Abstract
Introduction:
The demand for comprehensive data resources has surged due to advancements in research aimed at tackling complex questions. Data integration techniques have evolved significantly, moving from the integration of various omics data types for a single patient to multimodal integration leveraging machine and transfer learning. This evolution has given rise to the concept of digital twins to facilitate in silico prediction of treatment response. Furthermore, the emergence of data fusion and data fabric concepts highlights the growing need for broader data connectivity, advanced analysis workflows, and supportive architectures. We introduce an AI-enabled, molecular data fabric that incorporates these innovative concepts to define a molecular information exchange for multimodal molecular-informed clinical treatment decision support.
Materials and Methods:
A patient data fabric is constructed using transfer learning for context-based integration among multiple data sources, molecular data types, and patient-derived model systems, which are all centrally connected through digital twin communities to a patient's molecular profile. Utilizing a single case research study design and retrieval augmented generation from 14 National Institutes of Health/National Cancer Institute-funded resources, our molecular fabric consists of four interconnected digital applications designed for case-based learning with therapeutic implications. These applications focus on integrating existing with emerging (e.g., noncanonical DNA structures) DNA-based markers for treatment insights. Hosted primarily on Amazon Web Services, our fabric was applied to two cases discussed within a molecular tumor board.
Results:
Our fabrication analysis of an HER2-negative, metastatic gastric cancer patient with an eleven gene signature identified three potential trials and gene targets not previously considered, with potential novel targeting via triplex structure DNA motifs. Our fabrication analyses of an EGFR-mutated small-cell lung cancer patients' 19 gene signature that matched to 68 targeted trials narrowed focus upon a single gene that led to an expanded trial option outside of the molecular report, and a non-B perspective on treatment response. Both case fabrication analyses identified patient-derived cell line digital twins for experimental testing of hypothesis generating results.
Conclusion:
We have developed a prototype molecular treatment data fabric, with AI-enabled design capabilities through digitalization to advance precision oncology. Our case applications highlight its potential to augment treatment personalization in cancer care.
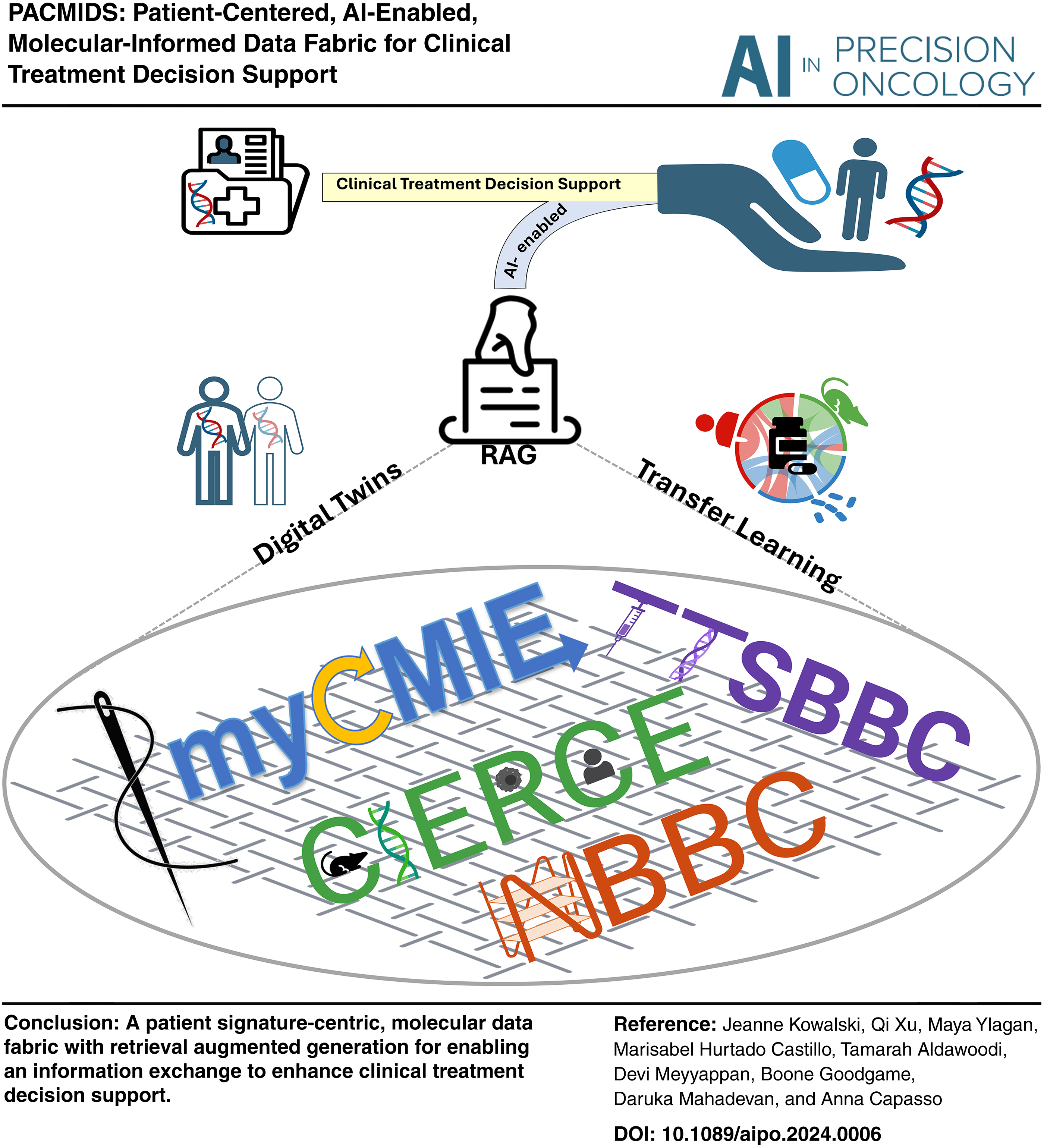
Introduction
In the fast-evolving field of health care, especially within oncology, medical professionals continue to struggle with the complexities of choosing treatment among several molecular-informed options. Traditional tools such as online repositories, clinical practice guidelines, and molecular tumor boards (MTBs), although invaluable, struggle to keep pace with the surge of new technologies, the vast amount of data, and inefficiencies in the electronic health record systems. This has spurred the development of computational tools aimed at making sense of these resources, yet their integration into clinical treatment decision support (CTDS) systems remains disjointed. In response, the concept of a “data fabric” has emerged as a revolutionary idea designed to seamlessly integrate various data sources and computational tools, enabling real-time support for CTDS. Despite the growing recognition of its potential, practical guidance on designing and utilizing this fabric for enhanced CTDS is scarce.
We introduce an AI-enabled molecular treatment data fabric, showcasing its application in creating patient-centered, computational workflows that augment CTDS. Our design incorporates four digital applications interconnected to facilitate the integration of molecular signatures through digital twins. Each application is tailored to harness specific knowledge from twin communities, enhancing resource integration and translating insights directly back to patient care. This digital transformation represents a significant leap forward in preparing resources for their AI capabilities across different health care facets within a unified data fabric. The National Cancer Institute highlights the importance of a data fabric, recognizing its potential to streamline clinical integration and navigate the uncertainties of AI implementation in cancer treatment. Our prototype data fabric with its novel applications marks a critical step toward this effort in terms of personalized, efficient, and integrated health care solutions.
Materials and Methods
This study was reviewed by the University of Texas at Austin Institutional Review Board who made the determination of non human subjects research. Written informed consent was obtained from the individual(s) for the publication of any identifiable images or data included in this article.
