Abstract
The infection of feline thymic lymphoma 3201 cells with a cell culture-adapted Petaluma strain of feline immunodeficiency virus (FIV) led to the establishment of survivor cells designated as 3201-S after a productive infection associated with extensive cell killing. 3201-S cells were free of FIV DNA, and were found to express CXCR4, a coreceptor for infection but not CD134, a primary receptor. When 3201-S cells were reinfected with FIV, viral DNA was transiently detectable for 5 days postinfection, indicating that 3201-S cells cannot support the FIV replicative cycle. Furthermore, comparative studies found that in contrast to SDF-1α-responsive 3201 cells, 3201-S cells did not show a flux of Ca2+ in response to SDF-1α, implying that CXCR4 is not functionally active on 3201-S cells. These results suggest that 3201 cells can be heterogeneous in the phenotype of the CXCR4 expressed, and this heterogeneity may account for the differences in susceptibility to FIV. Determining the mechanism(s) within 3201-S cells that restrict FIV could result in therapeutic strategies against FIV infection.
The development of appropriate animal models of acquired immunodeficiency syndrome (AIDS) is critical to better understand the pathogenesis of lentivirus-induced immune deficiency which is etiologically linked to human immunodeficiency virus type 1 (HIV-1). 1 Only two animal lentiviruses have thus far been implicated in AIDS-like diseases, feline immunodeficiency virus (FIV) 2 and simian immunodeficiency virus (SIV). 3 The use of SIV in macaques provides an opportunity to evaluate a primate lentivirus within a species closely related to humans. A major concern, however, is the relative shortage of non-human primates available for infectivity trials as well as for safety and efficacy studies on candidate vaccines and antiviral drugs. The advantages of a feline model of AIDS are not only the similarities to human AIDS, but also practicality and cost compared with the use of non-human primates. 4 Hence understanding more about lentiviruses in cats not only helps humans but also provides much needed information to help cats. As with HIV, FIV primarily targets CD4+ T cells, but it uses CD134, a T cell activation and co-stimulatory molecule, and not CD4 as a primary receptor, 5,6 inducing an AIDS-like immunodeficiency in its natural host species due to a progressive depletion of CD4+ T cells. 7 Through comparative studies there is the potential for development of safe and effective therapies for the natural species at risk, cats, and the feline animal model thus provides the research community with a well-established research species and an easily handleable animal. Moreover, FIV has the advantage that a high level of biocontainment at laboratory facilities is not required, thereby representing a valuable model for HIV/AIDS research. This model has already been used for the testing of antiviral drugs. 8–10
We have described earlier that the feline lymphoma cell line 3201 was highly susceptible to the cytopathic effects of FIV-Petaluma. Moreover, 3201 cells that survived FIV infection established a chronic FIV-producer cell line (3201/FIV) during a chase period of 2 months. 11 Recently, we repeated this experiment and found that the 3201/FIV cell line was established consistent with our earlier observations, but the frequency of FIV antigen-positive cells decreased gradually over time during further incubation. On day 86 postinfection, none of the FIV-surviving cells which were designated as 3201-S was any longer positive for viral antigens. ‘(Tochikura, TS, unpublished observations)’. This is of interest considering that 3201-S cells have not yet been characterised.
In the present study, we demonstrated that viral DNA was no longer detected in 3201-S cells. Although proviral DNA was detected in the FIV-reinfected 3201-S cells early in the infection, the viral genome was no longer detectable afterwards, suggesting that 3201-S cells were unable to support detectable viral replication. As it has been reported that 3201 cells express relatively high levels of CXCR4, a chemokine receptor whose natural ligand has been identified as the chemokine SDF-1 (stromal cell derived factor), necessary for the efficient infection of target cells, 12,13 we first assessed the 3201-S cells for CXCR4 expression by flow cytometric analysis using a panel of CXCR4 monoclonal antibodies recognizing antigenically distinct conformations of CXCR4. 14 We have found that both 3201 and 3201-S cells express this chemokine receptor, although the parental 3201 cells could be distinguished from 3201-S cells in that the former cells express higher levels of CXCR4 with respect to fluorescence intensity. Of particular interest to the present study, 3201-S cells were unable to support the viral replicative cycle after reinfection with FIV. Furthermore, SDF-1α stimulates calcium mobilisation in 3201 but not in 3201-S cells, suggesting that the receptor is not functional in 3201-S cells. These findings led us to hypothesise that phenotypic heterogeneity exists in the original 3201 cells and their differential expression of CXCR4 may influence CXCR4-dependent FIV entry and responsiveness to SDF-1α. Our data could have important implications for understanding intrinsic resistance to the virus in infected animals.
