Abstract
Early information on the metabolism and toxicity properties of new drug candidates is crucial for selecting the right candidates for further development. Preclinical trials rely on cell-based in vitro tests and animal studies to characterize the in vivo behavior of drug candidates, although neither are ideal predictors of drug behavior in humans. Improving in vitro systems for preclinical studies both from a technological and biological model standpoint thus remains a major challenge. This article describes how microfluidics can be exploited to come closer to this goal in combination with precision-cut liver slices (PCLS) as an improved organomimetic system. Recently, we developed a novel microfluidic-based system incorporating a microchamber for slice perifusion to perform drug metabolism studies with mammalian PCLS under continuous flow. In the present study, the viability and metabolism of human PCLS were assessed by the measurement of the leakage of liver-specific enzymes and metabolism of four different substrates: lidocaine, 7-hydroxycoumarin, 7-ethoxycoumarin, and testosterone. All experiments were verified with well plates, an excellent benchmark for these experiments. Clearly, however, human tissue is not readily available, and it is worth considering how to perform a maximum number of informative experiments with small amounts of material. In one approach, the microfluidic system was coupled to an HPLC system to allow on-line monitoring and immediate detection of unstable metabolites, something that is generally not possible with conventional well-plate systems. This novel microfluidic system also enables the in vitro measurement of interorgan interactions by connecting microchambers containing different organ slices in series for sequential perfusion. This versatile experimental system has the potential to yield more information about the metabolic profiles of new drug candidates in human and animal tissues in an early stage of development compared with well plates alone.
Keywords

Introduction
Only one in nine newly developed drugs entering clinical trial will be approved by the European and/or the U.S. regulatory authorities. 1 Many of the drugs that fail are withdrawn due to unforeseen toxicity or lack of efficacy. 2 This means that preclinical tests, which include both in vitro and in vivo methods, are not able to adequately predict how new drugs will act in the human body. There is thus a continuing demand to develop in vitro systems that are able to better mimic the in vivo environment. 3 Preclinical tests normally use animals (in vivo tests) and human and animal tissues, cells, or cell fragments (in vitro tests). 4 Taking into account the extrapolation of results obtained with animal material in vitro to those obtained in vivo for tested animals, a prediction of the human in vivo behavior of a drug can be made based on in vitro data obtained from human material. However, as interspecies differences are often evident with respect to the behavior of animal and human cells or tissues under culture conditions, each in vitro system developed with animal material should be verified for its applicability to human material. If an in vitro system based on human material with excellent correlation to the in vivo situation were to exist, animal experiments could be reduced and replaced. To develop an in vitro system that correlates well with the in vivo situation, attention should be paid to the selection of a biological system that represents the physiology of the organ as closely as possible. Although there are various models to determine absorption, distribution, metabolism, excretion, and toxicity profiles, only a few exhibit a good correlation to the in vivo situation, among them primary hepatocytes and precision-cut liver slices (PCLS). 5 The main advantage of PCLS compared with primary hepatocytes is that PCLS contain all liver cell types in their physiological environment. 6 This model is thus also very useful to study toxicity processes that are mediated via multicellular mechanisms.
Normally, PCLS are incubated in well plates or dynamic roller culture systems. 7 In both systems, the slices are cultured in an excess of medium that is refreshed only once every 24 h. This leads to an accumulation of waste products and a decline or even depletion of medium components. By applying microfluidic technologies, it becomes possible to maintain a constant environment around the slices by introducing a medium flow to perifuse the slices. (Use of the word “perifuse” rather than “perfuse” in the context of slices indicates that medium is assumed to flow around the slices rather than through them. Slices consist of tissue that is several cell layers thick; transport of fluids through these layers is known to take place primarily by diffusion, 7 making the assumption correct.) The presence of a constant flow over the tissue has also been suggested to be more consistent with the in vivo situation. 8 –10 We therefore recently developed a new in vitro system based on microfluidic technologies for the incorporation of precision-cut tissue slices. 11 In this system, rat liver slices remain viable up to 24 h, and metabolic activity is retained. The new microfluidic system for incubation of PCLS is compared with the more conventional well-plate system in Figure 1.
