Abstract
Background:
The suitability of artificial intelligence (AI) and large language models (LLMs) to assist in curating real-world data (RWD) from electronic health records (EHR) for research holds transformative potential. Programmed death-ligand 1 (PD-L1) biomarker testing guides cancer treatment decisions, but results are hard to access because lab reports are unstructured and require clinical expertise to interpret. Additionally, results vary by cancer type, and documentation patterns have changed over time. This study explored the ability of LLMs to rapidly extract PD-L1 biomarker details from the EHR.
Materials and Methods:
We applied open-source LLMs (Llama-2-7B and Mistral-v0.1-7B) to extract seven biomarker details relating to PD-L1 testing from the Flatiron Health US nationwide EHR-derived database: collection/receipt/report date, cell type, percent staining, combined positive score, and staining intensity. Two approaches were used: zero-shot experiments (no fine-tuning) exploring a range of prompts and fine-tuning on manually curated answers from 500, 1000, and 1500 documents. In both cases, we validated performance using 250 human-abstracted answers across 10 cancer types. Additionally, we compared the LLM’s ability to extract PD-L1 percent staining to a deep learning model baseline trained on >10,000 examples.
Results:
We successfully used LLMs to extract biomarker testing details from EHR documents. Fine-tuned outputs consistently conformed to the desired RWD structure. In contrast, zero-shot outputs were frequently invalid and exhibited hallucinations. Fine-tuning performance improved with additional training examples. F1 scores ranged from 0.8 to 0.95, and date accuracy (within 15 days) ranged from 0.85 to 0.9. Fine-tuned LLMs exceeded the performance of the deep learning model baseline (ΔF1 = 0.05) despite the significant difference in training data.
Conclusion:
LLMs, fine-tuned with high-quality labeled data, accurately extracted complex PD-L1 test details from EHRs despite considerable variability in cancer type, documentation, and time. In contrast, zero-shot prompt extraction was not effective at the model scale examined here. Validation required access to high-quality data labeled by experts with access to the source EHR.
Introduction
In precision oncology, the deployment of artificial intelligence (AI) and large language models (LLMs) holds transformative potential, particularly in extracting and structuring real-world data (RWD) from electronic health records (EHR) for research purposes. Yet, the effectiveness of these technologies in this domain remains an area of active exploration.
Tailoring the right cancer treatment for patients often relies on testing for important biomarkers, like programmed death-ligand 1 (PD-L1), which necessitates intricate data curation due to its unstructured reporting across various cancer types and evolving documentation practices over time. Recognizing this challenge, previous work by Sushil et al. 1 evaluated the efficacy of different LLMs (GPT-4, GPT-3.5-turbo, and FLAN-UL2) in zero-shot information extraction from clinical notes. However, their study did not address potential enhancements through model fine-tuning. 1
Building on this groundwork, our study investigates the capability of LLMs to swiftly and accurately pinpoint and extract PD-L1 biomarker details within the EHR. PD-L1 testing information is critical to physician management, drug selection, trial inclusion criteria, and outcomes analyses. It is also challenging to accurately extract due to variations in documentation over time, cancer type, and the multiple different relevant results that a test yields. By employing LLMs like Llama-2-7B and Mistral-v0.1-7B, we explored both zero-shot and fine-tuned approaches to data extraction with the goal of understanding if these tools can: (1) extract PD-L1 clinical details in a readily usable and analyzable format and (2) extract multiple PD-L1 clinical details accurately and at once. Additionally, we sought to understand how LLM data extraction compared with traditional deep learning methods, utilizing a large, annotated dataset from a comprehensive oncology-specific, EHR-derived database from approximately 800 sites across the United States. This comparative analysis sought to delineate the current strengths and limitations of LLM’s ability to curate complex and domain-specific information from real-world documentation in order to highlight opportunities and areas of needed improvement, paving the way for more nuanced and effective integration of AI in clinical research workflows.