Materials and methods
Cells
3201 cells, 15 kindly provided by Dr Lawrence Mathes (The Ohio State University, Columbus, OH, USA), are a feline thymic lymphoma cell line. Cells were cultured in a medium consisting of equal parts Leibovitz L-15 medium and RPMI 1640 medium supplemented with 10% heat-inactivated fetal bovine serum (FBS), penicillin (100 IU/ml) and streptomycin (100 μg/ml). 3201 cells were routinely passaged 1:5 every 3 to 4 days and incubated at 37°C. Feline peripheral blood mononuclear cells (PBMCs) were obtained from a specific pathogen-free (SPF) adult cat (Shimizu Laboratory Supplies, Kyoto, Japan). The PBMCs were initially stimulated with 5 μg of phytohemaglutinin (PHA) per ml for 3 days, maintained on 5 ng of recombinant human interleukin-2 (IL-2) per ml, and replenished with fresh PHA and IL-2-stimulated PBMCs in a complete RPMI 1640 medium with 20% FBS every 5 to 7 days.
Antibodies
The following antibodies were used in this study: mouse anti-feline/human CXCR4 monoclonal antibodies (MAbs) – clones 44708 and 44716 (NIH AIDS Research and Reference Reagent Program, Rockville, MD, USA), and clone 44717 (R&D Systems, Minneapolis, MN, USA); anti-CD134 MAb reacting with feline CD134 but only weakly with human CD134 (Affinity BioReagents, Golden, CO, USA); sera from FIV-infected cats (kindly provided by Dr Makoto Hitomi and Dr Haruo Konishi) and control plasma from an uninfected SPF cat (Shimizu Laboratory Supplies, Kyoto, Japan).
Preparation of virus
A cell culture-adapted strain of FIV-Petaluma was obtained from the NIH AIDS Research and Reference Reagent Program (Rockville, MD, USA). Tissue culture supernatant from 3201 cells persistently infected with FIV(3201/FIV) was used as a source of infectious virus. After more than 80% of the 3201 cells became positive for FIV antigens, as detected with an indirect immunofluorescence assay (IFA), the culture supernatant was filtered through a 0.45 μm Millipore membrane filter and stored at −70°C in small aliquots until used. The viral titre was 103.8 50% tissue culture infectious dose per ml assayed in 3201 cells. 11
Infection
Target cells were infected with FIV at a multiplicity of infection (MOI) of 0.04. After the adsorption of FIV at 37°C for 1 h, cells were cultured in complete medium at 37°C in 5% CO2. The cells were subcultured every 3 to 4 days. The efficiency of viral infection was analysed based on cell viability, monitored with the trypan blue dye exclusion test, and by IFA. 11
Immunofluorescence assay (IFA)
The expression of FIV antigen in FIV-infected cells was detected by indirect immunofluorescence. 11 Cells were smeared on microscope slides and air-dried. Smeared cells were fixed with cold methanol for 15 min and air-dried. The slides were incubated for 45 min at 37°C in a humidified atmosphere with a 1:400 dilution of serum from an FIV-infected cat (IFA titre to FIV; 1:2560) or control plasma from an uninfected SPF cat. The slides were then washed twice in PBS and incubated for 30 min at 37°C with fluorescein-isothiocyanate (FITC)-conjugated anti-cat IgG (Serotec, Raleigh, NC, USA). The slides were next washed twice with PBS, and mounted with 50% glycerol in PBS. Positive cells were examined by epifluorescence microscopy. For the calculation of percentage, over 500 cells were counted.
Flow cytometry
Flow cytometric analyses were performed as described previously 16 with minor modifications. Briefly, 2×105 cells were washed by centrifugation and resuspended in FACS buffer ([1% FBS−PBS]+0.1% sodium azide). Cells were then incubated with anti-CXCR4 MAbs or anti-CD134 MAb for 40 min at room temperature. After being washed with PBS, cells were incubated with FITC-conjugated anti-mouse IgG (Cappel) in the dark for 20 min at room temperature. After another wash with PBS, stained cells were resuspended in FACS buffer, examined with a FACScan flow cytometer (Becton Dickinson, San Jose, CA, USA), and then analysed using FlowJo software (Tree Star, Ashland, OR, USA). Data were collected from approximately 10,000 events for each experimental condition, and results are expressed as mean fluorescence intensity (MFI) and a single-parameter log fluorescence histogram.