Resource digitalization
For AI integration to be effective, it is essential to first digitize processes and systems, ensuring seamless interaction between human inputs and AI outputs. Recognizing this, we have digitized key resources, including databases, molecular profiling reports, and public data, all with therapeutic relevance (Fig. 1). To augment in real-time, informed treatment discussions, we developed my Cancer Molecular Information Exchange (myCMIE), 1 a digital companion for molecular profile reports (MPRs). To explore DNA motif-based emerging markers with therapeutic implications, we augmented existing databases with analysis platforms through non-B Burden in Cancer (NBBC) 2 and Triplex Target Sites Biomarkers and Barcodes in Cancer (TTSBBC). 3 Motivated by the NCI's Clinical Treatment Discovery and Development program's focus on exploring nontraditional targets, we digitalized molecular signatures across patient-derived models, leading to the creation of Cancer Information Exchange for Recurring, Co-Occurring, and Co-Enriched Signatures (CIERCE). 4
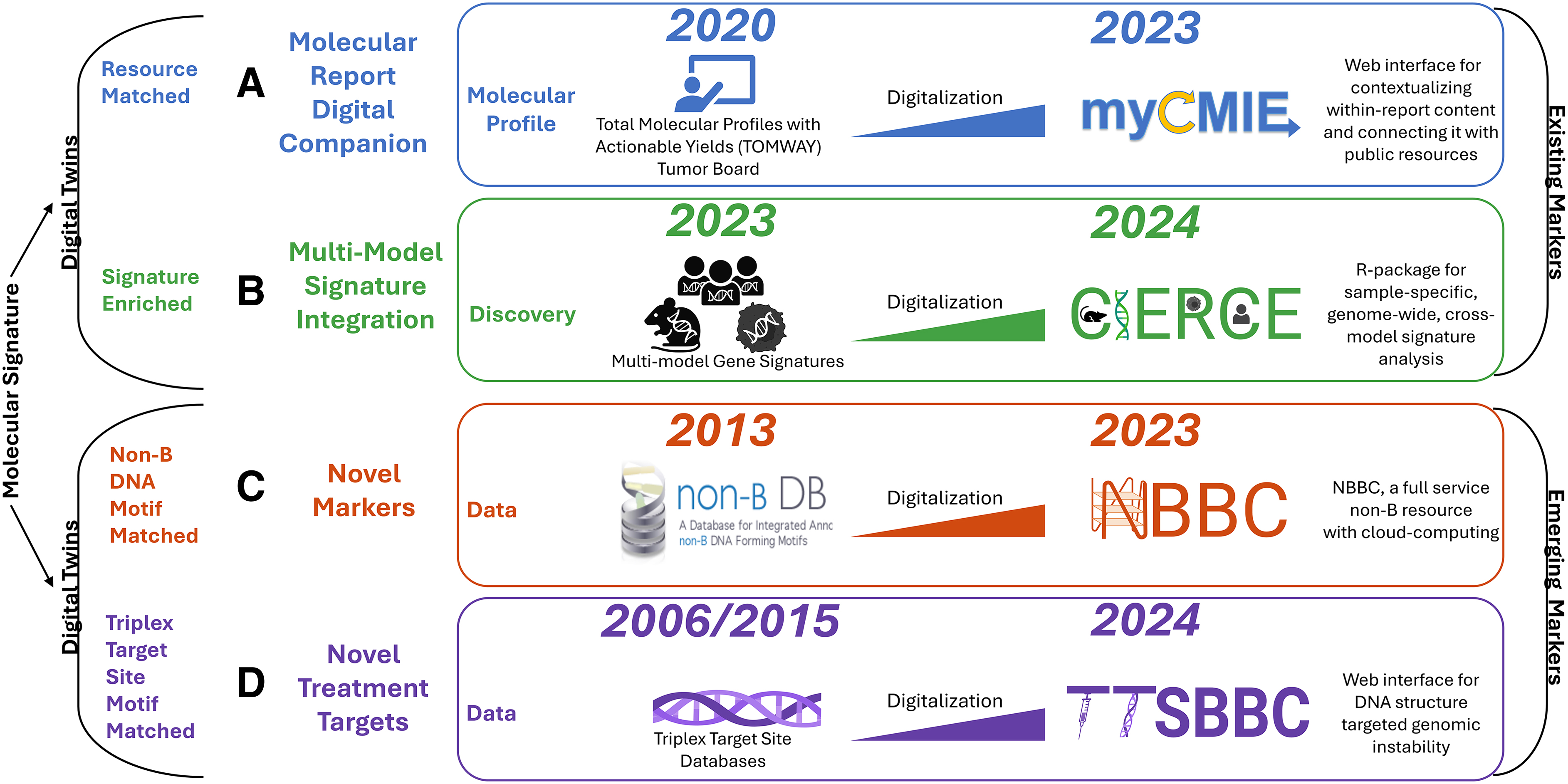
Digitalization of databases, report content, and public resources for exploration of existing and emerging markers with therapeutic implications. Each application includes date/data source initiated (on left) and its digitalization (on right).
For the purpose of our data fabric, all applications utilize the same standardized input: a patient's molecular gene signature, encompassing both primary genes and variants of uncertain significance to define their total molecular profile (Fig. 1). A patient's total molecular profile is then matched with samples and signatures using National Institutes of Health (NIH)/National Cancer Institute (NCI)-funded resources specific to each application (Fig. 2). Thus, each resource is designed to individually connect with others through patient profile matched, resource-specific digital twin communities to build a patient-centric molecular information exchange that facilitates transfer learning among resources by forming a comprehensive set of case profile-matched markers.
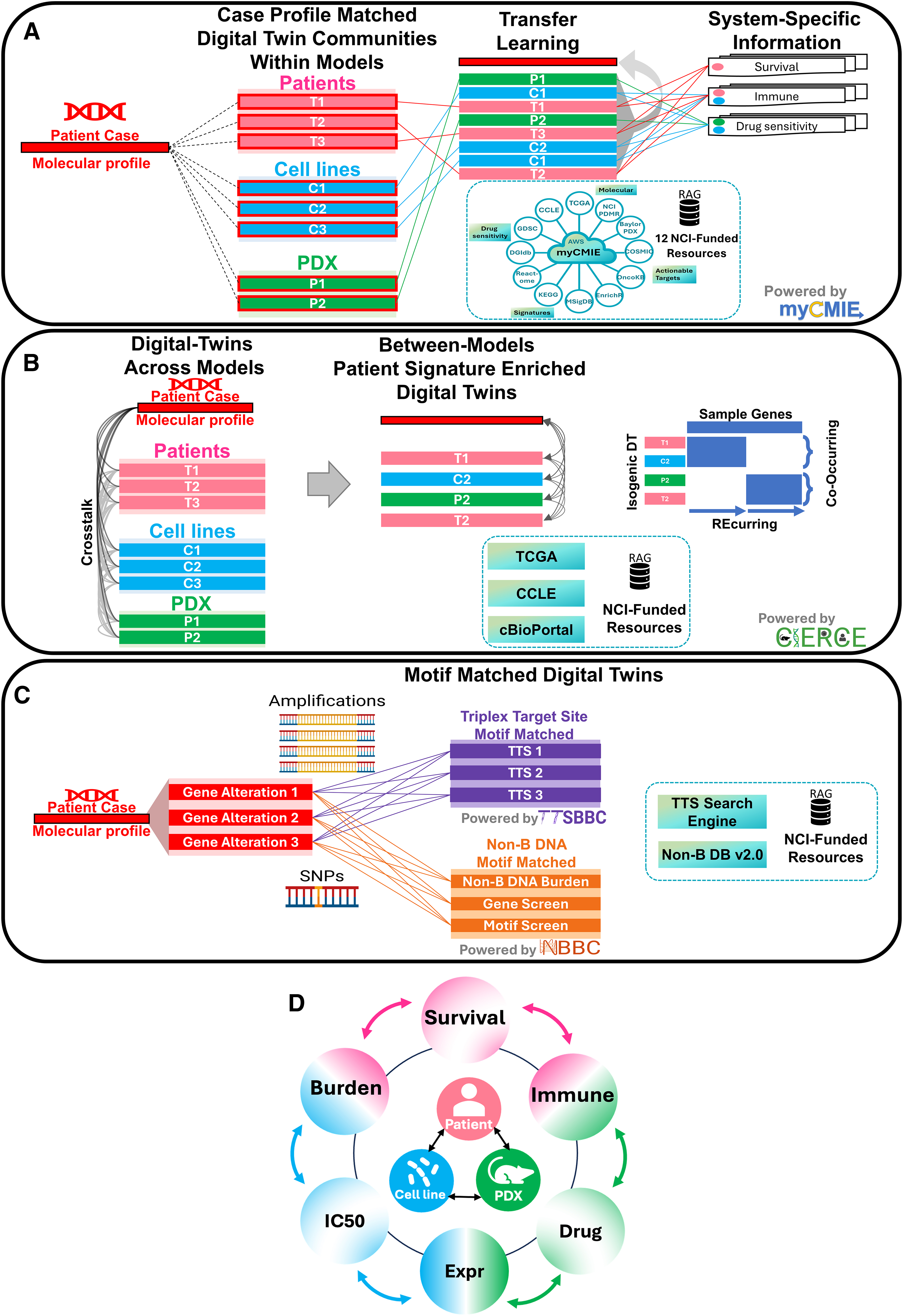
Four digital applications that define our molecular treatment data fabric to collectively form a patient profile molecular information exchange using RAG from several resources.
Resource-matched twins for context-based molecular report integration
myCMIE 1 is a platform designed as a digital companion to MPRs, aiming to connect patient case profiles with similar patient-derived models, such as other patient cases, cell lines, and xenografts. To accomplish this, myCMIE utilizes as input a patient's total molecular alteration profile and compares it with other samples' molecular alteration profile. Case profile matching is done by quantifying on a gene-wise basis, the percentage among the total profile, of gene alterations present in a comparison sample profile, and a set of digital twins defined by a user-selected threshold applied to the percentage.
Communities of digital twins that align with the input patient profile may be constructed for each populated resource (Fig. 2A) and connected through prevalence clustering that allows for the integration of multiple systems, each holding unique data on drug responses and genomic alterations. myCMIE leverages this information by initiating transfer learning from the digital twins to the patient profile, enriching the understanding derived from individual patient profiles. The transfer learning from the patient profile digital twins facilitates a patient-centered information exchange across different systems.