Materials and Methods
Data source
The data source for this study was the Flatiron Health Research Database: a US nationwide EHR-derived, de-identified database, which comprised patient-level structured and unstructured data originated from approximately 280 cancer clinics (∼800 sites of care) from both academic and community oncology practices (majority from community oncology settings).2,3 We included patients diagnosed with 1 of 10 cancer types between January 1, 2011, and September 31, 2023. The 10 cancer types included were nonsmall cell lung cancer (NSCLC; n = 146), gastric and esophageal cancer (n = 36), breast cancer (n = 20), bladder cancer, head and neck cancer, prostate cancer, renal cell carcinoma, melanoma, ovarian cancer, and hepatocellular carcinoma (all n < 20).
LLM extraction
We applied open-source LLMs (Llama-2-7B and Mistral-v0.1-7B) in this study.4,5 We used two approaches for LLM extraction of information from EHR. The first approach used zero-shot experiments to explore the effectiveness of a range of prompts. The second approach used LLMs that had been fine-tuned on a range of answers that were manually curated by clinical experts. Specifically, the fine-tuning was conducted for 500, 1000, and 1500 PD-L1-relevant documents (e.g., PDF result reports, pathology documents) to understand how the number of expert labels in fine-tuning affects model performance. Additionally, for the fine-tuned approach, three epochs (the number of times training data is shown to the LLM) were performed for each set of training documents (i.e., 500, 1000, 1500) to better understand LLM’s ability to improve in data extraction at a given number of training labels. Each individual evaluation was repeated once to assess for consistency in results. Fine-tuning was performed using low-rank adaptation, an established method for parameter-efficient fine-tuning of LLMs (training hyperparameters are summarized in Supplementary Table S1). 6
Evaluation
We evaluated the performance of LLM extraction in several ways. First, we assessed the ability of LLMs using both zero-shot and fine-tuned models to extract the relevant clinical details in a JavaScript Object Notation (JSON) format, even if the associated PD-L1 details were incorrect. As LLMs generate free-text outputs that are often not readily analyzable, we wanted to assess the ability of LLMs to parse clinical documents into a practical format that allows for evaluation, independent of getting the right answers or values for PD-L1 details. Second, we assessed the ability of LLMs using both approaches to not only extract PD-L1 details in a JSON format but also extract all seven clinical details accurately. For evaluation, we used expert human abstraction to create a test set consisting of 250 patients across 10 cancer types and a large network of practice sites in the United States. Human abstractors followed consistent policies and procedures and our published quality approach.3,7 To evaluate the performance of continuous variables such as PD-L1 percent staining, we bucketed the values into clinically meaningful groups and evaluated them as multiclass variables. Finally, we compared the performance of LLM extraction of PD-L1 percent staining to a deep learning model (LSTM) trained on >10,000 examples that is currently utilized in various Flatiron Health datasets.8,9 Performance was reported using precision, recall, and F1 score for multiclass variables and 15-day date accuracy for date variables. Performance was also stratified by cancer type and reported for NSCLC, gastric, esophageal, and breast cancer (for which there were at least 20 labels to assess performance) while the other cancer types were bucketed together for assessment. To ensure comprehensive reporting of methods and findings, we reviewed and fulfilled the components of the PALISADE Checklist from the ISPOR Task Force on Machine Learning Methods and Health Economics and Outcomes Research. 10
Ethics
The institutional review board of WIRB-Copernicus Group (WCG IRB) approved the study protocol prior to study conduct and included a waiver of informed consent.
Results
We successfully used LLMs to extract biomarker testing details from EHR documents. Figure 1 illustrates the workflow for unstructured EHR documents as inputs to the LLM, prompts for extraction of the test result details, and output from the models.