Polymerase chain reaction (PCR)
Petaluma 17 FIV gag and envelope (env) genes were amplified by nested PCR as described previously. 18,19 Briefly, a 329-base pair (bp) fragment corresponding to the region from 1036 to 1364 of the gag gene was generated with an outer primer set (sense, nucleotides 917-936, 5′-AATATGACTGTATCTACTGC-3′;antisense, nucleotides 1650-1628, 5′-TTTTCTTCTAGAGTACTTTCTGG-3′) and an inner primer set (sense, nucleotides 1036-1055, 5′-TATTCAAACAGTAAATGGAG-3′;antisense, nucleotides 1364-1345, 5′-CTGCTTGTTGTTCTTGAGTT-3′). An 873-bp fragment, corresponding to the region from 7207 to 8079 of the env gene, was generated with an outer primer set (sense, nucleotides 7032-7053, 5′-GATGGAATGAGACTATAACAGG-3′;antisense, nucleotides 8119-8098, 5′-CAACATAACATGAATAGCTGCC-3′) and an inner primer set (sense, nucleotides 7207-7230, 5′-CCAACAATTAAGTTATTGTACAGA-3′; antisense, nucleotides 8079-8058, 5′-TATTCCATGACTTCCTCTGGGA-3′). PCR was performed in 0.2 ml microfuge tubes in a total volume of 50 μl. A DNA sample (a DNA amount equivalent to 2×105 cells in 10 μl) was added to 5 μl of 10-fold concentrated reaction buffer (Takara, Kyoto, Japan), 5 μl of deoxynucleotide mix (Takara, Kyoto, Japan) containing 2.5 mM of each, 2 μl of 20 μM primer mix, 0.5 μl of Taq polymerase (5 unit/μl; Takara, Kyoto, Japan) and 37.5 μl of distilled water, and then mixed. The DNA was amplified with a DNA thermal cycler (Applied Biosystems, Foster, CA, USA). The amplification involved incubation for 5 min at 94°C as the first denaturation, followed by 30 cycles of denaturation for 1 min at 94°C, primer annealing for 1 min at 55°C and synthesis for 1.5 min at 72°C, and a final incubation for 5 min at 72°C. The nested PCR products were electrophoresed through a 2% agarose gel. The gel was incubated with ethidium bromide, and the bands were visualised by an ultra-violet transilluminator at 312 nm, and then photographed.
The real-time PCR assay was performed with a DNA Engine Opticon Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA) by using a QuantiTect SYBR Green PCR Kit (Qiagen KK, Tokyo, Japan) according to the manufacturer's instructions. Briefly, each sample was amplified in a final volume of 20 μl containing 10 μl of 2× Quantitect SYBR Green PCR Master Mix, 0.3 μM of inner primer for the gag gene whose sequence was the same as that used for the nested PCR, and 5 μl of extracted DNA dilution (sample) or a recombinant DNA plasmid encoding for a FIV gag protein derived from the Aomori 2 strain. 20 Cycling parameters were 15 min at 95°C for the hot start Taq polymerase activation followed by 55 cycles of 15 s at 94°C, 30 s at 55°C and 30 s at 72°C. The reaction was carried out testing all the samples and five standard plasmid dilutions in duplicate together with a no-template control.
Calcium flux measurements
Calcium flux in response to SDF-1α was measured using a modification of a method described previously. 16 Either 3201 or 3201-S cells were loaded with a fluorescent probe, Indo-1 AM (1 μM; Dojindo, Tokyo, Japan), in RPMI 1640 medium for 30 min at 37°C. The cells were washed twice with Hanks' balanced salt solution (HBSS), and then adjusted to 5×105 cells/ml in HBSS. After pre-heating for 5 min at 37°C, the cells were stimulated with SDF-1α (1 μg/ml, recombinant human SDF-1α; PeproTech, London, UK).
Fluorescent emissions at wavelengths of 405 and 485 nm were monitored upon excitation at 338 nm using a spectrofluorometer RF-1500 (Shimadzu, Kyoto, Japan). The calculation of the change in intracellular Ca2+ concentrations in combination with digitonin permeabilisation and EGTA treatment was done using the Intracellular Ca2+ Measurement Program Pack according to the manufacturer's instructions (Shimadzu, Kyoto, Japan).
