Abstract
Introduction
L
We have studied the transcriptome and redox proteome under NaOCl stress in the mycothiol (MSH)-producing model bacterium Corynebacterium glutamicum. Using liquid chromatography–tandem mass spectrometry (LC-MS/MS) analysis and redox proteomics we provide for the first time evidence that protein S-mycothiolation is a widespread protein protection mechanism under NaOCl stress in Actinomycetes. The S-mycothiolome includes proteins that function in the glycolysis, biosynthesis of methionine, glycogen, nucleotides, thiamine, protein translation, and antioxidant functions, including striking conservation of some S-thiolated proteins (e.g., Tuf, MetE, GuaB) across Gram-positive bacteria. We further show that the S-mycothiolated peroxidase Tpx is inactive and reactivated by the Mrx1/MSH/Mtr electron pathway in vitro.
In eukaryotes, protein S-glutathionylation has emerged as major redox-regulatory mechanism that controls the activity of redox sensing transcription factors and protects active site cysteine (Cys) residues from irreversible oxidation to sulfonic acids (8). Many eukaryotic proteins, like α-ketoglutarate dehydrogenase, glyceraldehyde 3-phosphate dehydrogenase, ornithine δ-aminotransferase, pyruvate kinase, heat specific chaperones, and regulatory proteins (c-Jun, NF-κB) are reversibly inactivated or activated by S-glutathionylation (8, 24). However, little is known about the regulatory role of protein S-thiolation for bacterial physiology. We have recently identified protein S-bacillithiolations as mixed BSH protein disulfides in response to NaOCl stress in Bacillus and Staphylococcus species (7). Protein S-bacillithiolation controls redox-sensing regulators and protects active site Cys residues of essential and conserved metabolic enzymes from overoxidation. The redox-sensing MarR-family OhrR repressor is inactivated by S-bacillithiolation after NaOCl stress resulting in upregulation of the OhrA peroxiredoxin (Prx) that confers NaOCl resistance (6, 29). Mass spectrometry identified 54 S-bacillithiolated proteins in different Bacillus and Staphylococcus species that include 29 unique proteins and 8 conserved proteins. The S-bacillithiolated proteins function as metabolic enzymes in biosynthetic pathways for amino acids (methionine, branched chain and aromatic amino acids), cofactors, nucleotides; or as translation factors, chaperones, redox, and antioxidant proteins (7). The methionine synthase MetE is the most abundant S-bacillithiolated protein in Bacillus species under NaOCl stress. S-bacillithiolation of MetE occurs in its active site Zn center leading to enzyme inactivation and methionine auxotrophy (6). Since amino acid biosynthetic enzymes and translation factors are conserved in the S-bacillithiolome among Firmicutes, we propose that S-bacillithiolation could function in redox-regulation of protein synthesis.
Protein disulfides and S-thiolations are reversible Cys oxidations that are reduced by the thioredoxin (Trx)/thioredoxin reductase (TrxR) system or by the glutaredoxin (Grx)/GSH/glutathione disulfide reductase system in Escherichia coli (15). The Trx system is involved in reduction of inter- and intramolecular protein disulfides and the Grx proteins catalyze de-glutathionylation of S-glutathionylated proteins. In Actinomycetes, a Grx-like mycoredoxin1 (Mrx1) with a CGYC active site motif has been structurally and biochemical characterized. Mrx1 reduces a MSH mixed ß-hydroxyethyl disulfide (HED) and exclusively uses the MSH/Mtr/NADPH electron pathway (57). Corynebacterium glutamicum serves as model for pathogenic Actinomycetes, such as Corynebacterium diphtheriae and Mycobacterium tuberculosis and is also of biotechnological importance. The disulfide stress response in Actinomycetes is controlled by the ECF sigma factor σR in Streptomycetes coelicolor or its σH homolog in C. glutamicum (26). σR or σH are sequestered at control conditions by their cognate redox-sensitive anti sigma factors, RsrA or RshA, respectively in S. coelicolor or C. glutamicum. Redox-regulation has been shown for the RsrA/σR system in S. coelicolor. RsrA senses disulfide stress by oxidation of the Zn-binding Cys residues leading to Zn release and free σR(1). The RshA/σHsystem of C. glutamicum controls genes for the Trx/TrxR system (trxB, trxB1, trxC), for MSH biosynthesis and recycling (mshC, mca) and for the mycothiol disulfide (MSSM) reductase (mtr) (3, 11). The regulation of the Trx/TrxR and MSH pathways by RsrA/σR or RshA/σH homologs is conserved among Actinomycetes (1, 26, 45).
However, nothing is known about proteins that are redox-controlled or protected by protein S-mycothiolation under ROS stress conditions in Actinomycetes. Thus, we applied here transcriptomics, redox proteomics, and mass spectrometry (MS) to analyze the NaOCl-stress response and proteome-wide protein S-mycothiolations in the MSH-producing C. glutamicum.
Results
NaOCl-stress elicits a RshA/σH disulfide stress response in C. glutamicum
Previously, we identified protein S-bacillithiolation by sub-lethal NaOCl concentrations in Firmicutes bacteria (7). To optimize the conditions for protein S-mycothiolation, the MSH-producing C. glutamicum ATCC13032 wild type was grown in minimal medium and treated with sub-lethal NaOCl concentrations that reduced the growth rate about half-maximal. Treatment of exponentially growing C. glutamicum cells with 180 μM NaOCl at an optical density at 500 nm (OD500) of 8.0 resulted in a slower growth compared to untreated cells (Supplementary Fig. S1; Supplementary Data are available online at
The changes in the C. glutamicum transcriptome in response to disulfide-stress provoked by diamide were previously published (3, 11). To visualize the overlaps in the gene expression profiles between NaOCl and diamide stress, we performed a hierarchical clustering analysis. The cluster analysis revealed a strong upregulation of the disulfide stress σH regulon by both diamide and NaOCl (Fig. 1, Supplementary Fig. S2, Supplementary Table S3). About 40 genes of the σH regulon were 2–80-fold induced by NaOCl stress that are labeled in the Heat map and Scatter plot in Figure 1. The rshA gene encoding the anti sigma factor for σH was most strongly upregulated by NaOCl stress (22–47-fold). In addition, NaOCl stress caused strong inductions of the σH-regulon genes for thioredoxins trxB1 (10–13-fold), trxC (3-fold), cg1375 (3-fold), the thioredoxin reductase trxB (4–10-fold), dsbA-like thiol-disulfide oxidoreductases cg2838 or dsba1(15–29-fold), cg2661 or dsba2 (2–4-fold), and two methionine sulfoxide reductases msrA and msrB (12–15-fold). The σH-regulon genes for MSH biosynthesis, recycling and reduction were also upregulated, including the Cys ligase mshC (6-fold), the MSH-S-conjugate amidase mca (3–10-fold) and the MSSM reductase mtr (2–5-fold). Most strongly induced were the σH-dependent phage-associated gene cg0378 (50–80-fold) and the tRNA-(5-methylaminomethyl-2-thiouridylate)-methyltransferase gene cg1397 (17–45-fold). The σH-regulon also integrates the ClgR and HspR heat-shock regulons that control chaperones and proteases of the protein quality control machinery (3, 11). The genes of the ClgR-regulon (clgR, clpP2, dnaJ2) and HspR regulon (hspR, dnaK, clpB) were 2–11-fold induced by NaOCl stress.