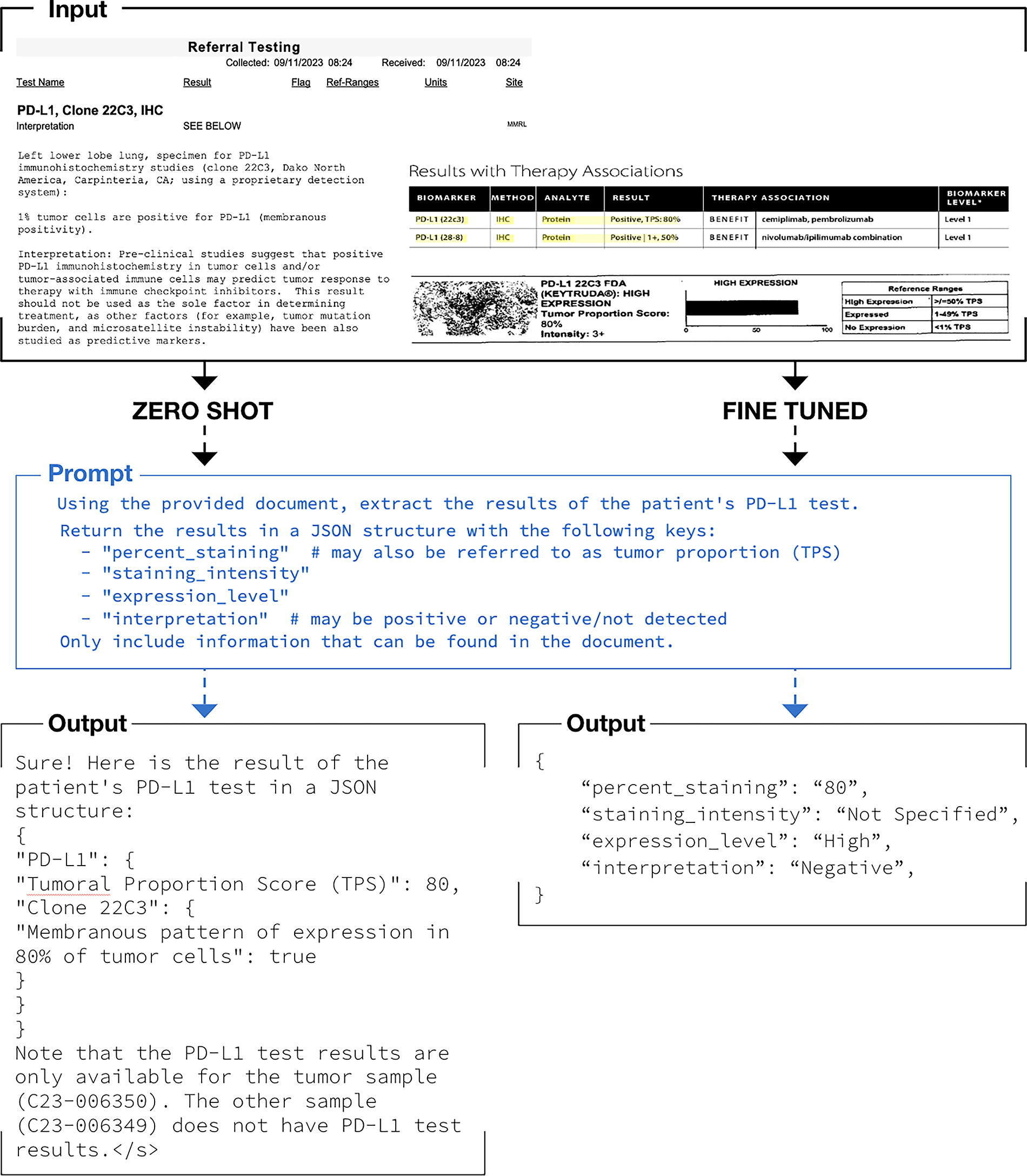

Workflow illustration for large language model extraction of oncology biomarker testing report details.
We found that the zero-show approach outputs (bottom left of Fig. 1) were largely free-text summaries of PD-L1 details and did not consistently conform to a valid JSON format. We also found the approach to be prone to hallucinations. Examples of hallucinations were the LLM repeating the prompt back, making up values, or referencing parts of the chart that were not related to PD-L1 at all. Given the inability of the 7B parameter LLMs to perform the desired task, we tested a larger LLM (Llama-2-70B) to see if the results improved. However, we again found that the model was similarly prone to hallucinations and invalid outputs (not in the desired JSON format). Additionally, the increased size of the model made computation slow. Because zero-shot LLMs did not successfully extract answers in the desired JSON format, we did not subsequently assess their ability to accurately extract the actual PD-L1 details. Conversely, the fine-tuned model output (bottom right of Fig. 1) consistently and accurately extracted all seven clinical variables in the desired JSON format. Figure 2 shows the ability of Llama-2-7B and mistral-7b-v0.1 to parse extracted details into a JSON format with valid—but not necessarily correct—PD-L1 details. Results in Figure 3 from early experimentation show that with few training examples (500 going to 1000 and then 1500), the F1 scores increase and then plateau quickly. This showcases the ability of LLMs to learn from very few labels in comparison to the many thousands of labels that are often necessary to train LSTM-based deep learning models. Running multiple epochs with the same number of training examples also resulted in improved performance, illustrating that the LLM can learn to perform the task better via increased computation without an increase in label number.