Results
Establishment of 3201 cells surviving FIV infection
After the exposure of 3201 cells to cell-free culture fluids derived from the 3201/FIV cell culture, continuous growth of the infected cells and production of FIV were observed as described previously, 11 however, the frequency of FIV antigen-positive cells decreased gradually over time during further incubation. When the surviving cells were evaluated at 86 days postinfection (dpi) for their ability to be a reservoir of FIV, neither infectious virus nor FIV antigen-expressing cells could be detected by IFA even after treatment with IUdR or phorbol myristate acetate, a well-known inducer of retroviruses (data not shown). 21,22 The nonvirus-producing survivor cells, here referred to as 3201-S, were indistinguishable in morphology and growth rate from mock-infected 3201 cells (data not shown).
To determine if FIV cDNA was present in the FIV-surviving 3201-S cells after exposure to FIV, samples were examined by nested PCR. DNA isolated from cell cultures derived from virus-producing 3201/FIV cells showed the 329-bp and 873-bp products specific to FIV gag and env, respectively, whereas no specific bands were detected in samples from 3201-S and mock-infected 3201 cells (Fig. 1).

Agarose gel showing the results of PCR amplification of DNA for the FIV gag-, and env gene regions obtained from the indicated cells. A nested PCR was used to amplify the FIV gag or env gene sequence as described in the materials and methods section. The positions of bands of 329 bp and 873 bp corresponding to gag and env, respectively, are shown. Lanes: M, DNA size marker (100-bp ladder); 1 and 2 (duplicate), mock-infected 3201 cells; 3–6 (quadruplicate), 3201-S cells; 7 and 8 (duplicate), 3201/FIV cells.
Absence of CD134 expression in 3201 cells
CD134 has been reported to be expressed in PBMCs and IL-2-dependent T cell lines, and to represent a primary target of FIV. 5 An analysis was undertaken next to determine whether CD134 expression might be involved in FIV infections among 3201 cell derivatives. Consistent with previous findings that 3201 cells lack CD134 expression, 6 the binding of anti-CD134 MAb, detected by flow cytometry and indicated as mean fluorescence intensity (MFI), was observed on activated PBMCs used as a positive control but not on 3201 or 3201-S cells (more than a fivefold difference in MFI, data not shown), confirming that CD134 is not likely to contribute to FIV infection in 3201 cells.
Presence of CXCR4 expression in 3201 and 3201-S cells
It has been shown that 3201 cells express relatively high levels of CXCR4, used by FIV for infection. 12,13 The cell-surface expression of CXCR4 on 3201, 3201-S, and 3201/FIV cells was next evaluated by flow cytometry using anti-CXCR4 44717 MAb. As shown in Fig. 2, a profound shift was seen in the fluorescence histogram of anti-CXCR4 stained cells when compared to the background (FITC-conjugated secondary antibody alone), indicating the presence of cell surface CXCR4 protein on 3201 cells. The amount of CXCR4 was markedly reduced in the FIV-infected 3201 cells, with more than 90% of the cells positive for FIV antigens. This finding is in agreement with a report that the surface expression of CXCR4 was markedly reduced on Crandell feline kidney (CRFK) cells after FIV infection, presumably due to either a down-regulation of CXCR4 expression or the elimination of CXCR4-expressing cells from the culture. 12 Of particular interest is that approximately 45% of the cells in the 3201-S culture were recognised by the MAb, albeit with a low intensity, compared to almost 100% and 2% (1.6-fold above the background level) of the cells in 3201 and 3201/FIV controls, respectively (data not shown). This will be discussed later.
