Abstract
Introduction
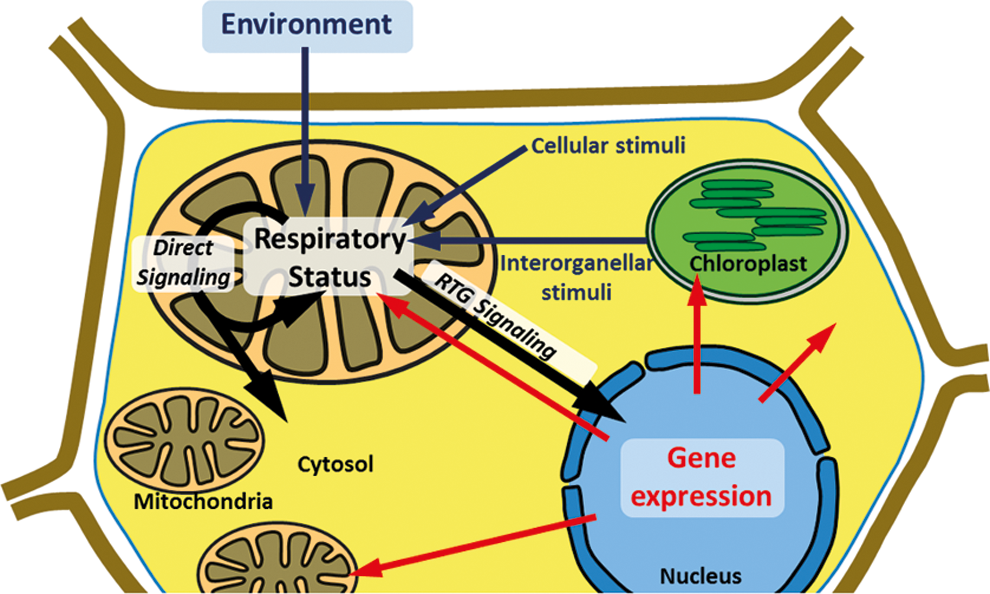
How Mitochondrial Physiology Can Create Signals: Candidates and Mechanisms
Respiratory metabolism is a dominating feature of plant mitochondria. Primary mitochondrial signals are therefore highly likely to be integral part of, or closely associated with, respiration. It is a general assumption that changes in respiratory status trigger signal transduction. Indeed, alterations in transcript abundances have been observed when respiration is compromised or modified (25, 32, 46, 62, 130, 135, 195). Changes in respiratory status impact on various associated mitochondrial parameters and processes, such as production of reactive oxygen and nitrogen species (ROS/RNS), redox status of NAD(P)H and antioxidant pools, ATP/ADP ratio, the proton-motive force, and total metabolite levels, all of which require consideration as potential stimuli to trigger mitochondrial signaling (Fig. 2).
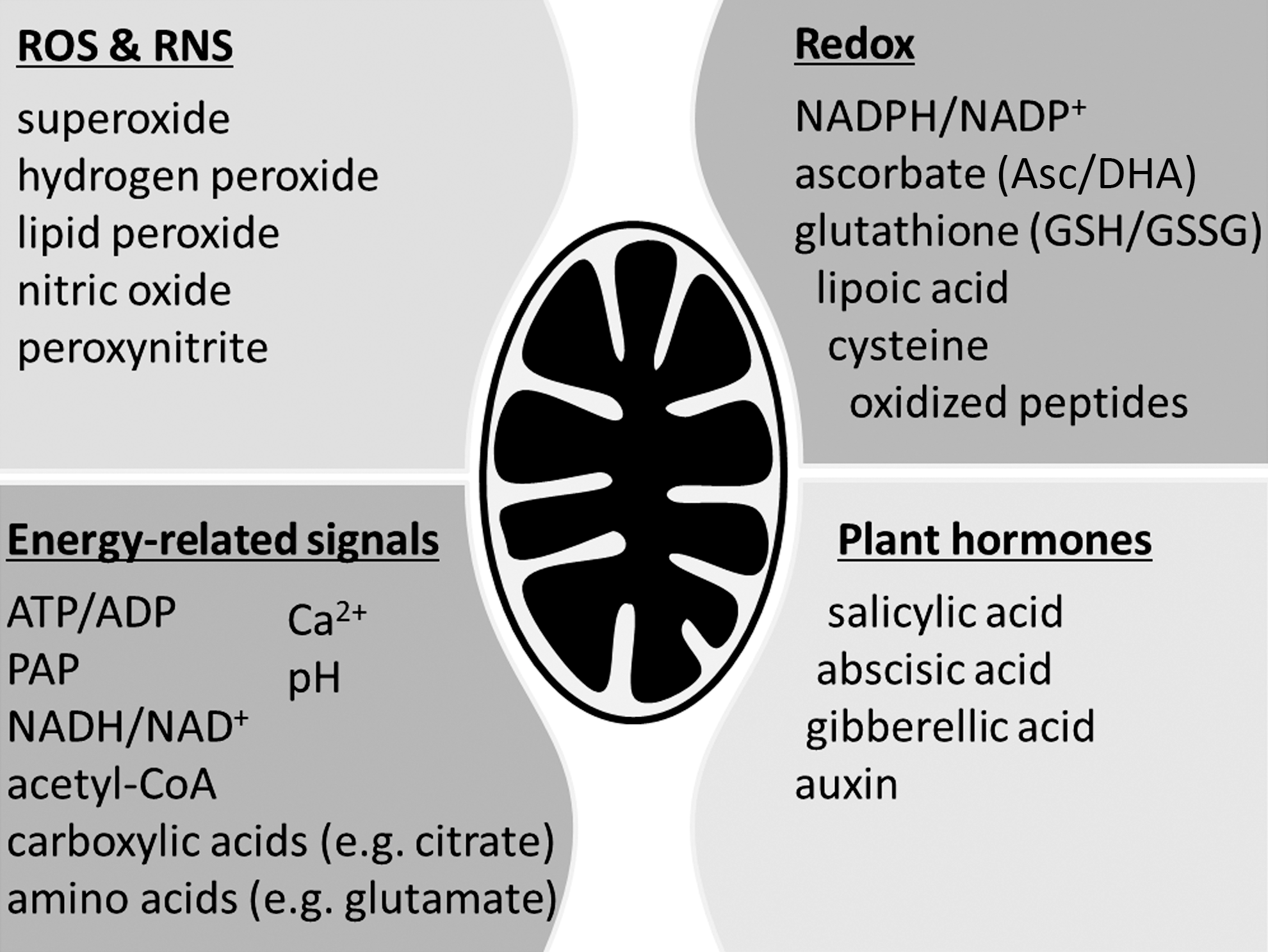
Signals derived from mitochondrial ROS and redox systems
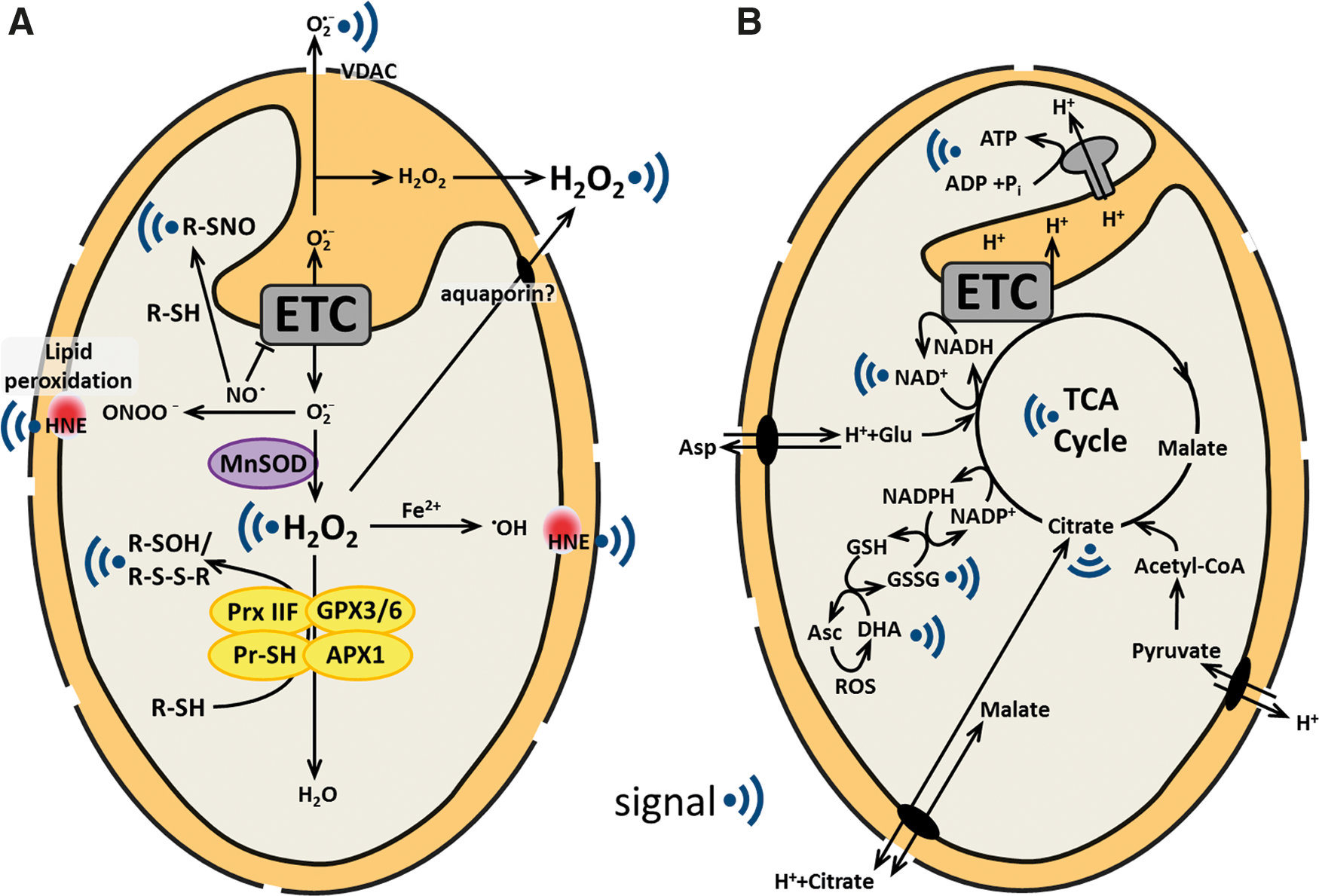
The mitochondrial electron transport chain (mtETC) is a major generator of ROS in mitochondria. Electron transport occurs via flavin, metal centers, and quinones. Depending on their redox state and location within a respiratory complex, they are able to pass single electrons directly to molecular oxygen, which is reduced to superoxide. Main sites of superoxide production are located at complex I (NADH dehydrogenase) and complex III (cytochrome b/c1 complex) (220). Flavin-containing enzymes, such as the 2-oxoglutarate dehydrogenase, may also contribute to ROS production depending on the metabolic status of mitochondria (148, 220). In animals, two respiratory modes are considered as major contributors toward mitochondrial ROS production, (1) when the matrix NADH/NAD+ ratio is high, and (2) during reverse electron transport back to complex I due to a highly reduced ubiquinone pool and high membrane potential (148). However, it is yet unclear whether these conditions have a biological relevance in plants, where the electron transport is more flexible due to the presence of additional alternative respiratory enzymes. External and internal alternative NAD(P)H dehydrogenases and the alternative oxidase (AOX) allow for electron transport and oxygen reduction without proton pumping (142). Uncoupling proteins can facilitate passive proton flux across the inner membrane, dissipating the proton gradient and leading to uncoupling of electron transport from ATP production (204). Although ROS can damage cellular constituents, they can also convey specific information about cellular status regulating growth, development, stress adaptation and, the initiation of a programmed cell death, for example, in pathogen defense and xylogenesis (141, 230). Each ROS has its individual biological effects due to differences in diffusion range, half-life, membrane permeability, and reactive specificity and is—in principle—able to convey specific information (57). Different ROS must therefore be individually appraised (Fig. 3A).