Signature-enriched twins for multimodel integration
CIERCE 4 is a method that includes two processes for identifying recurring and co-occurring (RECO) gene sets and samples with coenriched gene signatures (crosstalk), aimed at constructing a multimodel signature exchange. RECO, focused on the interconnected nature of genes, identifies gene groups that are recurrent across samples and co-occur within samples by using binary prevalence clustering. Crosstalk uses hypergeometric distribution-based significance testing to evaluate the enrichment of sample-level molecular signature information across multiple systems of samples. A signature-enriched digital twin community is defined by identifying samples across multiple resources significantly coenriched with the patient signature. A particular application of crosstalk used in our data fabric is to facilitate the cross-system sample-pair coenrichment testing to define patient-matched isogeneic digital twins across systems.
Motif-matched twins for non-B DNA signature exploration
Non-B DNA assessment involves two main platforms, TTSBBC 3 and NBBC, 2 to identify the presence and prevalence of non-B DNA motifs colocalized with a gene signature. To further augment these platforms, we have developed a statistical approach, MoCoLo (Motif Co-Localization), for significance testing of colocalization. 5 Triplex Target Sites (TTSs) have been demonstrated to impact transcription and DNA repair processes for targeting specific genes or regions.6,7 For a given input gene signature, TTSBBC 3 specializes in searching over 37 million TTS motifs by structured query language database retrieval to inform on site colocalization and sequence feature metrics for their prioritization. By comparison, for a given input gene signature, NBBC 2 screens over 11 million non-B DNA forming structure motifs, quantifying their presence as a non-B burden (a metric similar to tumor mutation burden [TMB]) by counting the number of non-B motifs present in a mutated gene region, normalized by non-B library size and region length.8,9
In cancers with low TMB, analyzing non-B TMB (nbTMB) could offer insights into varying patient outcomes, as demonstrated in our research on early-stage pancreatic cancer patients. 8 Together, these platforms support the exploration of noncanonical DNA structure motifs colocalized with a patient's molecular gene profile to broadly construct motif-matched twins.
Data sources
All data sources, their definitions, and greater details on resource capabilities are embedded within each published resource. Our retrieval augmented generation (RAG) system incorporates 14 NIH/NCI resources, including databases and knowledge bases (Fig. 2), as well as manual searches for articles, guidelines, and conference abstracts.
Figures
All figures were generated by, or in part, using biorender.com
Molecular tumor board
Each case was discussed during our continuing medical education-credited, Livestrong Cancer Institutes (LCI) MTB and joint, LCI-Mays Cancer Center (MCC) MTB, and in a real-time MTB during the 2024 University of Texas Health San Antonio Practical Applications of New Agents in Oncology conference.
Results
Case 1: HER2-negative, metastatic stomach adenocarcinoma
The National Comprehensive Cancer Network (NCCN) recommends 5-fluorouracil and oxaliplatin (FOLFOX) combined with nivolumab as a first-line treatment for HER2-negative, PD-L1-positive, metastatic stomach adenocarcinoma (mSTAD), with FOLFOX showing a 40–50% response rate in gastrointestinal cancers. 10 We report on a young adult with stage IV mSTAD who initially benefited from FOLFOX but later experienced recurrence. Given the high risk of treatment failure, a referral was made to the LCI MTB at the initial response stage for consideration of future treatment strategies.
The patient's clinical sequencing revealed 13 alterations, seven of which were variants of unknown significance (VUSs), with limited options for local trials. By integrating both key alterations and VUSs into a comprehensive molecular profile, we utilized the platforms myCMIE, 1 CIERCE, 4 and TTSBBC, 3 enhanced by RAG, for patient-centric fabrication analyses (Fig. 3). This approach led to identifying additional treatment options beyond those identified with existing CTDS, as contained within the MPR, in addition to potential options that require further testing of a single gene's expression and potentially novel targets for exploration.
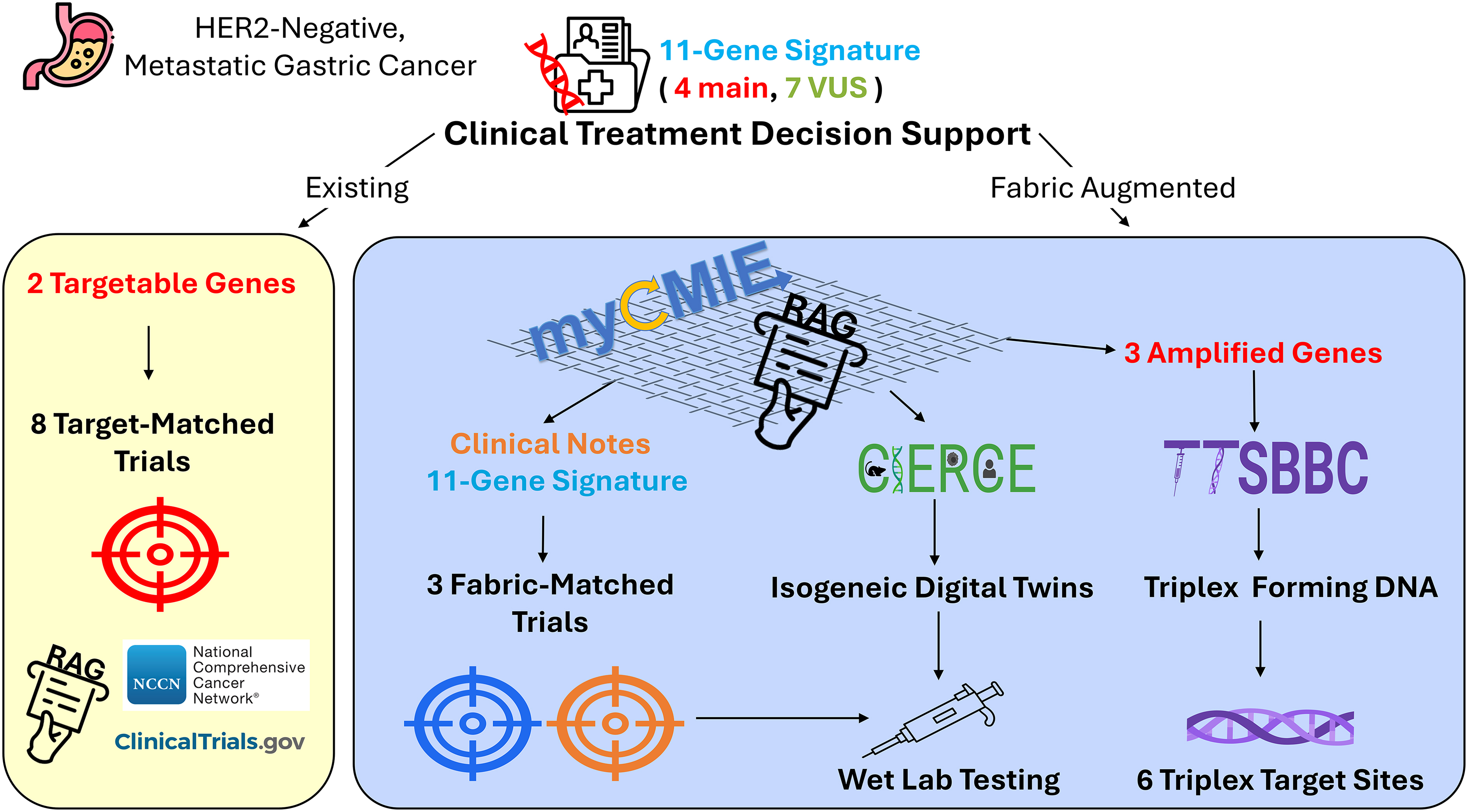
Overview of results from existing and fabric-augmented CTDS for an HER2-negative, metastatic gastric cancer case. The existing CTDS (in yellow) includes the two targetable genes based on the clinical-grade sequencing report that matched to eight clinical trials using ClinicalTrials.gov database and NCCN guidelines. Our fabric augmented CTDS (in blue) uses our data fabric consisting initially of myCMIE and RAG to streamline focus among the molecular signature and clinical notes resulting in the identification of three expanded trial options that could be tested using CIERCE, and potentially novel targets to explore based on the three amplified genes identified as part of the molecular report. CTDS, clinical treatment decision support; NCCN, National Comprehensive Cancer Network.
Presentation
A 45-year-old male with a history of seizure disorder and iron deficiency anemia presented with progressive dyspnea and fatigue, noting black stools since starting iron supplements. Despite no other bleeding symptoms, he was found to have critically low hemoglobin of 3.1 g/dL and severe iron depletion. After emergency treatment with blood transfusions and iron therapy, he was advised to stop using emoxypine succinate, a seizure prevention medication that chelates iron. Persistent symptoms led to further investigations, revealing left supraclavicular lymphadenopathy and a mass at the gastroesophageal junction, diagnosed as mSTAD, confirmed by a repeat left supraclavicular lymph node fine-needle aspiration biopsy. Immunohistochemistry testing revealed HER2-negative and PD-L1-positive status. The patient began FOLFOX treatment, augmented by palliative radiation and nivolumab, which was later discontinued due to severe dermatitis.
Initial chemotherapy showed a favorable response, but subsequent scans after chemoradiation indicated recurrence, with carboplatin and irinotecan later initiated. A tissue specimen preinitial FOLFOX treatment was sent for molecular testing (FoundationOne CDX). A post-FOLFOX treatment blood biopsy at progression (Guardant360) revealed a KIT exon 10 mutation. Tissue genomic profiling revealed microsatellite stability with low TMB.