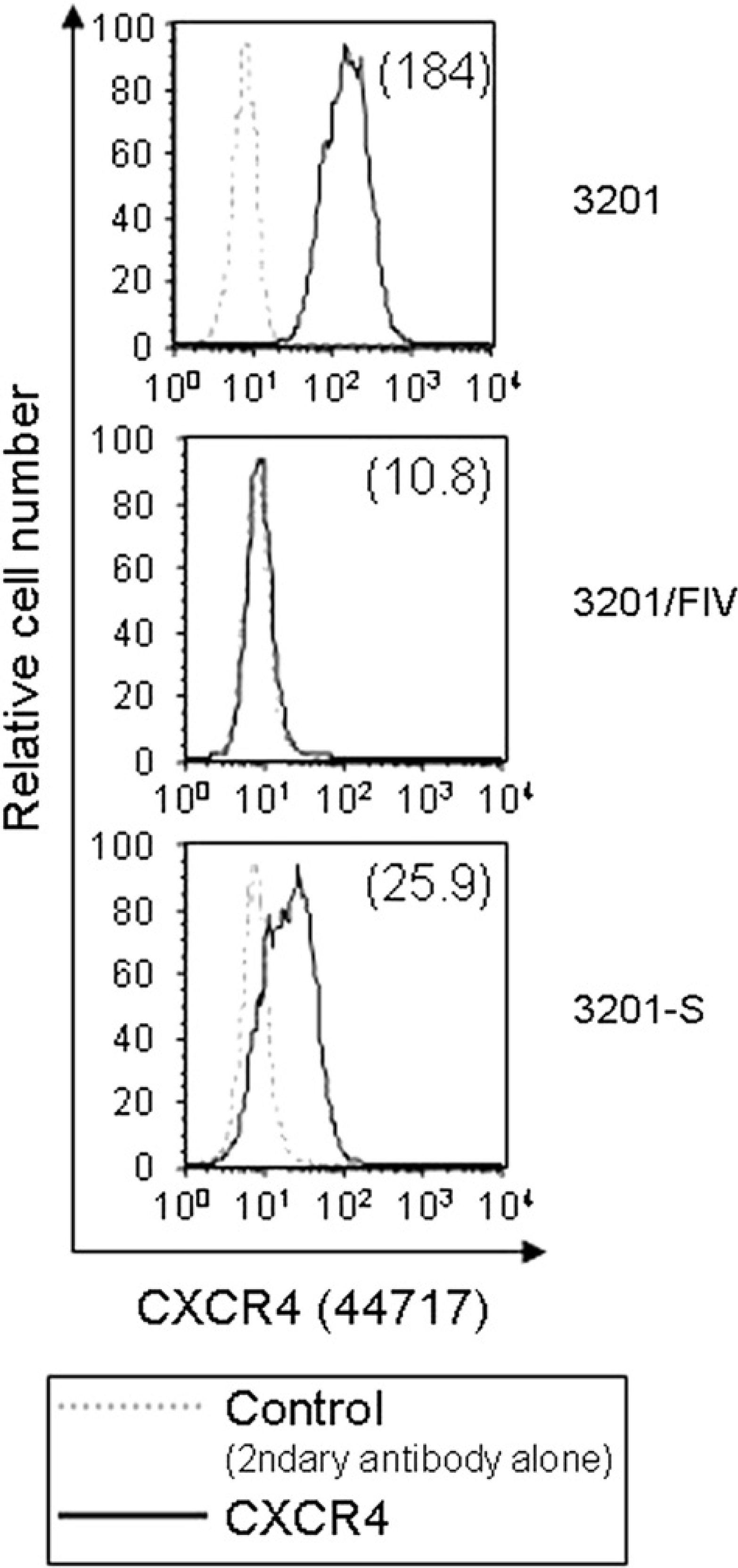
Flow cytometric analysis of CXCR4 expression on 3201, 3201-S, and 3201/FIV cells. Cells were stained with anti-CXCR4 44717 MAb, followed by FITC-conjugated anti-IgG (bold line). For all panels, data are shown as relative cell number (y-axis) plotted against relative fluorescence intensity (x-axis). The background staining is the signal derived from incubation of the cells with the FITC-conjugated secondary antibody only (dashed line). MFI is indicated in parentheses.
Given previous reports that CXCR4 existed in antigenically distinct states in most cell types, presumably due to conformational heterogeneity, 14 we investigated the ability of other two commercially available MAbs against CXCR4 (clones 44708 and 44716), compared with that of clone 44717, to recognise CXCR4 on 3201-S cells as well as on 3201 cells. In agreement with previous findings that clone 44717 reacted with feline CXCR4 most efficiently among the three MAbs, 14 3201 cells appeared to express CXCR4 at the highest levels when stained with clone 44717 (Fig. 3). We, therefore, selected clone 44717 for further studies.