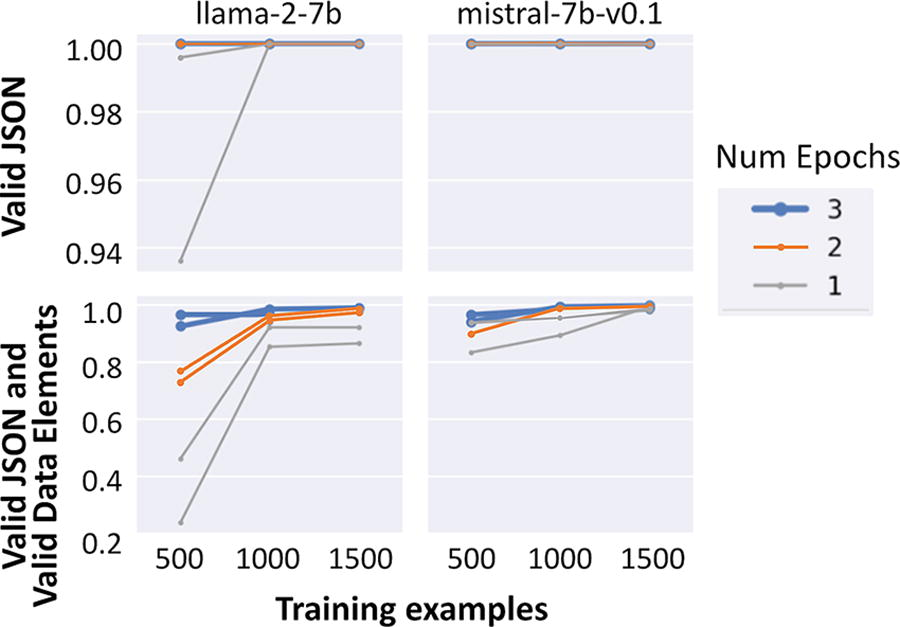
Ability to parse large language model response into JSON and valid data elements.
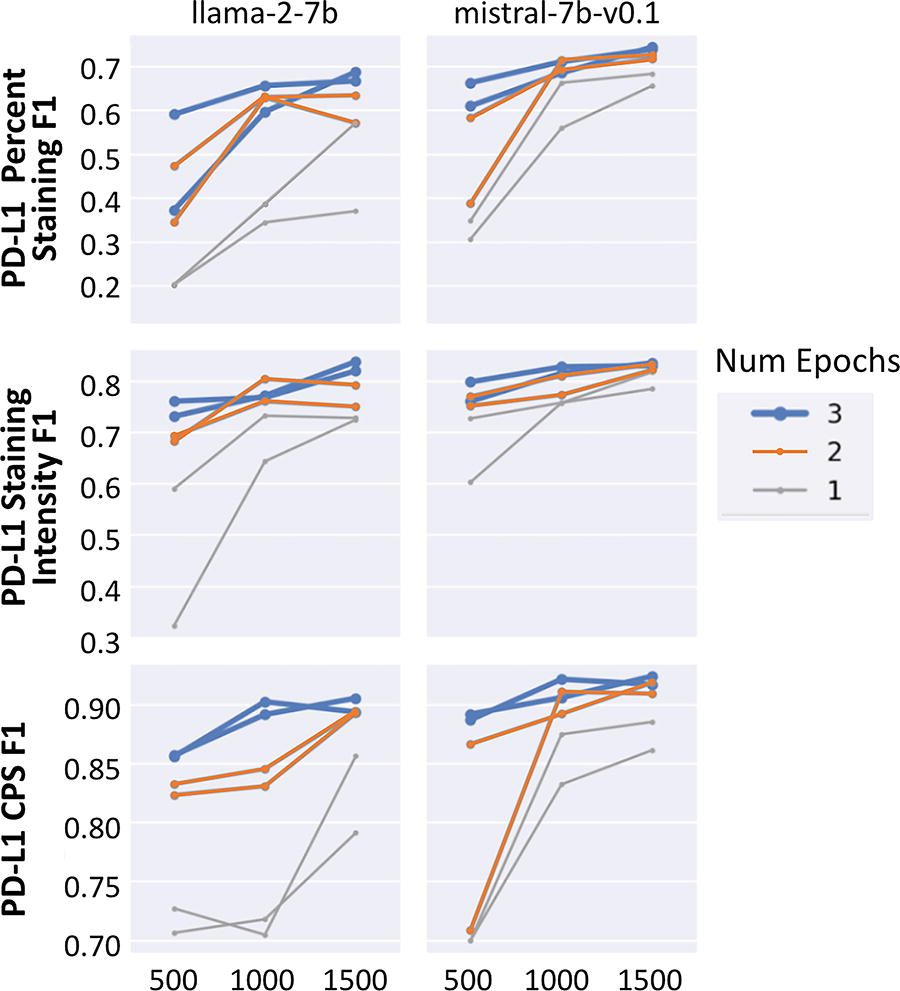
Accuracy of large language models in extracting biomarker details. CPS, combined positive score; PD-L1, programmed death-ligand 1.