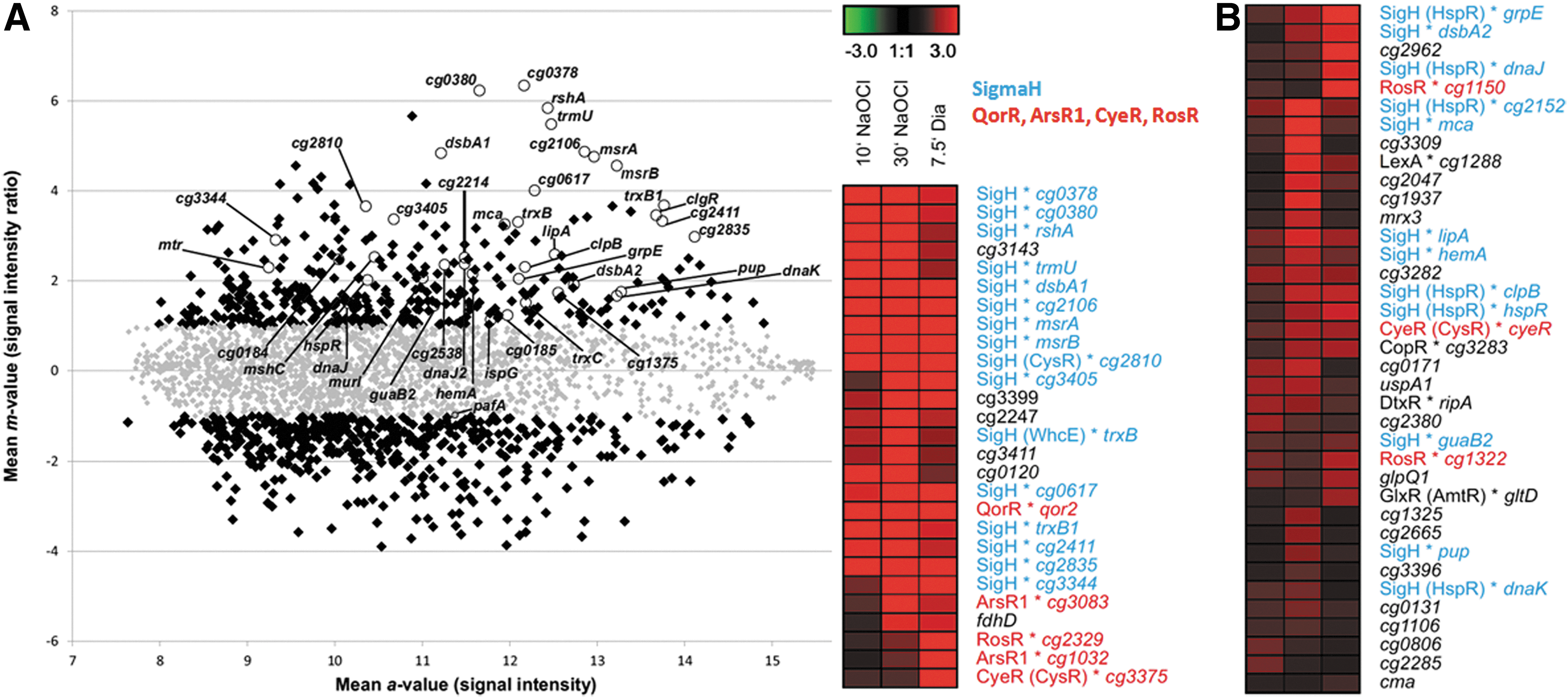
In addition, other redox-sensing regulators respond to NaOCl stress. The MarR/DUF24-family regulator QorR is a redox sensor for disulfide stress (10) and controls the quinone reductase-encoding qorA gene that was 10-fold induced by NaOCl stress (Fig. 1, Supplementary Tables S1 and S2). The ArsR-family transcription factors CyeR, ArsR1 and ArsR2 are three- to fivefold induced by NaOCl stress that control metal ion efflux pumps, arsenate reductases and NADPH-dependent flavin oxidoreductases. ArsR1 (Cg3082) controls cg3083, encoding a putative secondary Co2+/Zn2+/Cd2+ efflux transporter of the cation diffusion facilitator family that was 10-fold induced by NaOCl. ArsR2 regulates an arsenate reductase (arsX or cg0319) that reduces arsenate to arsenite and an arsenite efflux pump (arsC1 or cg0318) (60) that are 2-fold upregulated by NaOCl. RosR is a thiol-based peroxide-sensing MarR-family repressor (4) and the RosR-controlled genes for flavin oxidoreductases cg1150 and cg2329 are 2-fold induced by NaOCl. Furthermore, the hmp gene is 19-fold upregulated by NaOCl stress, which encodes a flavohemoglobin for nitric oxide detoxification. The hmp gene is under control of the NsrR regulator in E. coli and Streptomyces that senses nitrosative stress by an FeS cluster, but transcriptional regulation of hmp is unknown in C. glutamicum (59). It is further interesting to note, that no typical oxidative stress defense genes were induced by NaOCl stress, such as genes for the catalase (kat) or alkyl hydroperoxide reductase ahpC that are controlled by OxyR in the related C. diphtheriae (25).
Thiol-redox proteomics identifies 30 NaOCl-sensitive proteins in C. glutamicum
Next, we used our fluorescent-label redox proteomics approach to quantify the level of reversible thiol oxidation versus the protein amounts for 45 proteins after NaOCl stress in the C. glutamicum wild type and in the mshC mutant (Figs. 2 and 3, Supplementary Table S4). The wild type and mshC mutant were chosen for the comparison of the redox proteomes to identify possible S-mycothiolated proteins in the wild type. For redox proteomics, untreated and NaOCl-exposed cells were harvested in urea/thiourea buffer in the presence of 100 mM iodoacetamide (IAM) to alkylate all reduced thiols. Proteins with reversible thiol-oxidations, including S-thiolations were reduced with Tris-(2-carboxyethyl)-phosphine (TCEP) and labeled with the fluorescent dye BODIPY FL C1-IA. The fluorescent image was overlaid with the Coomassie-stained protein amount image and the fluorescence/protein amount ratios were quantified using the Decodon software as level of reversible thiol oxidations. Proteins with basal level oxidation include the Prx Bcp (3–5-fold redox ratios) and the thiol peroxidase (Tpx) (7–10-fold redox ratios) that form intramolecular disulfides during their peroxide detoxification cycle (36). Further conserved redox-sensitive proteins with basal oxidations include the dihydrolipoyl dehydrogenase Lpd (PdhD homolog) and the glyceraldehyde-3-phosphate dehydrogenase GapA in C. glutamicum that are also oxidized to intramolecular disulfides (6).
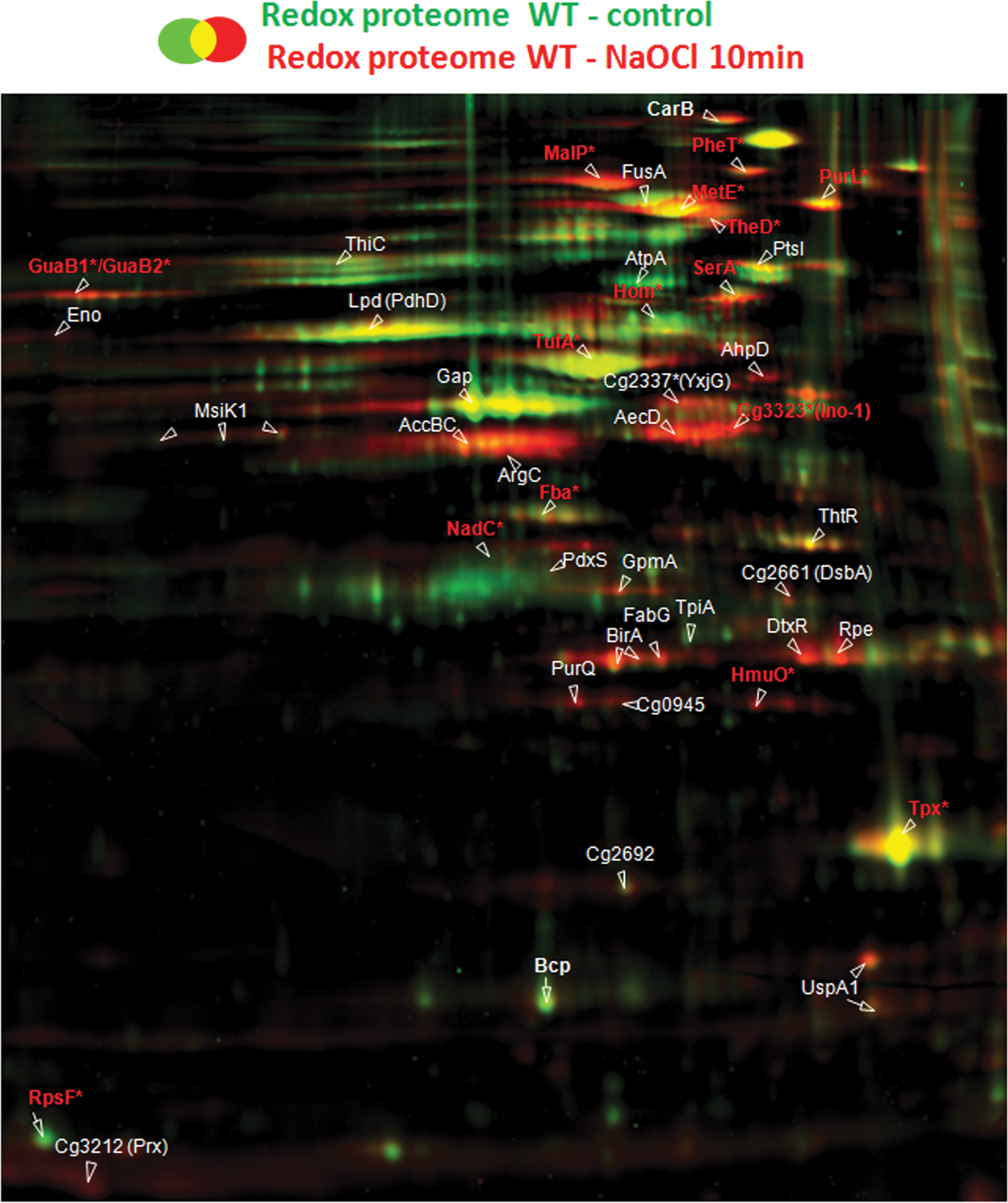
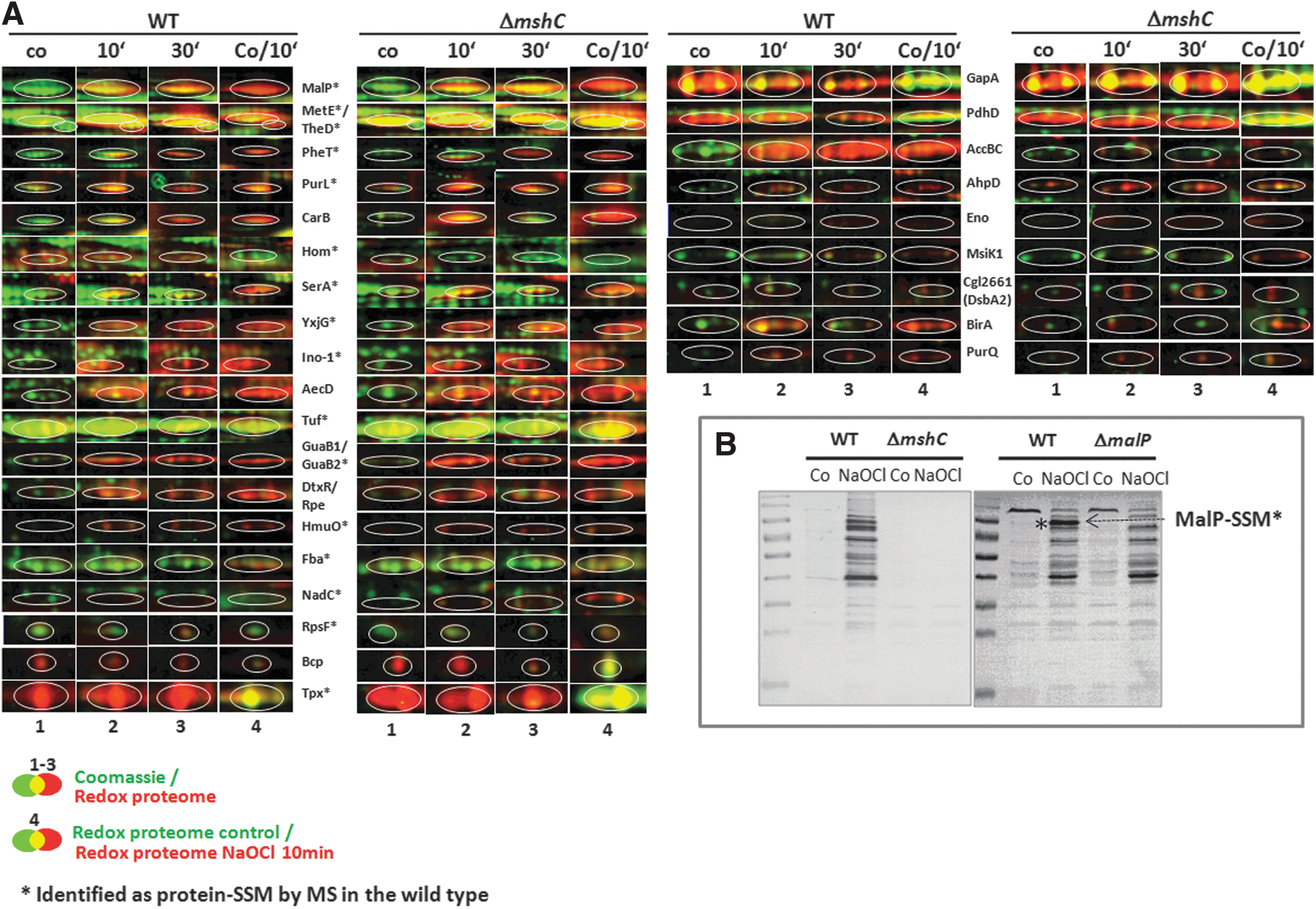
