Abstract
Purpose:
xDECIDE is a clinical decision support system, accessed through a web portal and powered by a “Human-AI Team,” that offers oncology health care providers with treatment options personalized for their cancer patients and outcomes tracking through an observational research protocol. This article describes the xDECIDE process and the artificial intelligence (AI)-assisted technologies that ingest electronic medical records, generate a structured personal health record, standardize clinico-genomic features, and produce ranked treatment options based on clinical evidence, expert insights, and real-world outcomes generated by the system itself.
Methods:
Patients enroll directly into the IRB-approved pan-cancer XCELSIOR registry (NCT03793088). Patient consent permits data aggregation, continuous learning from clinical outcomes, and sharing of limited datasets for research. xDECIDE aggregates and processes medical records with natural language processing and machine learning to generate a structured care summary with a standardized list of patient features. These features are utilized by an ensemble of AI-based models called xCORE (xCures Option Ranking Engine) to create a ranked list of treatment options. The output of xCORE is reviewed by molecular pharmacologists and oncologists in a virtual tumor board (VTB) setting to create a report of treatment options and supporting rationales, individualized to the patient. Treating physicians use an interactive portal to view these data and reports and to continuously monitor their patients' information.
Results:
At the time of writing, over 7000 patients have enrolled in XCELSIOR, including over 1000 with central nervous system cancers, over 800 with pancreatic cancer, and over 300 patients each with lung, breast, and colorectal cancers. Over 350 VTBs have been performed for patients across indications including glioma, pancreatic cancer, and fibrolamellar carcinoma, with >450 therapeutic options discussed and over 2000 consensus rationales delivered. Over 500 treatment rationale statements (“rules”) have been encoded to reference discrete patient features to improve algorithm decision-making.
Conclusion:
xDECIDE clinical decision support can aid oncologists in their practice of medicine. The system identifies potentially effective treatment options individualized for each patient from the integration of real-world evidence, human expert knowledge and opinion, and scientific and clinical publications and databases, and offers a platform to learn from the experience of every patient.
Introduction
The promise and problems of precision oncology
The goal of precision oncology is to offer cancer patients treatment options that are tailored to the particulars of their individual disease, as well as to contextual factors, such as their treatment preferences. 1 In oncology, this information, which we call the patient's “profile,” will almost always include a mix of demographics, clinical information, histopathology, and tumor biomarkers such as genetic variants that have diagnostic, prognostic, or treatment significance. Unfortunately, delivering precision oncology is exceedingly complex; thousands of unique combinations of variants have been observed, 2 each potentially requiring a different treatment approach. The knowledge about these variants is spread over an enormous volume of papers, guidelines, and databases.
Between 2017 and 2020, the number of genes and variants of significance in the CIViC database increased at compound annual growth rates of 11% and 35%, respectively. 3 The number, length, and complexity of National Comprehensive Cancer Network (NCCN) treatment guidelines have been increasing at a compound annual growth rate of ∼21% for the past 20 years. 4 Today, this equates to ∼3000 pages of new guideline information for a general medical oncologist to absorb each year (see Supporting analysis in Supplement S1). When there are no approved treatment options, there may be dozens of relevant clinical trials for an oncologist and patient to choose from. For example, there are currently >6000 actively recruiting interventional trials, and 36 active expanded access programs in cancer across the United States alone (data Based on a clinicaltrials.gov 5 search for recruiting interventional trials and expanded access programs in cancer, carried out on August 10, 2023.).
Researching trials for patients is a time-consuming activity and only a limited number of cancer patients qualify for interventional clinical trials. 6 It can be difficult to reliably assess patient eligibility for an oncology trial, especially for patients with advanced disease or poor performance status.7,8 While an enormous number of potential novel combination regimens could be utilized off-label, implementation of these approaches is limited by the additional time and funding required, and by a lack of information about their validity or applicability for specific patients.
The volume of actionable information, the speed of new information creation, and the need to quickly weigh the relative value of this information for a specific patient together demand computational support. Yet, in our experience of operating hundreds of virtual tumor boards (VTBs) in advanced cancer, physicians continue to rely on the opinions of trusted colleagues, and web searches that may or may not uncover the most relevant or recent knowledge.
To address this problem, described in detail by Shrager et al., we have created XCELSIOR, a pan-cancer, patient-centric outcomes registry (NCT03793088, Fig. 1 and Supplement S1) and the associated xDECIDE process. 9 The registry aggregates real-time, real-world clinical outcomes data in the context of observational research. We leverage this data, along with insights and recommendations from leading oncologists, in xDECIDE, an artificial intelligence (AI)-augmented clinical decision support system (CDSS) accessed via the web. xDECIDE is operated by human experts in cancer biology and clinical practice, supported by AI and machine learning algorithms that surface and rank evidence-based personalized treatment options for any cancer patient in the United States.