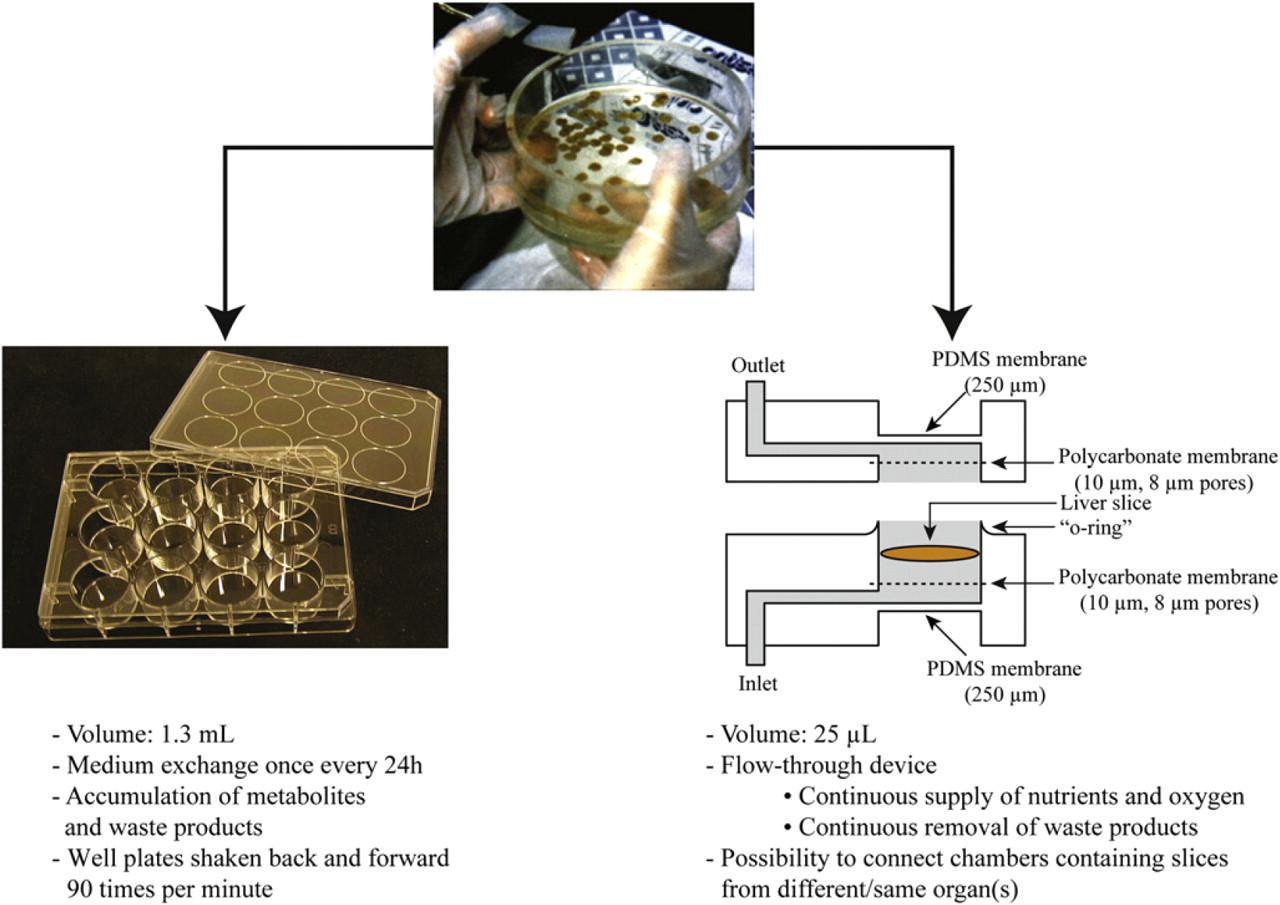
Illustration of the two tissue-slice incubation systems, the well plate and the microfluidic device, for performing metabolism and toxicity studies.
Development of the microfluidics-based system has made sophisticated experimentation possible. It enables on-line analysis of metabolites, since metabolite concentrations in the outflow of medium are high because of the low volume of the chamber and the low flow rate used. Interorgan interactions studies are made possible by the continuous flow of drug-containing medium through interconnected chambers. Two chambers, each containing a tissue slice from a different organ, can be sequentially perfused. How products produced by the first organ slice affect the second slice can be assessed in this way.
In the study reported here, the incorporation of human material in the microfluidic system is demonstrated for the first time. Although metabolism studies have been successfully performed with rat liver slices, 11 it is not a given that human liver slices will also be metabolically active in the microfluidic device because of the interspecies differences that exist. This study thus set out to verify that human tissue could also be tested in the microfluidic system, using well-plate experiments as controls, because these serve as an excellent benchmark. To assess the viability of the slices, the leakage of liver-specific enzymes was measured. Because the main purpose of this system is to perform metabolism studies, the ability of human material to convert substrates into metabolites was also determined. 7-Ethoxycoumarin (7-EC), 7-hydroxycoumarin (7-HC), testosterone (TT), and lidocaine (LI) were chosen as substrates because they are involved in various phase I (hydroxylation, oxidation, N-deethylation, and O-deethylation) and phase II (glucuronidation and sulfation) metabolic routes. 12,13 7-HC is a phase I metabolite of 7-EC, which is further converted into two phase II products, 7-hydroxycoumarin glucuronide (7-HC-G) and 7-hydroxycoumarin sulfate (7-HC-S). LI is converted by the liver mainly into monoethylglycinexylidide (MEGX). Finally, TT is converted into several hydroxytestosterone metabolites and into androstenedione (TT-A), which is the major metabolite of TT. In this study, only the metabolites 6β-hydroxytestosterone, 2β-hydroxytestosterone, and TT-A were measured.
Microfluidic Device
The microfluidic device is made of poly(dimethylsiloxane) (PDMS) and contains a 25-μL microchamber into which organ slices, such as PCLS, can be incorporated (Fig. 1). The fabrication process and verification of this device for metabolism studies with rat liver slices have been elaborated elsewhere. 11 In short, two porous polycarbonate membranes formed the floor and ceiling of the PDMS microchamber to create a well-characterized vertical flow of medium from the inlet around the tissue slice and up to the outlet. Two PDMS membranes serve as aeration windows for the exchange of oxygen and carbon dioxide into the chamber when the device is placed in a humidified plastic container filled with carbogen gas to maintain an optimal incubation environment.
The chamber is continuously perfused with fresh medium to ensure the stability of the slice environment during incubations. The system has been verified for rat liver slices with respect to the optimized conventional well-plate system in which slices retain their in vivo metabolic rate for at least 8 h. 14 Recently, we showed that rat liver slices embedded in hydrogel can retain their viability for 72 h when incubated in the microfluidic device. 15 The hydrogel used was Matrigel that prevented the slices from attaching to the upper filter, which occurred during long incubations. The slices completely retained their metabolic phase II activity for 72 h; however, phase I metabolism did decrease over time. This is probably because of the lack of endogenous compounds in the medium in which in vivo may be responsible for the continuous expression of the enzymes involved.