Treatment of C. glutamicum with 180 μM NaOCl stress resulted in strongly enhanced reversible thiol-oxidation of 30 NaOCl-sensitive proteins (Figs. 2 and 3; Supplementary Table S4). The search for conserved Cys residues using the CDD database (
Shotgun-LC-MS/MS identifies 25 proteins with S-mycothiolations in NaOCl-treated cells
Proteins that form MSH mixed disulfides after NaOCl stress were identified using LTQ-Orbitrap liquid chromatography–tandem mass spectrometry (LC-MS/MS) analysis. The specific S-mycothiolated Cys peptides were identified by searching the LC-MS/MS results for the 484 Da mass shift of MSH at Cys residues. We further observed that the precursor ions have lost myo-inositol during MS/MS fragmentation. These neutral loss precursor ions with a mass loss 180 Da were used as diagnostic abundant peaks for verification of S-mycothiolated peptides (Supplementary Fig. S5). In control cells, we identified two peroxidases, the Tpx S-mycothiolated at the active site Cys60 and the resolving Cys94 and the glutathione peroxidase (Gpx) S-mycothiolated at the active site Cys36. Interestingly, the spectral counts (quantitative value) of these Cys-SSM peptides of Tpx and Gpx increased strongly in NaOCl-treated cells. In addition, the Prx Bcp was identified with an Cys89-SS-Cys94 intramolecular disulfide peptide and the dihydrolipoyl dehydrogenase Lpd was oxidized to the conserved Cys42-SS-Cys47 disulfide in control cells (Supplementary Fig. S6I, J).
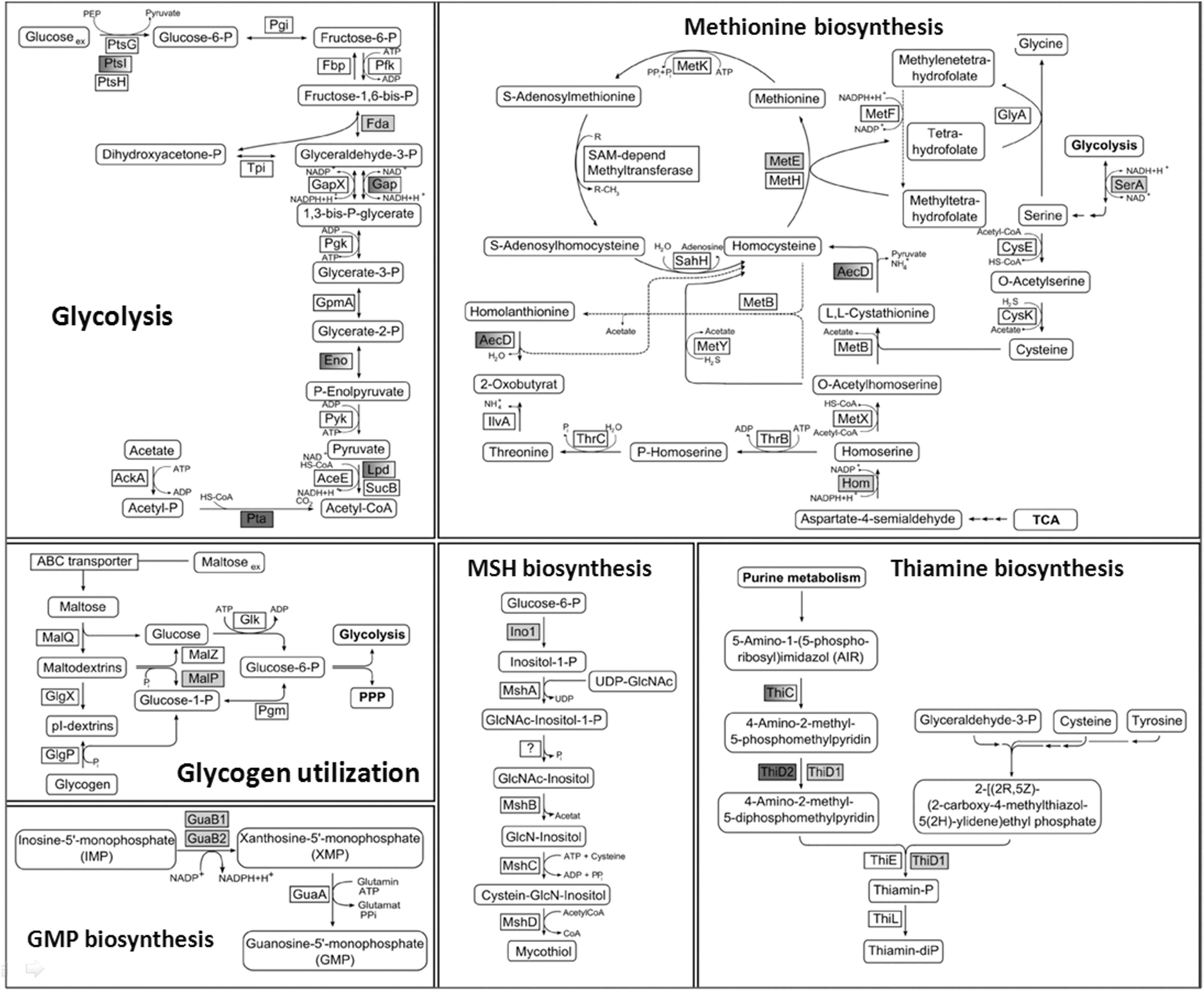
In NaOCl-treated cells, we identified 25 S-mycothiolated proteins in C. glutamicum with characteristic myo-inositol-loss precursor ions in their fragment ion spectra (Table 1; Supplementary Tables S5, Supplementary Fig. S5). These 25 S-mycothiolated proteins include 16 proteins identified also in the redox proteome as NaOCl-sensitive proteins, including Tuf, GuaB1, GuaB2, SerA, and MetE as conserved targets for S-thiolations across Gram-positive bacteria (7). The metabolic pathways, including the functions of the main S-mycothiolated proteins are shown in Figure 4. The S-mycothiolated proteins are involved in the metabolism of carbohydrates, for example, glycolysis (Fba, Pta, XylB), glycogen and maltodextrin degradation (MalP); amino acid biosynthesis pathways for serine, Cys, methionine (MetE, SerA, Hom); nucleotide and thiamine cofactor biosynthesis (GuaB1, GuaB, PurL, NadC, ThiD1 und ThiD2); redox and antioxidant functions for peroxide detoxification (Tpx, Gpx), methionine sulfoxide reduction (MsrA), heme degradation for iron mobilization (HmuO), and protein translation, such as ribosomal proteins (RpsF, RpsC, RpsM, RplM) and translation elongation factors (Tuf). The S-mycothiolated proteins MetE and MalP represent also very abundant proteins with strongly increased oxidation ratios after NaOCl stress in the redox proteome. The comparison of the S-mycothiolation pattern between wild type and malP mutant cells by nonreducing MSH-Western blots confirmed that MalP is one of the most strongly S-mycothiolated proteins in NaOCl-treated cells (Fig. 3B). Interestingly, also the first enzyme of the MSH biosynthesis pathways, the myo-Inositol-1-phosphate synthase (Ino-1 or Cg3323) was S-mycothiolated at Cys79 suggesting perhaps regulation of MSH biosynthesis under NaOCl stress.