Superoxide
The superoxide radical anion is not membrane permeable and has a very short half-life in the mitochondrion. Hence, a potential superoxide sensor would need to reside in close proximity to the location of its generation. In principal, superoxide can act specifically through disruption of iron–sulfur (Fe-S) clusters. Such a sensing mechanism is adopted for activation of the transcriptional regulator SoxR in Escherichia coli (38). Superoxide levels in the matrix are considered to be very low due to the high rate of dismutation by the manganese superoxide dismutase (MnSOD) (146). However, under strongly oxidizing conditions, MnSOD is itself modified and was found decreased in abundance in Arabidopsis (212), and post-translational modifications, such as tyrosine nitration and lysine acetylation identified on MnSOD in human and mouse mitochondria, may additionally modulate superoxide signals (56, 157). In plant and animal mitochondria, there are also several Fe-S cluster-containing enzymes, of which aconitase is particularly prone to inactivation by superoxide (143). In line with that, MnSOD suppression in Arabidopsis causes decreased mitochondrial aconitase activity and TCA cycle flux (146).
Nitric oxide
The volatile radical nitric oxide (NO) regulates several physiological and developmental processes in plants, from pathogen defense to root growth, stomatal closure, and flowering [reviewed in ref. (22)]. It was demonstrated that respiratory activity contributes directly to NO production, which is further stimulated under anoxic and hypoxic conditions (68, 166). However, no classical mitochondrial NO synthase has yet been identified in plants. NO production under hypoxic conditions results in inhibition of aconitase activity and in induction of AOX (68). Furthermore, mitochondrial NO levels are increased in the absence of AOX, pointing to a role of AOX in the regulation of NO production (36). Several NO targets have been identified in plant mitochondria, including glycine decarboxylase (GDC) and mtETC constituents (160). GDC activity is inhibited by S-nitrosylation, and appears to be involved in the regulation of NO-dependent signal transduction in Arabidopsis, although the underlying signaling pathway is yet elusive (160). NO-dependent nitrosylation of cysteine residues is an important physiological mechanism in the regulation of redox-active proteins (149), and S-nitrosoglutathione is a major storage and transport form of NO (183). NO readily reacts with superoxide to form peroxynitrite, which can interfere with tyrosine-dependent kinase signaling pathways through tyrosine nitration (227). It is thus far unknown which enzymes are able to detoxify peroxynitrite in mitochondria, but peroxiredoxins (PRXs) are likely candidates (183). Several studies have demonstrated NO-dependent regulation of nuclear-encoded mitochondrial transcripts in Arabidopsis (83, 162, 168). However, it remains currently unclear whether mitochondrial NO mediates signaling directly by diffusion into the nucleus or indirectly through modulation of redox signaling.
Hydrogen peroxide
Hydrogen peroxide (H2O2) can specifically oxidize reactive cysteine residues in proteins. This reactivity is actively used in biological systems to control redox regulation and signaling within the cell. The transcription factor (TF) OxyR from E. coli acts as such a cysteine-based H2O2 sensor, controlling the expression of antioxidant defense-related genes (38). H2O2-sensitive, cysteine-based TFs have also been identified in plants: Rap2.4a, which regulates the expression of plastidial antioxidant enzymes, bZIP16, a member of the G-group bZIP TFs, and TGA1, which interacts with NPR1 in the activation of salicylic acid (SA) induced pathogen defense genes (41, 118, 201, 202). The contribution of mitochondrial H2O2 to regulate the activities of such redox-sensitive TFs is however not yet fully understood. In plants, the interpretation of H2O2 signals is particularly complex, as H2O2 is generated in multiple compartments. H2O2 perceived by redox-active cysteine residues in the cytosol or the nucleus is therefore likely to induce a general, rather than a mitochondrion-specific, response (145). However, plant mitochondria contain many proteins with redox-active thiols that could sense H2O2 in situ and mediate specific mitochondrial H2O2 signaling.
Antioxidants: ascorbate and glutathione
Ascorbate and glutathione are the major redox buffers of plant cells, including mitochondria, and are essential to both plant growth and development [recently reviewed in ref. (55)]. Although both antioxidants are biochemically very different, they act together in the ascorbate–glutathione cycle, which is a core component of the antioxidant machinery in plant mitochondria (29). Regulation of the ascorbate–glutathione cycle on the protein level would be a suitable way for the cell to control peroxide levels; however, it is yet unknown whether post-transcriptional modifications can affect its activity (50, 53, 89, 90).
Ascorbate
The mtETC plays a key role in ascorbate biosynthesis. Inhibition of respiration at the level of complex III, IV, or V leads to a strong decrease in total cellular ascorbate content in plants (25, 138, 248). The effects of ascorbate on gene expression are well documented in Arabidopsis (94, 95, 163). A strong overlap of transcriptome signatures was reported between mutants of ABI4, VTC1 and VTC2 (94). The abi4 mutant is deficient in a TF that regulates sugar sensitivity of plants as well as mitochondrial and plastidial RTG signaling (64), whereas the vtc mutants are deficient in total cellular ascorbate. This suggests that ABI4 plays a role in low ascorbate signaling, most likely by modulating abscisic acid (ABA) levels (94). Respiration-dependent changes in mitochondrial ascorbate synthesis may modulate interorganellar communication between plastids and mitochondria (214) and could regulate RTG signaling as a common signal from both organelles. Further dissection of the signaling mechanisms upstream of ABI4 will need to elucidate the role of ascorbate in RTG crosstalk.
Glutathione
Mitochondrial glutathione redox regulation is largely autonomous from other cellular compartments in Arabidopsis (194), making changes in glutathione redox potential a suitably specific signal. The mitochondrial glutathione pool is kept highly reduced by glutathione reductase 2, which shows dual localization in mitochondria and plastids (29). Although glutathione is only synthesized in plastids and the cytosol, immunocytochemical staining suggested that mitochondria contain particularly high glutathione concentrations (246). Intriguingly, in pad2-1 mutants that show decreased glutathione content by 80%, only mitochondrial glutathione immuno labeling remained at similar levels as observed in wild type (WT) (246). A mitochondrial glutathione transporter has not been identified yet in plants, but in analogy to the mammalian system, the dicarboxylate/2-oxoglutarate transporters in the inner mitochondrial membrane (IM) are potential candidates (232). The mitochondrial glutathione uptake system may dynamically modulate thiol redox status and signaling. Several Arabidopsis mutants of mitochondrial complex I are insensitive to the glutathione biosynthesis inhibitor buthionine sulfoximine, and are able to maintain higher glutathione levels, providing a yet-unexplained connection between mitochondrial respiration and glutathione metabolism (102). Under normal conditions, the mitochondrial glutathione redox state is highly reduced [approximately −360 mV; (193)]. However, when mitochondrial function is impaired, like in mutants of the mtETC (46, 248) or mitochondrial antioxidant enzymes (49, 146), the glutathione pool can partially oxidize. The shifted redox glutathione potential is likely to oxidize specific protein thiols directly or indirectly. Several pathways have been described how redox information may be processed within the thiol/disulfide redox regulatory network of the cell (101).
Mitochondrial redoxins
Triggering thiol switches by redox changes is a likely mechanism of mitochondrial signaling (Fig. 3A). Given the highly reduced resting state of the thiol redox environment in the matrix, a shift in redox potential toward oxidation is typically regarded as a redox signal. However, in principle, also a reductive shift could act as a trigger, in a similar manner as reported during pathogen response in plants (41, 54). The mitochondrial thiol redox pool consists of glutathione, lipoic acid, free cysteine, as well as protein thiols. In mammals, protein thiols dominate mitochondrial thiol content by far implying a major role for the proteome in redox control (179). A similar situation is likely to also exist in plants. With respiratory metabolism acting as both electron donor (via NADPH) and acceptor (detoxification of H2O2), the mitochondrial thiol pool is perfectly positioned to play a major role in mitochondrial redox signaling.
Thiol-based peroxidases
Peroxides that are produced in the mitochondrial matrix are detoxified by thiol-based peroxidase systems, such as PRXs and glutathione peroxidases (GPXs), which are critical for the protection against oxidative stress (16, 49). The oxidized cysteines of both enzymes have to be regenerated during their catalytic cycle, which occurs mainly via the thioredoxin-/NADPH-dependent thioredoxin reductase (TRX/NTR) system or alternatively via the glutaredoxin (GRX) system (for PRX) as well as to some extent also directly by GSH (49, 105). Plant mitochondria contain an atypical type II PRX (PRX IIF) (49), and two predicted GPXs (GPX3 and GPX6). PRXs can be transducers in redox signaling in plants and animals acting as floodgates for H2O2 under strong oxidative conditions, through hyperoxidation of their catalytic cysteine residues (101). The inactivated sulfinic form of PRX can then be regenerated by sulfiredoxin (SRX) (96). Regeneration of PRX by SRX is dependent on ATP and TRX as a reducing agent. SRX was recently shown to reside both in mitochondria and chloroplasts of plants, and was able to regenerate the mitochondrial PRX IIF (86). Direct PRX-mediated regulation of transcription was demonstrated in yeast, where oxidized PRX mediates disulfide formation in the TF YAP1 and thus regulates its partitioning between the cytosol and the nucleus (155). Although direct interactions between plant PRX/GPX-proteins and TFs have not been described so far, thiol-based interaction with proteins involved in signaling or metabolic regulation might be of similar importance to modulate redox signaling. Such a mechanism allows the transfer of the oxidation state of PRX to proteins that are less reactive with H2O2 (180). The GPX3 protein of Arabidopsis, for example, has been suggested to inhibit the ABA-insensitive 2-protein phosphatase, and thus alters ABA signaling (137). Further research will need to address the specificities and in vivo interactions of mitochondrial thiol peroxidases to elucidate their role in peroxide-mediated mitochondrial redox signaling.