On-line Analysis System for Metabolism Studies
In initial experiments, medium fractions were collected at the outlet of the microfluidic device and analyzed off-line in a manner analogous to the procedure used for analysis in studies using conventional well plates. Sampling and analysis done in this way involves a number of manual steps and is inefficient. It is also very difficult, if not impossible, to accurately determine the formation rate of unstable metabolites that decompose during incubation in the time between formation and sampling. For these reasons, the direct coupling of the microfluidic system to an HPLC system was realized to allow for automated on-line monitoring of metabolites. Because of the low volume and flow rate involved, relatively high concentrations of metabolites can be achieved in the continuously flowing medium, enabling sensitive detection. A simplified schematic overview of the on-line analysis system is given in Figure 2 A. Medium is delivered to the tissue slices by the peristaltic pump at a flow rate of 10 μL/min. In addition, two computer-controlled syringe pumps have been incorporated into the system to deliver substrate at various concentrations or to test for drug–drug interactions (total flow rate of the two syringe pumps was 100 nL/min). The medium and substrate are mixed just in front of the chip entrance before introduction to the microchamber where the tissue slice is exposed to the substrate. The outlet of the chamber is connected to an injection valve equipped with a 75-μL injection loop. Once the loop is filled with medium containing substrates and metabolites from the chamber, its contents are injected onto the HPLC column by switching the injection valve to bring the injection loop in-line with the HPLC system. Three chambers per experiment are used in conjunction with three injection valves that are switched sequentially. Use of an automated system to switch between syringe pumps and injection valves means that much of the original manual labor associated with this type of analysis is circumvented. A detailed description and verification of the on-line analysis system is described elsewere. 16
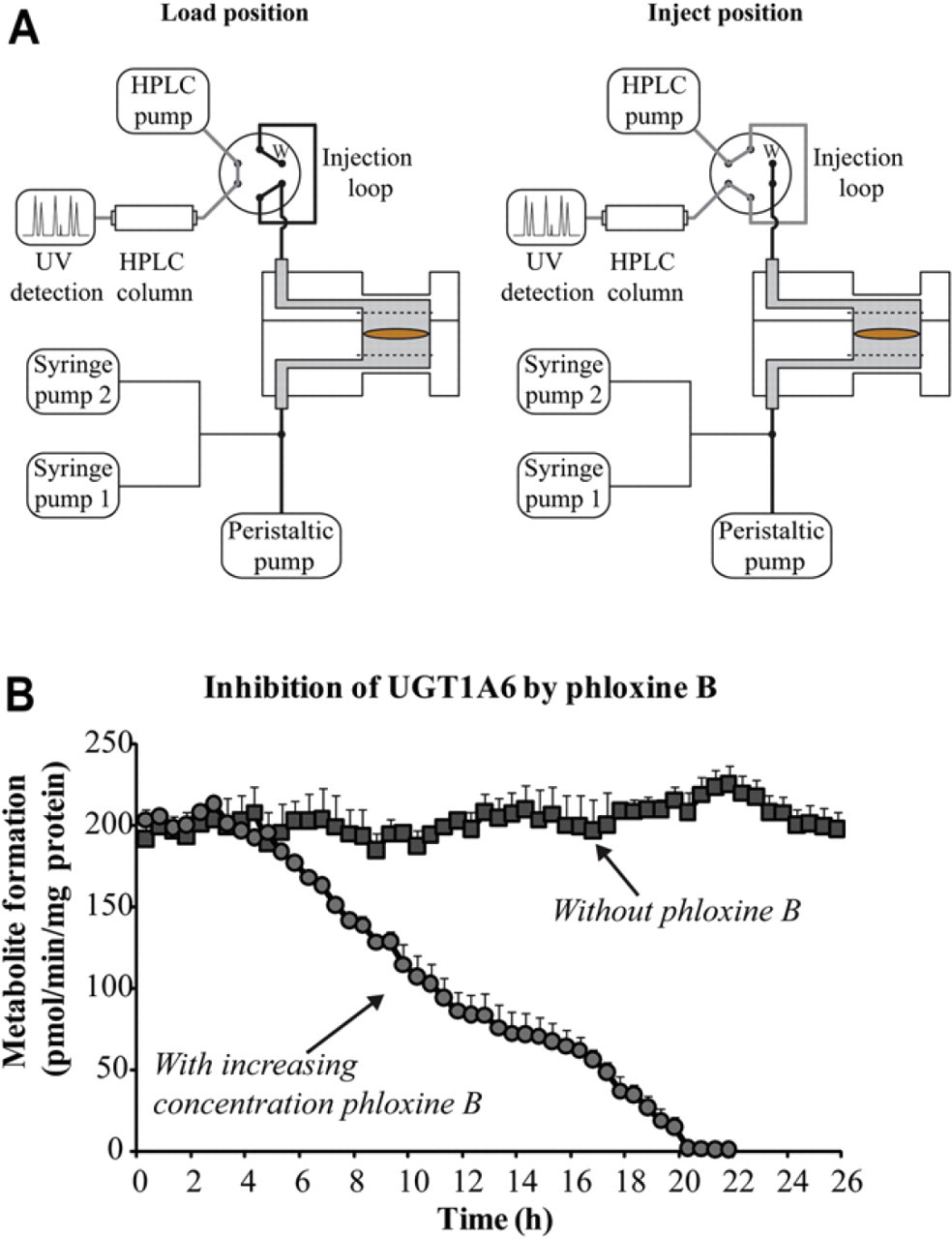
(A) Simplified schematic overview of the on-line analysis system for metabolism and inhibition studies; w = waste. (B) Results of an on-line inhibition study in which the concentration of the inhibitor, phloxine B, was varied to study its effect on the conversion of 100-μM 7-HC into 7-HC-G by the enzyme, UGT1A6. Reprinted with permission from van Midwoud et al. 16 Copyright 2011 American Chemical Society Publications.