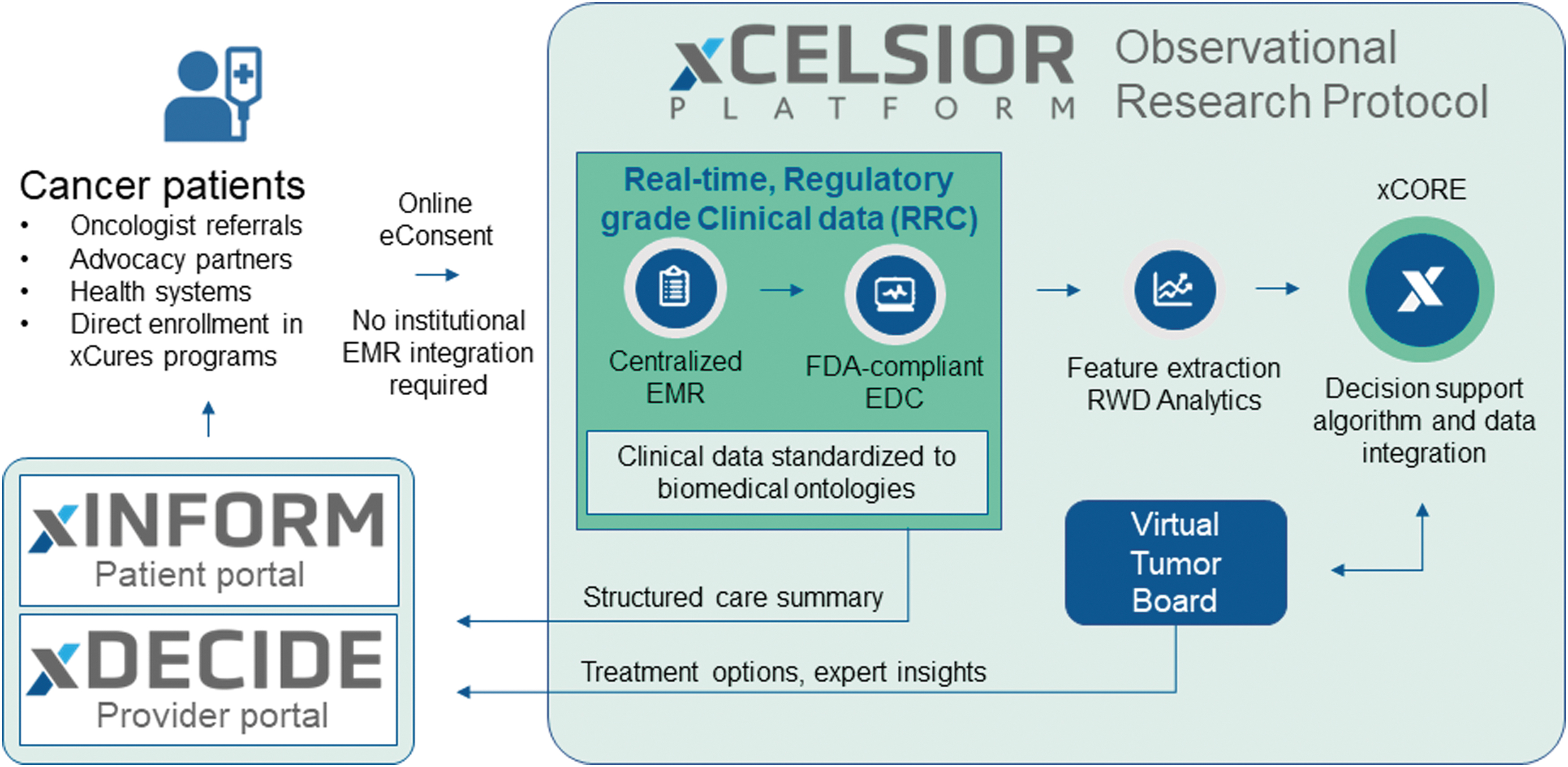
The XCELSIOR study schema. Patients with cancer or suspected cancer enroll in the XCELSIOR master observational protocol by online registration and eConsent via xINFORM, a personalized web portal. EMRs are gathered from all sites of care. Key data elements relevant to cancer are annotated in a central EDC system by study staff and normalized to a specialized OMOP-based ontology. At the level of an individual patient, these data elements offer features for ingestion into the treatment options ranking engine. At the cohort level, these elements offer RWD for patient feature-matched outcomes in the xCORE algorithm. A virtual tumor board offers expert review, training, and validation of the outputs of the algorithm. Aggregated EMR, longitudinal structured care summaries, and treatment option recommendations are available to patients and providers via the xINFORM and xDECIDE interactive web portals. EDC, electronic data capture; EMR, electronic medical records; OMOP, Observational Medical Outcomes Partnership; RWD, real-world data; xCORE, xCures Option Ranking Engine.
Methods
Overview of the xDECIDE CDSS
xDECIDE is a CDSS operated by a “Human-AI Team”—a collaboration between human experts and AI-based software tools—as defined by the FDA's “Good Machine Learning Practice for Medical Device Development: Guiding Principles.” This guidance reads, in part: “Focus Is Placed on the Performance of the Human-AI Team: Where the model has a ‘human in the loop,’ human factors considerations and the human interpretability of the model outputs are addressed with emphasis on the performance of the Human-AI team, rather than just the performance of the model in isolation.” 10
In xDECIDE, the human experts include leading clinicians, who review cases individually or together in a panel discussion called a “Virtual Tumor Board” (VTB), patient advocates, and scientists with expertise in molecular pharmacology, cancer biology, and genomics. Data analysts conduct medical record reviews, medical coding, and patient tracking to ensure the most accurate clinical data are captured for each patient. Computer scientists with expertise in knowledge management, cloud and web engineering, statistics, and machine learning develop the algorithms that support the human processes, making xDECIDE more accurate, efficient, and scalable.
Figure 2 depicts the overall xDECIDE workflow. Patients register and electronically consent to the XCELSIOR study via an interactive patient portal called xINFORM. Their electronic medical records are collected, aggregated, and processed using optical character recognition (OCR), natural language processing (NLP), and named-entity recognition (NER), as described in more detail below, to extract structured concepts in established coding systems. Discrete data elements (coded concepts) are highlighted in an annotation system and are then confirmed/refined by human annotators. This data extraction and coding receives multiple levels of human and machine review and results in the patient's records being coded to biomedical ontologies to create the patient's digital clinico-genomic profile. This profile is input into an ensemble AI algorithm to create a preliminary ranking of potential treatments which are reviewed by an internal team of PhD scientists with expertise in molecular pharmacology and cancer biology. A subset of cases are discussed in a VTB to gather expert opinion and link treatment rationales with patient clinico-genomic features.