Thioredoxins
TRXs in general regulate protein function by reduction of disulfide bonds and by modulating thiol switches in course. They provide reductant for the catalytic activity of PRXs, and have direct control of the activities of metabolic enzymes [recently reviewed in ref. (136)]. Intriguing new functions for mitochondrial TRXs and their reductases (NTR) in denitrosylation and activation of the apoptotic regulator caspase-3 have recently been identified in human lymphocytes (20). The plant mitochondrion also contains a functional TRX/NTR system, with TRXo1, TRXh2, NTRA, and NTRB as the major components (178). More than 50 in vitro TRX-targets have been identified in plant mitochondria, covering major metabolic pathways such as the TCA cycle, electron transport, ATP synthesis, and photorespiration (13). Poplar AOX is specifically regulated in vitro by Trxh1, which reduces a disulfide in the AOX dimer that is crucial for its activity (61). Trxo1 was shown to serve as an electron donor to Prx IIF of Arabidopsis and pea in vitro (15, 49). This indicates that plant mitochondrial TRXs may regulate respiratory metabolism, antioxidant activity, and redox signaling. However, their role in vivo has still to be demonstrated. The redox state of mitochondrial TRXs is dependent on the activity of the NTRA and NTRB, which are targeted to either the cytosol or mitochondria depending on alternative splicing in Arabidopsis (178). Deletion of either the cytosolic or the mitochondrial NTR in mouse resulted in embryonic lethality (34). In contrast to that, NTRA and NTRB double-knockout plants of Arabidopsis are viable and fertile, but have a wrinkled seed phenotype, slower plant growth, and pollen with reduced fitness (177). Complementation of the knockout plants showed that the phenotypes could only be rescued by the cytosolic, but not by the mitochondrial, isoforms (177). Furthermore, knockout lines for either of the two mitochondrial TRXs were reported to lack a particular phenotype (136). This raises the question to what extent the mitochondrial metabolism is regulated through TRXs and whether there are alternative mechanisms to compensate a loss. Candidates for such compensation may include GRX systems as shown for TRXh3 (177), or yet unknown players similar to the two-domain protein NTRC, which is localized in plastids (99).
Glutaredoxins and S-glutathionylation
GRXs are structurally related to TRXs, but function mostly independent of NTRs, using GSH as reductant instead, although some exceptions exist (58, 215). The functional roles of GRXs are broader than those of TRXs, reflected by variable active sites and the occurrence of monothiol, dithiol, and multidomain GRXs [recently reviewed in ref. (136)]. Generally, GRXs can reduce protein disulfides, de-glutathionylate proteins, and bind and incorporate Fe-S clusters into proteins (136). Despite the presence of many GRX genes in Arabidopsis [about 50 members from five different subgroups, (136)], only one monothiol GRX is predicted to reside in mitochondria [GRXS15 (187)]. The mitochondrial localization of GRXS15 was confirmed experimentally by proteome analysis and green fluorescent protein fusion (29, 100). The function of GRXS15 is yet unknown (14), and it remains an open question which enzymes mediate protein de-glutathionylation in plant mitochondria. S-glutathionylation is a reversible post-translational modification of proteins and plays a conserved role in protecting proteins from irreversible oxidation, regulating enzymatic activities (usually leading to inhibition) as well as redox signaling. Several mitochondrial target proteins for glutathionylation have been identified in Arabidopsis and Chlamydomonas, such as HSP60, fumarase, aconitase, malic enzyme, galactonolactone dehydrogenase, and GDC (42, 114, 160, 245). Metabolic changes that result from activity changes of these enzymes may in turn trigger mitochondrial signaling. A role for human 2-oxoglutarate dehydrogenase as a possible redox sensor was recently proposed, as glutathionylation occurs at the lipoic acid moiety of the E2 subunit only in specific cell types (134). Further research will be needed to clarify the functional role of glutathionylation in plant mitochondria and to identify potential mitochondrial GRXs in metabolic regulation and redox signaling.
Mitochondrial NADPH pool
Respiratory metabolism transfers reductant from carbon skeletons to the mitochondrial pools of redox cofactors NADH, but also NADPH. Both can serve as substrate for the ETC via the different specific dehydrogenases; however, NADPH is likely the sole source of the reductant to maintain mitochondrial thiol-redox homeostasis. It is not certain where exactly the reductant to maintain the NADPH pool originates from in the plant mitochondrion in vivo, but an obvious source is NADP+-dependent isocitrate dehydrogenase (144). Since NADPH can only serve as a substrate for electron transport under elevated free calcium levels in the matrix (174), it will mainly act as a reductant source for the mitochondrial thiol-redox systems, thus linking respiratory metabolism and redox regulation. A drop in NADPH availability, due to either consumption by alternative electron transport or insufficient reduction rate by respiratory metabolism, could accordingly act as a signal that is transduced by the thiol-redox network as a consequence of decreased disulfide re-reduction.
Colocation for redox regulation
Mitochondrial redox signaling from the mtETC is further complicated by the split location of mitochondrial genes in both the nuclear and the mitochondrial genome. The Colocation for Redox Regulation (CoRR) hypothesis has first been suggested two decades ago and provides an elegant theoretical framework of how signaling between the ETC and the organellar genome may be organized (2, 3). According to CoRR, a redox imbalance in mitochondrial electron transport can act as a signal that modulates organellar gene expression adjusting the capacity or composition of the mtETC, thus relieving the imbalance in a negative feedback loop on the level of a single mitochondrion or plastid. For such a mechanism to function, a two-component redox regulation system consisting of a redox sensor and a redox-response regulator is the minimum requirement. However, a more complex signal transduction mechanism to link the status of electron transport with expression of the organellar genome is also accommodated by the CoRR hypothesis: The redox status of a certain complex could be read out by a specific sensor protein such as a signaling kinase that interacts physically with the complex; alternatively, its redox status could be reported more indirectly, for example, by its degree of ROS release or the status of its direct membrane lipid environment. Recently, evidence for the former possibility was found in chloroplasts by the identification of a sensor kinase, which was reported to link the redox status of the plastoquinone pool to plastidial transcription of the psaA gene and thus photosystem stoichiometry adjustment (171). For mitochondria, similar components of the postulated CoRR mechanism are yet to be discovered. In plants, redox-response regulation may be organized by specific ancestral sigma factors, such as SIG5 (219) and associated factors, for example, ANK6 (243), as well as gene-specific RNA polymerase activities (108) [see Liere et al. (117) for a recent review of organellar transcription in plants]. As a complication, mitochondrial DNA (mtDNA) is distributed highly heterogeneously across the individual discrete mitochondria of a plant cell that undergo frequent fusion and fission. Often, mitochondria only contain a substoichiometric amount of the full genome, and some mitochondria contain no mtDNA at all (6, 170). For these mitochondria, CoRR requires additional features: upon a signal, fusion with a DNA-containing mitochondrion needs to occur, so a change in gene expression can be achieved; alternatively, CoRR could act post-transcriptionally on mtRNA, which may be stored in mitochondria that do not contain any mtDNA.
Signals derived from mitochondrial bioenergetic processes
Mitochondrial energy conversion is highly regulated to allow flexible adjustment to imbalances. This requires a sophisticated energy-signaling machinery (51). Central energy metabolites, such as NAD+, ATP, or carboxylic acids, are well suited to be sensed as signals, as their concentrations in the matrix and their import or export rates strictly depend on the metabolic state of the mitochondrion (Fig. 3B).
NAD+
The mitochondrial respiratory activity is intimately connected with NADH production by the TCA cycle and associated reactions, as well as by photorespiratory glycine oxidation in leaves. Hence, the redox state of the NADH/NAD+ pool is set by the balance of electron influx into and efflux out of the pool. Shifts in its redox potential can occur as a result of imbalances between metabolic flux and electron transport. This balance is even more complex in plants compared to animals due to the presence of alternative NADH dehydrogenases that can be upregulated upon stress [reviewed in ref. (175)]. Furthermore, NAD+ levels can be regulated by import into mitochondria by the recently identified NAD+ transporter, NDT2 (159). While this uptake can adjust the overall pool size, it is unlikely to have any major impact on mitochondrial NADH/NAD+ ratio, due to its modest capacity compared to that of the dehydrogenase and reductase systems in the matrix. The amount of free NADH in isolated plant mitochondria has been found to be relatively constant (93), while there is evidence for an up to fivefold change in extractable matrix NADH concentration of protoplasts depending on the metabolic situation (with NAD+ levels remaining unchanged) (85). Mitochondrial dehydrogenases might be involved in NADH/NAD+ redox sensing; the tobacco CMSII mutant shows altered redox homeostasis across the whole cell as a result of lacking complex I activity (46). In mammalian mitochondria, sirtuin-type (SIRT) protein deacetylases act as metabolic sensors for NAD+ and are activated under conditions of calorie restriction (228). They regulate ATP production, metabolism, apoptosis, and cell signaling by NAD+-dependent deacetylation of target proteins involved in these processes (228). Acetylated mitochondrial proteins, such as cytochrome c, have been identified in Arabidopsis recently and may be involved in NAD+-dependent signaling (50, 234). Arabidopsis contains two sirtuin homologs (78), but only AtSIRT2 has a predicted mitochondrial targeting peptide like its mitochondrial homolog in rice (31). Although it is tempting to hypothesize about an NAD+ sensing function of AtSIRT2 in plant mitochondria, AtSIRT2 is most closely related to mammalian SIRT4 (161), which only possesses weak ADP-ribosyltransferase activity (1).