With this fully automated system, it was possible to determine the formation rate of the acyl glucuronide metabolite of diclofenac, which is thought to be responsible for serious hepatotoxicity. Diclofenac acyl glucuronide is, like many other acyl glucuronides, chemically unstable, 16 which makes it extremely difficult to detect in conventional systems because of chemical decomposition during incubation. Testing of drug–drug interactions, such as the determination of IC50 values for inhibition of enzyme activity (concentration in which 50% of the enzyme is inhibited), has also been demonstrated with the on-line analysis system. Inhibition of UGT1A6, the enzyme responsible for conversion of 7-HC, by phloxine B was the selected case study. Syringe pumps were used to increase the concentration of phloxine B in the medium every 2 h in steps of 20 μM from 0 to 200 μM, and the metabolism of 7-HC into 7-HC-G by the enzyme UGT1A6 was measured. The metabolite formation remained unaffected until a concentration of 40-μM phloxine B was reached (at 4 h, Fig. 2B). Thereafter, a gradual decrease in metabolite formation occurred until hardly any metabolites were formed at 200-μM phloxine B (at 20 h, Fig. 2B). An IC50 value of 81 ± 5 μM was calculated from these data, obtained using only three slices, as compared with 24 slices in the conventional well-plate equivalent experiment. Therefore, this system has great potential for the testing of scarce tissue such as human material.
Interorgan Interactions
The flow in the developed microfluidic device facilitates not only on-line analysis but also enables the in vitro assessment of interorgan effects through sequential perfusion of microchambers connected in series and containing different organ slices. To study interorgan interactions in an in vitro study remains a great challenge in preclinical testing, although it is known that these interactions do occur in the human body. In our study, the influence of the intestine on the production of bile acids in the liver was studied as an example. The physiological situation was mimicked by coupling the outlet of a microchamber containing a rat intestinal slice to the inlet of a microchamber containing a rat liver slice (Fig. 3 A). The interplay between intestinal and liver slices was then assessed by introducing medium containing the primary bile acid, chenodeoxycholic acid (CDCA). On exposure to this bile acid, the intestine produces growth factor FGF15, a protein known to cause downregulation of the enzyme, cytochrome P450 (CYP) 7A1, which is responsible for bile acid synthesis in the liver. 17,18 This regulation was reproduced in vitro using our microfluidic system. Intestinal and liver slices were sequentially perifused with 50-μM CDCA, and the expression of CYP7A1 in the liver slices was assessed using real-time PCR (Fig. 3B). 19 Single liver slices were incubated with 50-μM CDCA for comparison.
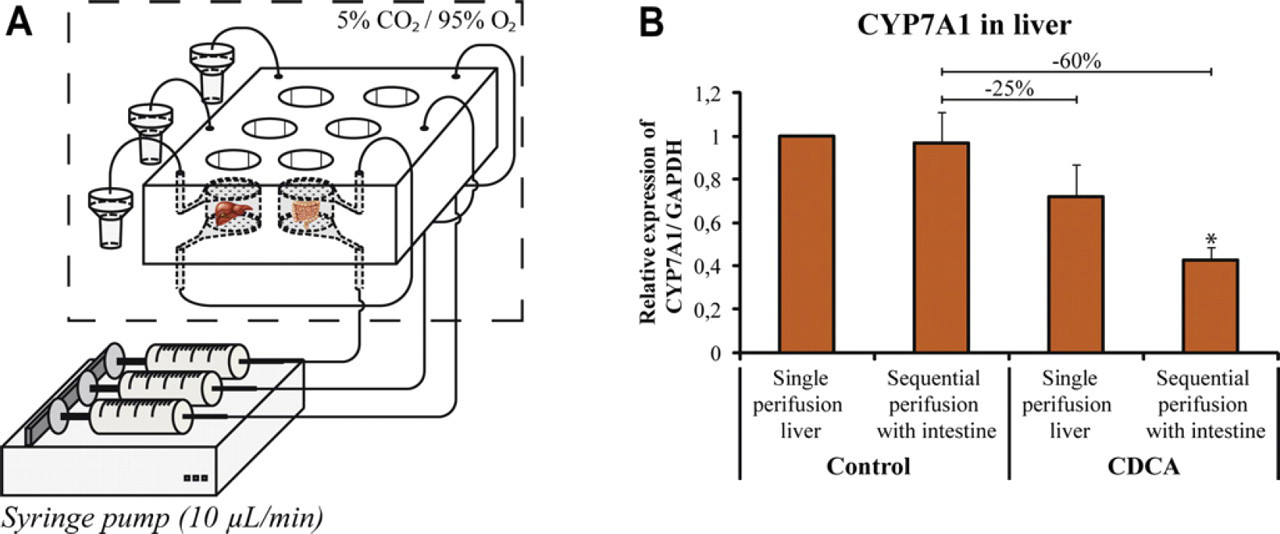
(A) Schematic view of the sequential perifusion of two tissue slices to assess interorgan interactions between intestinal and liver slices. (B) Effect of perifusing a liver slice with medium from a CDCA-treated intestinal slice on the CYP7A1 expression in the liver. GAPDH, glyceraldehyde 3-phosphate dehydrogenase, the housekeeping gene. Reprinted with permission from van Midwoud et al. 19 Copyright 2010 Royal Society of Chemistry.