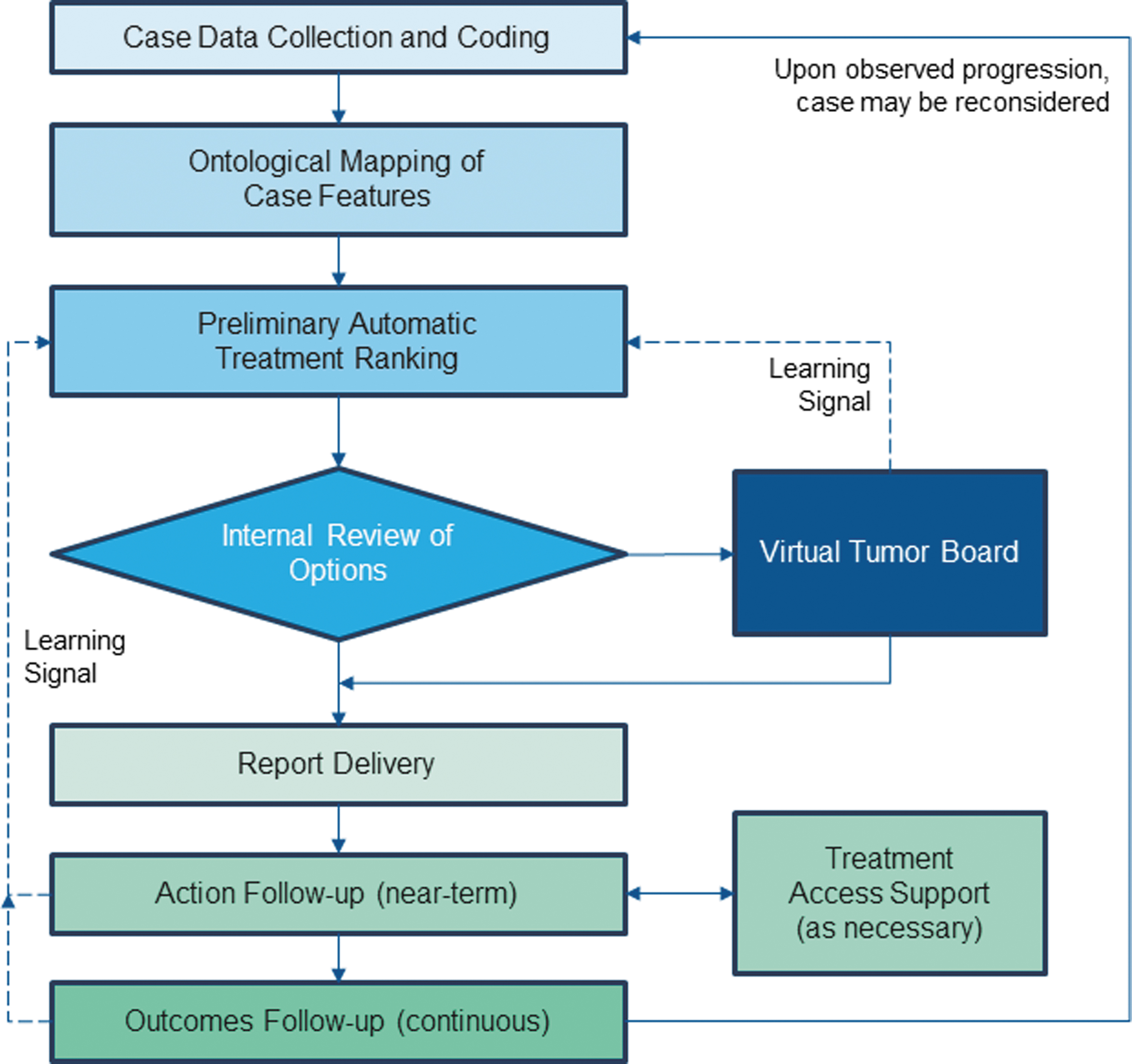
Overview of the xDECIDE CDSS workflow. Simplified process diagram of each stage of the xDECIDE CDSS workflow. Dashed lines represent learning signals fed back to the algorithm to tune hyperparameters. CDSS, Clinical Decision Support System.
An interactive online report of the resulting options, including scientific/medical rationales and relevant clinical citations, is simultaneously delivered to the oncologist and patient via the web-based xDECIDE and xINFORM portals, respectively. Oncologists and patients can interact via their respective portals to indicate their treatment decision. If they have chosen an expanded access or off-label treatment, treatment access support by xCures staff, patient advocacy organizations, or other groups can be triggered. Oncologists and patients can continue to interact with the platform to update preferences and history and to request updates or additional treatment options for consideration. Performance feedback at a number of points serves as data for machine learning, which continuously improves the AI-based treatment-ranking algorithm (dashed lines in Fig. 2).
In what follows, we describe the xDECIDE subprocesses in more detail, identifying instances where human decision-making is augmented by machine learning, automation, and other technologies. The detailed algorithms and performance analyses are described in separate articles, in preparation.
Clinical data collection and ontological coding of case features
After a patient has consented into the XCELSIOR protocol, their electronic medical records are gathered and classified by document type and visit date. This includes diagnostic pathology, clinic notes, radiology reports, and molecular profiling reports, among other records. This and all subsequent analyses take place in a 21 CFR Part 11-compliant database.
Fax and paper documents are first run through an OCR process that results in machine-readable text. An NLP layer parses this text, and a NER module extracts clinical concepts, mapping relevant text fragments to clinical codes. Unrecognized codes and codes with low confidence scores are mapped by human data abstractors, and the NER module is periodically retrained with training sets incorporating these problematic mappings. This facilitates the identification, coding, and mapping of information from the raw medical records into discrete data elements that populate custom Case Report Forms (CRFs), modeled on standards including mCODE 11 and CDISC CDASH. 12 CRF customization includes fields for primary and secondary cancer diagnoses, tumor stage/grade, therapy regimens and their event dates (e.g., start/stop), circulating tumor marker values, and biomarker findings, among others. Data elements are source-verified, and provenance links to the source medical records are maintained, including the precise coordinates in the source image file.
Data collection from medical records is overseen by human data abstractors trained in using these OCR and NER systems. The record collection and coding process extracts two primary kinds of data: structured data elements, and natural language from unstructured components of medical records (patient interval history summaries, physician treatment plans, radiology findings and impressions, among others). The structured content is standardized using the Observational Medical Outcomes Partnership (OMOP) vocabulary, which incorporates a variety of coding systems, including CiVIC, 3 NCIT, 13 MedDRA, 14 LOINC, 15 SNOMED, 16 NCBI Gene, 17 UniProt, 18 augmented by other coding standards such as mCODE, and additional custom xCures ontologies, to provide a harmonized set of patient clinico-genomic features suitable for algorithm ingestion. The NER system automatically extracts features that are then reviewed by human data abstractors who accept or correct the system's suggestions. The coded, structured data elements are then put through a secondary review and Source Data Verification to ensure accuracy. A final medical review is conducted to ensure case completeness, identification of any missing data elements, or key facets of the individual case. If an appropriate coding does not yet exist for a term, it is added in a mapping table. If no suitable code for a concept resides in the existing ontologies, the xCures custom ontology is extended to accommodate it. Finally, these new mappings and codes are incorporated into subsequent iterations of the NER system's training. Supplement S2 shows a screenshot from one of the internal case review dashboards.
The result is a complete and accurate longitudinal summary of the patient's cancer history from diagnosis to present in a format that is easily ingested into algorithms and graphically displayed to patients, providers, and experts involved in VTBs and treatment option curation and report preparation. See Figure 3 for graphical summary of the process.