Metabolites of the TCA cycle
Different types of organic acids from various metabolic processes are taken up by mitochondria and are oxidized by respiration (4). After import into the matrix, the flux through the mitochondrial metabolic pathways determines the steady-state concentrations of the individual metabolite pools. Sensing individual pool sizes and/or flux through individual reactions provides a suitable way to monitor respiratory status. For instance, different flux modes of the TCA cycle could be detected depending on the relative pool sizes (211). Furthermore, metabolites such as pyruvate, citrate, isocitrate, 2-oxoglutarate, and malate/oxaloacetate, as well as glutamate and glycine/serine are particularly important in intercompartmental metabolite exchange in plants (172) and thus well placed to act in signaling. In human cells, succinate, fumarate, and 2-hydroxyglutarate have signaling functions as oncometabolites, as they are able to inhibit several chromatin modifiers and thus affect transcription (239). In plants, chloroplastic malate has been discussed as a redox signal, since excess reducing equivalents produced by the photosynthetic ETC can be exported by the malate valve (190), which helps to regenerate NADP+ as an electron acceptor of chloroplast metabolism. After its export, malate is oxidized to oxaloacetate in the cytosol, and the NADH produced can be used for metabolic processes such as nitrate reduction or in mitochondrial respiration through external NADH dehydrogenases (175). TCA-cycle organic acids have long been discussed to be of particular importance in RTG signaling in yeast as well as in plants (66, 120). TCA cycle dysfunctions strongly correlate with increased levels of citrate, isocitrate, and 2-oxoglutrate, which are thought to control the expression of TCA cycle genes citrate synthase, aconitase, and isocitrate dehydrogenase through the ‘RTG pathway’ in yeast (120). Various mitochondrial enzymes such as aconitase, pyruvate dehydrogenase, and GDC are susceptible to inactivation by oxidation. Inhibition of these enzymes by increased ROS production will result in slowdown of the TCA cycle flux (146). AOX1 transcript levels are commonly used as a readout marker for mitochondrial RTG signaling in plants. In tobacco cell culture, the AOX1 transcript is known to be induced (up to 24-fold) after exogenous feeding with citrate, isocitrate, malate, or 2-oxoglutarate, as well as by inhibition of aconitase, resulting in increased internal citrate levels (66). It was recently demonstrated in Arabidopsis roots that citrate accumulates after inhibition of aconitase upon hypoxia, and that citrate, but not NO, mediates the induction of AOX (transcript as well as activity) (68). However, it is yet unclear how carboxylic acids may induce a specific RTG response, as they are usually highly abundant in various subcellular compartments. One possible mechanism would be that a carboxylic acid is sensed by specific metabolite receptors or transporters at the IM. In bacteria, organic acids such as citrate are sensed by high-affinity sensor kinases, such as the two-component histidine sensor kinases CitA/B and DcuS/R (191). However, homologous proteins have not been identified in plants thus far.
ATP and derivatives
Sensing changes in mitochondrial ATP/ADP ratios would be a particularly efficient way to communicate mitochondrial bioenergetic status to the rest of the cell. A recent transcriptomic study indeed demonstrated that a gradual decrease in cellular ATP/ADP ratio by inhibition of the mitochondrial ATP synthase in Arabidopsis induces RTG signaling and results in an induction of alternative respiratory pathways (60). The underlying signaling pathways that report changes in the ATP/ADP ratios and that might involve the mitochondrial ATP/ADP translocator are however still unknown. Mitochondrial kinases that might respond to changes in ATP levels are discussed to play a role in RTG signaling in animal mitochondria (80). Furthermore, animal mitochondria are able to control cell cycle progression through regulation of ATP production (156). Plants also contain the energy-sensing kinase SnRK1, which belongs to SNF-1-related kinases. However, SnRK1 does not sense AMP levels, and although it is activated during starvation, the identity of the signal that is perceived still needs to be elucidated (10). SnRK1 is a likely candidate to interact with mitochondrial RTG signaling pathways, as its target genes are also regulated in mutants with defects in mitochondrial functions (195). Recently, evidence was found for 3′-phosphoadenosine 5′-phosphate (PAP), a precursor of AMP, to act as a plastidial RTG signaling metabolite in Arabidopsis. Usually, PAP is dephosphorylated to AMP by the enzyme SAL1, which is dual-localized to mitochondria and plastids. Under drought and high light stress, PAP accumulates in the nucleus and regulates nuclear gene expression possibly through inactivation of XRN-type exoribonucleases (47). Hence, it will be interesting to elucidate whether PAP also plays a role in ‘mitochondria-to-nucleus signaling’ in plants.
Calcium
Until relatively recently, calcium in plant mitochondria was assumed to be of little regulatory importance (44). This view has gradually changed as a result of the discovery of several mitochondrial processes in plants that are controlled by calcium, including alternative electron transport (174), protein import (109), and permeability transition (8). Introduction of an aequorin probe, and recently of a cameleon sensor, into plant mitochondria showed in vivo calcium dynamics with signatures different to those in the cytosol (124, 128). Mitochondrial calcium uptake and storage in plants may in course play an active role in modulating and adjusting cytosolic calcium signals. Additionally, calcium is likely to fulfill regulatory functions inside the mitochondrion. Plant mitochondrial proteomes contain several proteins with calcium-binding motifs (30), which typically couple changes in calcium activity to changes in protein structure and function. In mammals, calcium is a central regulator of mitochondrial energy metabolism. TCA cycle flux is controlled by calcium on the level of different dehydrogenases, coupling energy output to demand (67). Although the required machinery for calcium sensing appears to be present in plant mitochondria, it is yet unclear how mitochondrial calcium dynamics link to physiological regulation. Free calcium levels are modulated by release and sequestration. Respiratory intermediates, such as citrate and isocitrate, have very different binding affinities to calcium (18); changing steady-state levels of those metabolites can therefore impact on free calcium levels in the matrix, and convey information about respiratory status. pH changes also affect calcium binding. Like in mammalian cells, calcium flux across the plant IM is mediated by a transport system for calcium (207). Recently, the genetic identity of two components of the mitochondrial calcium uniporter was identified: a channel-forming subunit and a regulatory subunit (17, 40, 165). Homologs are present in plants with predicted mitochondrial targeting (207). Rapid changes in free matrix calcium referred to as pulses coincide with a rapid bioenergetic remodeling in single mitochondria of Arabidopsis (197). Excessive matrix calcium load can trigger cell death signaling in plants (122). Calcium is expected to act as a central hub in plant mitochondrial signaling, and in-depth dissection of its regulatory properties is a very active field of current research.
Mitochondrial pH
Mitochondrial pH is intimately linked to respiratory status. The pH gradient across the IM is a constituent of the proton-motive force that drives ATP synthesis. Matrix pH depends on both, the pH in the cytosol and the pH gradient across the mitochondrial membrane, and either can change (167). If cytosolic pH changes, for example, before guard cell movement (88), matrix pH will follow. While the pH gradient across the IM has long been assumed to be relatively small and constant, this view is currently changing (189, 198). In Arabidopsis roots, matrix pH can transiently increase, due to an increased pH gradient (196, 197). Similar events have been observed in mammalian mitochondria (9, 189). While the pH gradient per se is generated by respiratory metabolism, it can be shaped by ion movements across the IM, including potassium and calcium (167, 197, 218). Dynamic changes of matrix pH in vivo, depending on cytosolic pH, energetic status, and ion movement, make pH a very suitable mitochondrial signal. A pH shift in the matrix will cause a change in respiratory metabolism depending on the pH optima of the different mitochondrial enzymes. pH changes also impact on the structural organization of respiration, which has recently been shown in isolated potato mitochondria, in which acidification induced the dissociation of supercomplexes (173). Cellular acidification also causes activation of the alternative NAD(P)H dehydrogenases in Arabidopsis (59). Changes in the pH gradient have direct impact on the matrix concentrations of respiratory substrates, many of which are in dynamic equilibrium with the proton gradient that drives their import. This includes concentration not only of phosphate, but also of organic acids such as pyruvate. Changes in the pH gradient are therefore closely linked to metabolic regulation and signaling.
Plant hormones and mitochondrial signaling
The regulation of plant mitochondrial functions by plant hormones and their possible connection to RTG signaling has only begun to be explored. Mitochondrial respiration is inhibited by SA, which may provide a mechanistic link to mitochondrially regulated biotic stress responses (131). In an Arabidopsis mutant of the complex II gene SDH1-1, SA signaling is disrupted and linked to disturbed mitochondrial ROS production (65). Gibberellic acid (GA) might also have a role in the regulation of mitochondrial functions, as growth defects of complex I mutants can be partially rescued by GA treatment (135, 164). Furthermore, mutants with defects in pre-mRNA splicing of the complex I subunit nad2 as well as mutants with defects in editing of nad4 were recently identified as hypersensitive to ABA (119, 147). The identification of ABI4 as a mediator of RTG signaling suggests an intimate linkage between mitochondrial regulation and ABA signaling (64). In addition, crosstalk of mitochondrial ROS with auxin and ABA signaling has also been reported recently (74), further pointing to a central role of mitochondria in orchestrating cellular hormone signaling.