The expression of CYP7A1 was not changed when an intestinal slice was perifused in front of a liver slice in the absence of CDCA (control). This indicates that under control conditions, the intestinal slice does not produce products that affect CYP7A1 expression in the liver. By introducing CDCA, the CYP7A1 expression in a singly perifused liver slice decreased (∼25%), a well-known effect of bile acids. 17 However, by inserting an intestinal slice in front of the liver and perifusing both slices sequentially with CDCA, a further decrease of 60% was observed. This is most probably because of the excretion of the FGF15 protein by the intestine, as increased expression of FGF15 messenger RNA was also measured in the intestinal slice.
By using our microfluidic system, we were able to show this interorgan interaction with in vitro testing. In vivo testing of this interaction is possible in animals; however, in vivo testing of such interorgan interactions in humans is virtually impossible. With the incorporation of human material in our device, we will be able to determine such human interorgan interactions in vitro.
Human Liver Slices
The studies with mammalian slices described above have shown that the use of microfluidics makes more sophisticated experimentation possible when compared with conventional well-plate formats. However, it is not a given that human liver slices will also function optimally in the microfluidic device. It was thus imperative that studies assessing the viability and metabolic activity of human liver slices in the microfluidic device were undertaken. The slices were produced basically as described previously, 11 the only difference being the use of human tissue instead of rat tissue. Pieces of human liver tissue were obtained from redundant donor tissue as surgical waste remaining after split-liver transplantation, in accordance with the procedures approved by the Medical Ethics Committee of the University Medical Center in Groningen, The Netherlands. The pieces of human liver were stored in ice-cold Viaspan organ preservation solution (University of Wisconsin solution) until slices were prepared from this tissue. 20 The microfluidic incubation system used for the perifusion of human liver slices is similar to the microfluidic system presented previously, with fraction collection performed off-line. 11 Slice viability was determined by monitoring the leakage of the enzymes, lactate dehydrogenase (LDH), alanine aminotransferase (ALAT), and aspartate aminotransferase (ASAT). When cells are damaged, their contents are released into the medium and can easily be measured. Analysis was performed according to a routine laboratory procedure at the Clinical Chemistry Laboratory, University Medical Center of Groningen, using the Roche/Hitachi Modular System (Roche, Mannheim, Germany). Metabolic activity was assessed by incubating the slices with 7-EC (500 μM), 7-HC (500 μM), LI (5 mM), or TT (250 μM) and comparing the resulting metabolite formation with slices incubated in well plates as control, as described previously. 11,19
Viability of Human Liver Slices
The leakage of the enzymes LDH, ALAT, and ASAT is generally considered to be a good measure of liver slice viability. Although ALAT and LDH are present in the cytosol of hepatocytes, ASAT is present in the mitochondria. 21 –23 The leakage of these enzymes as a percentage of the total amount measured over time was very comparable in well plates and microfluidic device, as shown in Figure 4. After 4 h, only ∼5% of the enzyme had leaked out of the tissue slices for all three enzymes and in both incubation systems. This means that 95% of cells remained viable after 4 h. Even after an incubation period of 24 h, the cumulative leakage of enzymes was still low. Less than 10% of the total ALAT had leaked out in both well plates and microfluidic device, whereas about 15% leakage of ASAT in both well plates and microfluidic device had occurred. About 10% LDH had leaked from the slices in the microfluidic system, whereas the percentage leakage appeared higher in well plates (15%); however, this difference was not significant. According to these data, about 85% of cells present in the human slices were still intact after 24 h of incubation, so that it may be concluded that liver slices remain viable in both systems. It was demonstrated previously that human liver slices maintained their viability when incubated for 24 h in well plates. 24 These experiments have verified that liver slices also remain viable when incubated in the microfluidic system under continuous-flow conditions.