Cytoplasmic protein extracts of Corynebacterium glutamicum ATCC13032 wild type and the isogenic mshC mutant were harvested at 15 min after NaOCl stress in urea-IAM-buffer, tryptic in-gel-digested and the peptides were analyzed using a LTQ Orbitrap-Velos™ mass spectrometer as described in the Methods section. 25 Proteins with S-mycothiolations were identified by the additional mass of 484 Da at Cys residues and with the diagnostic neutral myo-inositol loss precursor ions that appeared as abundant ions in the MS/MS spectra of each mycothiolated peptide. Peptides with S-cysteinylations were identified with a mass difference of 119 Da at Cys residues. The table includes the protein names, Cg-accession numbers and protein functions as derived from the UniprotKB database (
16 proteins have been identified as NaOCl-sensitive proteins in the Thiol-redox proteome (Figs. 2 and 3) with increased redox/protein ratios after NaOCl stress in this study. The redox/protein amount ratios are given in Supplementary Table S5.
The S-mycothiolated proteins MetE, GuaB and Tuf are also conserved targets for S-bacillithiolation in Bacillus or Staphylococcus species according to our previous study.
The S-mycothiolated peroxiredoxins Tpx and Gpx were identified in untreated and NaOCl-treated cells. All other S-mycothiolated proteins were only identified in NaOCl-exposed cells.
Cys, cysteine; IAM, iodoacetamide; LC-MS/MS, liquid chromatography–tandem mass spectrometry; LTQ, linear trap quadrupole; MW, molecular weight; ROS, reactive oxygen species.
We were interested if proteins are S-thiolated by alternative redox buffers, such as ergothioneine (EGT) or Cys in C. glutamicum in the mshC mutant. In previous studies, we identified few proteins that are S-cysteinylated in the absence of BSH in B. subtilis, but the bshA mutant was more sensitive to NaOCl stress indicating that S-cysteinylation cannot fully compensate for S-bacillithiolation (6, 7). In Mycobacterium smegmatis, the histidine-derived thiol EGT has been suggested to compensate for the loss of MSH in mshA mutants (56). However, we were unable to detect proteins that form mixed disulfides with EGT in the mshC mutant. Instead, we identified eight S-cysteinylated proteins in the mshC mutant, which overlapped with only six proteins detected as S-mycothiolated in the wild type (MalP, MetE, Ino-1, PurL, NadC, and RpsM) (Table 1; Supplementary Table S5, Supplementary Fig. S6). The S-cysteinylated Cys-peptides identified in the mshC mutant were identical with the S-mycothiolated Cys-peptides of the wild type. However, compared to S-mycothiolated peptides the number of spectra (spectral counts) was significantly lower for S-cysteinylated petides and even for 19 S-mycothiolated proteins of the wild type no S-cysteinylated peptides were detected in the mshC mutant (Table 1; Supplementary Table S5B). These results indicate that Cys can be used for alternative S-thiolations in the absence of MSH, but S-cysteinylation cannot fully compensate for S-mycothiolation.
The roles of mycothiol, σH and MalP in protection against disulfide stress
Next, we analyzed tpx, gpx, malP, mshC, and sigH mutants for their sensitivity to NaOCl stress in growth phenotype assays. Previously, the sigH and mshC mutants were shown to have decreased oxidative stress resistance in C. glutamicum (27, 35). In our growth phenotype assays, the mshC and sigH mutants are also very sensitive to NaOCl stress confirming previous results (Fig. 5).
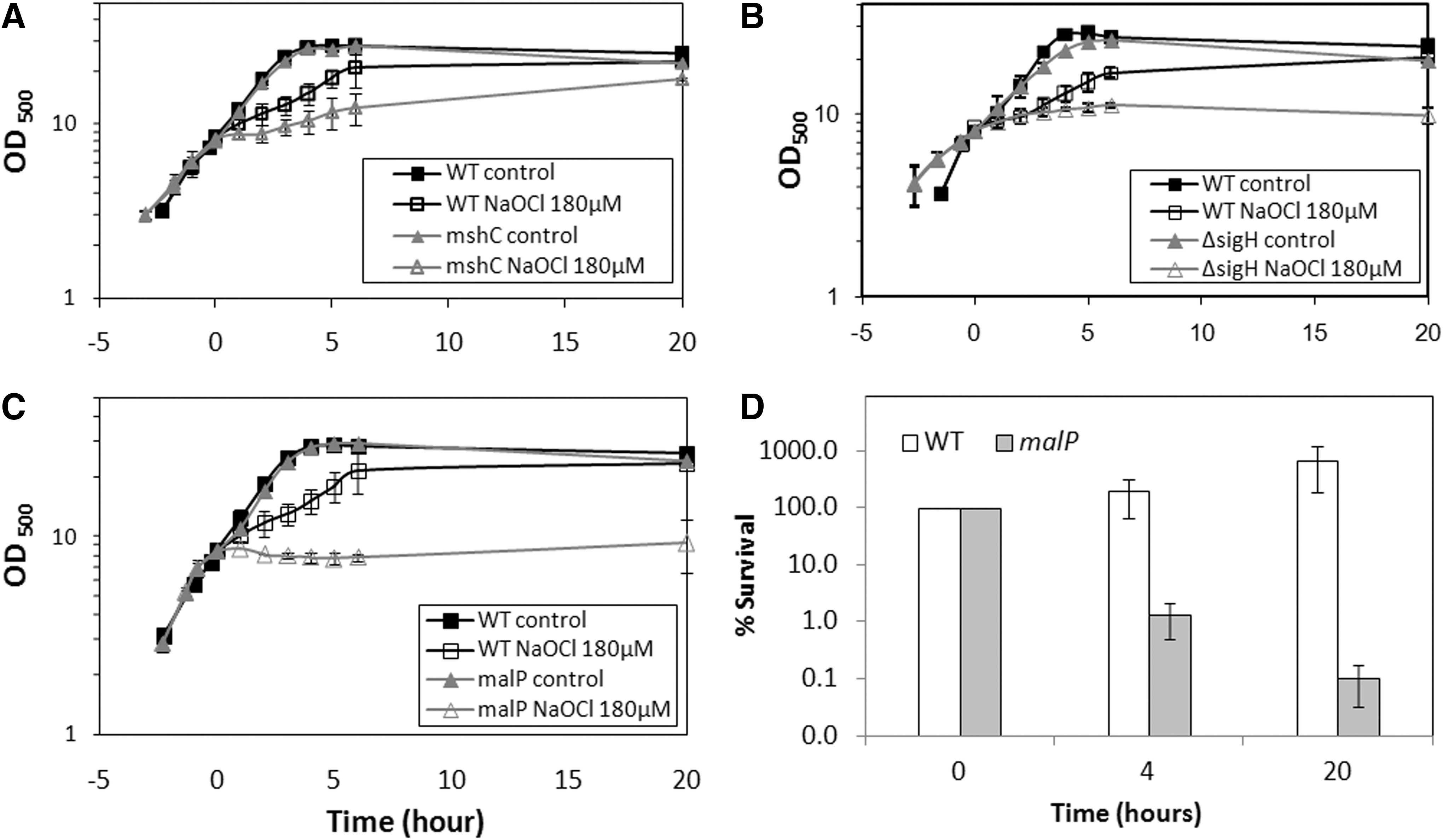
Growth phenotype assays further revealed an increased sensitivity of the malP mutant to NaOCl-stress indicating that S-mycothiolation of MalP confers protection against NaOCl stress (Fig. 5). The sensitivity of the malP mutant to NaOCl stress is caused by an increased killing effect of NaOCl stress as determined in survival assays. No NaOCl-sensitive growth phenotypes were detected for tpx and gpx mutants (data not shown). In conclusion, the σH/RshA disulfide stress system, MSH and MalP confer protection against NaOCl stress in C. glutamicum.