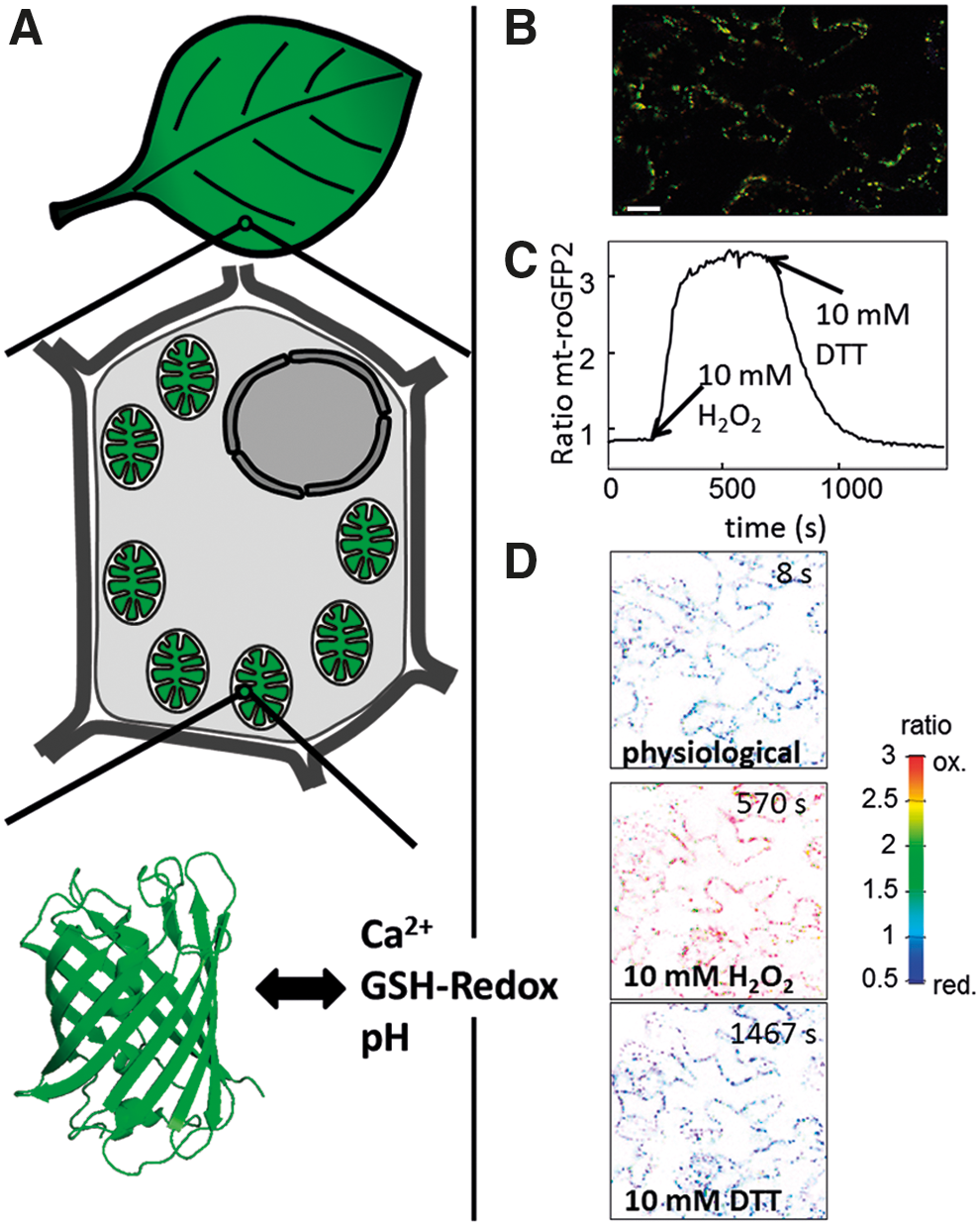
In vivo sensors to probe for the origin of signals
Monitoring dynamic changes in mitochondrial physiology specifically in planta is a prerequisite for detecting potential signals. Technically, this has remained challenging; however, much about mitochondrial status changes has been inferred from measurements in isolated mitochondria (65, 208), whole-tissue extracts (218), or different staining methods—histologically (240) or in vivo (132, 237). Many of the problems associated with those approaches, such as lack of cellular context or lack of accurate spatial and temporal resolution, have been overcome by the introduction of fluorescent protein sensors (Fig. 4). These probes can be targeted to mitochondria, solving the problem of insufficient spatial specificity. Additionally, they are largely nontoxic and can be monitored in living plant cells or tissues, by microscopy or fluorimetry (185). An important feature of these sensors is the rapid and reversible equilibration with their direct environment, allowing live monitoring of physiological changes. Similar to the more traditional techniques, detailed knowledge about sensor specificity is paramount for meaningful measurements and significant efforts must and have been undertaken to address this issue. A variety of different fluorescent protein sensors that cover central mitochondrial parameters and potential signals have been developed and used over the recent years, including pH (121, 167, 196), calcium (107, 150), H2O2 (19, 70), glutathione redox potential (69, 71, 193), ATP levels (87) and ATP/ADP ratio (21), and NADH levels (247) and NADH/NAD+ ratio (84). A subset of those sensors has been introduced into the matrix of plant mitochondria and has allowed dynamic measurements of pH, calcium, and glutathione redox potential in vivo (Fig. 4) (128, 194, 196). The use of redox-sensitive green fluorescent protein redox sensors that interact with GRXs and thereby sense glutathione redox status has established that the mitochondrial glutathione pool is highly reduced under resting conditions (92, 193), but reacts dynamically to inhibitors of respiratory metabolism and several abiotic stresses (Fig. 4) (115, 186, 194, 197). These changes can occur independently from the cytosol, suggesting a large degree of physiological autonomy of the mitochondria, which is consistent with the presence of a complete set of enzymes for redox maintenance in the matrix (see above). Measuring matrix calcium dynamics of guard cells in response to osmotic stimuli and of root tips upon extracellular ATP using a cameleon probe showed that mitochondrial calcium dynamics correlate with those in the cytosol and the nucleus, but show different kinetic properties, with mitochondrial responses occurring more slowly, but lasting for longer (128). This cameleon sensor appears technically very promising to open a door to dissect plant mitochondrial calcium signaling. A circularly permuted yellow fluorescent protein sensor reported to respond to mitochondrial superoxide has also been used in plants (74, 196). However, this probe has been shown to actually report pH and not superoxide in mitochondria (198, 231). Application of this pH sensor enabled the detection of transient alkalinization events in single Arabidopsis mitochondria. Similar alkalinization events have since been reported using SypHer, another protein-based sensor for pH, in single mitochondria of mammalian cells (9, 189). The growing number of available sensors for important mitochondrial parameters in combination with dedicated imaging approaches will facilitate an integrated understanding of different mitochondrial changes as suitable signals. This will not replace, but complement and increase, the effectiveness of other techniques, such as genetic screening or cell-specific profiling, which will be required to dissect mitochondrial signaling pathways and to identify their components.
How to Send Out the Message: Mitochondrial Signals Passing the Membranes
A mitochondrial signal involved in RTG regulation has to pass the mitochondrial boundaries; thus, the signaling cascade has to reach the cytosol by propagation from the respective mitochondrial microcompartment that it originates from.
Passing the outer membrane
The outer mitochondrial membrane (OM) is generally considered highly permeable for the physical passage of messengers due to the presence of porins, which form large channels and allow diffusion of small molecules, including most metabolites. However, the view of free and uncontrolled diffusion across the outer membrane has been challenged with transport through porins (voltage-dependent anion channel/mitochondrial porins [VDACs]) likely to be regulated. The Arabidopsis genome contains four VDAC genes, and their mutants show differential phenotypes, suggesting specific functions of the different VDAC proteins (113, 216, 238). Additionally, the intermembrane space (IMS) has been reported to maintain its own physiological microenvironment with respect to pH and glutathione redox potential (82, 169), which requires at least limitation of diffusion. If regulated or not, VDACs are able to mediate transit of a diverse set of small molecules. VDACs are excellent candidates to allow passage of signals that originated from the inner side of the OM, the IMS, the crista space (CS), and the outer side of the crista membrane (CM) and the inner boundary membrane. The release of factors from the IMS and CS, including cytochrome c, into the cytosol for cell death signaling in plants requires additional OM permeabilization. Physical rupture of the OM due to permeability transition pore (mtPTP) opening and matrix swelling has been reported as a mechanism. Similar to mammalian BAX, other death-promoting, pore-forming plant proteins are likely to also mediate OM permeabilization in the absence of rupture (242).
Getting through the intermembrane space
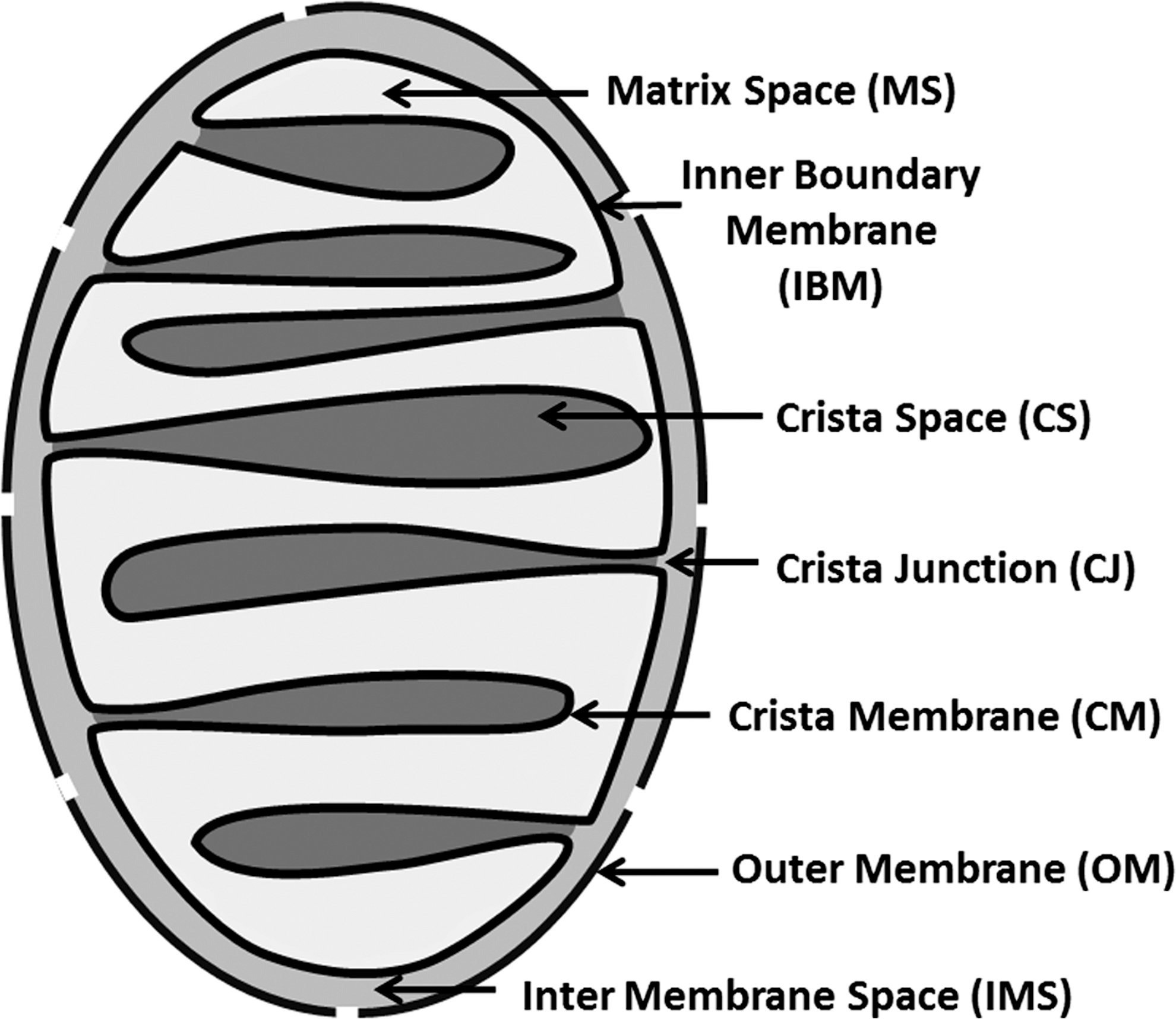
Recent advances in the understanding of mitochondrial architecture have led to a renaissance of the old concept that the IMS and the CS require careful distinction (Fig. 5) (152). The distribution of respiratory complexes is compartmentalized in the inner membrane with a preference for the CM (39, 225). Depending on the nature and regulation of crista junctions (CJs), the CS may not even always be continuous with the IMS. A signal generated in the CS needs to enter the IMS via a CJ. Ongoing research into the regulation and dynamics of CJs will reveal their potential role in mediating transport and transduction of signals.