The ability of fine-tuned LLMs to extract the correct PD-L1 details was then assessed. Table 1 shows the performance of extraction for PD-L1 percent staining, staining intensity, cell type, and combined positive score (CPS) and for specimen collected, received, and result date. F1 scores ranged from 0.80 to 0.95 for multiclass variables, and date accuracy (within 15 days) ranged from 0.85 to 0.90. A confusion matrix of errors shown in Figure 4 shows the PD-L1 percent staining bucketing details and reveals that the LLM’s most common failure mode is extracting 0% when the patient has evidence of a greater percent staining. We hypothesize that these errors stem from a combination of optical character recognition errors in documentation and not surfacing the correct set of contexts in the model. We also looked at the breakdown of performance across cancer types in Figure 5 and saw that while there was some variance in performance by cancer type and PDL1 variable, it was largely consistent across diseases.
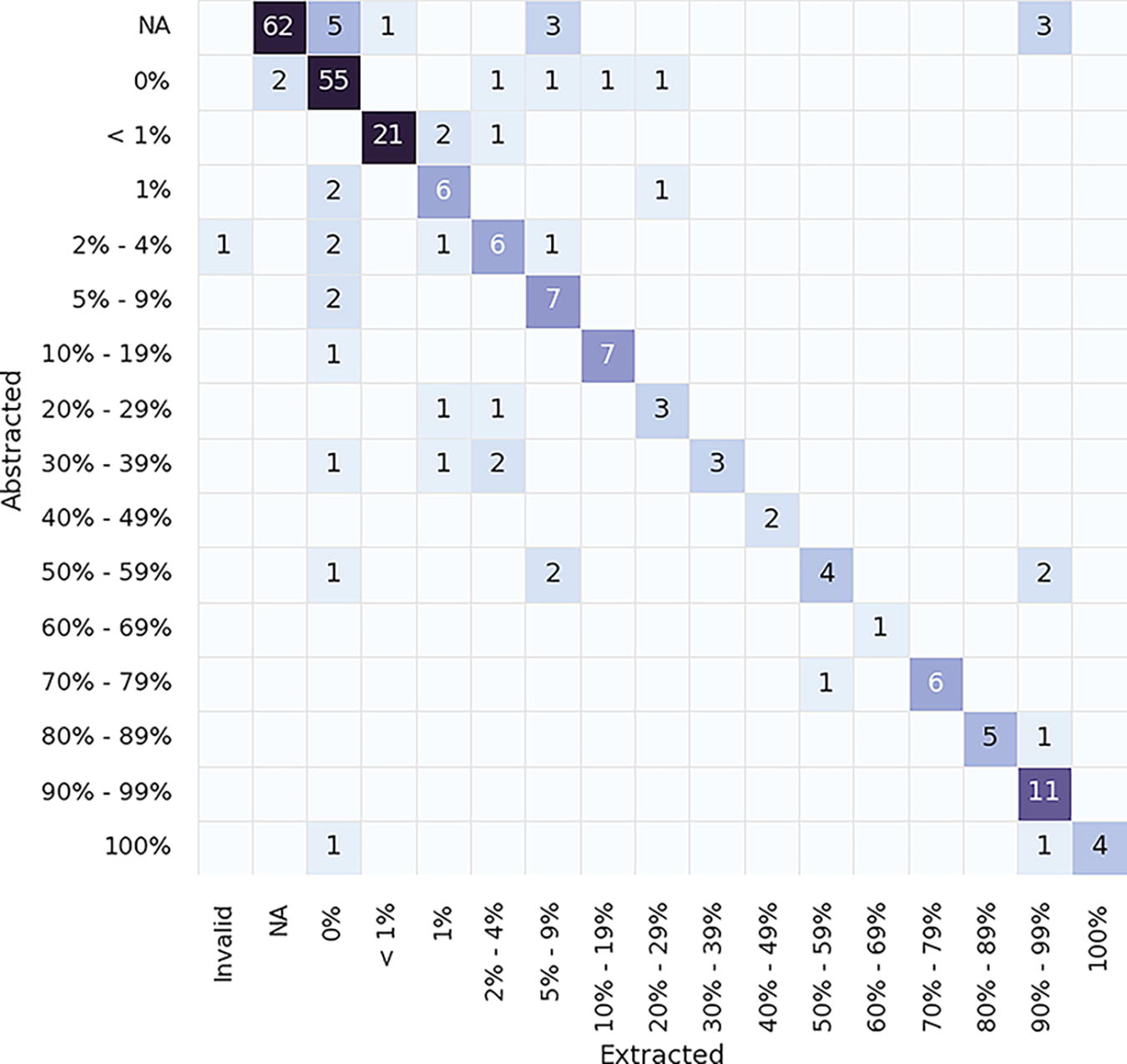
Confusion matrix showing errors between extracted LLM values (mistral-7b-v0.1) and abstracted values for PDL1 percent staining variable. LLM, large language model.
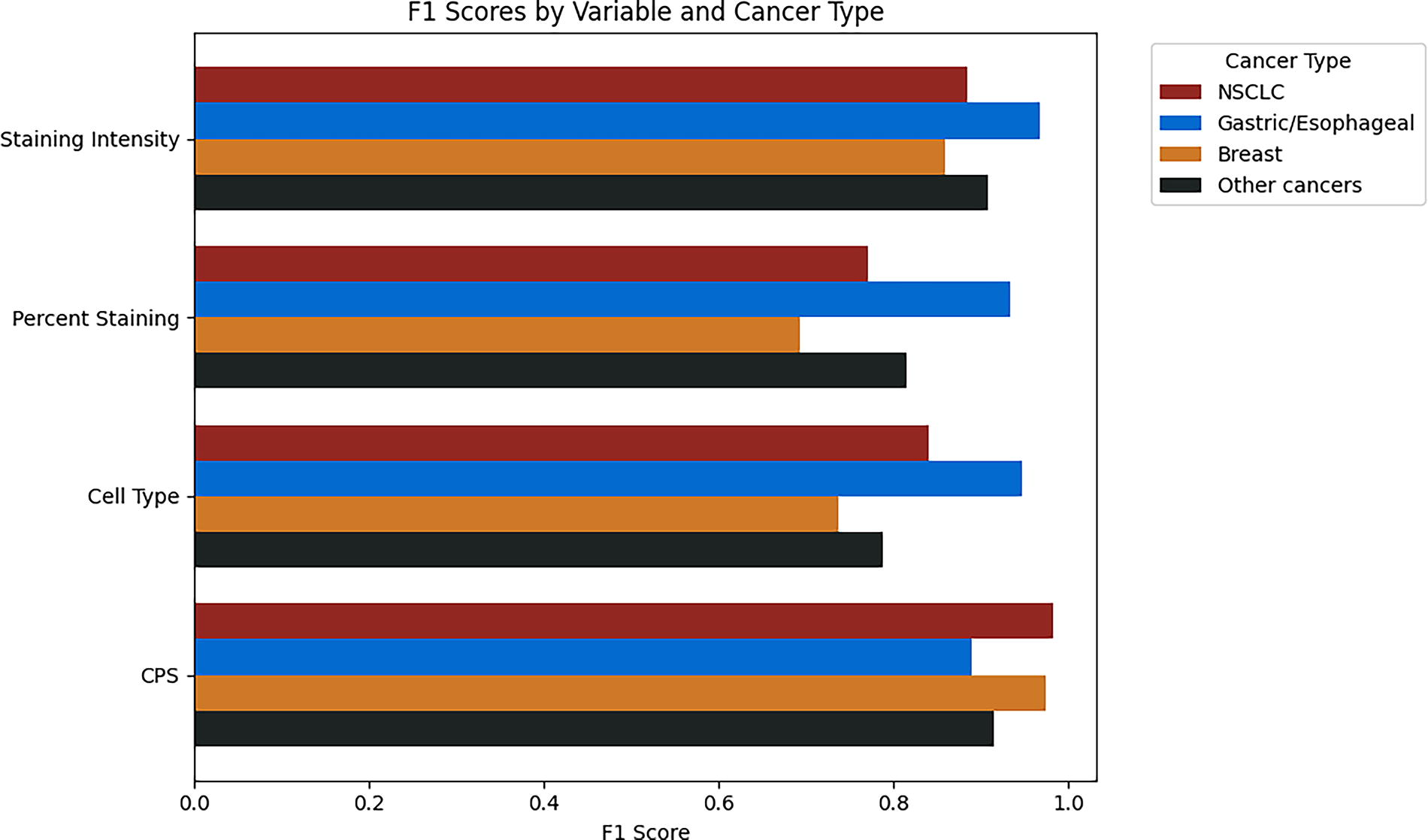
Comparison of LLM (mistral-7b-v0.1) extraction F1 performance across cancer types. NSCLC, nonsmall cell lung cancer.
Comparison of Metrics for Data Variables Between Llama2-7B and Mistral-7b-v0.1 Models