NaOCl stress leads to reduced glycogen degradation which could involve S-mycothiolation of MalP in vivo
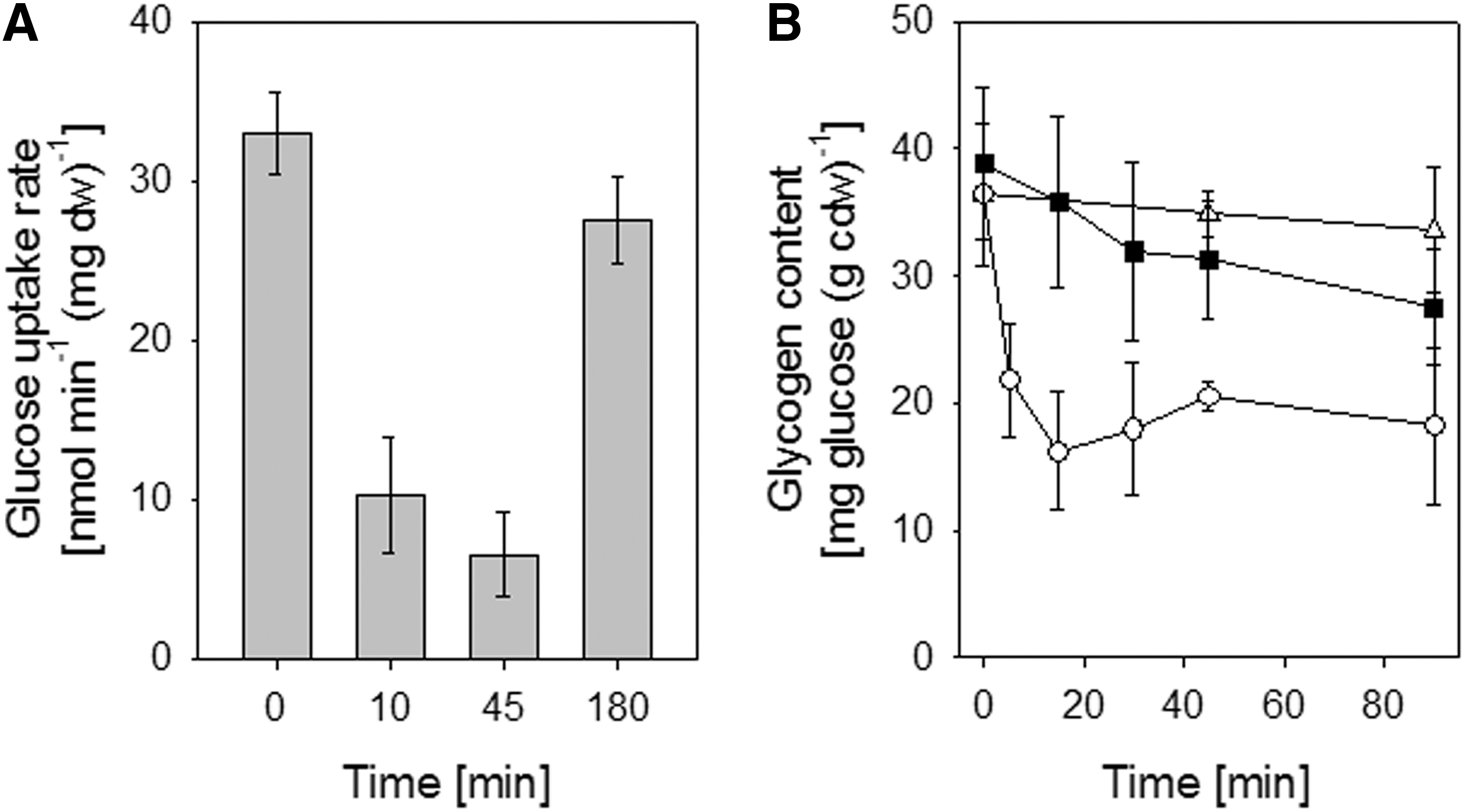
In C. glutamicum, glycogen transiently accumulates during the exponential growth. Glycogen is degraded by MalP either slowly before the onset of the stationary growth phase or rapidly after osmostress when glucose uptake is reduced (52). It could be possible that S-mycothiolation of MalP prevents glycogen degradation to save the energy source under oxidative stress. To test an effect of S-mycothiolation of MalP on glycogen degradation in vivo, the glycogen content and glucose uptake rates (GURs) were determined after NaOCl stress. The GUR decreased about threefold 10 min after addition of NaOCl (Fig. 6A). For comparison, osmostress caused by NaCl leads to a twofold decrease of the GUR to 15.6±0.7 nmol/min/mg dw (52). The glycogen content was quickly decreased about twofold within 15 min after osmostress (Fig. 6B). However, the glycogen level of NaOCl-treated cells was only slightly reduced within 45 min of stress exposure. Since the glycogen content remains stable in NaOCl-treated cells, despite the drastically reduced glucose uptake, glycogen degradation could be inhibited by S-mycothiolation of MalP. Future studies are underway to determine the effect of S-mycothiolation on MalP activity in vitro.
S-mycothiolation of Tpx in vitro abolishes its peroxidase activity which can be restored by the Mrx1/MSH/Mtr electron pathway
Next, we were interested to see whether the identified S-mycothiolated proteins are substrates for Mrx1. However, redox proteome comparison between the wild type and mrx1 mutants did not show a difference in the oxidation ratios of NaOCl-sensitive S-mycothiolated proteins (data not shown). Thus, we tested if S-mycothiolation affects the peroxidase activity of Tpx in vitro and if Mrx1 is able to reactivate Tpx. Tpx was expressed and purified from E. coli and the peroxidase activity of Tpx was analyzed in a coupled enzyme assay composed of Trx, TrxR, and NADPH. Purified Tpx has peroxidase activity when coupled to the Trx electron pathway, which is abolished when Tpx is omitted in this reaction (Fig. 7A, B). To test the effect of S-mycothiolation on the peroxidase activity, reduced Tpx was preincubated with reduced MSH before the addition of 1 mM hydrogen peroxide (H2O2). LC-MS/MS analysis revealed strong S-mycothiolation of all three Cys residues of Tpx, the peroxidatic Cys60, the resolving Cys94 and also the nonconserved Cys81 (Fig. 7, Supplementary Table S6). S-mycothiolation of Cys81 in vitro (but not in vivo) might be due to the much higher MSH amount applied in the in vitro reaction for purified Tpx.
