Abstract
Objective
To investigate the relationship between microsatellite polymorphism in the Heme oxygenase-1 (HMOX1) gene promoter and high-altitude pulmonary edema (HAPE) in Han Chinese.
Methods
Eighty-three construction workers who developed HAPE 2 to 7 days after arrival at Yushu (3800 m) in Qinghai, China, and 145 matched healthy subjects were included in this study. The amplification and labeling of the polymerase chain reaction products for capillary electrophoresis were performed to identify HMOX1 genotype frequency. The alleles were classified as short (S: <25 [GT]n repeats) and long (L: ≥25 [GT]n repeats) alleles.
Results
Patients with HAPE have significantly higher white blood cell count, heart rate, and mean pulmonary artery pressure, but lower hemoglobin and arterial oxygen saturation than healthy subjects without HAPE. The numbers of (GT)n repeats in the HMOX1 gene promoter show a bimodal distribution. However, there is no significant difference in the genotype frequency and allele frequency between patients with HAPE and healthy subjects without HAPE. Chi-square test analysis reveals that the genotype frequency of (GT)n repeats is not associated with HAPE.
Conclusion
The microsatellite polymorphism in the HMOX1 gene promoter is not associated with HAPE in Han Chinese in Qinghai, China.
Introduction
High-altitude pulmonary edema (HAPE) is a rare life-threatening condition that often occurs in healthy individuals who rapidly ascend to altitudes above 2500 m. 1 It is characterized by unequal hypoxic pulmonary vasoconstriction and regional overperfusion of capillaries in areas of least arterial vasoconstriction, leading to increased capillary pressure and leakage.2,3 Previous epidemiologic studies have shown a significant ethnic difference in HAPE risk, and individual risk may vary based on the genetic factors.2,3 It was reported that Han individuals were more susceptible to HAPE than others when exposed to the same altitude.
Heme oxygenase-1 HMOX1 as a transcriptional activator may be involved in the etiology of HAPE. 4 The HMOX1 gene is located on human chromosome 22 at 22q12 5 and is a rate-limiting enzyme in the catabolism of heme into bilirubin, free divalent irons, and carbon monoxide (CO). Hypoxia can transiently increase the transcriptional activity of the HMOX1 gene, resulting in increased production of CO and bilirubin. CO has potent vasodilatory, anti-inflammatory, antioxidative, anti-apoptotic, and antiproliferative activity,6,7 while bilirubin is an antioxidant. 8 The overexpression of HMOX1 can prevent hypoxia-induced inflammatory responses and the development of pulmonary hypertension. 8 Wang et al have shown that the reduction of CO production by ZnPP (HMOX1 inhibitor) significantly enhanced the proliferation of hypoxic smooth muscle cells. 9
The (GT)n short allele frequency (<25 repeats) of a panel of polymorphisms in the HMOX1 gene is associated with significantly increased HMOX1 promoter activity when compared with the long allele frequency (≥25 repeats). 10 The (GT)n repeat polymorphism in the promoter region has important functional implications. 11 Thus, we hypothesized that the (GT)n microsatellite polymorphisms in the HMOX1 gene promoter were associated with susceptibility to HAPE. In this study, we attempted to identify key HMOX1 genes regulating HAPE susceptibility.
Materials and Methods
HAPE and Non-HAPE Healthy Subjects
A total of 228 subjects were recruited in this study, including 83 HAPE patients (age 38.3±10.0 [mean±SD] years) and 145 male non-HAPE subjects (age 38.7±9.6 years) from May 2012 to July 2013. All subjects were low-altitude residents and ascended rapidly to an altitude of 3800 m during the Yushu earthquake in Qinghai Province. All HAPE patients were hospitalized at Yushu Prefecture People’s Hospital and evaluated by an experienced physician (RLG). HAPE was diagnosed based on the presence of flake or cloud infiltrate shadows at unilateral or bilateral pulmonary hilar on chest x-ray. Other clinical symptoms included dyspnea, cough, and white or pink foamy sputum. Cardiovascular diseases, such as myocardial infarction and heart failure, and pneumonia were ruled out by clinical and electrocardiogram examination. Clinical parameters, such as age, oxygen saturation, heart rate, and mean pulmonary artery pressure, were also measured. All subjects signed an informed consent for participation in this study, and all protocols were approved by the Medical College of Qinghai University Medical Ethics Committee.