CPS, combined positive score.
A final analysis compared the fine-tuned LLM’s ability to extract PD-L1 percent staining to an LSTM deep learning model. Despite a large discrepancy in training data (deep learning trained on >10,000 labels as opposed to 500–1500 LLMs), the LLM was able to exceed the performance of the deep learning model (F1 0.82 vs. 0.77) when compared using the same dataset.
Discussion
Our findings underscore the substantial role that LLMs, particularly when fine-tuned, can play in the domain of precision oncology. This study marks a significant advance in the utilization of AI to extract complex biomarker data such as PD-L1 testing details from EHR. The comparison between zero-shot and fine-tuned LLM approaches reveals clear distinctions in performance, with fine-tuning considerably enhancing the extraction accuracy.
LLMs fine-tuned with high-quality, expert-labeled data from the EHR-derived database were able to consistently and accurately extract PD-L1 details. There are several implications of these results. First, fine-tuned LLMs demonstrated robustness to changes and variabilities in cancer type as well as documentation and results over time. Given the speed at which the standard of care in oncology changes and the way care is documented in the EHR evolves, this adaptability is crucial. Additionally, the results demonstrated that fine-tuned LLMs could (1) tackle challenging extraction tasks, such as dates, with high performance, (2) efficiently extract multiple details at once, and (3) extract in a readily analyzable JSON format. The ability of fine-tuned LLMs to extract complex and nuanced clinical details accurately and efficiently allows real-world data to be utilized for important applications, including clinical decision making at the point of care, assessing for clinical trial eligibility, better understanding disparities in testing or outcomes, and improved data management in general. Ultimately, high-quality and efficient data extraction has the potential to help clinicians provide better and more personalized care and help researchers and data scientists glean insights that were previously infeasible.
Another important finding from the study was that relatively few training labels were needed to improve the performance of data extraction. Increases of ≥500 labels led to dramatic improvements in performance. This is important as there may not be many training labels available for some applications, particularly for clinical details that are rare, such as some biomarkers. Further, this study illustrated that even in the absence of additional training labels, running multiple epochs with the same number of labels allowed the LLM to learn to perform the task better, again demonstrating that good extraction results can still be achieved when additional training labels are not available.