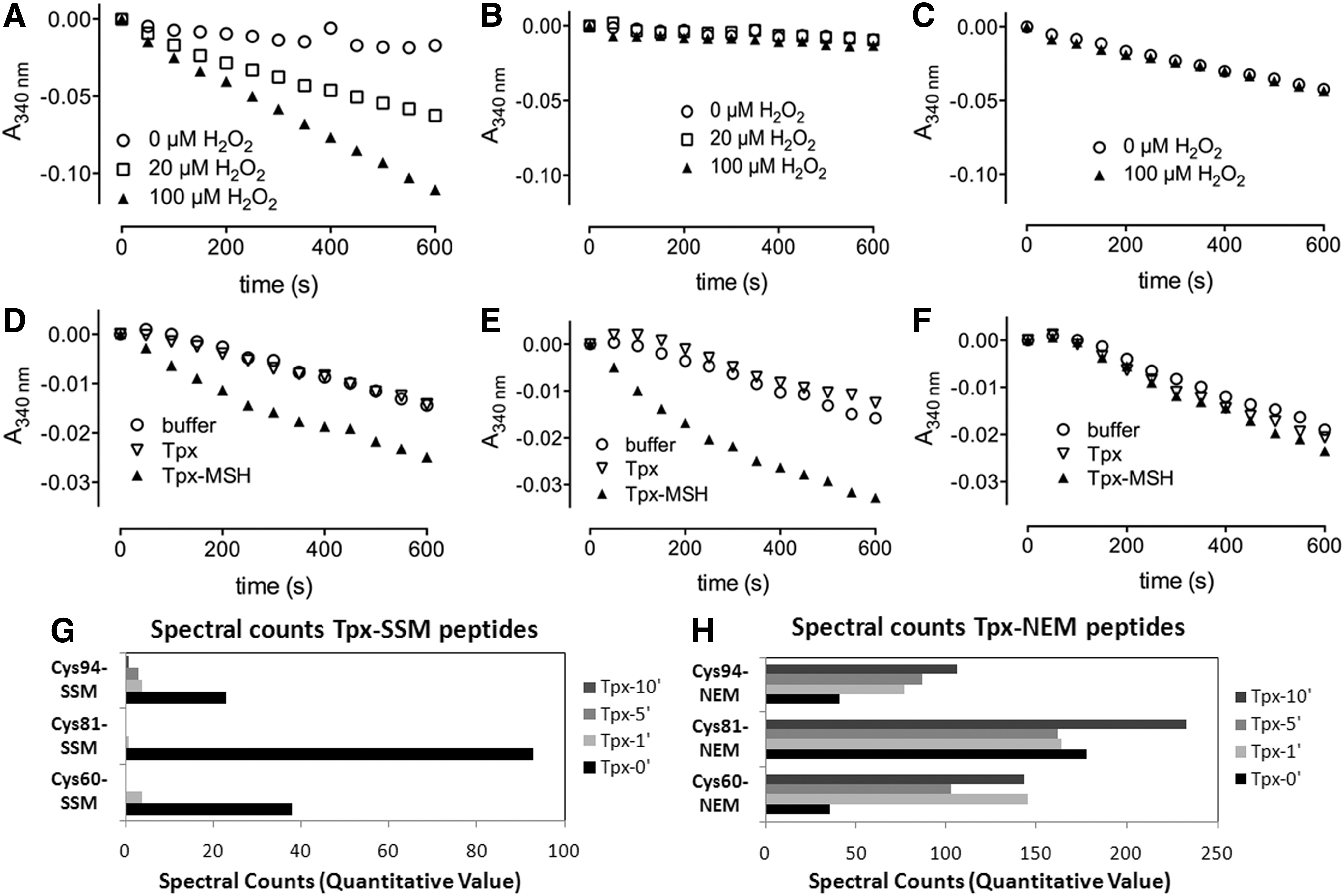
The S-mycothiolation of Tpx most likely protects the active site from overoxidation to sulfinic and sulfonic acids during NaOCl stress, and might change its peroxidase activity. To test this, S-mycothiolated Tpx was used in this Trx/TrxR-coupled assay and no peroxidase activity was measured confirming that S-mycothiolation of the active site Cys60 inhibits the peroxidase activity (Fig. 7C) (48). To reactivate S-mycothiolated Tpx, we reconstructed the Mrx1/MSH/Mtr electron transfer pathway in vitro with S-mycothiolated Tpx as substrate. Control samples without Tpx or with nonmycothiolated Tpx were included in the assay. Only the sample with 50 μM Tpx-SSM showed consumption of NADPH and the rate of NADPH consumption increased when 75 μM Tpx-SSM was applied in this assay (Fig. 7D–F). No reduction of Tpx-SSM was observed when Mrx1 was omitted, indicating that the high MSH concentration (400 μM) is not sufficient to reduce Tpx-SSM without Mrx1. MS confirmed that the S-mycothiolated Cys60, Cys81, and Cys94 peptides are completely reduced within 5–10 min by Mrx1 (Fig. 7G, H, Supplementary Tables S6A, B). During Tpx-SSM reduction, Mrx1 gets S-mycothiolated at its active site Cys12 as transient Mrx1-SSM intermediate that was also detected by MS (Supplementary Table S6D). However, most Mrx1 protein was oxidized to the Cys12-SS-Cys15 intramolecular disulfide during Tpx-SSM reduction (Supplementary Table S6E).
Discussion
Mycothiol is the major LMW thiol in Actinomycetes and functions in detoxification of oxidants, electrophiles, toxins and several antibiotics (13, 39). In C. glutamicum, MSH plays very broad and essential roles in the detoxification of alkylating agents, toxins (methylglyoxal), oxidants (H2O2, diamide), antibiotics, glyphosate, ethanol, and toxic heavy metals (Zn, Co, Mn, Cr) (35). MSH is also an essential cofactor for the maleylpuruvate isomerase in the degradation of aromatic compounds, such as gentisate and 3-hydroxybenzoate and required also for detoxification of naphthalene and resorcinol (14, 35). Mutants in the MSH biosynthesis pathways were unable to grow in the presence of gentisate and 3-hydroxybenzoate, which indicates that MSH cannot be replaced by Cys as thiol-cofactor. In addition, MSH is required for conjugation of arsenate to a MSH-S-conjugate that is detoxified by two MSH-dependent arsenate reductases ArsC1 and ArsC2 in C. glutamicum (43).
In this article, we present for the first time evidence that MSH also functions in widespread protein S-mycothiolation as important thiol-protection mechanism under hypochlorite stress in C. glutamicum. In addition, we provide evidence that S-mycothiolation inhibits the peroxidase activity of Tpx in vitro which can be restored by Mrx1. Protein S-thiolation was induced in our previous study in B. subtilis by NaOCl stress, which causes a disulfide stress response and induction of the Spx, PerR and OhrR-regulons in the transcriptome (6). Here we confirmed that NaOCl stress elicits a disulfide stress response also in C. glutamicum that is controlled by the σH/RshA regulatory system (3). In addition, similar thiol-based redox regulators respond to NaOCl stress in both bacteria, belonging to the MarR/DUF24-family that control quinone oxidoreductases or to the ArsR-family regulating metal ion or arsenate efflux systems (6). The results that NaOCl stress causes a common disulfide stress response in bacteria are in agreement with the chemistry of NaOCl. Hypochlorite is defined as reactive chlorine species that leads to chlorination of thiol groups to form sulfenyl chloride (Cys-SCl) as unstable intermediate (20, 21). Cys-SCl reacts further with proximal thiols to form disulfides or is overoxidized to Cys sulfonic acid. Thus, S-mycothiolation is particularly important under NaOCl stress to prevent lethal overoxidation of Cys residues.
Using shotgun-LC-MS/MS analysis, we identified 25 cytoplasmic proteins that are S-mycothiolated under NaOCl stress conditions in the wild type and 16 of these are also major NaOCl-sensitive proteins in the thiol-redox proteome. In contrast, only 6 of these S-mycothiolated proteins were identified as S-cysteinylated in the absence of MSH with significantly lower spectral counts (Supplementary Table S5B). This indicates that S-cysteinylation can only partly compensate for alternative S-thiolations in the mshC mutant. However, due to the instability of S-thiolation modifications we cannot exclude that other redox buffers, like CoASH or EGT are also involved in S-thiolations in the absence of MSH. However, phenotype analyses of the mshC mutant support clearly that MSH is important for protection under NaOCl stress. The number of S-mycothiolated proteins is higher in C. glutamicum compared to S-bacillithiolated proteins identified in Bacillus or Staphylococcus in our previous work (7). This may be related to the higher MSH amounts present in Actinomycetes compared to BSH in Firmicutes (38). The proteins identified in the S-mycothiolome are functionally very similar to the identified S-bacillithiolated proteins in Firmicutes and S-glutathionylated proteins identified in eukaryotes (7, 33). Except for the thiol peroxidases Tpx and Gpx, all S-mycothiolated proteins were exclusively identified under NaOCl stress conditions, not in untreated cells, which confirms our previous results in Firmicutes bacteria. Thus, S-mycothiolation functions as redox switch and protects redox-sensitive thiols against overoxidation mainly under oxidative stress conditions.
The S-mycothiolated proteins were identified within the metabolic pathways for methionine biosynthesis (MetE, SerA, Hom), glycolysis (Fba, Pta, PckA), glycogen utilization (MalP), nucleotide biosynthesis (GuaB1, GuaB2, PurL, NadC), MSH biosynthesis (Ino1) and thiamine biosynthesis (ThiD1, ThiD2). Further S-mycothiolated proteins function in protein translation (Tuf, PheT, RpsC, RpsF, RpsM, RplM) or have antioxidant functions, such as the peroxidases Tpx and Gpx. In our previous study, we identified 8 conserved S-bacillithiolated proteins in B. subtilis and other Firmicutes bacteria, including the methionine synthase MetE, the inosine 5′-phosphate (IMP) dehydrogenase GuaB, the translation elongation factor Tuf, the inorganic pyrophosphatase PpaC, the phosphoglycerate dehydrogenase SerA, the chorismate mutase AroA, the ferredoxin-NADP+ reductase YumC and the putative bacilliredoxin YphP (6, 7). Interestingly, five of these conserved S-bacillithiolated proteins are also abundantly S-mycothiolated proteins in C. glutamicum, including MetE, GuaB1, GuaB2, Tuf and SerA. Thus, we hypothesize that these proteins are probably conserved targets for S-thiolations across bacteria. GuaB and MetE possess also conserved active site Cys residues, including the active site Zn center of MetE (Cys713 in Cg-MetE) or the thioimidate intermediate-forming active site in GuaB (Cys302 in Cg-GuaB1 and Cys317 in Cg-GuaB2). This active site of GuaB is buried in the GuaB structure from Vibrio cholerae (PDB code 4FEZ) (Supplementary Fig. S7). S-glutathionylation of GuaB has been also reported in human T-lymphocytes and endothelial cells (18, 32). It is further interesting that different enzymes of the thiamine biosynthesis pathway are S-thiolated in Firmicutes and C. glutamicum. For example, ThiG and ThiM were S-bacillithiolated in Bacillus and Staphylococcus (7), whereas the multifunctional ThiD1 protein with thiamine phosphate synthase and hydroxymethylpyrimidine (HMP) kinase domains was S-mycothiolated in C. glutamicum at Cys451 that is the conserved HMP substrate binding site.