Passing the inner membrane—transporters, pores, diffusion, and relays
According to the chemiosmotic theory, the inner membrane needs to be highly impermeable, to allow oxidative phosphorylation to occur in any efficient manner (140). Any required transport of charge or protons must occur in a specific and controlled manner. This is achieved by a comprehensive set of transporters, which mediate passage of metabolites, cofactors, metal ions, as well as nucleic acids (103) and polypeptides. As described above, these compounds are suitable mitochondrial signals and in turn may be exported from the matrix space by appropriate transporters. While most information on carriers in plant mitochondria is based on biochemical data, the transport system is only partly characterized genetically. The genetic and functional conservation of plant mitochondrial transporters has been recently covered by a review (158). Since then, the genetic identities of the calcium uniporter and the pyruvate transporter have been elucidated (17, 24, 40, 76).
The mtPTP has been suggested to act as a mediator of matrix signals, and in support of this transient opening of the mtPTP under physiological conditions has been reported [e.g., ref. (79)]. In such a case, a brief opening of an unspecific channel would allow passive export of matrix contents to the cytosol. This however implies a drastic impact on mitochondrial functionality, and the evidence for such scenarios is currently inconclusive. In plants, there is evidence for opening of an mtPTP under certain conditions such as oxidative stress and mitochondrial calcium overload. However, its properties are not well defined, although they are often assumed to be parallel to those in animal systems [e.g., refs. (8, 116, 199); see McInerney et al. (133) for a review covering mtPTP]. Nuclear gene expression changes induced by interference with mitochondrial function in tobacco were suppressed by treatments with bongkrekic acid, an inhibitor of the mtPTP in animals (131). The mechanism of bongkrekic acid is however binding and inhibition of the mitochondrial ATP/ADP carrier, a putative component of mtPTP in mammals, thereby severely interfering with mitochondrial and cellular ATP homeostasis.
Passive diffusion across the IM does not require any transporters. This mode of membrane passage is only efficient for small, largely nonpolar compounds. Products of lipid peroxidation, such as 4-hydroxy-2-nonenal, may cross membranes in such a way and modify proteins and cofactors inside and outside of the mitochondrion (233).
The relay of a signal by a membrane component, rather than the physical passage of the signaling compound itself, represents an alternative pathway to pass information across the IM. Transmembrane signaling proteins, such as receptor kinases, can operate in such a way. Signaling substrates could be bound, or modify the protein on the matrix side by specific physiological changes, such as calcium, pH, or thiol redox status, to transmit the information on to the other side. A small set of transmembrane kinases has indeed been identified in the Arabidopsis proteome, raising the possibility of phosphorylation-based signaling across mitochondrial membranes (75).
Getting ROS across the membrane
Physical passage of ROS signals across the IM deserves a closer look. In isolated animal mitochondria, complex I-derived superoxide is released into the matrix, while complex III generates superoxide on both sides of the IM (148). This is also assumed to be true for plants, although detailed experimental confirmation is lacking. In the matrix, negatively charged superoxide cannot pass the IM by diffusion and is efficiently converted to H2O2, which is then assumed to readily diffuse across the IM due to its biophysical properties (small, neutral, and long lifetime). Additionally, it was shown that aquaporins can mediate membrane passage of H2O2, including aquaporins from Arabidopsis (23, 139). At least one Arabidopsis aquaporin localizes to mitochondria in pollen and is regulated in a pH-dependant manner (206). Membrane passage of superoxide mediated by a chloride channel has been a surprising finding and deserves in-depth follow-up (73).
Mitochondrial unfolded protein response and export of peptides
In mammalian mitochondria, unfolded proteins induce RTG signaling by the mitochondrial unfolded protein response (mtUPR) (188). Although the signal that initiates mtUPR is not known, specific TFs are activated by this pathway to regulate the expression of mitochondrial heat shock proteins (HSPs) and proteases. A recent study in Caenorhabditis elegans demonstrated that the TF ATFS-1 is targeted to mitochondria and subsequently degraded under control conditions (151). Under stress, the import efficiency of ATFS-1 is reduced and allows its accumulation in the nucleus to activate mtUPR. TFs localized in mitochondria have also been identified in plants; however, their function in RTG signaling is yet unclear (106). Mitochondrial proteins that cannot be repaired are degraded in all mitochondrial subcompartments (110). The peptides resulting from protein degradation are discussed to play a role in mtUPR in C. elegans and yeast (52). In yeast, the inner-membrane AAA protease Yme1 is responsible for the generation of peptides that were shown to regulate nuclear gene expression (7). There are currently little experimental data whether a similar mechanism exists in Arabidopsis. Recently, it was proposed that oxidatively modified peptides may be able to transduce specific spatial and chemical information from plant organelles; this elegant theory however still requires experimental testing (145).
How the Mitochondrion Controls the Cell
After mitochondrial signals have been generated and transduced they can control a wide range of cellular functions. This broad range of signaling targets reflects the significance of mitochondria in the cell. Mitochondrial signals can either feed back to adjust mitochondrial function or trigger changes in other parts of the cell. Control can be subdivided into direct regulation without the requirement of gene expression changes and indirect regulation of cellular functions by adjusting expression of nucleus-encoded genes by RTG regulation (Fig. 1).
Targets of direct regulation
Regulation of individual mitochondria
A major target of mitochondrial regulation is the determination of mitochondrial status and fate itself. By regulating protein import from the cytosol, mitochondria can control their own composition, and ultimately their structure and function. During yeast mitochondrial biogenesis, function of the translocase of the outer membrane complex is regulated depending on respiratory activity by cytosolic kinases, which thereby control protein import (192). This provides an elegant feedback mechanism that adjusts, for example, the import of mitochondrial metabolite carriers to respiratory demand. Similarly, the thiol-redox status in the IMS may impact on protein import regulation via the disulfide relay system. In plants, components of this import mode are conserved, although the system has been reported to show mechanistic differences to yeast (28). In principle, mitochondrial control of protein import provides an elegant mechanism of readily adjusting the proteome of individual mitochondria according to need in a decentralized fashion and of maintaining mitochondrial homogeneity across the cell.
Regulation of mitochondrial populations
Mitochondrial dynamics, including fusion, fission, motility, and autophagy, must be, at least partly, controlled by signaling from the respective individual mitochondrion. Mitochondrial dynamics are central to mitochondrial quality control, to maintain a homogeneous, functional set of mitochondria in a cell, and to prevent accumulation of mitochondria that disrupt rather than support cellular homeostasis, for example, by releasing excessive amounts of ROS or consuming ATP [see ref. (123) for a recent review]. Although only a few proteins of the dynamics machinery have been identified in plants, they are good candidates to act as targets for direct control in single mitochondria. For instance, size, morphology, and number may be regulated by directly modulating the activity of components of the mitochondrial fission and fusion machinery, including the Arabidopsis dynamin-like proteins 2A and B (ADL2A and B; also referred to as DRP3A and B), NETWORK1/ELM1, and BIGYIN (5, 126, 200). Yeast, slime mold, and fly share a homologous gene (CLU1, cluA, and clu) critical in cellular distribution and positioning of mitochondria that is also present in plants (friendly mitochondria [FMT]) (35, 48, 123). A mutant of FMT shows a very similar mitochondrial phenotype to mutants of the homologous genes in other organisms, including highly clustered mitochondria (125). This makes FMT a potential target for mitochondrial control of their dynamics. How such regulation may be achieved in plants is currently unknown, however, and requires further research attention.
Mitochondrial regulation of intercellular protein transport
Recently, mitochondrial redox status has been implicated in the regulation of transport through plasmodesmata (209, 210). Silencing of ISE1, a predicted mitochondrial helicase, led to an oxidative shift in mitochondrial thiol-redox state as well as increased intercellular protein transport. A similar correlation was observed in response to pharmacological intervention with mitochondrial redox potential. Further research will need to establish if and how the reported correlations link mitochondrial status and intercellular exchange mechanistically.
Mitochondrial regulation of cell death
Mitochondrial control of necrosis and apoptosis has been intensely studied in animals [see ref. (213) for a review], but a close link between mitochondria and cell death induction also exists in plants (98, 122, 226). Several principles such as ROS involvement, loss of electrical potential across the IM, and release of IMS proteins, such as cytochrome c, are conserved components of mitochondrial death signaling (11, 12, 222), although many protein players appear not to be present in plants (33, 111, 176). Mitochondrial cell death signaling has been implicated in pathogen response (37, 112) and developmental processes such as formation of the vasculature and leaf shape (127, 244). Physical targets of mitochondrial death signaling are thought to include cellular dismantling processes, such as tonoplast rupture and DNA degradation, although mechanistic detail remains sketchy.
The retrograde response
A major target for mitochondrial signaling is the nucleus, where cellular gene expression is regulated. This allows constant transcriptional synchronization between mitochondrial and cellular status and is likely to be of particular relevance during mitochondrial biogenesis and during profound changes in environmental conditions [recently reviewed in ref. (181)]. Most commonly, the nuclear-encoded AOX1a transcript has been used as a read-out marker for plant mitochondrial RTG signaling (64, 77). The role of AOX in the acclimation of mitochondrial respiration under stress has been covered by several reviews [e.g., refs. (142, 223)]. A further set of 26 nuclear-encoded and stress-responsive transcripts was identified in a recent transcriptome analysis that specifically analyzed nuclear genes encoding mitochondrial proteins in Arabidopsis (224). Several transcripts of alternative respiration were present that appear generally more responsive to stress treatments than the standard mtETC components (Fig. 6A) (241). Furthermore, transcripts encoding mitochondrial HSPs are specifically induced by mitochondrial dysfunctions as a part of the RTG response (182, 224). However, mitochondrial RTG signaling not only regulates the expression of mitochondrial proteins but also more generally reprograms nuclear gene expression affecting various subcellular compartments and functions (Fig. 6B) (221). Transcript groups encoding proteins involved in photosynthetic light reactions, organellar and cytosolic ribosome composition, as well as pathogen response were recently identified as common targets of regulation under mitochondrial respiratory dysfunction in Arabidopsis (195). This suggests that mitochondrial signaling engages in transcriptional crosstalk with the chloroplast, via control of nuclear genes, in addition to direct metabolic interaction. Controlling translation by regulating ribosomal abundance and composition may allow overall adjustment of the cellular proteome as acclimation response to changing respiratory activity and energy availability. The regulation of pathogen-related transcripts may imply that interference with mitochondrial respiration is a general feature of biotic stress. Several mitochondrial inhibitors originate from bacteria and fungi (e.g., oligomycin, myxothiazol, and antimycin), and pathogen-induced SA signaling may affect respiration directly (see above). The transcriptomic evidence further supports the concept of the mitochondrion as a central integrator of biotic stress signaling, a part of which is mediated by mitochondrial signaling.