This is particularly noticeable when fine-tuned LLMs outperformed an LSTM deep learning model despite a large difference in training data. The LSTM deep learning model has previously been shown to have good performance and help unlock clinically important research questions, so the ability to improve upon it with fewer available labels is a promising finding that adds to the ongoing discourse regarding AI applications in healthcare and better understanding which tools and approaches are best for specific use cases.11–13
However, our study also highlights the challenges associated with AI applications in this field. The zero-shot extraction methods, although useful in broad applications, fell short in the context of detailed and specific oncological data extraction, where precision is paramount. These results are consistent with prior studies and underscore the need for tailored AI solutions that can address the specific needs of precision oncology.1,14 Future work will be needed to explore and understand how these results may change as newer and more powerful LLMs are created. This is particularly true given LLMs propensity to hallucinate while sounding confident, which highlights the importance of performing validation of these evolving capabilities. Validation tools and efforts will need to be able to keep pace with newer and more powerful AI tools, which will require access to high-quality data labeled by experts and access to the source EHR information for error analysis if possible.
There were a number of limitations of this study. The first includes the number of documents and the context size that we could input into the LLM. Given the constraints of the LLM model size, training and evaluation were performed using only specific PD-L1-relevant documents. As newer versions of LLMs are developed with larger context sizes, it will be important to understand how increased size affects performance, whether these models can accurately extract information across the entire patient chart, and the number and types of labels needed to fine-tune them appropriately. Another limitation is cost. While LLMs outperformed the deep learning model, it cost approximately 40x more to extract the information using the LLM. It will be important to continue monitoring the costs associated with using LLMs for data extraction as well as understanding the use cases that require more comprehensive and potentially better performing tools at an increased cost versus a lower one. Additionally, as larger and more powerful LLMs are able to achieve similar performance in a zero-shot manner, fine-tuning of smaller LLMs could become a more cost-efficient alternative.
While there is little reason to think that PD-L1 report documentation differs from what is captured within the Flatiron Health Research Database, it is possible that these results do not generalize to other datasets. Instead, it is more likely that the generalizability of our findings may not extend to other types of biomarkers or molecular tests, including ones that may not be standard of care. Related to this, while the test set size was adequate for assessing performance overall and across a few individual cancer types, it was not large enough to understand performance across other strata such as additional cancer types, different biomarkers, or different care sites. The breast-specific metrics were more variable than those observed across other cancer types, which may be related to the low number of examples (20) in the test set. Future work should explore how LLM performance may differ across different datasets and rarer cancer types. This will also be important for different biomarkers and their associated prevalences (particularly rare ones), testing approaches (IHC, NGS, etc.), and vendors (Caris, Guardant, etc.). Finally, the ethical implications of these powerful tools must be considered and remain top of mind as they become more widely used and performant. Processes will need to be in place to assure and maintain patient privacy. Additionally, as these tools are increasingly used for patient-facing applications such as clinical trial matching and clinical decision support, accurate and comprehensive validation approaches will be needed to ensure model performance remains fair across different patient populations, particularly those that may have been previously underrepresented in research.
Conclusion
In conclusion, to fully realize the promise of precision medicine and to truly improve care and outcomes for patients, it will be essential to have access to important clinical details that have been captured in an unstructured format as part of real-world practice. Our study lays the groundwork for future research to explore how LLMs may be leveraged to extract the information that ultimately will drive this mission.
Footnotes
Acknowledgments
Darren Johnson (
Authors’ Contributions
A.B.C., B.A., J.K.L., and G.A. contributed to the conceptualization, validation, and investigation of the study and review and editing of the article. J.K.L. and G.A. contributed to the study’s methodology, software, formal analysis, and resources. A.B.C., J.K.L., and G.A. contributed to data curation. A.C. and B.A. wrote the original draft of the article and provided project administration. A.B.C. and J.K.L. contributed to the visualization of the work. A.B.C. and G.A. provided supervision during the study.
Author Disclosure Statement
During the study period, A.B.C., B.A., J.K.L., and G.A. reported employment with Flatiron Health, Inc.—an independent member of the Roche Group—and stock ownership in Roche.
Data Availability Statement
The data that support the findings of this study were originated from and are the property of Flatiron Health, Inc., which has restrictions prohibiting the authors from making the data set publicly available. Requests for data sharing by license or by permission for the specific purpose of replicating results in this article can be submitted to
Funding Information
This study was sponsored by Flatiron Health, Inc.—an independent member of the Roche Group.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