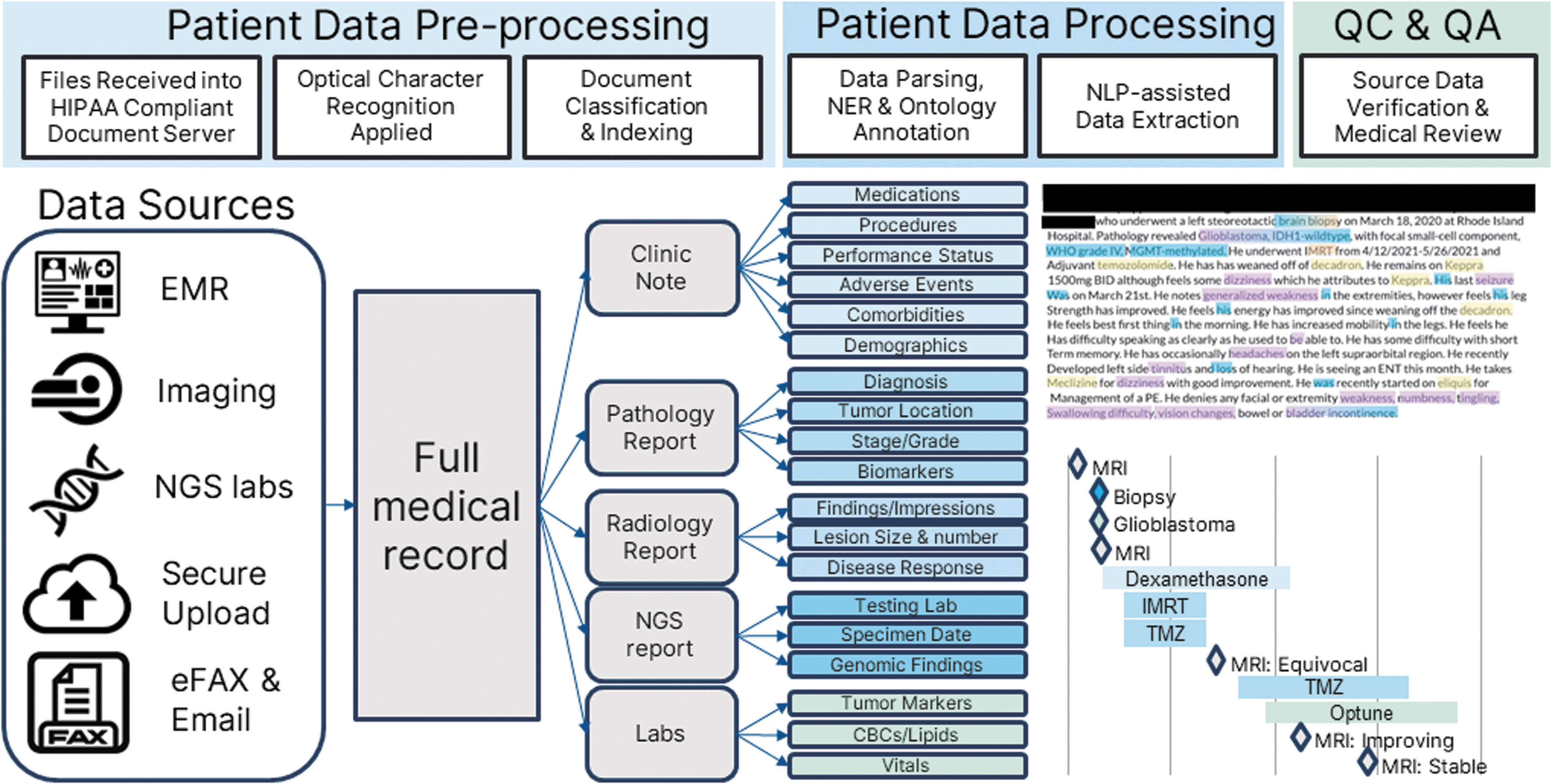
Clinical data ingestion, extraction, and coding workflow. The overall data processing pipeline has three stages: patient data preprocessing, patient data processing, and QC/QA. First, patient medical records are aggregated from diverse sources and centralized in a HIPAA-compliant document server. OCR is applied, and documents are classified into specific encounter types. Second, NER is utilized to identify key cancer-relevant data elements within unstructured text. Data associates leverage a custom data curation software to annotate a central EDC system and code to biomedical ontologies in one step, maintaining source-data provenance. Lastly, QC, QA, and medical review offer a fully longitudinal and source-verified care summary for use by the xCORE treatment ranking algorithm. HIPAA, Health Insurance Portability and Accountability Act of 1996; NER, named-entity recognition; OCR, optical character recognition; QA, quality assurance; QC, quality control.
The options library and the xCORE ranking algorithm
The principal output of xDECIDE is a ranked listing of treatment options selected based on the extracted patient features and preferences. They are selected from the xDECIDE Options Library, a comprehensive list of plausible cancer treatment regimens that include a standardized coding of abbreviations and aliases of interventions and treatment combinations. For example, “gem/abrax” is coded to the combination of “gemcitabine” and “nab-paclitaxel.” At a minimum, any option in the library has been used previously for treatment of human cancer patients. The sources for this library include (1) individual treatments and combination regimens that form the arms of all interventional cancer clinical trials abstracted from clinicaltrials.gov, 5 (2) published case reports, (3) patient treatments observed in the XCELSIOR registry, and (4) novel regimens proposed during tumor board reviews by clinicians with real-world experience. Each option consists of three components: (1) a normalized nomenclature of the component interventions, for example, “ipilimumab, nivolumab, radiation therapy”; (2) scientific/medical rationale(s) derived from past cases and/or published literature; and (3) an access mechanism being one of: standard-of-care (SOC), clinical trial, off-label, expanded access, or diagnostic. For options based in clinical trials, locations and recruiting status are maintained up to date according to clinicaltrials.gov. 5
The Options Library consists of ∼12,000 therapeutic options involving over 4000 discrete interventions alone or in various combinations. These options are derived from the arms of over 15,000 interventional oncology clinical trials over the past decade, of which ∼5000 are currently recruiting and available as options to present in reports, as well as regimens observed in the clinical care of patients in the XCELSIOR registry.