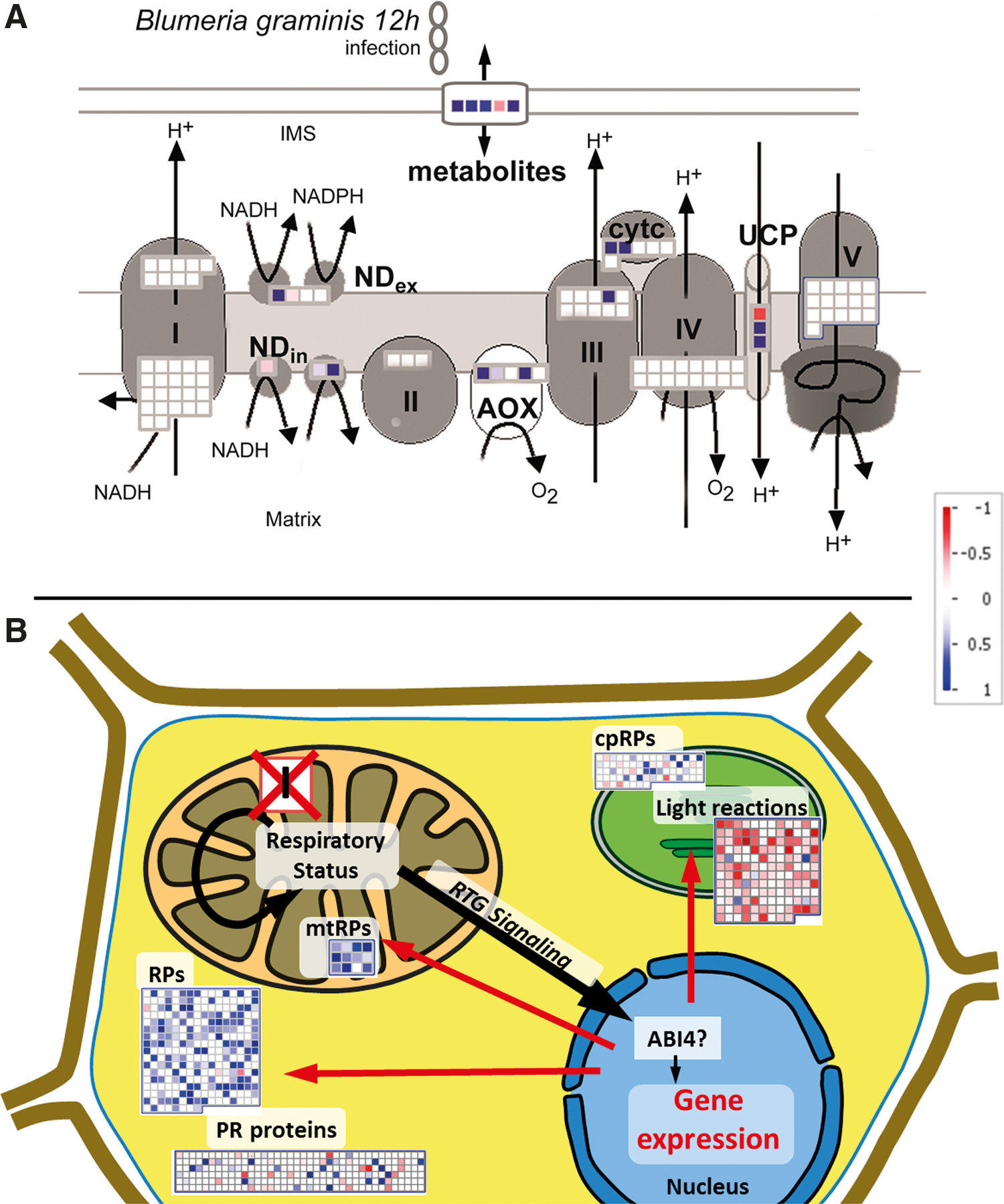
Although no cytosolic components of mitochondrial RTG signaling are known in plants, a small number of TFs have been identified in the nucleus, regulating the expression of nuclear-encoded mitochondrial proteins. Not all of them are responsive to RTG signaling, however, but respond to other stimuli such as diurnal rhythm in case of TEOSINTE BRANCHEDI, CYCLOIDEA, and PCF (TCP) TFs (63), and seed maturation in case of ABI3, FUS3, and bZIP53 (184). Only, ABI4 was identified as the first endpoint of a mitochondrial RTG signaling pathway, and acts as a repressor of AOX1a transcription (64). Furthermore, ABI4 also acts as a repressor of photosynthetic gene expression and is a regulator of plastid RTG signaling (104). However, AOX1a expression is usually derepressed upon mitochondrial stress while photosynthetic transcripts are repressed (195). Hence, ABI4 is unlikely to regulate both pathways single handedly, and other downstream transcriptional regulators must exist.
Mitochondrial dysfunction phenotypes
A large number of different phenotypes have been associated with defects in mitochondrial function in plants. While many are mutant specific, it is possible to extrapolate a reoccurring pattern of typical mitochondrial dysfunction phenotypes in plants (Fig. 7). Their diversity sheds light on the importance of mitochondrial function and signaling in the WT plant cell.
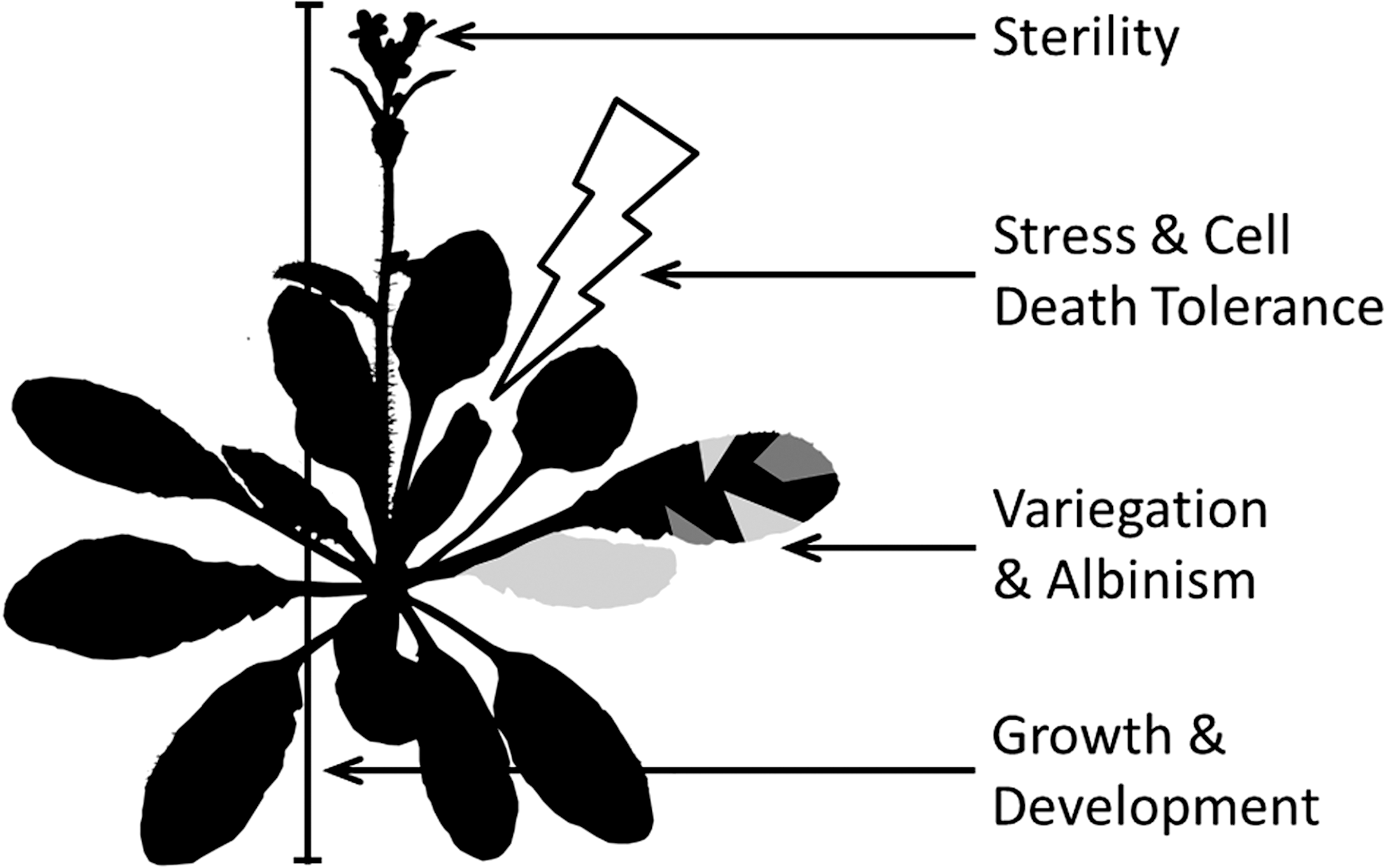
Stress and cell death tolerance
A common feature of different mitochondrial respiratory mutants is a modified susceptibility to stress. Mutants of complex I showed increased tolerance to NaCl, hyperosmotic, and cold conditions in Arabidopsis (135) as well as to ozone and tobacco mosaic virus infection in tobacco (45). A recA1×msh1 double that suffers severe disruption of mitochondrial genome integrity was particularly tolerant to heat stress (203). Shot1, an Arabidopsis mutant lacking a mitochondrial transcription termination factor-related protein, was also found to be thermotolerant (97). By contrast, an Arabidopsis line lacking AOX1a displays increased sensitivity to a combination of high light and drought (62), while overexpressors are more tolerant to salinity (205). Taken together, these phenotypes may be accounted for by mitochondrial dysfunction, generating internal stress, and triggering a cellular acclimation response, mediated by RTG regulation. This can make plants either more susceptible, if normal signaling is disrupted, or more tolerant, by internally priming the cell for an upcoming stress impact. Such a model implies that in WT, the mitochondrion functions as a stress sensor and is involved in mounting an effective cellular response (Fig. 1). Modified susceptibility to cell death is considered another mitochondrial phenotype, potentially affecting both stress responses and development (98, 229). Targeting mitochondrial-mediated cell death by cyclosporine-A treatment of lace plant causes striking changes in leaf morphology (127).
