechanism that drives monomerization of NPR1. A search for NPR1 interacting redox enzymes identified the SA-inducible oxidoreductase TRX-h5 as a catalyst for redox-regulated NPR1 conformation (61, 129). Mutant trx-h5 plants accumulated far less NPR1 monomer on SA treatment compared with wild-type plants. Moreover, both trx-h5 and ntra mutants failed to induce SAR against P. syringae, indicating that the cytosolic TRX/NTRA system is critical for the establishment of SAR. Thus, it is plausible that the TRX system is regulated indirectly by the circadian clock through the cyclic accumulation of SA or perhaps even directly, as is the case for the chloroplastic TRX system, in which expression of TRX-f and TRX-m genes is controlled directly by CCA1 binding to their promoters (5, 65). In addition to SA, establishment of SAR may involve the sequential action of the hormones JA and auxin whose accumulations are also controlled by the circadian clock (20). While JA triggers potent cellular oxidation (123), aspects of auxin signaling may be controlled by the action of the TRX/NTRA-NTRB systems (106), demonstrating further overlap between redox processes in immunity and the clock.
JA-dependent defense responses
Defence responses against necrotrophic pathogens and insects depend on the production of the defence hormones JA and ET. JA and ET function synergistically to regulate the expression of a large set of defence genes that is distinct from those controlled by SA. Consistently, the circadian peak phase for JA and ET levels coincides around midday, totally antiphasic with SA (Fig. 3), which is a major antagonist of JA signaling (124). Recent evidence indicates that redox enzymes are involved in regulating JA and ET accumulation. Silencing of GSNOR expression in wild tobacco plants led to decreased JA and ET accumulation on attack by the insect herbivore Manduca sexta, suggesting that high levels of GSNO or SNO proteins suppress JA synthesis (145). Reduced GSNOR activity also led to decreased accumulation of defence-related secondary metabolites on methyl-JA application, indicating that GSNOR1 function is required not only upstream but also downstream of JA/ET. Consequently, GSNOR-silenced tobacco exhibits enhanced susceptibility to M. sexta feeding, indicating that GSNO homeostasis is vital to appropriate regulation of JA/ET-dependent responses against herbivores. Recalling our discussion on the intricate role of the circadian clock in JA and ET production to anticipate insect feeding behavior (40), the data suggest that circadian and redox-based pathways converge on JA/ET-dependent defence responses against insects.
In addition to GSNOR, GRX enzymes are also involved in JA signaling. GRXs were found to interact with TGA transcription factors that play important roles in the regulation of SA- and JA/ET-responsive genes (92, 148). As discussed earlier, intramolecular disulphides are thought to preclude some TGA factors from interacting with the SA-responsive transcription coactivator NPR1, while SNOs and S-glutathione may regulate DNA binding (23, 66). Ectopic expression of several GRXs suppressed the activation of JA/ET-responsive genes. In case of GRX480, this required interaction with TGA factors and its GSH binding domain, implying GRXs as chief candidates for the regulation of redox-based modifications of TGA factors (92, 148).