MTB discussion
Among the eight existing CTDS, MPR listed clinical trials, six were local that exclusively targeted MDM2 (Fig. 4B). By performing an updated trial search and expanding the search for targeted trials to include VUS findings, we were able to expand treatment options to an additional 12 (one overlapped with the MPR), for a total of 17 actively recruiting trials (data not shown). Among them, five were Phase 2 local trials that were further vetted and included the TAPUR trial (NCT0269355), offering several treatment options based on VUS's (PDGFRB, ATM, STK11) and KIT mutation. While not local, NCT05694715 was the most recent trial to open for treatment with niraparib and irinotecan in metastatic solid malignancies with ATM, BRCA1/2, or PALB2 mutations.

Clinical-grade comprehensive genomic profiling results using existing CTDS augmented with myCMIE analysis identified eight, main-finding targeted trials that based on their profile-matched digital twin interrogation reveals an opportunity for improved treatment response by further exploration of therapies.
We explored MPR- and VUS-expanded trial options alongside the patients' NCCN-guided treatment history by creating STAD cell line digital twin case profile matches and interrogating compounds among them (Fig. 4C). Overall, cell line case profile matching showed co-occurring GNAS and ZNF217 amplifications primarily separate from co-occurring VUSs. Our case-matched IC50 analyses showed the patient treatment history with moderate to high IC50s relative to all available compounds tested, which, considering the patient's recurrence, supports the viability of our digital twin approach. Compared with the patient's NCCN guideline-based treatment history, experimental treatments with poly-ADP ribose polymerase (PARP) inhibitors failed to show a marked improvement in response. Among the current relevant targeted treatment options from the TAPUR trial (sunitinib/regorafenib for mutated PDFGRB, regorafenib for KIT, talazoparib for ATM, and temsirolimus for STK11), sunitinib showed promise but with an IC50 similar to irinotecan, which the patient did not respond, motivated the need to explore additional treatment options.
Our case total profile context analyses with myCMIE 1 revealed enrichment of PI3K-Akt and FAK pathways connecting main and VUS genes (Fig. 5D). We interrogated the expression of case profile genes alongside select genes in these pathways and up- and downstream from them (data not shown). A cluster analyses revealed several coexpressed genes among the case-matched cell lines that included SRC, integrins, AKT, RHOA, PTK2, and FGFR2 genes, supporting potential FAK/Src complex activation that plays a critical role in the epithelial–mesenchymal transition associated with solid tumor metastasis. 11 We interrogated multityrosine kinase inhibitors (TKIs), separated by their targeting, which overall showed an improved response compared with both the patient history and case-matched trials (Fig. 5B). Two drugs in particular, ponatinib that includes FGFR targeting and dactolisib for PI3K-Akt, had the lowest IC50s among all the interrogated compounds.
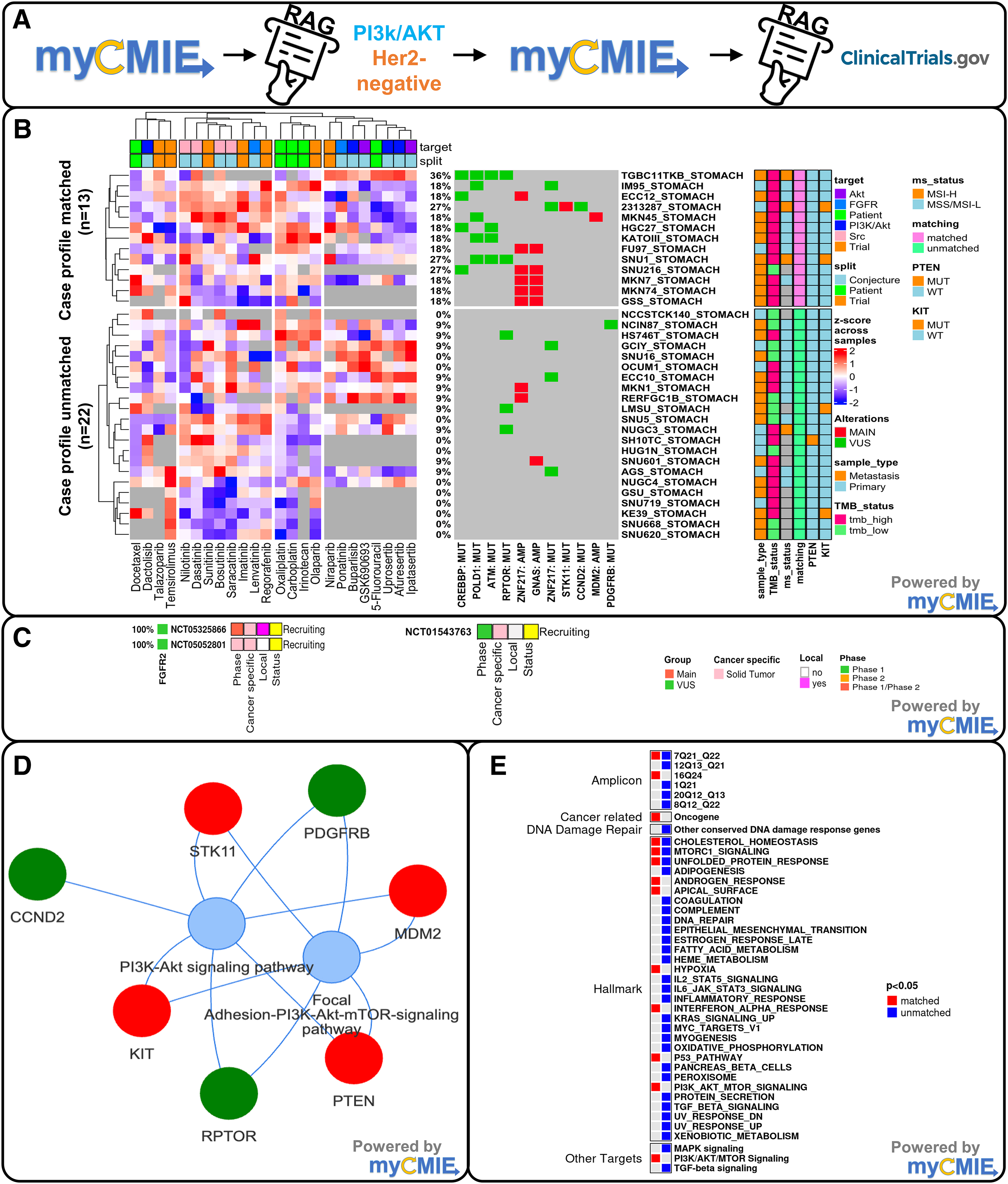
Fabrication analyses led to the identification of expanded trial matching that revealed compounds with increased sensitivity compared with both patient history and patient profile-targeted trial matching between the case profile matched versus unmatched STAD cell lines using an RAG-augmented, myCMIE platform.
Among the genes examined, FGFR2 expression testing may be considered, since if positive, further expands treatment options to include the study of bemarituzumab combined with mFOLFOX6 in gastric or gastro-oesophageal junction cancer (FIGHT) trials (Fig. 5C). These trials are expansions of the initial phase II FIGHT trial, 12 which reported that adding bemarituzumab, a first-in-class anti-FGFR2b monoclonal antibody to mFOLFOX6, improved outcomes for patients with untreated unresectable locally advanced or metastatic HER2-negative gastric/GEJ adenocarcinoma and FGR2b (an FGFR2 splice isoform) overexpression or gene amplification. Additional case-matching profile analyses applied to all and metastatic pancan cell lines showed a similar relative order of compound sensitivity (data not shown). Interestingly, case profile-matched alterations showed significant enrichment of hypoxia and PI3K-Akt.
As a member of the tyrosine kinase receptor (TKR) family, KIT activation initializes downstream signaling of several pathways, including JAK-STAT3 and PI3K-Akt-mTOR, which were significantly enriched in the case profile, and the latter, in the case-matched digital twins (Fig. 5E). KIT mutations that encode TKR functional domains have shown to be effective with imatinib. 13 In patients with KIT exon 10 mutations, imatinib has not been formally studied and remains divided.14–17 The phase III INTRIGUE trial of advanced gastrointestinal stromal tumor patients who had progressed on or developed intolerance to imatinib reported the emergence of heterogeneous secondary KIT mutations occurring in adenosine triphosphate (ATP)-binding pocket or activation loop, of which the former had a clinical benefit to sunitinib and the latter to ripretinib. 18
Our pancan analysis of 14 KIT mutated cell line analyses showed ∼70% with mutations in TKR functional domains, with two patients having a KIT exon 10 mutation, and while imatinib sensitivity varied among KIT-mutated locations, ponatinib IC50 remained consistently low (data not shown). The rarity of the KIT exon 10 mutation adds to a seemingly unique case molecular profile overall.