A case feature vector representing a patient's digital clinico-genomic profile, generated during case preparation and standardized as described above, is used to query the Options Library via the xCures Option Ranking Engine (xCORE), as summarized in Figure 4. xCORE is an ensemble machine learning system comprising several submodels and a weighted integrator. Submodels are based on multiple platform inputs: (1) An NLP submodel trained on all trials listed in clinicaltrials.gov, using NER to extract disease states relevant to specific treatment, plus inclusion and exclusion criteria which help limit the treatment to relevant patient populations—importantly, not only the treatment arms are used (which link the new therapy to the appropriate patient features) but also active-comparator arms (which describe the SOC against which the new therapy is being compared); (2) A biomarker-drug submodel, which links disease states to structured alterations from molecular profiling (using annotated databases such as CiViC 3 ) and drug–gene interaction databases, such as DGIdb 19 ); (3) a submodel that analyzes consensus option rankings (numerical data) from VTBs, which captures the knowledge of leading practitioners regarding expected efficacy and safety; (4) an NLP submodel that extracts treatment information from the rationale statements supporting or opposing the use of specific treatments based on discrete patient features, as captured in the interactions between VTB members while discussing a case; and (5) actual clinical outcomes of patients collected according to the XCELSIOR study, employing standard metrics such as overall survival, progression-free survival, and time on treatment, and using techniques such as propensity score matching to overcome potential selection bias. The case feature vector is independently distributed to these submodels. Each submodel scores each treatment, and these scores are subsequently combined into an integrated vector of treatment scores. The integration algorithm and each submodel have parameters that are adjusted through learning, described below and in more detail in the article in preparation (Wasserman et al., in prep).
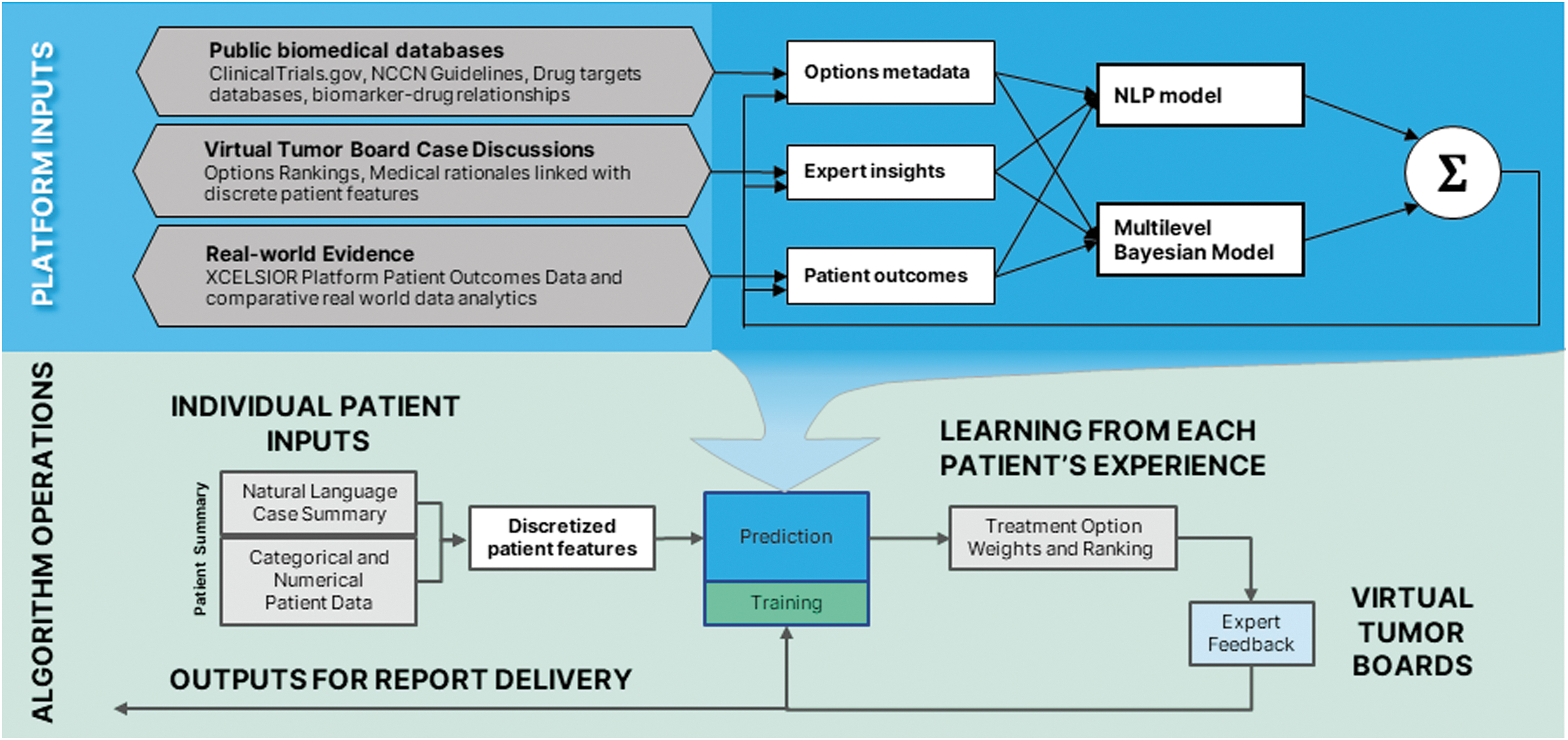
The xCORE algorithm. The xCORE algorithm utilizes submodels based on multiple public biomedical databases, expert opinion, and real-world evidence collected in the platform. Case data from incoming patients is encoded to model-specific feature vectors by separate encoders for each of the three models. These feature vectors are passed separately through each model, producing separate treatment predictions. These are then combined by an integrator, resulting in the final treatment option ranking. These are then reviewed by the internal and external expert team. The feedback from this review, as well as from patient choice and outcomes data, is used to tune the models, thus learning from a wide range of feedback based upon each patient's case, expert review, and clinical experience.
Virtual tumor boards
The output of the xCORE algorithm—patient features and ranked treatment options—is next reviewed by internal PhD scientists and a subset are discussed in a VTB. VTBs consist of at least three leading clinicians and two to three internal PhDs with expertise in molecular pharmacology and cancer biology. Clinicians are practicing medical oncologists or specialists in a particular field of oncology, including medical oncology, surgical oncology, radiation oncology, interventional radiology, or neuro-oncology/neurology, who hold an MD and/or PhD. These physicians are regarded as experts in their field based on their current appointment at top U.S. academic research institutions, generally as division leaders, and/or participation or leadership with national committees on cancer treatment guidelines (e.g., NCCN). They are actively involved in clinical research, serve as investigators on numerous clinical trials for novel therapeutics or treatment regimens, have a strong publication record in clinical and/or preclinical research, and hold expertise in one or more aspects of treatment for their disease specialty, for example, immunotherapies, targeted therapies, or signaling pathways.