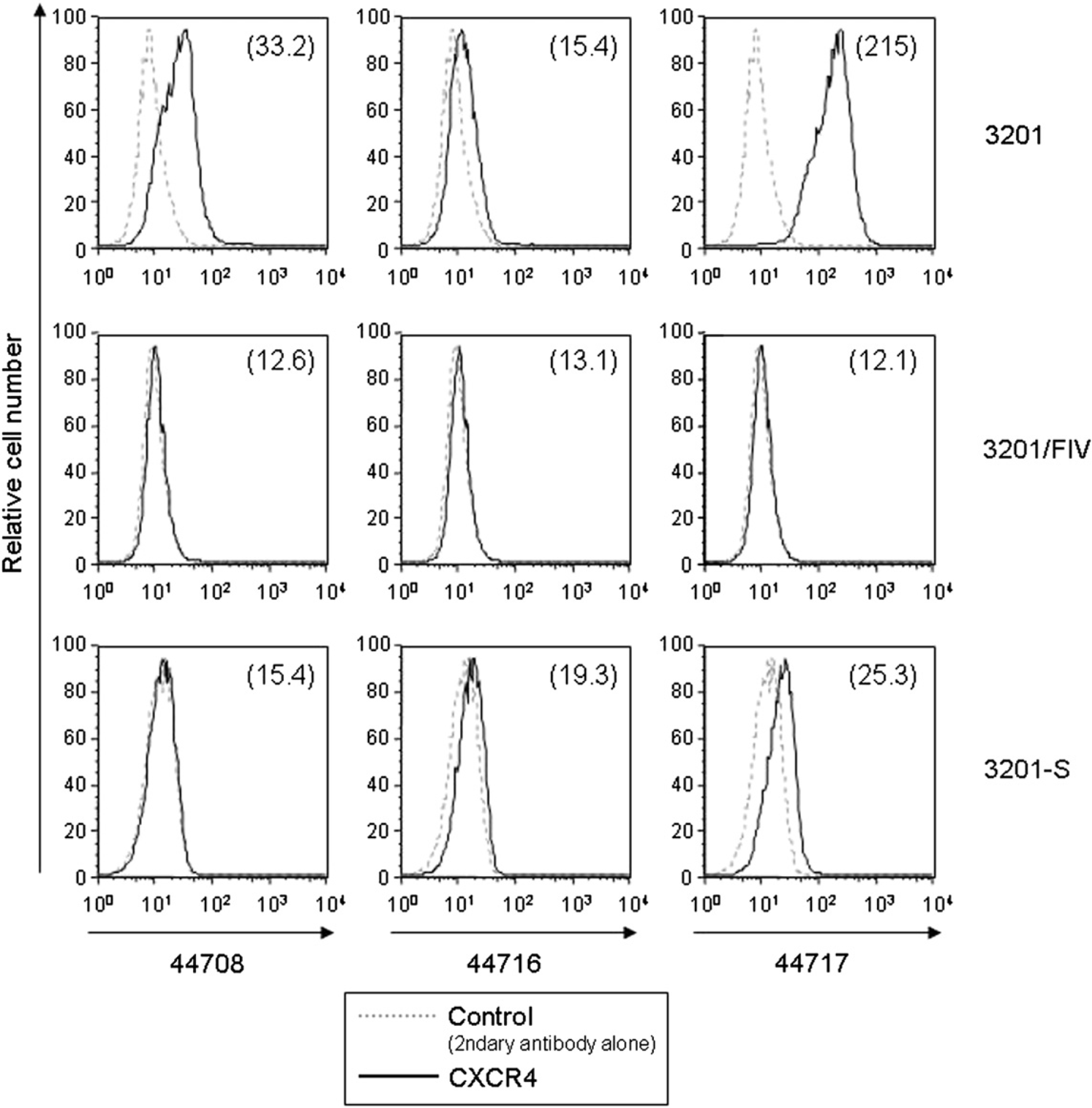
Comparative CXCR4 expression on 3201, 3201-S, and 3201/FIV cells assessed by flow cytometry using a panel of MAbs. Cells were incubated with anti-CXCR4 MAb clones 44708, 44716 or 44717, followed by staining with FITC secondary antibody. Designations are the same as in the legend to Fig. 2.
Demonstration of the presence of FIV DNA in FIV-reinfected 3201-S cells
To study the susceptibility of 3201-S cells to FIV reinfection, the cells were exposed to the culture supernatant of 3201/FIV cells at a MOI of 0.04. Viral antigens of FIV were first detected in 3201 cells by IFA as early as 3 dpi, and the frequency of detection increased gradually with time reaching more than 90% within a week (data not shown), whereas 3201-S cells remained negative indistinguishably from mock-infected 3201 cells during a 3-month culture period (data not shown). To further confirm the failure of FIV to reinfect 3201-S cells, DNA was extracted 5, 15 and 30 dpi from 3201-S cells that had been reinfected with FIV and analysed for FIV sequences by PCR amplification. Interestingly, FIV DNA was detected in the 3201-S cells reinfected with FIV 5 dpi by nested PCR, the copy number determined by real-time PCR being 4 to 7 per 106 cells, whereas FIV-infected 3201 cells tested positive for FIV throughout the experiment (Table 1). However, the viral genome in the FIV-reinfected 3201-S cells was no longer detectable at the subsequent testing points, indicating that FIV infection of 3201-S cells is inefficient compared to that of parental 3201 cells.