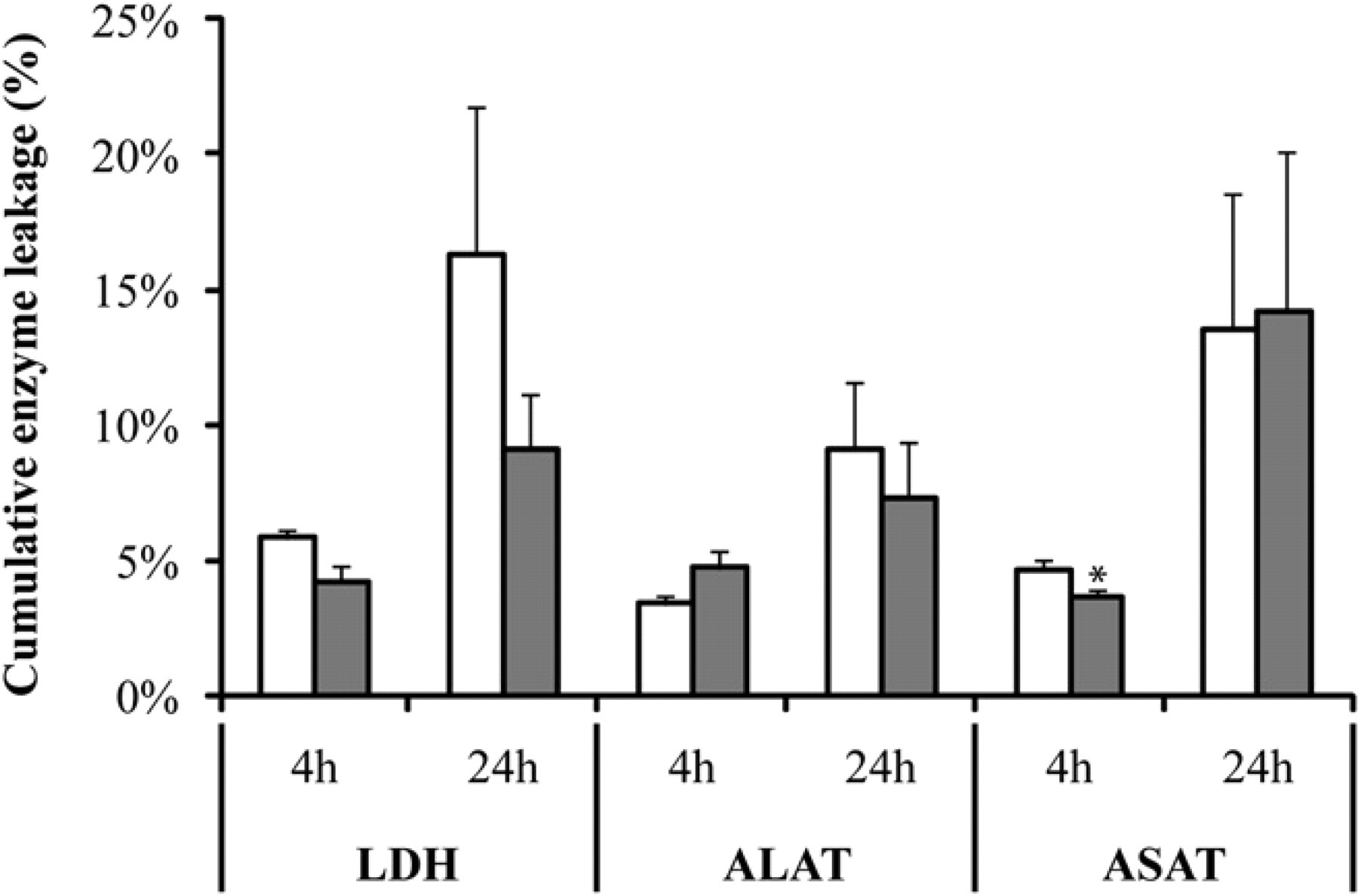
Cumulative enzyme leakage of LDH, ALAT, and ASAT from human liver slices after 4- and 24 h incubation of human liver slices in well plates (white bars) and biochip (gray bars), as a percentage of the total amount of each enzyme present in the slices. Results are mean ± standard error of the mean of three organs with four slices per incubation system. Significant differences with respect to well plates are indicated with an asterisk (P < 0.05).
Drug Metabolism in Human Liver Slices
As expected, large differences in metabolic rates were observed between livers from different patients, which is probably due to different diets and medications. Because rat liver slices incubated in well plates exhibit a good correlation to the rat situation in vivo, 14,25 results obtained for human slices in the microfluidic device were compared with those obtained for human slices from the same human livers in well plates. Figure 5 presents the metabolism of four substrates in fresh human tissue slices in both incubation systems during 2 h for TT and LI and 3 h for 7-EC and 7-HC.
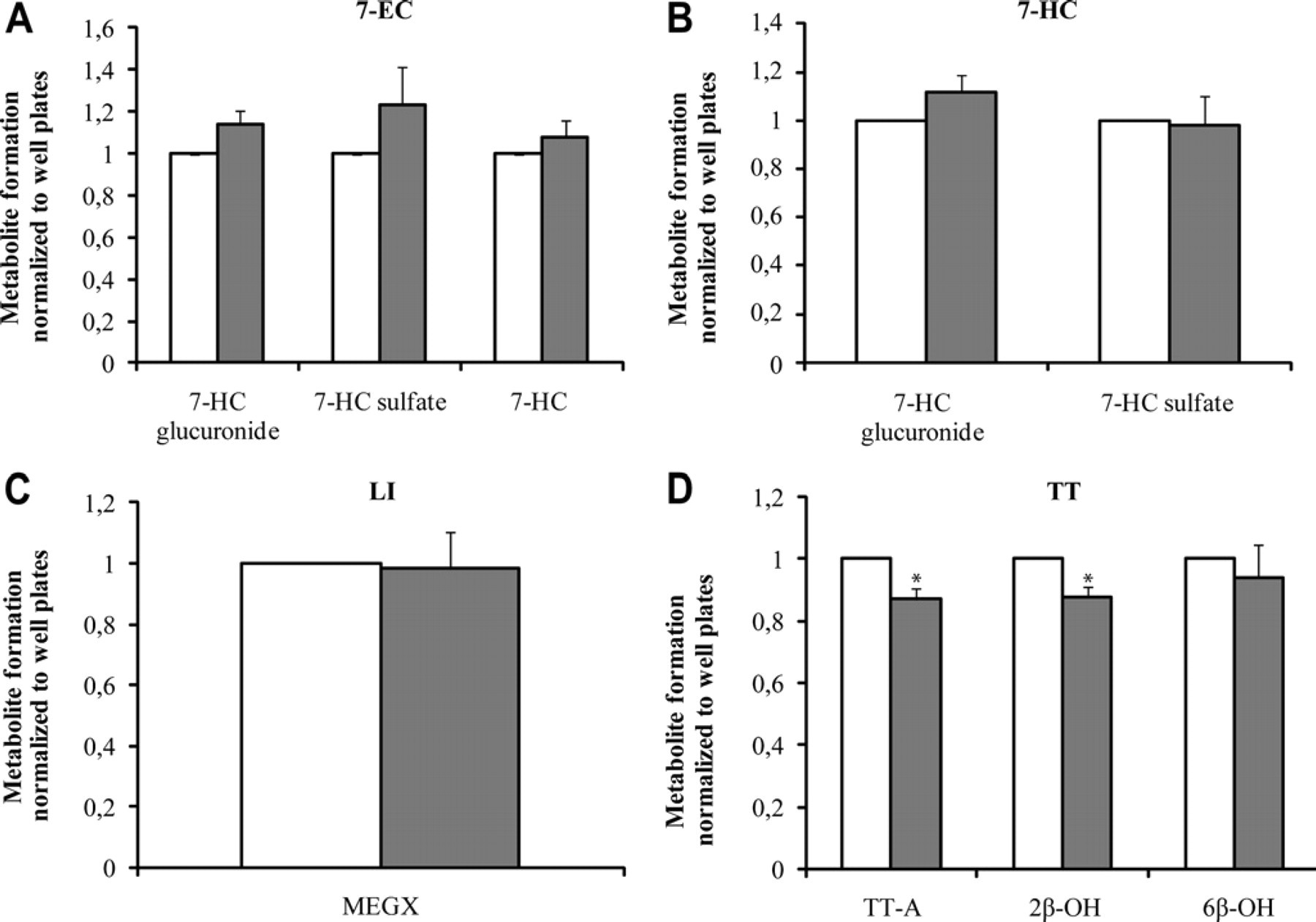
Metabolism of (A) 500-μM 7-EC (3 h), (B) 500-μM 7-HC (3 h), (C) 5-mM LI (2 h), and (D) 250-μM TT (2 h) in fresh human liver slices incubated in well plates (white bars) and in the microfluidic system (gray bars). Results for each liver (three slices per liver for each incubation system) are expressed relative to the activity in well plates as mean of three organs ± standard error of the mean. Significant differences with respect to the well plates are indicated with an asterisk (P < 0.05). 6β-OH = 6β-hydroxytestosterone; 2β-OH = 2β-hydroxytestosterone.
Human liver slices were able to form both phase I and phase II metabolites from 7-EC (Fig. 5A), as shown previously. 24 These results indicate the activity of CYP, sulfotransferase, and UDP-glucuronosyltransferase in human liver slices. Although the metabolite formation in the microfluidic device appeared to be slightly higher for all three metabolites than in the well plates, the differences were not significant. The metabolism of 500-μM 7-HC into 7-HC-G also appeared to be slightly higher in the microfluidic device than in the well plates (Fig. 5B); but again, this difference proved to be insignificant. Sulfation rates were very similar. The formation of MEGX from LI (Fig. 5C) demonstrated the capacity of the human liver for N-deethylation of the substrate and was very comparable both in well plates and microfluidic device for all three livers tested. Only in the case of TT, metabolite formation was a significant difference between well plates and microfluidic device observed (Fig. 5D). The measured formation rates of TT-A and 2β-hydroxytestosterone were significantly higher in the well plates. These are also the two most hydrophobic metabolites formed from TT and are more hydrophobic than any of the other metabolites studied for 7-HC, 7-EC, and LI. One of the well-known disadvantages of using PDMS as a substrate material in microfluidics is its tendency to absorb and adsorb hydrophobic compounds. 