Using the myCMIE platform, 1 we additionally explored insights from case profile matching to TCGA STAD patients (data not shown). Within these 29 Digital Twins (DT)+ patients, TMB-high, microsatellite instability high patients, characterized mainly by co-occurring case VUSs, 6 had significantly (p = 0.04) longer overall survival compared with TMB-low, microsatellite stable (MSS) patients characterized by dual GNAS and ZNF217 amplifications. The case presented with TMB low and MSS, in addition to three key finding gene amplifications. Among them, we focused upon ZNF217 based on RAG informed pathways downstream, many of which were significantly enriched in the case total profile (Fig. 6B). Using our TTSBBC platform, 3 we identified six triplex forming oligonucleotide sites to explore ZNF217 targeting (Fig. 6C). Using our computationally identified DNA signature similar cell lines that differ by enrichment of the case profile (Fig. 6D), TTS and other fabric-informed treatment option discoveries may be efficiently translated back to bench for their further study.
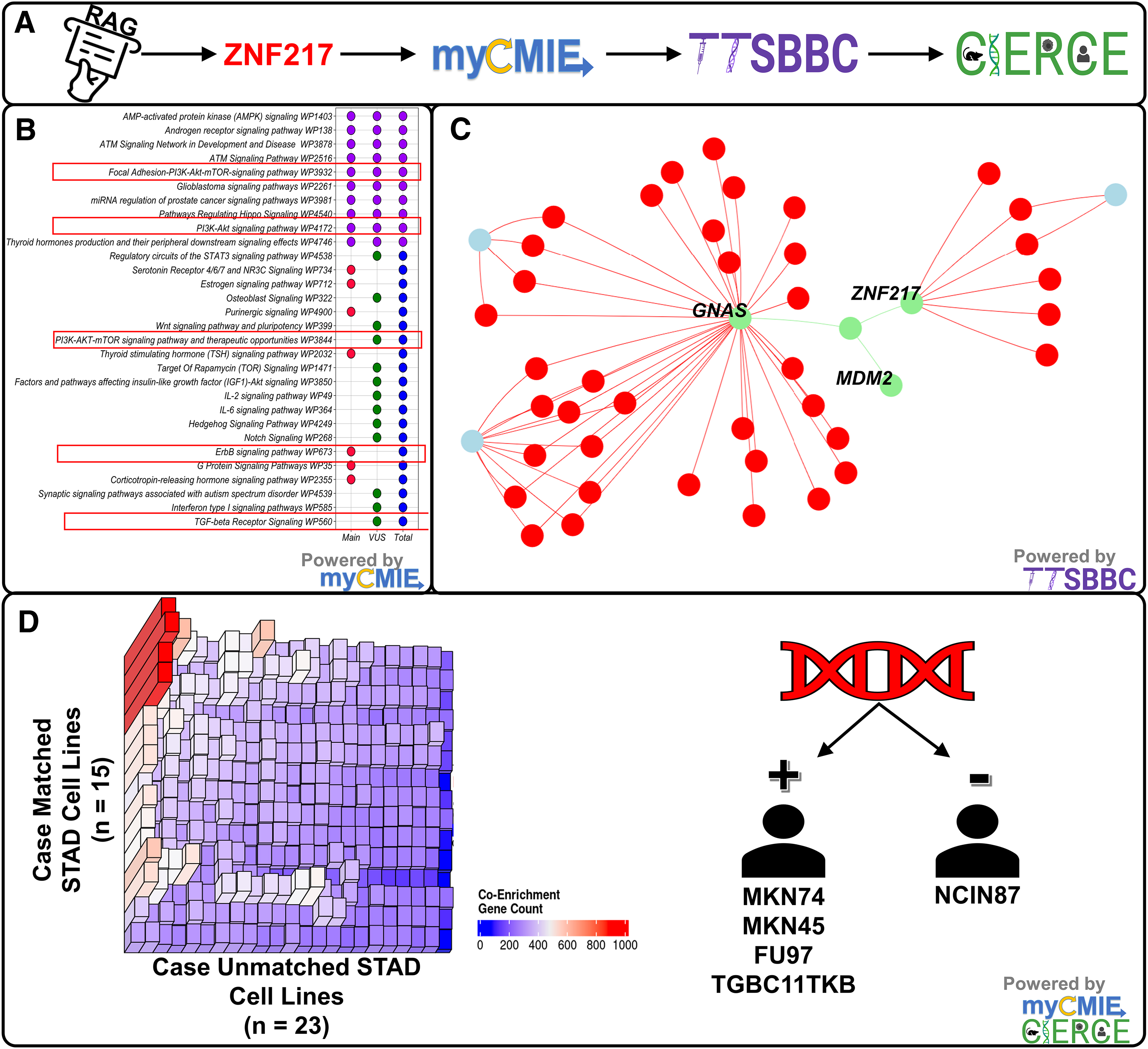
Fabrication analyses led to the focus upon a main gene finding, ZNF217, revealing several downstream pathways that are enriched in the case profile, which may be explored for targeting using DNA triplex structures and tested with treatment using in silico, digital twin isogeneic cell lines matched and unmatched with the case profile.
In addition, we identified a relatively high inferred, within-sample abundance of M2 macrophages across DT+ TCGA STAD patients with corresponding high CD4 memory resting and CD8 T cell abundance, depending upon respective key and VUS occurrence (data not shown). Using TCGA pancan to case profile match, co-occurring VUSs depicted matches of 50% or greater, while the separate co-occurrence of key or VUS findings described metastatic patient profile matches. Among the ∼11,000 TCGA pancan patients, we were unable to identify samples with representative co-occurrence of both key and VUS case gene alterations. Thus, the patient tumor molecular profile is seemingly unique to TCGA STAD and TCGA pancan.
Our case profile context-based integrated analyses offer novel conjectures with therapeutic implications. First, M2 macrophage abundance among TCGA case-matched samples supports a possible M2 phenotype. M2-type macrophages play a significant immunosuppressive role that includes expression of surface ligands, PD-L1, which the patient tested positive. Considering the proinflammatory properties associated with M2 macrophages, this phenotype is further supported by the patient not tolerating immunotherapy.
Hypoxia can be a key driver of macrophage recruitment and polarization in the tumor microenvironment (TME) and is common in most solid tumors 19 and was enriched in the case-matched cell lines (Fig. 5E). HMGB1 has been reported with hypoxia-induced macrophage polarization 20 and was expressed in 18 of the 29 TCGA STAD case-matched patients. If supported through additional studies, inhibition of M2 polarization by a CSF1R inhibitor may warrant further study. Second, FAK and TME are intricately linked, with FAK signaling implicated in TME remodeling and the progression of most cancer types that with an M2 phenotype may affect treatment choice. 21
Outcome
The patient achieved a complete response after completing FOLFOX that was followed by recurrence. The profile-linked analyses offered expanded treatment options not otherwise included in their MPR. Since testing, the patient was hospitalized, and subsequently placed on second-line therapy with FOLFIRI (leucovorin calcium (folinic acid), fluorouracil, and irinotecan hydrochloride) and is not responding. Consideration is being given to a multi-TKI, testing for FGFR2b expression, and a biopsy for combined DNA and RNA comprehensive genomic profiling.
Case 2: EGFR-mutated, small-cell lung carcinoma
Small-cell lung cancer (SCLC) accounts for <15% of lung cancer cases, with EGFR mutations being uncommon. 22 While EGFR TKIs are standard first-line therapy for metastatic non-SCLC, their effectiveness in SCLC is less clear, with chemotherapy (platinum and etoposide) showing response after non-SCLC transformation to SCLC. 23 We present a case of stage IV, metastatic SCLC with an EGFR mutation. Given the uncertain efficacy of EGFR TKIs in treating EGFR-mutated SCLC, a referral was made to the LCI-MCC MTB to evaluate treatment options. Clinical sequencing identified 19 alterations, including 12 VUSs, presenting 73 trial options primarily based on EGFR TKIs and PARP inhibitors. Leveraging a comprehensive analysis through platforms, myCMIE, 1 CIERCE, 4 and NBBC, 2 enhanced by RAG, we identified an expanded treatment beyond conventional targeting, suggesting a tailored approach for this patient's care (Fig. 7).