Clinical Tests
Height, weight, blood pressure, and heart rate of all subjects were measured by standard methods. Hemoglobin concentration, white blood cell count, and hematocrit were measured from venous blood samples using the Mindray Hematology Analyzer (BC-2300, Shenzhen, China). Oxygen saturation and heart rate were recorded using a pulse oximeter (Ohmeda 3700 Pulse Oximeter; Datex-Ohmeda, Boulder, CO). Pulmonary arterial pressure was measured by echocardiography (EchoPac, version 6.3; GE Healthcare, Chicago, IL).
Reagents and Instruments
The reagents and instruments used in this study included polymerase chain reaction (PCR) thermocycler (ABI 9700, ABI, Foster City, CA), sequencer (ABI 3730XL), low-speed centrifuge (Eppendorf Centrifuge 5415R, Hamburg, Germany), ultra-low temperature freezer (Forma 725), gel scanning imaging system (UVP, Upland, CA; Labworks, Alpine, UT), ultraviolet spectrophotometer (Beckman DU800, Brea, CA), steady flow electrophoresis apparatus (DYY-6B-type, Beijing Sixty-one Instrument Factory, China), refrigerated centrifuge, small desktop centrifuge, electrophoresis tank, Gentra Puregene Blood Kit (158389, Qiagen), D2000 DNA-Marker (Tiangen, Beijing, China), agarose gel powder (Formatas), gene amplification reagents (Capitalbio Technology, Beijing, China), sequencing reagents (ABI), and GeneMapper 4.1 software (ABI).
Genomic DNA Extraction and Genotyping of HMOX1 Promoter Microsatellite Polymorphism
Genomic DNA was extracted from peripheral blood leukocytes using Gentra Puregene Blood Kit. HMOX1 promoter polymorphisms were analyzed by PCR on an ABI 9700 thermocycler using the following fluorescein amidite–labeled primers: sense primer, 5׳-TTCTGGAACCTTCT-GGGACG-3; and anti-sense primer, 5׳-GGCCGGTCACATTTATGCTC-3. The PCR cycles consisted of predenaturation at 94°C for 5 minutes, 35 cycles of 94°C for 30 seconds, 60°C for 30 seconds, and 72°C for 20 seconds, and a final extension at 72°C for 60 minutes. Amplified fragments were resolved on 1.5% agarose gels. The amplification length was 373 bp. Primers were synthesized in the Bioassay Laboratory of CapitalBio Corporation. The PCR products were analyzed by standard capillary electrophoresis on an ABI 3730 XL sequencer (Applied Biosystems). Data were imported into GeneMapper 4.1 (ABI) to identify the sequencing.
Statistical Analysis
The between-group differences in the genotype frequency and allele frequency were compared using t test or χ2 test in SPSS statistical software (version 17.0; SPSS Inc, Chicago, IL). P <.05 was considered statistically significant. All polymorphisms were examined for deviation from Hardy-Weinberg equilibrium using PowerMarker 3.25. The distribution of HMOX1 (GT)n genotype frequency in non-HAPE subjects conforms to the Hardy-Weinberg equilibrium (P =.07), indicating that these subjects are representative of the overall population.
Results
Physiological Characteristics of the Subjects
The physiological characteristics of HAPE and non-HAPE healthy subjects are shown in Table 1. There is no significant difference in age between the groups (P =.766). However, HAPE patients have significantly higher white blood cell count, heart rate, and mean pulmonary artery pressure, but lower hemoglobin and arterial oxygen saturation than non-HAPE healthy subjects (P <.01).
Physiological characteristics of HAPE and non-HAPE healthy subjects
Data are expressed as counts and percentages, mean±SD.
P <.05 was considered statistically significant.