Circadian Redox and Abiotic Stress
In addition to biotic threats, plants are continuously exposed to different abiotic stresses such as exposure to high light, extreme temperatures, drought, and salt stresses. The circadian clock and both the production and signaling roles of ROS and RNS play intricate roles in the response of plants to abiotic stress. ROS are produced in different organelles, with the chloroplasts making high contributions during the light phase. Photosynthesis in the chloroplasts always generates some ROS, but under conditions of high light, electrons may bypass the normal electron flow of the photosynthetic chain, thus generating high levels of oxidative stress. In addition to chloroplasts, the peroxisomes and mitochondria are significant sources of intracellular ROS production, while NADPH oxidases generate apoplastic ROS involved in cellular signaling and systemic signaling in response to stress (83). In contrast, RNS production is less well understood, but advances have been made recently in understanding the synthesis of NO in the green lineage. While in animals NO is produced by NO Synthase (NOS) through the NADPH-dependent oxidation of
Transcriptional profiling has clearly shown that different ROS and RNS induced by abiotic stimuli regulate the expression of thousands of genes (35, 138) through transcription factor up-regulation (35). Biochemically, ROS may function through modulating the activity of MAP kinase signaling pathways. Components of the cold- and salt-responsive MEKK1-MKK1/2-MPK4/6 pathway are activated by hydrogen peroxide, which also induces NO production by a nitrate reductase through the phosphorylative action of MPK6 during lateral root development (131, 139). It is also likely that reactive cysteines of key transcriptional regulators are subject to and regulated by oxidative modifications, but few explicit examples are currently known. In analogy to mammals, plant heat shock factors are thought to be activated by oxidative modification of redox-sensitive cysteines in their DNA-binding domain (1). As a testimony of oxidative modified transcriptional regulators in plants, however, several antioxidant enzymes with cysteine-reducing activities have been implicated in responses to abiotic stresses. The TRX/NTR antioxidant system has been implicated in oxidative stress that is the result of developmental processes (105, 106, 115). Moreover, chloroplastic NTRC is involved in abiotic stress responses, as a knock out mutant showed increased sensitivity to ROS, drought, and salt stress. NTRC is of particular interest, as this unusual protein contains both functional NTR and TRX domains, but, surprisingly, does not function as a TRX/NTR system in a single polypeptide (116). Furthermore, a screen for mutants with elevated sensitivity to high temperatures identified GSNOR1. NO-overproducing mutants were similarly impaired in thermotolerance, and pharmacological assays show that wild-type plants treated with NO donors became susceptible to temperature increases, implying a wider role for NO donors and thus S-nitrosylation in the regulation of thermotolerance (62).