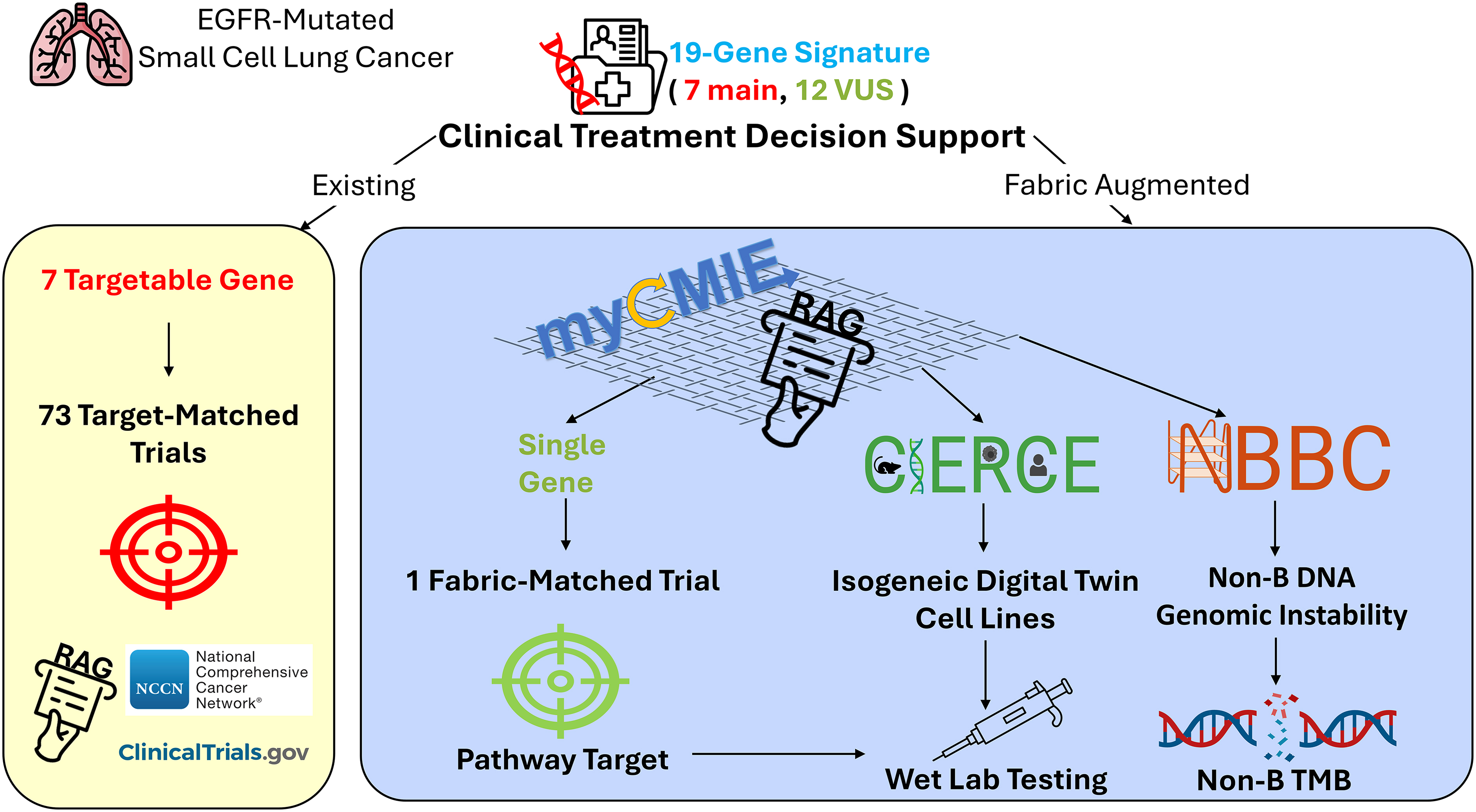
Overview of results from existing and fabric-augmented CTDS for an EGFR-mutated, SCLC case. The existing CTDS (in yellow) includes seven targetable genes based on the clinical-grade sequencing report that matched to 73 clinical trials using ClinicalTrials.gov database and NCCN guidelines. Our fabric-augmented CTDS (in blue) uses our data fabric consisting initially of myCMIE and RAG, led to the focus upon a single gene VUS, resulting in an expanded trial option that may be further explored using CIERCE for selection of computationally derived isogenic cell lines with and without the case-matched profile, and interrogation of non-B DNA burden for treatment response. EGFR, epidermal growth factor receptor; SCLC, small-cell lung cancer.
Presentation
A 73-year-old Caucasian female presented to the emergency department with persistent cough without sore throat, chest pains, and shortness of breath. She had reported no loss of appetite or weight loss. A positron emission tomography (PET) scan showed a hypermetabolic left lung superior segment 4.0 × 3.3 × 3.3 cm lobulated mass extending from the perihilar region to peripheral lung, with suspected extension to the pleura and extensive mediastinal hypermetabolic lymphadenopathy. A biopsy indicated poorly differentiated carcinoma with neuroendocrine differentiation that favors high-grade neuroendocrine carcinoma, likely pulmonary. Endobronchial ultrasound and mediastinal sampling and node 7 and 4R showed poorly differentiated carcinoma of neuroendocrine origin. Bronchoalveolar lavage from left lower lobe was negative. Patient was treated with cisplatin plus etoposide plus durvalumab. Imaging showed a favorable treatment response. Postconsolidation radiation to the lung, the patient resumed maintenance immune checkpoint therapy.
A PET scan showed progressive disease with a new lesion and was treated with lurbinectedin until progression. Patient was rechallenged with carboplatin plus etoposide plus atezolizumab due to prior good response. Subsequently, the patient received radiation therapy (5 fractions) to the left parietal lesion with extraosseous extension with bulky epidural and scalp components. Patient enrolled in the Dialectic Trial (BCLXL-Protac- DT2216, NCT04886622) and completed 2 cycles of therapy. A cervical lymph node specimen postinitial treatment with cisplatin + etoposide + durvalumab, and lurbinectedin at disease progression was sent for molecular profiling (Caris Life Sciences). A subsequent circulating tumor DNA (ctDNA) biopsy postplatinum and was sent for molecular profiling (Guardant360). An EGFR L858R mutation was detected in both tissue and ctDNA, with an additional EGFR R776C mutation detected in ctDNA. Tissue genomic profiling revealed microsatellite stability with low TMB.
MTB discussion
The 19 gene signature case molecular profile resulted in 73 targeted trials based on existing CTDS, as contained in the MPR. Among the trials, EGFR TKIs and PARP inhibitors were the main findings. We explored these inhibitors alongside the patients' NCCN-guided treatment history by creating SCLC cell line digital twin case profile matches (DT+) and interrogating compounds among them (Fig. 8C). Notably, carboplatin is shown with the highest IC50, both among the patient's treatment history and overall, among all compounds with available IC50s. EGFR-TKIs vary in terms of their IC50s, with osimertinib displaying as the most sensitive among those interrogated on the SCLC DT+ cell lines. In a case report of an EGFR-mutated SCLC patient treated with osimertinib, the patient failed to show meaningful improvement and the disease continued to progress, although the TKI was not used in combination with cytotoxic chemotherapy. 24
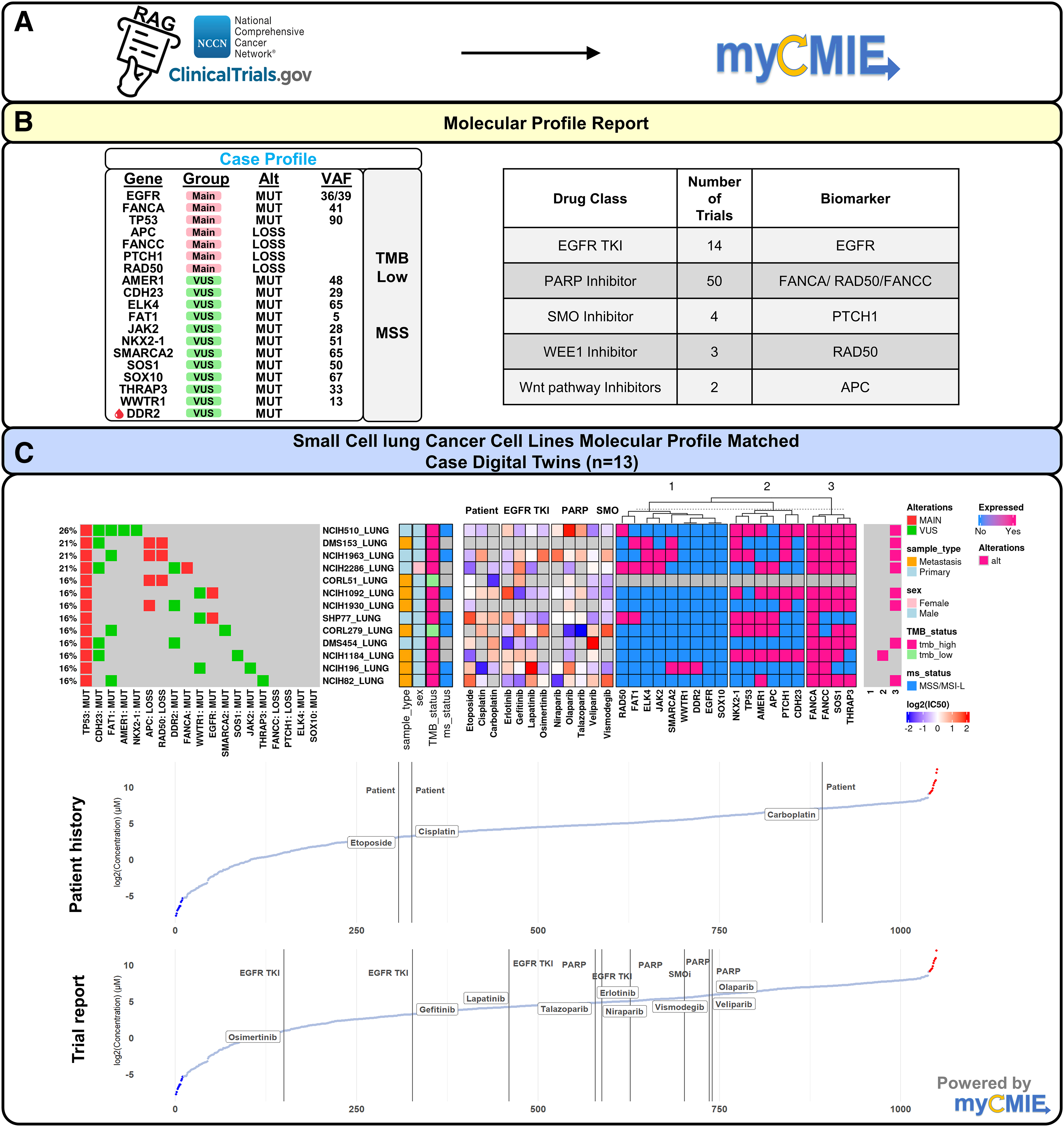

Clinical-grade comprehensive genomic profiling results using existing CTDS augmented with myCMIE analysis identified 73, main-finding targeted trials that based on their profile-matched digital twin interrogation reveal an opportunity for improved treatment response by further exploration of therapies.