Growth and development
Another common feature of many mitochondrial mutants is a growth phenotype. Reduced or retarded growth has been observed under various conditions for both shoots (46, 97, 135, 203, 236) and roots (49, 62), which is often associated with a prolonged vegetative phase. Mitochondrial metabolism is intimately linked with photosynthetic carbon assimilation via several biochemical processes, including photorespiration, nitrogen assimilation, and the dissipation of excess energy. Hence, mutations in mitochondrial TCA cycle enzymes of tomato have been found to either positively (in case of aconitase and malate dehydrogenase) or negatively impact on photosynthetic carbon assimilation (in case of fumarase), which ultimately affects growth or fruit yield [reviewed in ref. (154)]. Interestingly, it has been reported that retarded growth could be partly rescued by GA treatment (135, 236), implying a degree of active mitochondrial growth control, rather than pure passive acclimation, for example, to compromised mitochondrial energy conversion.
Variegation and albinism
Variegation or albinism can also be a striking phenotype of mitochondrial dysfunction. In such cases, disrupted mitochondrial function leads to impaired plastid development. The maize nonchromosomal stripe mutants (NCS2-6) have different distinct mtDNA defects affecting expression of complex I and IV, respectively, and disrupting mitochondrial translation (129, 153). White-green sectoring, as a result of compromised chloroplast development, is determined by the relative abundance of mutant to WT mtDNA variants. Mutants and suppressors of a MutS homolog in plants, MSH1 (also called CHM1), also show leaf variegation in various species. Although this has been assumed to be the result of mtDNA rearrangement, a recent study has shown that in this case, variegation is actually the result of lack of MSH1 in the plastid (236).
Sterility
A classical mitochondrial phenotype is cytoplasmic male sterility (CMS), which has attracted major interest due to its agronomic significance in breeding. Various independent mitochondrial mutations have been demonstrated to cause CMS in several species, resulting in incompatibility of organellar gene expression with the nuclear genome and disrupted pollen development in turn. The role of the mitochondrion in flower development and CMS has recently been reviewed (27). Also, female gametophyte development can be disrupted by genetic disturbance of the mitochondrial ATP synthase (60). CMS may be regarded as an extreme of the mitochondrial impact on fertility and implies that sexual reproduction is more generally regulated by mitochondrial function and signaling, also in WT plants.
Outlook
Toward an integrated view of mitochondrial signals
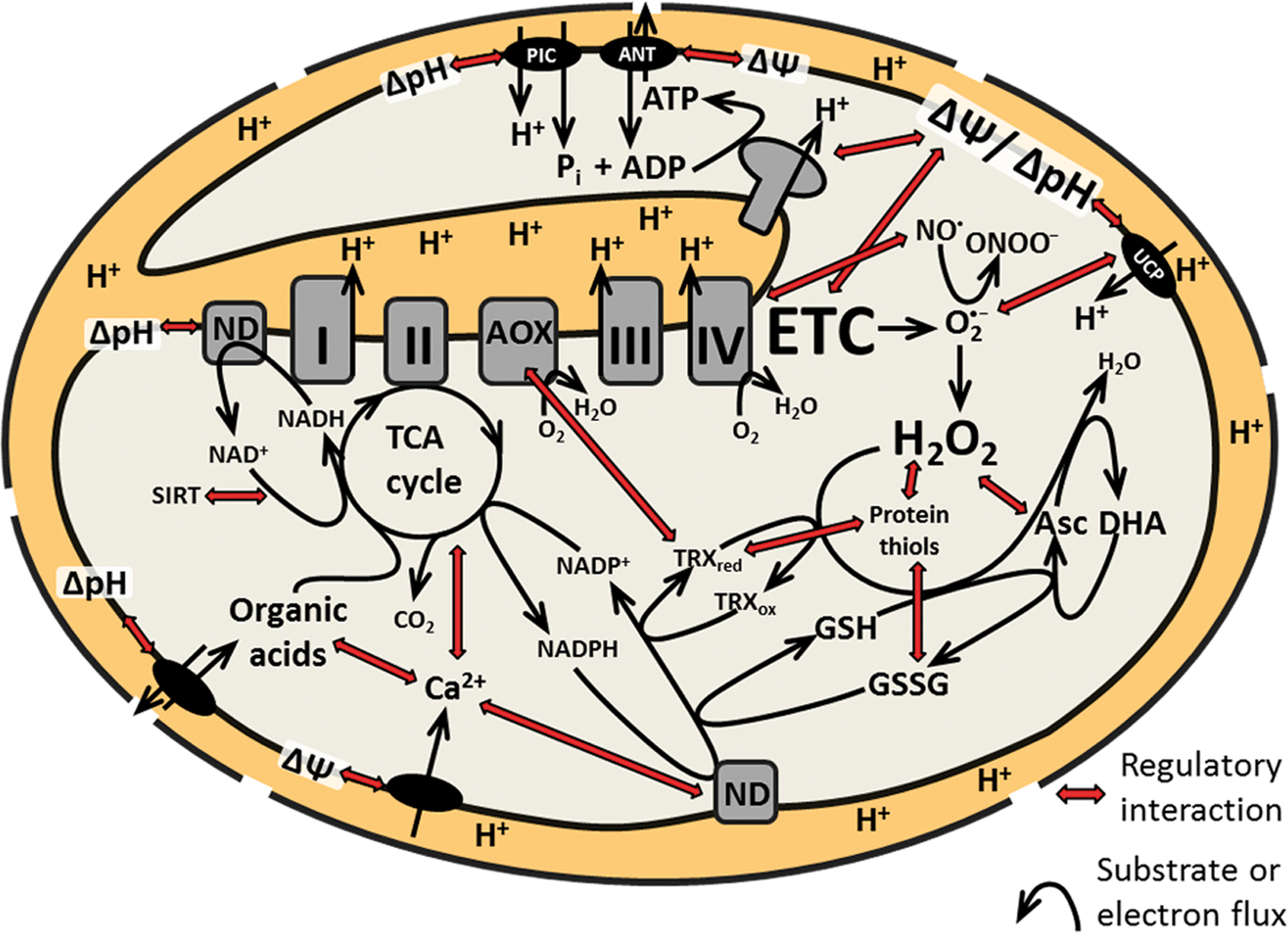
The current understanding of potential signals from different levels of mitochondrial function is incomplete. No single signal-sensor couple is described in plant mitochondria. Instead, the available data reveal the highly complex and dynamic inter-relation of the physiological parameters of candidate signals. Different parameters can change as a part of the same effect, because they are physiologically linked (Fig. 8). This principle that one change triggers many modifications with several potential signals as output appears to be the typical situation, and thus is critical for the dissection of mitochondrial signaling in vivo. As an example, calcium influx into the matrix may trigger a calcium signal, but it may also trigger matrix alkalinization and a pH signal. An increased pH gradient may lead to changed metabolite levels in the matrix and modified metabolite signaling and so forth. As another example, inhibiting complex I may reduce proton pumping and the rate of ATP synthesis, but at the same time lead to a shift in the matrix NADH/NAD+ ratio, modifying metabolic flux that also sets the NADPH/NADP+ ratio and in turn determines reductant delivery to the TRX system and the glutathione pool involved in redox control and ROS detoxification. The signaling effects of complex I inhibition could then be the result of any of the modified parameters acting as a stimulus. Mitochondrial complexity is unlikely to accommodate specific changes of individual parameters, such as calcium or H2O2 levels, and a more suitable view may be that of a highly interconnected network of potential mitochondrial signals. Such a network would then be able to adopt different discrete signaling states (similar to the flux modes of the metabolic network). Such a view would account for the yet-limited success in the dissection of mitochondrial (RTG) signaling, not only in plants. Experimental interference with a certain mitochondrial parameter would change the status of many others and thereby cause a shift in signaling state. While this allows studying the signaling response, for example, as a change in transcript levels, it makes it virtually impossible to pin the signal down to an individual physiological parameter. Instead, the concerted change of multiple parameters mounts the signal. Even if a sophisticated experimental system managed to cleanly isolate a specific signal, this would then be unlikely to reflect any physiologically relevant situation. Understanding the mitochondrial physiological network in detail would be a critical step forward. Similar to the research into metabolic networks, this could be addressed by a modeling approach. Before such a framework is available, it will be desirable to monitor a comprehensive set of mitochondrial physiological parameters, as well as the dynamics in mitochondrial post-translational modifications of proteins that might be involved in signaling [recently reviewed in ref. (72)]. Suitable plant systems for such a type of analysis include promoter–reporter lines [e.g., AOX1a::GUS (77)] that have been successfully used to monitor the activity of mitochondrial signaling pathways in Arabidopsis. Promoter–reporter lines have also turned out to be valuable tools for genetic screens that led to the identification of first components of plastidial RTG signaling pathways [e.g., refs. (104, 235)]. The outcome of such screens for mitochondrial RTG signaling components is still awaited in Arabidopsis (43). Claims about specificity of a single mitochondrial signal that will be identified in the future will need to be viewed in the context of the mitochondrial physiological network. Concerted activation of multiple signaling pathways by a shift in mitochondrial signaling state and downstream crosstalk between pathways allows the required fine-tuning to guarantee efficient and flexible cellular acclimation.
Footnotes
Acknowledgments
We would like to thank Karl-Josef Dietz (Bielefeld University, Germany) for critical reading of the manuscript, and David C. Logan (Université d'Angers, France), Wilson de Paula, and John F. Allen (Queen Mary, University of London, United Kingdom) for critical reading of manuscript sections on their expert areas. M.S. is very grateful to Andreas J. Meyer (University of Bonn, Germany) for providing the time and freedom to write this article jointly with I.F. We both thank Lee J. Sweetlove (University of Oxford, United Kingdom) for open exploration and critical discussion of mitochondrial signaling over the last decade that has significantly shaped the views presented in this article. We apologize to all colleagues whose work could not be cited due to space restrictions. This work was supported by the Deutsche Forschungsgemeinschaft, Germany (Emmy Noether Programme FI-1655/1-1 and Research Unit 804).