MetE has been first shown as target for S-glutathionylation in E. coli where it is modified at the nonconserved Cys645 located at the entrance to the active site (22). The methionine synthase MetE is the major S-bacillithiolated protein in Bacillus species (6, 7) and its S-thiolation occurs in its active site Zn center at Cys730 and also at the nonconserved Cys719 in B. subtilis. This leads to MetE inactivation and Met auxotrophy in NaOCl-treated cells. Here we confirmed that MetE is also a major target for S-mycothiolation in C. glutamicum and is oxidized at the conserved active site Cys713. This Zn-binding active site Cys is buried in the MetE structures of Thermotoga maritima and Streptococcus mutans (PDB code 2NQ5) (Supplementary Fig. S7), but the elastic and flexible nature of this catalytic Zn center has been demonstrated (28). However, in contrast to E. coli and B. subtilis, NaOCl stress did not cause methionine starvation since growth was not resumed by addition of Met to the growth medium after 30 and 60 min of NaOCl treatment (Supplementary Fig. S1B). Thus, the growth defect after NaOCl stress could be caused by multiple amino acid auxotrophies or by a general translation defect caused by S-mycothiolation of translation elongation factors and ribosomal proteins. Similar translation proteins have also been detected as reversibly oxidized in the redox proteome of E.coli after NaOCl stress (31), and as S-glutathionylated in oxidatively stressed human endothelial and yeast cells (18, 33, 54). The inhibition of protein synthesis during the time of oxidant removal is perhaps one of the conserved consequences of protein S-thiolation as verified in yeast cells (54).
Besides the identification of many conserved S-thiolated proteins across Firmicutes and Actinobacteria, there are also unique S-thiolated proteins in C. glutamicum. For example, the myo-inositol-1 phosphate synthase Ino-1, the first enzyme of the MSH biosynthesis pathway was S-mycothiolated at Cys79 suggesting that MSH biosynthesis itself might be feed-back redox-controlled by oxidative stress. Interestingly, the Ino-1 homolog of M. tuberculosis was identified as pupylated at K73, which determines its degradation by the proteasome (16), but K73 is not conserved in C. glutamicum Ino-1. It will be interesting to analyze the activity of S-mycothiolated Ino-1 and a possible cross-talk with the Pup proteasome in C. glutamicum in future studies. The main target for S-mycothiolation was the maltodextrin phosphorylase MalP in C. glutamicum that is involved in glycogen and maltodextrin utilization in C. glutamicum (53). MalP is the major reversibly oxidized protein in the thiol redox proteome of C. glutamicum and S-mycothiolated at the nonconserved Cys180, which is the only surface-exposed Cys residue in the MalP crystal structure of Corynebacterium callunae (PDB code 2C4M). In addition, a malP mutant was very sensitive to NaOCl stress suggesting that S-mycothiolation of MalP confers resistance to oxidative stress in C. glutamicum. C. glutamicum has the capability to accumulate transiently glycogen as energy and carbon source during the exponential growth when grown on glucose. Glycogen is degraded during the stationary phase or during osmostress by the glucan phosphorylase GlgP, the debranching enzyme GlgX that produces maltodextrins and the maltodextrin phosphorylase MalP (Fig. 4) (53). Thus, S-mycothiolation of MalP could protect and inhibit the enzyme and prevent glycogen degradation to save their energy source under oxidative stress. Indeed, the glycogen content decreased only slightly in NaOCl-treated cells despite the reduced GUR. This inhibition of glycogen degradation could be caused by a loss of MalP activity due to S-mycothiolation (Fig. 6). Further studies are underway to characterize the effect of S-mycothiolation on MalP activity. We further identified several glycolytic enzymes as targets for S-thiolation (e.g., Fba) or reversible thiol-oxidation (e.g., GapA, Eno) in C. glutamicum that were also reversibly inactivated by S-thiolation in yeast cells (54). It is proposed that the inhibition of the glycolytic flux by S-thiolation redirects glucose into the pentose phosphate pathway for NADPH regeneration to provide the reducing power for the Trx and Mrx1 systems (54). The inhibition of glycolytic enzymes by S-thiolation could be also a consequence of the reduced GUR under NaOCl stress.
As most interesting proteins, we identified the peroxidases Tpx and Gpx as S-mycothiolated at their active sites. Tpx and Gpx have been classified as atypical 2-Cys Prxs with a conserved peroxidatic active site Cys (CP) and a resolving Cys (CR). Atypical Prxs react with peroxides to a transient CP-SOH intermediate that forms an intramolecular disulfide with CR in the same subunit that is reduced by the Trx pathway during recycling (36, 47). Since MSH-deficient mutants of C. glutamicum and M. smegmatis showed peroxide-sensitive phenotypes, the existence of MSH-dependent peroxidases was suggested earlier (13, 35). It is possible that Gpx might be the MSH-dependent peroxidase that utilizes MSH as electron donor for peroxide detoxification resulting in MSSM generation that could be recycled by Mtr. For M. tuberculosis Tpx (Tpx-Mtb) the catalytic cycle and structure has been resolved (PDB code 1XVQ)(17, 48). Tpx was S-mycothiolated at Cys60 and Cys94 in vivo that both are solvent exposed in the Tpx-Mtb structure in contrast to Cys81 that is more buried and not conserved (Supplementary Fig. S7). Tpx-Mtb shares 56.6% sequence identity to C. glutamicum Tpx (Tpx-Cg) and the Cys60-Cys94 intramolecular disulfide was confirmed here using MS (Supplementary Table S6C). Tpx-Cg showed also NADPH-linked peroxidase activity and reduced H2O2 in a Trx/TrxR-coupled assay. For E. coli Tpx, it was shown that the CP-SOH was sensitive to overoxidation (2), which explains the protection of the Cys60 active site against overoxidation by S-mycothiolation in our study. We could show that S-mycothiolation of CP and CR inhibits the peroxidase activity which can be restored after reduction by the Mrx1/MSH/Mtr electron pathway. Thus, S-mycothiolation controls Tpx activity and protects CP against overoxidation. S-glutathionylated Prxs have been observed as reaction intermediates in the catalytic mechanism of peroxidase activity of 2-Cys Prx from Schistosoma mansoni, poplar Prx, and human Prx-I (5, 46). S-glutathionylation of Prx-I induced a structural change that leads to dissociation from oligomers to dimers with a loss of chaperone activity and an increased peroxidase activity of the active dimer (5, 44, 46). For 1-Cys D-Prx, it is been shown that S-glutathionylation induced the dissociation of noncovalent homodimers to monomers that are reduced by the Grx/GSH system followed by dimerization and reactivation of the peroxidase activity (42). Also Tpx is active as homodimer and it will be exciting to unravel the structural changes of Tpx by S-mycothiolation that perhaps also regulates the quaternary structure by dimer to monomer dissociation.
Experimental Procedures
Bacterial strains and growth conditions
Bacterial strains, plasmids, and primers are listed in Supplementary Tables S7 and S8. For DNA manipulation and plasmid isolation, E. coli was cultivated in Luria Bertani (LB) medium at 37°C. C. glutamicum ATCC 13032 wild type and sigH, mshC, malP, tpx and gpx mutant strains were grown in CGC minimal salt medium containing 1% glucose as carbon source at vigorous agitation at 30°C (12). When appropriate, nalidixic acid (50 μg/ml for C. glutamicum) or kanamycin (20 μg/ml for C. glutamicum and 50 μg/ml for E. coli) were added to the media. Sodium hypochlorite (15% stock solution) and H2O2 were purchased from Sigma Aldrich. For NaOCl stress experiments, C. glutamicum cells were grown in CGC minimal medium to an OD500 of 8.0 and exposed to 180 μM NaOCl that was freshly diluted in distilled water. The survival of the malP mutant was determined after different times of NaOCl stress by plating serial dilutions on LB plates and determining colony forming units (cfu) after 2 days. The cfu of the untreated culture was set as 100% and three replicate experiments were performed.
Construction of defined deletions in the C. glutamicum chromosome
Gibson assembly was applied to construct pK18mobsacB derivatives to perform allelic exchange of the gpx and tpx genes in the chromosome of C. glutamicum ATCC 13032 (50). The primers used to construct the pK18mobsacB derivatives are listed in Supplementary Table S8, that includes the gpx and tpx genes with internal deletions (Δgpx [480 bp] and Δtpx [447 bp]). The pK18mobsacB derivatives were cloned in E. coli JM109 (Supplementary Table S7). The pDN2 plasmid containing the ΔsigH deletion was constructed previously (3) and transformed into C. glutamicum ATCC 13032. The gene replacement in the chromosome of C. glutamicum ATCC 13032 resulted in the ΔsigH, Δgpx, and Δtpx deletion mutants that were confirmed by PCR using the primers in Supplementary Table S8.