Our SCLC DT+ cell line analysis indicated a moderately high IC50 for such therapies of which the patient did not respond, lending support for our approach. Collectively, these observations motivated the need for improved exploration of therapies that ideally display IC50s lower than osimertinib.
In an effort to streamline the 19 gene signature case profile, we utilized our CIERCE method to identify potential novel subsignatures enriched between TCGA pancan EGFR-mutated progressors and SCLC cell lines (Fig. 9B). This crosstalk analyses identified SMARCA2 and JAK2 as frequently occurring together among these two patient-derived model systems. Using myCMIE to further contextualize this result, we found these two genes, along with FANCC and PTCH1, to reside on chromosome 9. SMARCA2 is part of the ATPase catalytic subunit in the SWItch/Sucrose Non-Fermentable (SWI/SNF) pathway that has been shown to affect several downstream pathways, 25 some of which are enriched in our case signature (Fig. 9C). RAG-informed research revealed the potential for SWI/SNF to promote TKI resistance in EGFR-mutant lung cancer. 26 The study showed that resistant tumors rely on mutated SWI/SNF complexes and that pharmacologic inhibition of mSWI/SNF ATPase activity (SMARCA2/4 inhibitor) sensitized a patient-derived tumor to osimertinib.
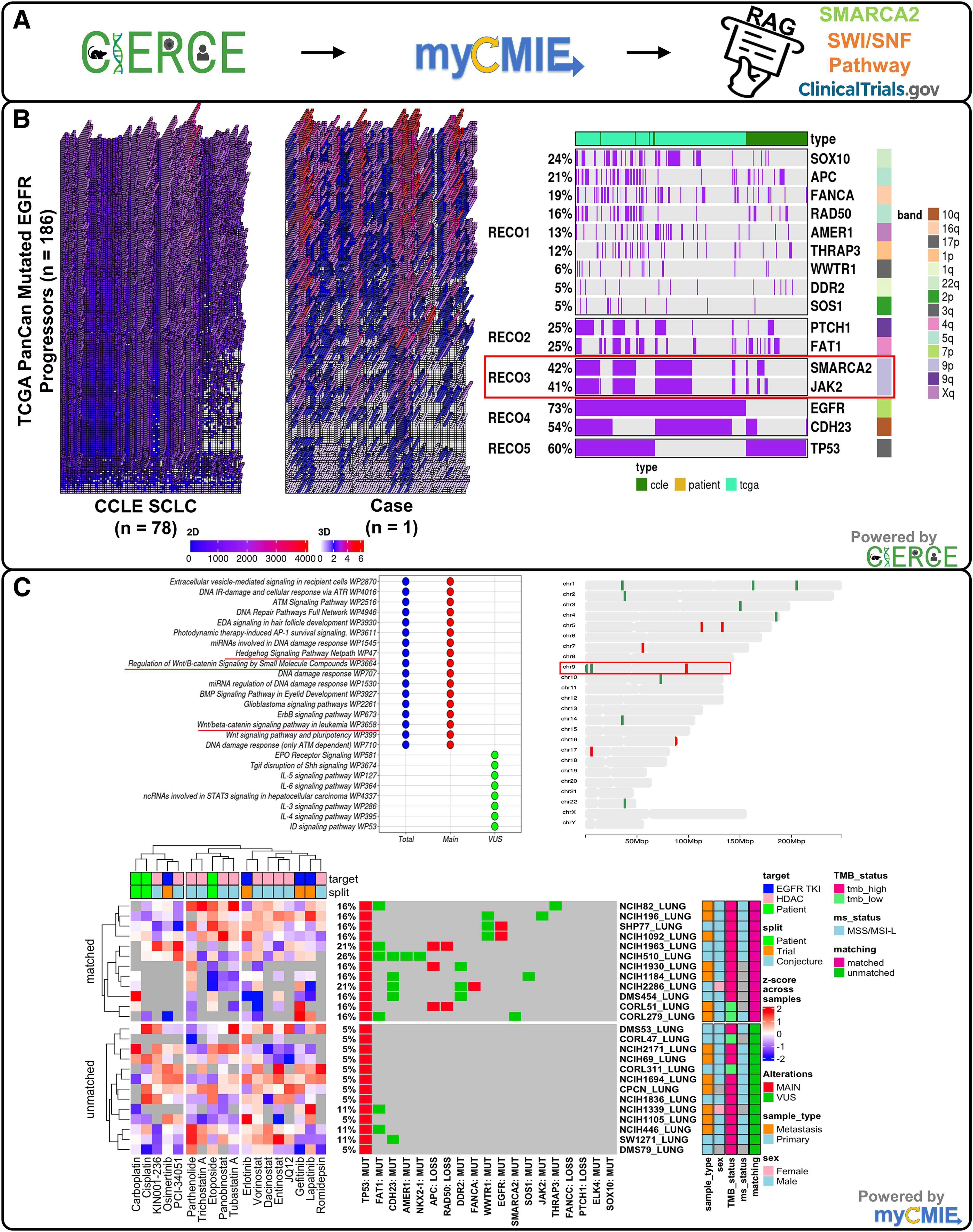
Fabrication analyses led to the focus on a single VUS gene (SMARCA2) that was further supported by myCMIE analyses of enriched pathways and spatial chromosome location hotspots revealed HDAC inhibitors as potential treatment options under consideration.
Altogether, our RAG-informed research led us to postulate that mutated SMARCA2 is likely to incur a mutated SWI/SNF complex, leading to chromatin accessibility changes and acetylation, resulting in EGFR resistance to EGFR TKI. To explore this context in the case profile, we compared targeting of EGFR TKIs, histone deacetylases, and NCCN-guided patient treatment history IC50s in a cluster analysis between SCLC case-matched and unmatched cell lines (Fig. 9C) that revealed fabric-informed, select HDAC inhibitors (e.g., panobinostat) with lower IC50s in the matched compared with the unmatched SCLC cell lines. In an RAG-informed clinical trial search, we identified an actively recruiting Phase I/II trial (NCT05053971) testing the drug combination of entinostat with a bromodomain inhibitor.
We next explored whether nbTMB could provide insights on both the lack of response based on the patient's treatment history and fabric-informed treatment options by clustering SCLC cell lines using the percentage of mutations colocalized to non-B DNA motifs (nbTMB-p) among the total mutation burden (Fig. 10B). With the exception of one cell line, the high nbTMB cluster contained metastatic cell lines. Upon comparing median IC50s among nbTMBp-derived clusters, carboplatin had a significantly higher IC50 in the high nbTMBp group compared with the other clusters. Carboplatin was shown with the highest IC50, both among the patient's treatment history and overall, among all compounds with available IC50s (Fig. 8C). Genomic instability can lead to mutations in genes responsible for DNA repair, drug transport, and apoptosis pathways, contributing to resistance. Non-B DNA structures can induce genomic instability by creating regions of the genome that are difficult to repair or by promoting mutations that confer resistance traits.