The circadian clock provides an adaptive fitness advantage by delivering anticipation of daily or seasonal changes (25). It is, therefore, perhaps not surprising that links between the circadian clock and abiotic stress responses have been strengthened in recent years. Microarray data analyses revealed a large overlap between genes controlled by the circadian clock and those regulated by heat, cold, salt, and osmotic stresses, and the clock controls many genes that are responsive to or involved in the biosynthesis of ABA, an indispensable regulatory hormone in abiotic stress responses (20, 73, 84). Consequently, ABA levels exhibit circadian oscillations (19), and many arrhythmic clock mutants and transgenics display abiotic stress phenotypes [reviewed in (113)]. The production and scavenging of ROS, critical to abiotic stress signalling, are regulated by the circadian clock and ROS- and clock-regulated genes significantly overlap (20), primarily via CCA1-mediated regulation (60), revealing an intimate relationship between the circadian clock, ROS, and responses to abiotic stress.
Future Perspectives
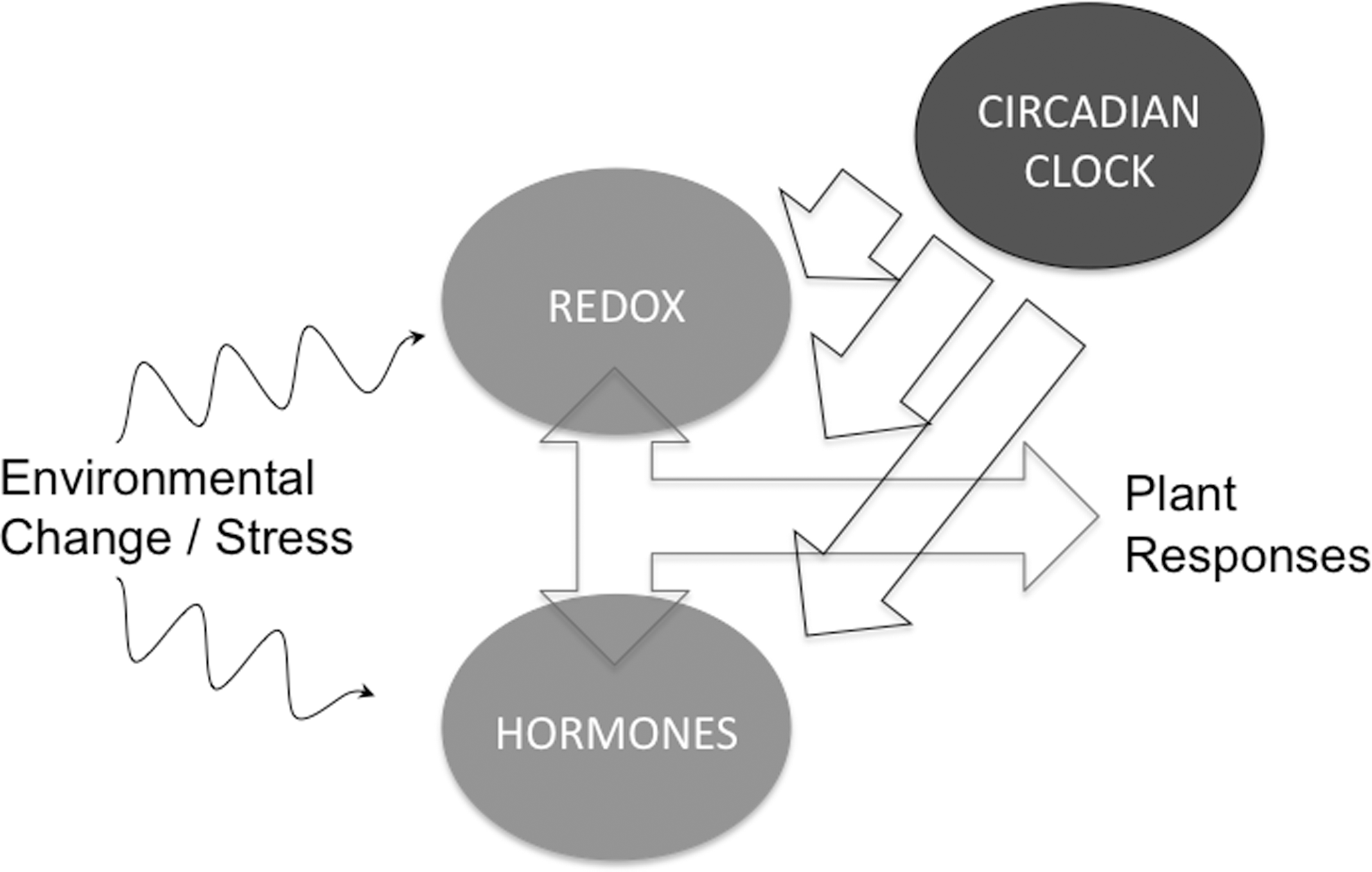
The identification of broadly conserved circadian PRX S-sulfination highlights that oxidative/reductive cycles could be the driving force behind the evolution of circadian rhythms. It is clear that both TTFL rhythmicity and the uncharacterized NTO are associated with oscillations in cellular redox states. Recent advances reveal intimate interplay between clocks, redox and hormone signaling in both healthy and diseased plants (Fig. 8). The molecular mechanisms of this interplay are, however, not yet clear. An analysis of circadian gene expression in mutants of redox enzymes compromised in biotic and abiotic stress responses could pave the way to a better understanding.
Innovation
Plant stress negatively affects agricultural production, and basic understanding of cellular mechanisms behind stress is, therefore, of high importance. Recently, the circadian clock has been shown to influence the cellular redox state and hormone signalling. These two factors are the main pathways which plants use to negate the effects of stress, and, therefore, a clear implication would be that the circadian clock helps plants deal with stress. Mechanistic bases behind this cross-talk remain unclear, and this review aims at combining current knowledge of the relevant parts of all these interconnected networks to advance our understanding of this emerging research area.
Footnotes
Acknowledgments
S.H.S. and G.v.O. are Royal Society Research Fellows (Fellowships UF090321 and UF110173, respectively), and this work was additionally supported by Royal Society grants to S.H.S. (RG110495) and G.v.O. (RG120372).