The VTB reviews the structured and unstructured patient case data and the xCORE output of potential treatment options in either a synchronous virtual meeting or an asynchronous dialogue in a HIPAA-compliant chat software and determines the final recommendations. The patient clinical care summary is available to the VTB, including the patient's past cancer diagnostic and treatment history, tumor genetics/biomarkers, results from any additional diagnostic testing, comorbidities, concomitant medications, and family history, as well as the most recent imaging reports. In some instances, additional inputs are available, for example, from alternative genomics-based algorithms or cell- and organoid-based functional profiling assays. Importantly, statements (“rules”) supporting or opposing specific options, together with the clinico-genomic patient features that entered into that decision, are solicited and coded for future learning in the xCORE algorithm. These are usually in the format: Patient feature X supports/opposes Option Y for Condition Z. 18 Ultimately, the case reviewers—whether this is one expert or a panel in a VTB—create a consensus recommendation of ranked therapeutic options.
Report finalization, delivery, and follow-up
Internal PhD-level staff utilize various dashboards to examine, manipulate, and combine all of the foregoing into a report. Supplement S3 provides a view of the Treatment Options Curation Dashboard within the admin portal. Staffs review the suggested treatment options from xCORE and combinations thereof, identify clinical trials that involve the suggested options, and perform additional research on published clinical data and outcomes data within the XCELSIOR registry from past cases. Thus, utilizing the algorithm outputs, previous reports delivered, real-world outcomes data, and potential new breaking research, PhD-level staff curate a final list of treatment options.
In this stage, treatment recommendations are augmented with rationales that help the recipient understand the logic of the recommendations. Rationales reference specific patient features that relate to the recommendation of the option presented, as well as the most relevant published clinical evidence supporting the use of that option. This clinical evidence is found by review of the internal databases and supplemented by web searches. The report is delivered simultaneously to the oncologist and patient through the xDECIDE and xINFORM interactive web portals, respectively. Supplements S4 and S5 provide screenshots of sample reports as viewed through the xDECIDE and xINFORM portals.
After report delivery, patients and oncologists can interact with the platform to indicate their chosen treatment options and provide their reasoning for pursuing particular options. This provides a rapid learning signal for the clinical utility of xDECIDE and provides natural language input that can be mined for supporting rationales and associated patient features.
If a selected option is available via off-label or expanded access, outreach is initiated to the treating physician to offer access support, including clinical evidence citations for inclusion in prior approval requests, interaction with pharmaceutical companies for expanded access requests, and/or regulatory support for single patient Expanded Access protocol preparation or FDA Form 3926.
The patient's medical records are periodically requested and processed to close the learning loop but can also be manually uploaded by the patient or their oncologist via the xINFORM or xDECIDE portals. If a change in the patient's dataset alters the xCORE algorithm's output, a new series of treatment options can be created, reviewed, and presented to the patient and their oncologist.
Machine learning and continuous process improvement
Many of xDECIDE's internal processes are self-improving through supervised learning, in which the outputs of the process are checked by either human experts or through patient outcomes metrics, and these signals are used to improve the processes. The following signals are actively and continuously integrated into the learning system:
Unmapped or missing terms/concepts identified during intake of electronic medical records, or identified by VTBs, are added to the medical record pipeline and coding system, either as new terms or as aliases of existing terms. Final consensus treatment recommendations arising from the VTB process are used to directly train the ranker. Treatment choices indicated by the patient or physician within their respective portals (xINFORM and xDECIDE) provide a rapid clinical utility signal. Rationales derived from expert statements in VTBs can suggest novel terms or rules. (More detail on this appears below.) Clinical outcomes and real-world evidence are among not only the most important learning signals but are also the most delayed. These include changes in tumor load, clinically relevant biomarkers, patient or clinician-reported functional status, emergence of adverse effects, and progression, treatment-changes, and survival.
The special role of treatment rationales in learning
Rationales commonly arise in VTB discussion and sometimes from patient choices. Rationales are important because they provide a potential explanation regarding why a particular option was chosen or ranked higher, or why a particular option was rejected (e.g., contraindicated) or ranked lower.20,21 By introducing rationales from human experts into machine learning processes, the dimensionality of the learning problem is reduced, requiring fewer observations to train an algorithm. 1 For advanced cancer patients, rationales are often derived from the discussion of biomarkers, comorbid conditions, positive or negative responses to prior therapies, or other patient features such as the presence or number of metastases in a particular organ. Some of these concepts can be found in existing biomedical dictionaries and ontologies, but many novel concepts have been identified and added to the extraction and ontology mapping processes. Rationales are initially collected in verbatim plain text and then operated on to extract novel features. This approach for precision-oncology decision-making mirrors the inclusion/exclusion of clinical trials, but rather than reducing the process to a set of binary rules, xDECIDE is dynamically learning the relative weights and interactions between features supporting or excluding treatments, applying them at the individual level, and learning to generalize these to other patients.
Recent advances in Large Language Models (LLMs) have significantly improved the ability to extract and manage natural language such as that found in clinic notes, treatment plans, and treatment rationales collected in the course of VTB discussions and are now being employed within xDECIDE. In addition, LLMs can rapidly integrate and succinctly summarize dense biomedical language from publications and treatment rationales. Internal users of xDECIDE are leveraging LLMs to create consensus composite rationales based on all relevant clinical evidence and past treatment rationales for discrete options in the context of individual patient cases. Importantly, LLMs used in this way are simply providing plain-language summaries that are constrained by features or interactions derived from statistically sound and interpretable models. We expect that LLMs will become a central feature in the xDECIDE platform at all levels of operation, but operated within the guardrails of explainable AI models rather than as a core element of our CDSS.