HAPE, high altitude pulmonary edema; non-HAPE, non-high altitude pulmonary edema; WBC, white blood cell; Hb, hemoglobin; SpO2, arterial oxygen saturation estimated by pulse oximetry; PAP, pulmonary artery pressure.
HMOX1 (GT)n Microsatellite polymorphisms
Alleles are classified as short (S: <25 [GT]n repeats) and long (L: ≥25 [GT]n repeats), as described in previous studies. The frequency of the HMOX1 (GT)n microsatellite in HAPE and non-HAPE subjects is shown in Table 2. The frequency of 2 short allele (SS), 1 short allele and 1 long allele (SL), and 2 long allele (LL) genotypes is 13.3%, 51.8%, and 34.9% in HAPE patients and 21.4%, 47.6%, and 31.0% in non-HAPE subjects, respectively. The frequency of S and L alleles is 39.2% and 60.8% in HAPE patients and 45.2% and 54.8% in non-HAPE subjects, respectively. There is no significant difference in the genotype frequency (P =.312) and allele frequency (P =.211) between the HAPE and non-HAPE subjects. Similar results can be obtained when these alleles are grouped into short (S: <27[GT]n repeats), intermediate (M: 27–32 [GT]n repeats), and long (L: ≥32 [GT]n repeats) alleles based on the literature. Table 3 shows that the frequency of SS, 1 short allele and 1 intermediate allele (SM), SL, 2 intermediate alleles (MM), 1 intermediate allele and 1 long allele (ML), and LL genotypes is 19.3%, 44.6%, 6.0%, 44.6%, 12.1%, and 0% in HAPE patients, and 27.6%, 32.4%, 9.0%, 19.3%, 10.3%, and 1.4% in non-HAPE subjects, respectively. In addition, the frequency of S, M, and L alleles is 44.6%, 46.4%, and 9.0% in HAPE patients, and 44.6%, 40.7%, and 11.0% in non-HAPE subjects, respectively. Again, no significant difference is observed in the genotype frequency (P = .357) and allele frequency (P = .468) between the 2 groups.
HMOX1 (GT)n microsatellite distribution in HAPE and non-HAPE subjects
HAPE, high altitude pulmonary edema; non-HAPE, non-high altitude pulmonary edema; S, short allelic frequency (<25 GT repeats); L, long allelic frequency (≥25 GT repeats); SS, two short alleles; SL, one short allele and one long allele; LL, two long alleles.
HMOX1 (GT)n microsatellite distribution in HAPE and non-HAPE subjects
HAPE, high altitude pulmonary edema; non-HAPE, non-high altitude pulmonary edema. S, short allelic frequency (<27 (GT)n repeats); M, middle allelic frequency (27≤(GT)n repeats<32); L, long allelic frequency (≥32 (GT)n repeats); SS, two short alleles; SM, one short allele and one intermediate allele; SL, one short allele and one long allele; MM, two intermediate alleles; ML, one intermediate allele and one long allele; LL, two long alleles.
It can be concluded that there is no significant difference in the allele frequency and genotype frequency between HAPE and non-HAPE subjects, thus suggesting that the (GT)n repeat polymorphism in the HMOX1 gene promoter may not be associated with the risk of HAPE.
The number of (GT)n repeats ranges from 16 to 38 and exhibits a typical bimodal distribution, with 1 peak located at 23 (GT)n repeats and the other peak at 30 (GT)n repeats, as shown in the Figure.