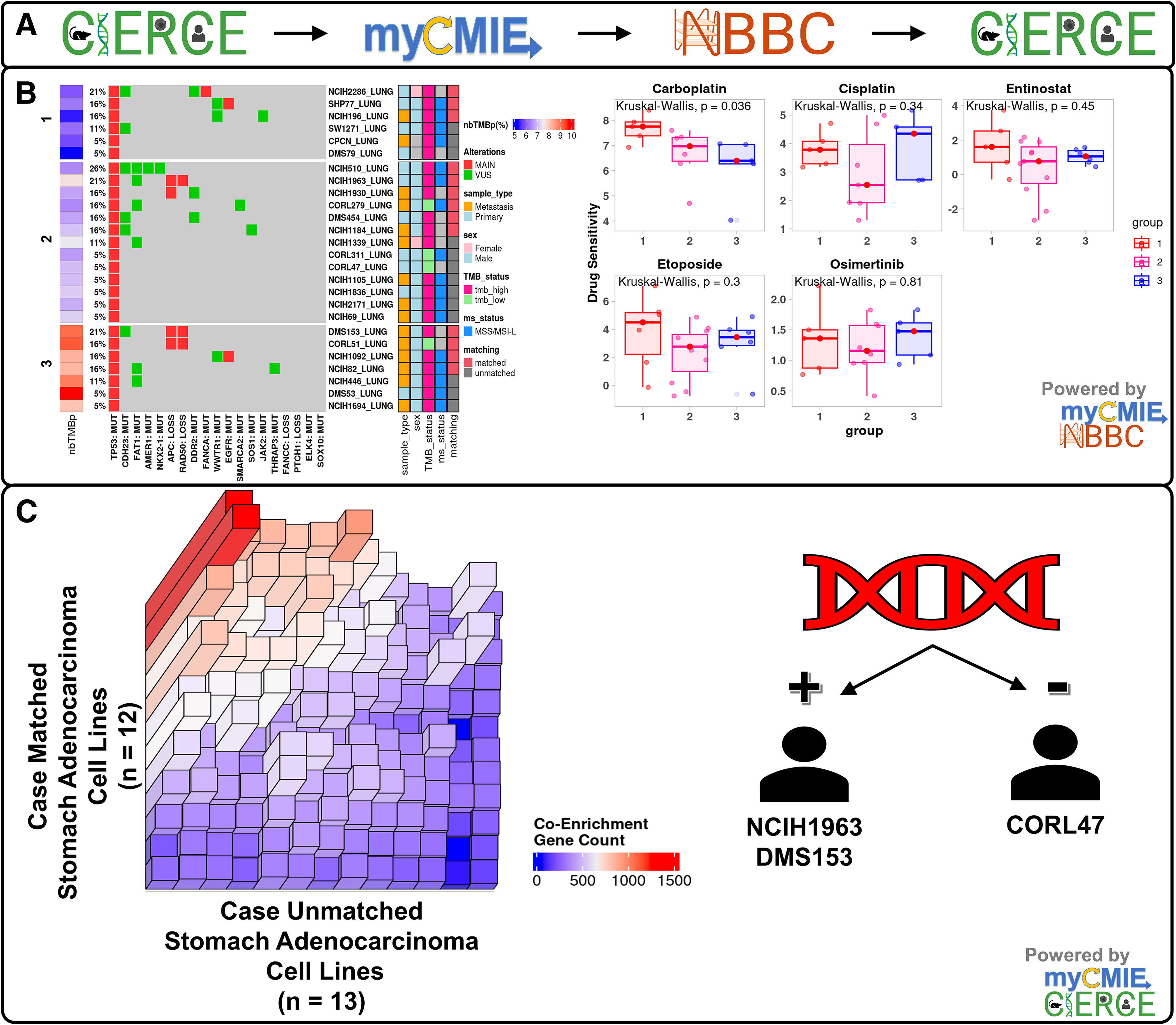
Fabrication analyses revealed insights into patient treatment history and non-B DNA tumor mutation burden that may be further explored using in silico, digital twin isogenic cell lines matched and unmatched with the case profile.
Understanding the interplay between genomic instability, non-B DNA structures, and carboplatin resistance is crucial for developing new strategies to prevent or overcome drug resistance in cancer treatment. Research in this area may focus on targeting specific DNA repair pathways, modulating the formation of non-B DNA structures, or identifying biomarkers of resistance, such as nbTMB, to guide treatment selection.
Outcome
The patient was offered compassionate use for entinostat and ZEN003694 as part of the identified, fabric-matching clinical trial (NCT05053971). She declined further treatment and opted for hospice care.
Discussion
Leveraging our molecular treatment fabric computational platforms, enhanced with RAG, we have connected insights from each case molecular profile to provide novel hypotheses for treatment responses and biological insights worth exploring. This integrated analysis with our data fabric facilitated a deep dive into biological and genetic pathways, identifying potential targets for new therapeutic interventions. Oncology's current reactive stance, often dictated by a herd mentality, underscores the need for a shift toward proactive, comprehensive profiling at diagnosis. This would enable a quicker pivot to more effective treatments upon the failure of first-line therapies, as per NCCN guidelines. In addition, the limited accessibility to targeted therapies in trials and logistical barriers to patient participation emphasize the necessity for improved coordination for their compassionate use.
Limitations of the study
Our analysis is limited to the availability of compound testing on cell lines, sample types, the degree of matching with the patient molecular profile, and a comprehensive literature search, among other factors, that should be taken into consideration. Cell line profiling data, while not always directly applicable to individual patient profiles for identifying optimal drug combinations, can be valuable, especially for advanced cases. This is because most cell lines are derived from patients with advanced diseases, where comprehensive profiling is common, often revealing likely drug matches. For instance, in SCLC, despite the absence of patient data in TCGA, there are SCLC cell lines with compound testing data available. Furthermore, the practice of testing compounds in cell lines, particularly in combination as seen in efforts such as Genomics of Drug Sensitivity in Cancer dataset 2, has gained renewed interest.
Targeting DNA quadruplexes presents a challenge due to their repetitive sequences throughout the genome, making it difficult to create specific inhibitors. Triplex target sites, when used in combination with chemotherapy, have demonstrated enhanced effectiveness compared with chemotherapy alone, but their use requires several gene copies to target with unknown off-target impact. However, we advocate for the use of non-B DNA forming structures as adding novel insights into genomic instability, potentially as markers that augment existing ones to inform on the heterogeneity in treatment response.
Our data fabric is intentionally patient-centric, treating each case as an individual study to prioritize depth of insight over a wide patient base. However, we recognize the importance of expanding our scope and are developing an AI-enabled system to automate workflows for input into each application, thereby increasing our capacity to handle a broader range of cases. This expansion is facilitated by the latest advancements in deidentification, text extraction, and large language models, aiming to enable multistakeholder use. We further acknowledge the dependency of our digital application's effectiveness on the quality and timeliness of the included resources. To maintain the cutting-edge status of our applications, we plan to engage students for support in identifying and integrating new and enhanced resources related to cancer into the fabric. This process involves evaluating these resources for Application Programming Interface availability, licensing, and their applicability to our work.
Furthermore, we recognize a challenge in applying our data fabric in clinical settings, where the focus is currently on developing AI-enabled infrastructures for processing, summarizing, and querying clinical notes with molecular content. The integration of our data fabric within existing clinical decision support systems will require additional efforts to develop an API that could be used with large language models for use by multistakeholders in care, and in accommodating additional case studies on a larger scale.
Conclusion
The shift toward digitalized systems represents a comprehensive overhaul, not just in adopting cutting-edge technologies but also in redefining the operation, innovation, and integration of existing resources such as databases, reports, and public data. This digital foundation is crucial for AI preparedness, enhancing efficiency, fostering innovation, and ensuring agility amidst the evolving landscape of research, characterized by complex questions and the need for sophisticated analyses involving diverse data sources. We have embraced digitalization by merging new and established markers with therapeutic relevance within our systems. This has enabled us to create a whole, canonical and noncanonical, DNA inclusive molecular data fabric designed for seamless AI integration. Our data fabric paves the way for personalized, AI-augmented CTDS systems by establishing patient-specific computational workflows.
Ultimately, our initiative culminates in a personalized care model driven by a bespoke computational workflow that originates from a cohesive molecular treatment data fabric, setting the stage for effective AI integration and signifying progressive thinking on AI in precision oncology.
Future Directions
Our future direction aims to expand our digital molecular treatment data fabric, which centrally connects patient profile-matched digital twin communities across resources for case-based learning with therapeutic implications. This digital foundation facilitates AI-driven development for CTDS. We are progressing on the design and implementation of an AI-enhanced, molecular information system that can process and summarize medical documents, while also working toward developing an API that will integrate our data fabric within it and with other digital health applications. This expansion will enhance the system's utility for stakeholders in cancer care by including large language models and conducting thorough testing in real-world settings. Our goal is to make this technology accessible to a wider range of patients and health care professionals, improving the molecular connectivity between digital health tools and patient data.
While many cancer centers have dedicated their efforts to constructing data lakes, lakehouses, and AI-powered data processing workflows, we have taken a different approach by prioritizing the ultimate objective in cancer care from the start. By first establishing our end goal, we are now strategically developing the means to achieve it, ensuring every step we take is purposefully aligned with improving patient outcomes.
Footnotes
Acknowledgments
The websites for myCMIE, TTSBBC, and NBBC are hosted on an Amazon Web Services serverless Fargate environment that is supported by UT-Austin IT Campus Solutions Emerging Technologies and Architecture team (Mr. Eric Weigel, Mr. Alex Knox, and Mr. Ladd Hanson) and Dell Medical School at UT-Austin.
Authors' Contributions
Conception and design: J.K. Data collection and assembly: M.H.C., T.A., B.G., A.C., D.Mahadevan, and D.Meyyappan. Data analysis: J.K., Q.X., and M.Y. Initial draft: J.K. Article edits: All authors. Accountable for all aspects of the work: All authors.
Author Disclosure Statement
The authors declare no conflicts of interest.
Funding Information
The University of Texas Dell Medical School Research Funds [to J.K.].