Results
At the time of writing, over 7000 patients have enrolled in XCELSIOR from >3000 cancer centers and oncology practices across the United States. This includes over 1000 patients with central nervous system (CNS) cancers, over 800 with pancreatic cancer, and over 300 patients each with lung, breast, ovarian, and colorectal, and breast cancers. The XCELSIOR study database contains over 60,000 unique verbatim terms abstracted from >15 million individual medical records. This data have been coded to ∼30,000 standardized terms across the domains of disease, treatments, labs, and genomics in a structured CRF database. In partnership with patient advocacy groups, over 150 VTBs of CNS cancer patients and ∼100 VTBs each of pancreatic cancer and fibrolamellar carcinoma patients have been performed. In the course of these discussions, >450 therapeutic options have been discussed and over 2000 consensus rationales have been delivered. Across all CNS VTBs that have taken place to date, patient-level data elements have been translated to 1859 features (977 unique, 882 inferred). Principal Components Analysis reduces these to 153 patient feature dimensions for ingestion by the xCORE clinical decision support algorithm. Over 500 treatment rationale statements (“rules”) have been encoded to improve algorithm decision-making between similar therapeutics or regimens in the context of individual patient features.
The following is a case example of how a community physician utilized xDECIDE (Fig. 5). A patient at the age of 78 first presented to the clinic with newly diagnosed, stage IV adenocarcinoma of the lung with diffuse, multiorgan high burden disease. His tumor at the time of presentation was EGFR exon19-deleted, to which he responded well to standard first-line therapy with osimertinib. He unfortunately progressed 1 year later, and over the next 32 months would see multiple SOC regimens with varying durations of response, which included combinations of or monotherapy with platinum chemotherapies, two different VEGF-inhibitors, a taxol derivative, and an anti-metabolite. Through the course of his progressions, the patient was treated with stereotactic radiation for single colony progression and symptomatic bone mets. At time of enrollment into XCELSIOR, the patient was 3-years postinitial metastatic diagnosis status post six lines of therapy with a retained ECOG of 0. Case data were annotated in a full longitudinal Care Summary (Fig. 5A). Normalized features derived from the structured data were utilized by xCORE to provide potential treatment options for review (Fig. 5B). The algorithm outputs were reviewed by PhD level staff who consulted local clinical trials and scientific and clinical literature, considering combinations of suggested interventions, and wrote short rationale statements for final options in the report (Fig. 5C). The physician and patient reviewed the outputs and both opted to try combination therapy with erlotinib and everolimus, an option related to an xDECIDE recommendation of osimertinib and everolimus. The three literature citations clearly outlined the reasoning, based on molecular characteristics of the patient's most recent biopsy, as well as the presumed escape mechanisms given the patient's mechanistic treatment history. The citations provided by xDECIDE were used as justification for authorization of the off label use of the combination, and resulted in an automatic approval without the requirement of peer-to-peer approval. The patient responded to the combination provided by xDECIDE and was able to enjoy a remission of 11 months and, given the complete response received, was in optimal and preserved functional status and low disease burden to then see an eighth line of off-label therapy, on which he continues to remain on at the time of publication (Fig. 5A).
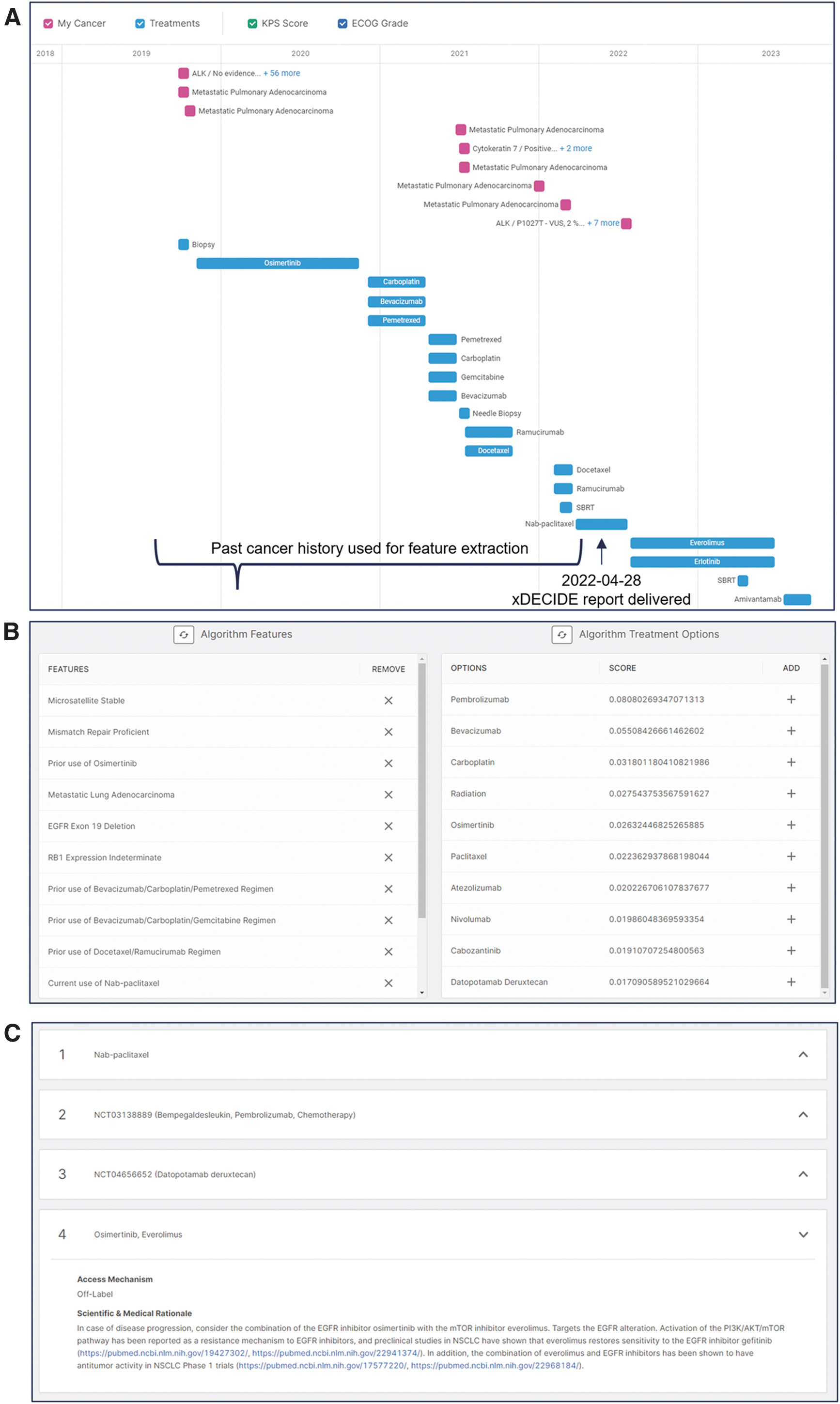
Case example of xDECIDE utility.