8,26 To verify if adsorption or absorption occurred, medium containing TT metabolites was flushed through a PDMS device that did not contain a liver slice. As expected, 10%–20% of the hydrophobic metabolites appeared to be absorbed in the device, and recoveries of only ∼80% for the most hydrophobic compounds were obtained. This correlates well with the lower amounts of metabolites observed in the microfluidic devices than in the well plates. Therefore it was concluded that the metabolite formation for TT was similar in well plates and microfluidic devices, but that 10%–20% of the metabolites formed were adsorbed or absorbed onto or into the PDMS, respectively. The hydrophobicity of PDMS limits the applicability of the microfluidic device somewhat and dictates that the possible adsorption or absorption of substrates and metabolites be taken into account when using this system.
To investigate the metabolic competence of slices after 24 h of incubation in well plates and the microfluidic system, the metabolism of 7-EC (phase I) and 7-HC (phase II) (Fig. 6 A and B, respectively) was determined after this time. The results are given in Figure 6 after normalization with respect to data obtained for fresh slices in well plates. After 24 h, the human slices were still able to form all three metabolites from 7-EC (Fig. 6A). However, the metabolite formation rates were only 35%–40% of the formation rates measured for fresh slices. This decrease in metabolism is probably due to the decrease in CYP activity over time, as has been described frequently. 7,27,28 The loss in CYP activity does not affect every isoenzyme equally. However, VandenBranden et al. 27 found a decrease in coumarin 7-hydroxylase activity (CYP2A6) of 60%–90% after 24 h in slices stemming from three human livers. This is in-line with our results, which reveal a decrease of 60%–65% in 24 h. The reduction in CYP enzyme activity is not limited to slices, as primary hepatocytes also exhibit this drop in activity during culture. 29 When metabolism in well plates and microfluidic device are compared after 24 h of incubation, no significant difference can be observed. Therefore, it can be concluded that the lower metabolic rates are not specifically caused by the incubation system but are a more general phenomenon exhibited by the tissue in this culture medium. Edwards et al. 30 showed that this decrease can be minimized by adding the proper medium supplements, such as the inducer, dexamethasone, to the medium. In contrast, phase II metabolism is well maintained in human slices after 24 h, at levels that are not significantly different from fresh slices in both well plates and microfluidic devices (Fig. 6B). This also indicates that human liver slices are viable for 24 h, independent of the incubation system. Because the conversion of 7-HC into 7-HC-G and 7-HC-S was comparable after 24 h of incubation with that observed in fresh slices in well plates, the decrease in 7-EC metabolism after 24 h is likely to be due to lower CYP activity.