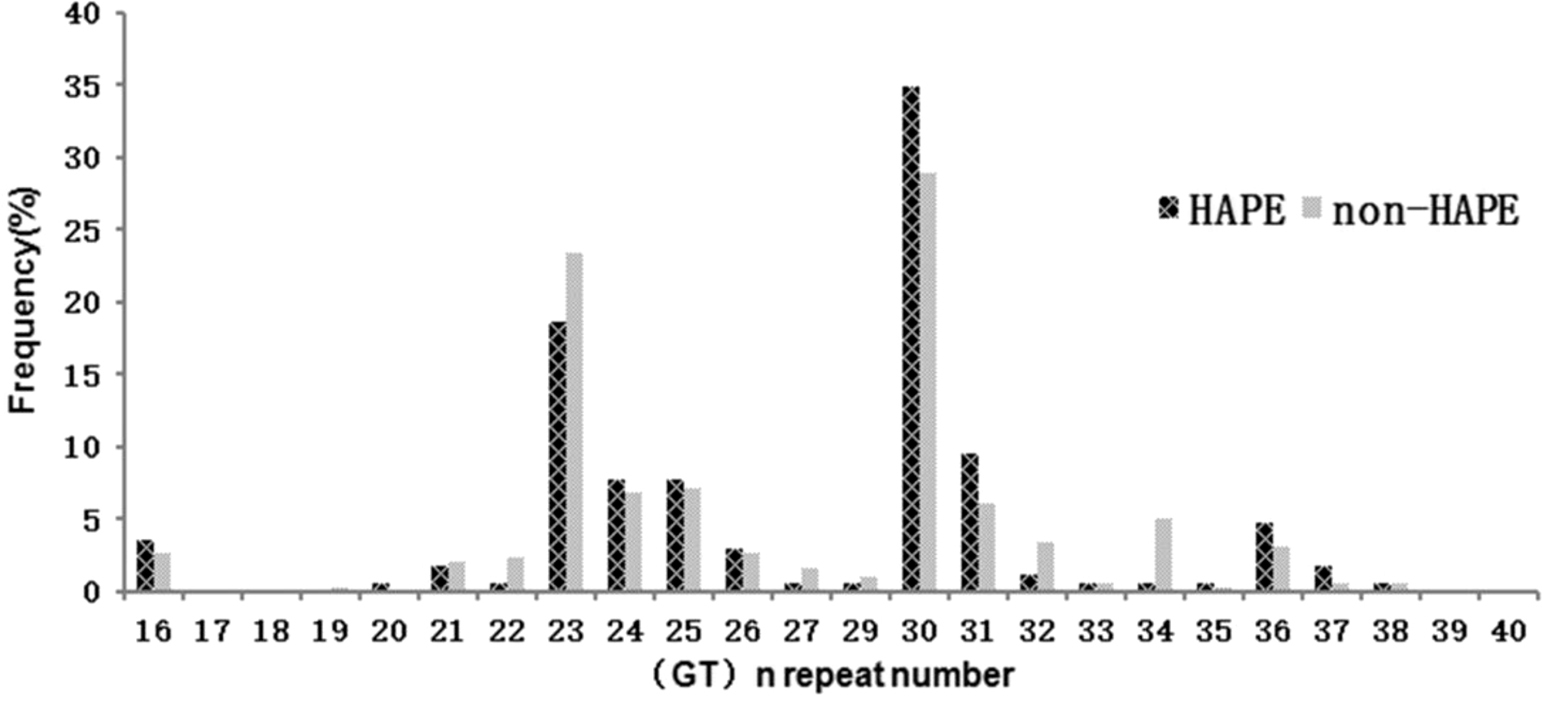
The frequency distribution of the number of (GT)n repeats in HAPE (n = 83) and non-HAPE (n=145) subjects. HAPE, high altitude pulmonary edema; non-HAPE, non-high altitude pulmonary edema.
Discussion
The major purpose of this study is to investigate the relationship between (GT)n microsatellite polymorphism in the HMOX1 gene promoter and HAPE in Han Chinese. However, we found no association between them in this study. HAPE is a noncardiogenic pulmonary edema that may occur in unacclimatized healthy lowlanders who rapidly ascend to altitudes above 2500 m. 12 It often occurs 2 to 4 days after arrival at high altitudes due to exaggerated hypoxic pulmonary vasoconstriction and pulmonary arterial hypertension. HAPE can be effectively treated with nifedipine, a pulmonary vasodilator, indicating that pulmonary hypertension is critical for the development of HAPE. 13
HAPE is characterized by oxidative stress and nonbacterial inflammatory response. Thus, hypoxia-induced oxidative stress plays an important role in the pathogenesis of HAPE, resulting in pulmonary vasoconstriction, pulmonary arterial hypertension, and eventually the development of HAPE. Under hypoxic conditions, the expression of erythropoietin (EPO) is activated via binding of hypoxia inducible factor-1. Increased secretion of EPO in response to hypoxemia has been reported in HAPE patients. However, our results revealed that the hemoglobin concentration is lower in HAPE patients (166.9±26.7) than in non-HAPE subjects (174.1±14.5) in Table 1 (P <.01), which is in contrast to previous results. One possible reason is that HAPE patients ascend high altitude rapidly and insufficient of EPO expression.