Detection of FIV DNA in the indicated cells after infection or reinfection with FIV.

Experiments were carried out in duplicate. ND=not done.
+=Positive result; −=negative result in nested PCR assay.
Result of real-time PCR assay is expressed as copy number per 106 cells.
Effect of FIV reinfection on the expression of CXCR4 on 3201-S cells
As FIV DNA was detected in the 3201-S cells at 5 dpi (Table 1), we investigated the effect of FIV reinfection on CXCR4 expression on 3201-S cells (Fig. 4). While approximately 20% of 3201-S cells expressed CXCR4 before FIV reinfention compared to 99% of 3201 cells, a similar proportion of CXCR4+ cells, ranging from 10% to 30%, was maintained over the course of the culture up to 30 dpi on the FIV-reinfected 3201-S cells (data not shown). Newly infected 3201 cells exhibited a profound decrease in CXCR4 cell surface expression, consistent with Figs 2 and 3, in which the frequency of FIV antigen-positive cells in the culture was 81%, 95%, and 90% at 5, 15, and 30 dpi, respectively (data not shown). These results indicated that the level of CXCR4 expression on 3201-S cells was not likely to be affected by FIV infection.

Effect of FIV infection or reinfection on the expression of CXCR4 on 3201 cells and 3201-S cells. Samples of mock-infected 3201, and FIV-infected 3201 or FIV-reinfected 3201-S cells were collected at the indicated time points, and processed for analysis by flow cytometry. Cells were incubated with anti-CXCR4 44717 MAb, followed by staining with FITC secondary antibody. Designations are the same as in the legend to Fig. 2. Cells were processed in parallel for analysis by the PCR (Table 1).
CXCR4 is not functional on 3201-S cells
The activation of calcium flux in CXCR4-expressing human cells is related to HIV infection and as reported, stimulation of CXCR4 by SDF-1α increases calcium flux in several HIV- susceptible cell types. 16,23 Thus, we became interested in whether 3201-S cells will respond to SDF-1α through calcium flux, and conducted a calcium mobilisation experiment to determine whether the CXCR4 expressed by 3201-S cells is a functional receptor. As shown in Fig. 5, SDF-1α, whose concentration used in this study has previously been shown to stimulate CXCR4 to near maximal levels, 23 clearly increased the intracellular Ca2+ level in the original 3201 cells. In contrast, SDF-1α failed to stimulate a response in 3201-S cells, suggesting that CXCR4 is not functionally active on 3201-S cells.

Comparison of SDF-1α responsiveness in 3201 and 3201-S cells. Cells were loaded with Indo-1 AM (1 μM) and incubated in RPMI 1640 medium for 30 min at 37°C in 5% CO2, followed by stimulation at the indicated time point (arrow) with 1 μg/ml SDF-1α. The intracellular Ca2+ concentrations were calculated as described in the materials and methods section.
Discussion
The primary receptor for FIV is CD134, a member of the tumour necrosis factor receptor superfamily, which is specifically expressed on activated CD4+ T cells. 5,6 FIV infection requires the expression of a coreceptor, the chemokine receptor CXCR4, 24 and the virus binds these receptors via its surface glycoprotein gp95. Some primary and cell culture-adapted strains of FIV such as Petaluma, however, can infect in the absence of CD134. 5 CD134-independent FIV infection is mediated by direct interaction with CXCR4, 24 analogous to infection with CD4-independent strains of HIV. 25 In research on FIV, the 3201 cell line is often used as an in vitro model. 6,11,18 Our present observation is consistent with the previous finding that 3201 cells abundantly express CXCR4 but not CD134. 6,12,13 It can thus be concluded from these studies that CD134 is not likely to be involved in the infection of 3201 cells by FIV.
We focused on CXCR4 expression in 3201-S cells, and examined whether its down-regulation may account for the inability of FIV to reinfect 3201-S cells. Interestingly, we demonstrated by flow cytometric analysis that 3201 cells vary in their affinity for feline CXCR4 staining with three MAbs (clones 44708, 44716, and 44717) in the following order: 44717>44708>44716 (Fig. 3). CXCR4 possesses four extracellular domains: an N-terminal region and three extracellular loops (ECL1, ECL2, and ECL3). 14 Based on epitope mapping using a panel of MAbs including clones 44708, 44716, and 44717, it seems likely that conformational differences in the CXCR4 ECL domains, in particular in ECL2, are responsible for the CXCR4 antigenic heterogeneity. Our finding that clone 44717 exhibited the strongest affinity for feline CXCR4 among the three MAbs was consistent with that reported for a transformed human T cell line, 14 confirming that CXCR4 on 3201 cells similarly exhibits considerable conformational heterogeneity. Feline CXCR4 displayed 94.9% amino acid sequence identity with human CXCR4, 24 and as the MAbs used in the present study react specifically not only with human and non-human cells expressing human CXCR4, but also with cells expressing feline CXCR4, 14 it is conceivable that this may also be related to the level of cross-reactivity between human and feline CXCR4 at the particular epitope recognised by the MAbs.