Conclusions
This manuscript provides a component-level description of the xDECIDE precision oncology platform. The xINFORM Patient Portal and the xDECIDE Provider Portal assist patients and providers with clinical decision support in the form of structured longitudinal case summaries abstracted from health records and personalized, ranked treatment options for their consideration. The platform utilizes OCR, NLP, and machine learning to augment human case review and therapeutic option selection, providing a scalable computational solution to real-world precision medicine applications. The goal of the xCORE algorithm is to deliver treatment option recommendations whose outcomes are equivalent to or better than those of molecular tumor boards at leading institutions. Operating under the XCELSIOR protocol, in the context of observational research, xDECIDE has the potential to reach the scale necessary to undertake long-term comparisons of the efficacy of treatments more efficiently than randomized controlled trials, or even than current adaptive trial technologies, as suggested in 2013 by Shrager, 22 and recently validated by Wasserman et al. 23
Discussion
The rapidly expanding and extremely complex oncology knowledge base, the explosion of novel therapeutic options and potential combinations, and an increase in worldwide cancer incidence and improved detection methods have led to a crisis of overburdened and exhausted clinicians. Medical AI, which has seen exponential maturation, streamlines complex processes, aggregates vast amounts of data, and provides insights beyond the grasp of unaided human cognition. The xDECIDE platform leverages these capabilities to provide robust clinical decision support for oncology treatment.
xDECIDE adopts a multifaceted approach, combining AI-driven technologies and human expertise. By employing NLP and machine learning, xDECIDE transforms the bulk of patient data into a structured and standardized format that feeds into the downstream personalization computations; the system's ability to aggregate and process patient medical data and create a vectorized set of features for each individual patient helps to fulfill the goal of personalized medicine. Within xDECIDE, the xCORE AI ensemble model facilitates decision-making and ensures that the treatment options provided are ranked, thereby aiding clinicians in their ultimate choice of therapy. In some regards, the system may be thought of as learning to identify subtle factors that support or refute various treatments from patient health records, but rather than learning a binary function, the system is learning to rank treatment options based on how well they the goodness of fit the characteristics of individual patients (i.e., of patient feature vectors). In practice, this means that the system can easily identify 5–10 plausible treatment options from a space of >10,000, and rank order those options for an individual patient, based on a large and diverse evidence base. The inclusion of a VTB brings molecular pharmacologists and oncologists into the decision loop to balance xDECIDE's AI computational advantages with human intuition and experience.
In VTBs reviewing xCORE outputs, oncologists typically find that some options were “obvious,” while others were unexpected or surprising, but logical when considered with the associated rationale. We view this as fundamental to the human-AI team in precision oncology and consistent with the benefits observed wherever an electronic marketplace for information with search technology becomes available. Specific benefits include significant time savings over manually searching for relevant information or publications, greatly reduced cognitive load associated with evaluating a list versus creating a set of options from scratch, and more optimal allocation. Indeed, the xDECIDE architecture is based on extensive research in choice theory. 24 Specifically regarding the number and nature of returned results, designed to reduce the cognitive load on busy oncologists and the use of rationales to make nonobvious options sensible to an expert. One area of future research is the application of patient-utility functions to help physicians further refine sensible options based on the patient's individual goals and risk tolerance. Utility functions also provide an easily visualized individual reranking of options, as described by Perini et al. 25
The rapid expansion of oncological knowledge and of new cancer treatments is unlikely to abate. To meet this evolving challenge, hybrid human-AI systems like xDECIDE will be required to improve the level of clinical care for most patients. They can assist with the immediate needs of decision support and personalized treatment options, but can also offer a framework for training, validation, and deployment of new AI models. As the integration of real-world evidence, human expertise, and clinical databases evolves, platforms like xDECIDE will be central to improving patient outcomes and setting new standards in oncology care.
Footnotes
Acknowledgments
Substantial contributions to the creation and operation of xDECIDE were made by Nick Arora, M. Shaalan Beg, Nicholas Blondin, Zac Cole, Colin FitzGerald, Keith Flaherty, Ekokobe Fonkem, Mike Freed, Sarah Ginn, Chezney Hinton, Sabrina Irizarry, Fabio Iwamoto, Vlod Kalicun, Paul Kent, Conan Kinsey, Amy Langville, Hope Lee, Jillian Hattaway Luttman, Al Musella, L. Burt Nabors, Chris Porter, Jameson Quinn, Lola Rahib, Jeff Rapp, Charles Redfern, Alanis Sabates, Kamalesh Sankhala, Davendra Sohal, Srikar Srinath, Tom Stockwell, Andrew Swain, Kristina Tadic, Marty Tenenbaum, Matthew Warner, Eric Wong, and Alayna Wood.
Authors' Contributions
M.A.S.: Conceptualization, Supervision, Methodology, Writing—Review and Editing. T.J.S.: Conceptualization, Writing—Review and Editing, Supervision, Data curation, Investigation, Visualization. A.W.: Conceptualization, Methodology, Formal Analysis, Software, Validation, Writing—review and editing. G.A.K.: Conceptualization, Supervision, Software, Writing—review and editing. B.F.: Conceptualization, Supervision. W.H.: Conceptualization, Supervision. Z.K.: Data Curation, Project Administration. D.C.: Data curation, Software. W.M.: Data curation, Software. M.E.N.: Conceptualization, Supervision. Z.O.: Data curation, Software. K.K.W.: Supervision, Writing—review and editing. S.K.: Supervision, Investigation, Validation. J.S.: Conceptualization, Formal Analysis, Software, Writing—original draft.
Data Availability
Data described herein are available through partnership with xCures, Inc., Contact: medical-affairs@xcures.com.
Author Disclosure Statement
M.A.S., T.J.S., B.F., A.W., G.A.K., Z.K., D.C., W.M., M.E.N., and K.K.W. are current employees of and have a personal financial interest in xCures, Inc. W.H., Z.O., and J.S. are former employees of and have a personal financial interest in xCures, Inc. S.K. has a personal financial interest in xCures, Inc.
Funding Information
No funding was received for this article.
References
Supplementary Material
Please find the following supplemental material available below.
For Open Access articles published under a Creative Commons License, all supplemental material carries the same license as the article it is associated with.
For non-Open Access articles published, all supplemental material carries a non-exclusive license, and permission requests for re-use of supplemental material or any part of supplemental material shall be sent directly to the copyright owner as specified in the copyright notice associated with the article.