HMOX1, a rate-limiting enzyme in the catabolism of heme, is transcriptionally upregulated by hypoxia inducible factor-1 14 and may play a protective role as an antioxidating gene. It can indirectly reduce vasoconstriction during vascular stress. Expression of HMOX1 in pulmonary arteries stimulated vascular relaxation, mediated by guanylate cyclase and cGMP, independent of nitric oxide. 15 Overexpression of HMOX1 using the endogenous agonists of HMOX1 can prevent the development of hypoxia-induced pulmonary hypertension and vascular remodeling in rats. 16 However, the pathogenesis of HAPE remains poorly understood. Previous research has revealed that a rapid ascent to high altitudes results in hypoxemia in a majority of Han recruits who traveled to the Lhasa plateau (3658 m). 17 HMOX1 is considered a vascular protective factor in humans due to its anti-inflammatory and provasodilatory properties. 18 The (GT)n repeat polymorphisms in the HMOX1 gene promoter are closely related to the transcription activity of HMOX1. 19 HMOX1 transcriptional activity is high when (GT)n repeats <25 and low when (GT)n repeats ≥25. 20 In our study, the frequency of SS, SL, and LL genotypes is 13.3%, 51.8%, and 34.9% in HAPE patients, and 21.4%, 47.6%, and 31.0% in non-HAPE healthy subjects, respectively. The frequency of S and L alleles is 39.2% and 60.8% in HAPE patients, and 45.2% and 54.8% in non-HAPE subjects, respectively. There is no significant difference in the genotype frequency and allele frequency between HAPE and non-HAPE subjects. Thus, it can be concluded that the (GT)n repeat polymorphism in the HMOX1 promoter may have no effect on the susceptibility to HAPE in Han Chinese.
There is a general consensus that HAPE is a multifactorial disease resulting from the complex interaction of environmental factors and multiple genetic polymorphisms. Thus, it is unlikely that a polymorphism in a single gene would profoundly influence the risk of HAPE. A wide variety of genes have been suggested to be involved in the development of HAPE, including EPO, genes encoding glycolytic enzymes such as aldolase A, enolase I, lactate dehydrogenase A, phosphofructokinase L, inducible nitric oxide synthase, vascular endothelial growth factor, angiotensin-1 convertingenzyme, tyrosine hydroxylase, serotonin transporter, 21 prolyl hydroxylase, surfactant protein A2, and endothelin-1 (a potent vasoconstrictor). 22 In addition, important risk factors of HAPE include male sex, cold, vigorous exercise, and poor physical health. However, the exact molecular mechanism of HAPE remains to be elucidated.
Our study is limited in several respects. First, the subjects included in this study were largely young men enrolled from a single center. Second, the negative result of this study may be related to a small sample size and the gene−gene interaction. Third, we were unable to detect HMOX1 promoter and the coding region single-nucleotide polymorphic sites. This is the focus of our future studies.
Author Contributions: Study concept and design (XFC, RLG); acquisition of the data (XFC, LM, SM, JX); analysis of the data (XFC); drafting of the manuscript (XFC); critical revision of the manuscript (RLG); and approval of final manuscript (XFC, RLG, LM, SM, JX).
Financial/Material Support: This study was supported by grants from National Program on Key Basic Research Project of China (No.2012CB518200); National Natural Science Foundation of China (No.31571231); Qinghai-Utah Joint Research Key Lab for High Altitude Medicine. (No. 2014-ZJ-Y39).
Disclosures: None
Footnotes
Submitted for publication July 2016.
Accepted for publication December 2016.