RNA isolation and DNA microarray hybridization
For RNA isolation, C. glutamicum ATCC 13032 cells were harvested before and 10 and 30 min after exposure to NaOCl stress from three replicate experiments and disrupted with a Precellys 24 homogenizer (Bertin Technologies). Total RNA was isolated using the RNeasy Mini Kit and treated with DNase-I as described (23). The hybridization of whole-genome oligonucleotide microarrays was performed using 15 μg of total RNA that was used for cDNA synthesis as described previously (49). The normalization and evaluation of the hybridization data was done with the software package EMMA 2 (9) using a signal intensity (a-value) cut-off of ≥7.0, a signal intensity ratio (m-value) cut-off of±1 (technical variance) for hierarchical clustering and±1 (significance threshold) for the single analysis. Genes with at least twofold expression changes and p-values<0.05 were considered as significantly changed genes.
Hierarchical clustering analysis
Clustering of gene expression profiles was performed using GENESIS (55). The transcriptome data sets included log2-fold expression changes of all genes with at least twofold expression changes (p-values<0.05) at 7.5 min after 2 mM diamide stress (Busche and Kalinowski, unpublished data), 10 and 30 min after 180 μM NaOCl, respectively.
Thiol redox proteomics and MALDI-TOF-TOF MS/MS
The thiol redox proteome analysis was performed as described (6). Cells were harvested at control and NaOCl stress conditions, sonicated, alkylated in 8 M urea/1% chaps/100 mM IAM, and acetone precipitated. The pellet was resolved in urea/chaps buffer without IAM and 200 μg of the protein extract was reduced with 10 mM TCEP and labelled with BODIPY FL C1-IA [N-(4,4-difluoro-5,7-dimethyl-4-bora-3a,4a-diaza-s-indacene-3-yl)-methyl)-iodoacetamide] (Invitrogen). The fluorescence-labeled protein extract was separated using two-dimensional (2D) polyacrylamide gel electrophoresis (PAGE), scanned for BODIPY fluorescence and stained with Coomassie as described (6). Quantitative image analysis was performed with the DECODON Delta 2D software (
LTQ-Orbitrap Velos MS
IAM-alkylated cell extracts were separated by 15% nonreducing sodium dodecylsulfate (SDS)-PAGE and tryptic in-gel digested as described (6). Tryptic peptides were subjected to a reversed phase column chromatography and MS and MS/MS data were acquired with the LTQ-Orbitrap-Velos mass spectrometer (Thermo Fisher Scientific) equipped with a nanoelectrospray ion source as described (6). Post-translational thiol-modifications of proteins were identified by searching all MS/MS spectra in “dta” format against a C. glutamicum ATCC13032 target-decoy protein sequence database extracted from UniprotKB release 12.7 (UniProt Consortium, Nucleic acids research 2007, 35, D193-197) using Sorcerer™-SEQUEST® (Sequest v. 2.7 rev. 11, Thermo Electron, including Scaffold 4.0; Proteome Software, Inc.). The Sequest search was carried out with the following parameters: parent ion mass tolerance 10 ppm, fragment ion mass tolerance 1.00 Da. Two tryptic miscleavages were allowed. Methionine oxidation (+15.994915 Da), Cys carbamidomethylation (+57.021464 Da), S-cysteinylations (+119.004099 Da for C3H7NO2S), S-mycothiolations (+484.13627 Da for MSH) and S-ergothionylations (+227.07284 for EGT) were set as variable post-translational modifications in the Sequest search. Sequest identifications required ΔCn scores of >0.10 and XCorr scores of >2.2, 3.3 and 3.75 for doubly, triply and quadruply charged peptides, respectively. Neutral loss precursor ions characteristic for the loss of inositol (−180 Da) served for verification of the S-mycothiolated peptides.
Nonreducing MSH specific Western blot analysis
Twenty-five microgram IAM-alkylated protein extracts were used for nonreducing 15% SDS-PAGE and MSH-specific Western blots as described (7). Polyclonal MSH antiserum was kindly obtained from Mamta Rawat and used at 1:500 dilution.
Analyses of the glycogen content and [14C]-glucose uptake studies
Analyses of the intracellular glycogen content were performed enzymatically as described (51). [14C]-glucose uptake studies were performed as described (34). After different times of stress exposure, the glucose uptake reaction was then started by addition 50 μM [14C]-glucose (specific activity 250 μCi μmol−1; Moravek Biochemicals). At given time intervals (15, 30, 45, 60, and 90 s), 200 μl samples were filtered through glass fibre filters (Typ F; Millipore) and washed twice with 2.5 ml of 100 mM LiCl. Radioactivity of the samples was determined using scintillation fluid (Rotiszinth; Roth) and a scintillation counter (LS 6500; Beckmann).
Cloning and purification of Tpx, Trx, TrxR, and Mrx1
Recombinant Trx, TrxR, and Mrx1 were cloned and purified as described by Ordóñez et al. (43). Tpx was cloned into plasmid pET28a (using primers tpx_pETfor and tpx_pETrev and the restriction sites: NcoI, XhoI). The resulting Tpx-pET28 vector, containing Tpx with a N-terminal HIS6-tag and a PreScission protease cleavage site (GE Healtcare) was transformed into E. coli BL21(DE3). The transformed cells were grown overnight at 37°C in LB medium with 50 μg/ml kanamycin. The cultures were 100-fold diluted into LB medium with 50 μg/ml kanamycin, induced with 1 mM IPTG at an OD600 of 0.7 and grown overnight at 30°C. Cells were harvested and resuspended in 20 mM Hepes (pH 7.5), 5 mM imidazole, 1 M NaCl, 1 mM DTT, 0.1 mg/ml 4-(2-aminoethyl) benzene sulfonyl fluoride hydrochloride and 1 μg/ml leupeptin to an OD600 of 200. After cell crack disruption, 50 μg/ml DNase I and 20 mM MgCl2 were added to cell lysate, left for 30 min at 4°C before centrifugation. The supernatant was loaded on a Ni2+-Sepharose (GE Healthcare) immobilized metal affinity chromatography (IMAC) column equilibrated with 20 mM Hepes (pH 7.5), 5 mM imidazole, 1 M NaCl, 1 mM DTT, and eluted with a linear gradient to 1 M imidazole in the same buffer. The eluted fractions were analyzed on SDS-PAGE (15% Tris-glycine) and the fractions with Tpx were pooled and further purified on a Superdex75PG (16/90) SEC column equilibrated in 20 mM Tris/HCl (pH 8.0), 250 mM NaCl, 1 mM DTT.
In vitro S-mycothiolation of Tpx
Tpx was incubated with a 6 M excess of reduced MSH before the addition of 1 mM H2O2. After an incubation of 5 min, the sample was loaded on an IMAC column and MSH and H2O2 were removed by a washing step with 50 mM HEPES pH 8.0, 500 mM NaCl. S-mycothiolated Tpx was eluted in the same buffer containing 300 mM imidazole.
Coupled Trx/TrxR and Mrx1/MSH/Mtr electron transfer assays
The Trx/TrxR and Mrx1/MSH/Mtr coupled enzyme assays described by Van Laer et al. were modified to include Tpx (57). Briefly, Tpx was reduced by 10 mM DTT for 15 min at room temperature and the excess of DTT was removed by size exclusion chromatography in 50 mM Hepes, pH 8.0, 500 mM NaCl. The Tpx/Trx/TrxR electron transfer pathway was reconstructed by incubating 500 μM NADPH, 3 μM TrxR, and 1500 nM Trx with 500 nM Tpx at 37°C in the assay buffer containing 50 mM Hepes pH 8.0, 500 mM NaCl. Increasing concentrations of H2O2 (20–100 μM) were added as substrate to the reaction mixture. The oxidation of NADPH due to H2O2 reduction was monitored as a decrease in the absorption at 340 nm in function of time. The Mrx1/MSH/Mtr assay contained 500 μM NADPH, 5 μM Mtr, 400 μM MSH, and 2 μM Mrx1. Mycothiolated Tpx (50 or 75 μM final concentration) was added and the absorption at 340 nm was monitored in function of time.
Purification and reduction of MSH
MSH was purified as described in Ordóñez et al. (43). Oxidized MSH was incubated with 25 mM TCEP and incubated for 20 min at room temperature. The sample was loaded on an anion exchange chromatography (Isolute PE-AX column) and MSH was eluted in water. As a quality control, a 1D proton NMR spectra in D2O was recorded and compared to the spectra obtained by Lee and Rosazza (30).
Footnotes
Acknowledgments
We thank Dirk Albrecht for the MALDI-TOF-MS analysis; Dana Clausen and Eva Glees for excellent technical assistance and Eva Schulte-Bernd for performing the microarray hybridizations. We wish to thank Mamta Rawat for providing polyclonal MSH antibodies and Johanna Milse for the mshC mutant. G.M.S. and L.C. thank Reinhard Krämer (Cologne) for continuous support. The support of the BMBF (grant 0315589F “FlexFit”) is gratefully acknowledged. This work was supported by a grant from the Deutsche Forschungsgemeinschaft (AN746/3-1) to H.A.
K.V.L. and J.M. thanks the agentschap voor innovatie door Wetenschap en technologie (IWT), Vlaams Instituut voor Biotechnologie (VIB), and the HOA project of the Vrije Universiteit Brussel (VUB) for their support. J.M. is group leader for Redox Biology of the VIB. We acknowledge the BMBS COST action BM1203 (EU-ROS).
Author Disclosure Statement
No competing financial interests exist.
Abbreviations Used
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