The presence of FIV DNA in the 3201-S cells at 5 dpi, albeit at an extremely low frequency, suggests that 3201-S cells are reinfectable by FIV as determined by the PCR assay, however, FIV failed to spread in 3201-S cells as demonstrated by the lack of any persistent presence of viral DNA (Table 1). Calcium flux experiments showed that 3201 cells but not 3201-S cells responded to SDF-1α with a rapid mobilisation of calcium, suggesting that CXCR4 is functionally active and is able to couple to G-protein signaling mechanisms, 26 as opposed to 3201-S cells whose CXCR4 receptor is likely to be dysfunctional. Our interpretation of these results is that 3201 cells grow as heterogeneous populations that are present in the starting culture prior to FIV infection. One population, representing the majority of the culture, expresses CXCR4 which supports FIV infection, but the others, comprising an extremely small number of cells, do not confer susceptibility to FIV infection possibly due to insufficient levels of CXCR4 expression. After the productive infection of 3201 cells, FIV-infected cells accompanied by viral antigens were probably eliminated from the culture due to cytopathic effects, and a small percentage of the cells no longer susceptible to FIV infection survived to replace the former population. Although little is known about why 3201-S cells are unable to serve as a reservoir for FIV, at least two models can be considered. In the first model, CXCR4 is impaired, though it remains to be seen whether this would be due not only to its relatively low constitutive levels, but also the extremely small percentage of cells involved in the expression, or CXCR4 differs either structurally or functionally between 3201 and 3201-S cells, with FIV infection and responsiveness to SDF-1α. The role of chemokines such as the β-chemokines RANTES, MIP-1α, and MIP-1β in natural resistance to infection by HIV-1 has been established, with highly exposed persistently seronegative individuals found to be homozygous for a 32-bp deletion in the CCR5 gene encoding a receptor for those β-chemokines, and another co-receptor for HIV-1. 27 If this should be the case with the CXCR4 observed in 3201-S cells, the dysfunctional CXCR4 may contribute to protection from FIV. Alternatively, in the second model, 3201-S cells carry restriction factors that confer resistance to replication by FIV. Recent in vitro studies showed that the existence of resistance factors (eg, Lv1, Ref1, and TRIM5α, etc) in various species of animals can inhibit a range of retroviruses including lentiviruses by targeting the capsid of incoming viruses to block infection, indicating susceptibility to retroviral infection to be determined, in part, by host genes with antiviral activity. 28,29 Our results thus suggest that the post-entry pathway followed by FIV could differ in 3201-S cells compared to parental 3201 cells. It remains to be elucidated whether similar restriction factor(s) participate in the inhibition of FIV replication in 3201-S cells.
In conclusion, it would appear that 3201 cells can be heterogeneous in the phenotype of the CXCR4 expressed, and this heterogeneity may account for the differences in susceptibility to FIV. Thus, 3201-S cells represent the first unique subpopulation of CXCR4+ T cells that are resistant to FIV replication, and may emerge as a consequence of selection during infection. Taken together, these findings may provide the basis for further studies to understand the role of 3201-S cells during FIV infection and identify host factors that could restrict the virus.
Footnotes
Acknowledgements
We are grateful to Dr Nobuyuki Ito, Dr Nobutaka Fujii and especially the late Dr James R Blakeslee Jr for their constant support and encouragement, and Dr Hiromu Takematsu and Masakazu Ohashi for their technical assistance. We also wish to thank Dr Ulrike Ziegner and Dr Karin Nielsen for critically reviewing the manuscript and helpful discussions. The following reagents were obtained through the NIH AIDS Research and Reference Reagent Program, Division of AIDS (DAIDS), NIAID, NIH as indicated (Reagent Program catalog numbers are shown in parentheses): FIV-Petaluma (236) from Dr Niels Pedersen and Dr Janet Yamamoto; CXCR4 Monoclonal 44708 (4084) from DAIDS, NIAID, produced by R&D Systems; CXCR4 Monoclonal 44716.111 (4085) from DAIDS, NIAID.