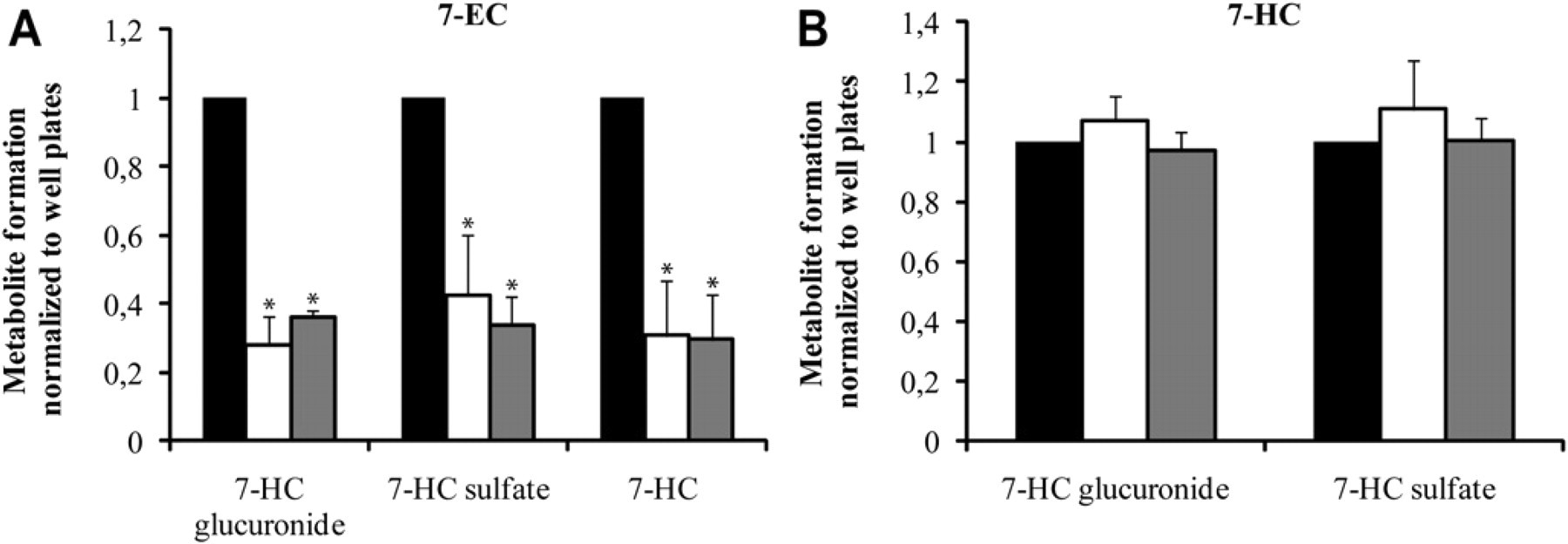
Phase I and phase II metabolism in human liver slices after 24 h of incubation in well plates (white bars) and microfluidic system (gray bars). (A) 7-EC was used to assess phase I metabolism and (B) 7-HC to assess phase II metabolism. Results are the mean of three organs ± standard error of the mean, with three slices per incubation system per experiment. Results are expressed relative to fresh slices in well plates (black bars). Significant differences with respect to fresh slices incubated in well plates are indicated with an asterisk (P < 0.05). No significant differences were observed between well plates and microfluidic system after 24 h of incubation.
Conclusion
Previously, we developed a microfluidic-based incubation system for the perifusion of PCLS. 11 This system was verified with respect to the well-plate system for the incubation of rat liver slices. However, the ultimate goal is to incubate human tissue to predict the metabolic pathway of xenobiotics without the need to sacrifice animals and with better extrapolation to the in vivo situation in humans. The microfluidic-based perifusion system was therefore also tested with human tissue in this study. We have shown that PCLS from human tissue remain viable in the microfluidic system while maintaining metabolic function. Slice viability was tested after 24 h of incubation by analyzing for leakage of liver-specific enzymes. The results showed that this leakage was low, indicating viable slices. The metabolic function was also maintained in the microfluidic device for up to 24 h at a level similar to the well-plate system, the benchmark for these kinds of experiments, as revealed by tests with four different model substrates (7-EC, 7-HC, LI, and TT). It can therefore be concluded that the microfluidic system is appropriate for metabolism studies using both rat and human liver slices. However, compared with conventional well plates, the use of microfluidics makes more sophisticated experimentation possible. An on-line analysis system has been developed, which makes it possible to measure unstable metabolites. Moreover, it enables easy variation of substrate concentrations over time in an automated fashion during an experiment. Experimentation with scarce tissue can be made more efficient using this approach. Another advantage is the ability to create a constant environment for the slices, both in terms of removing metabolites and waste products and in terms of flow. The effect of flow on the metabolic pathways of xenobiotics can be investigated, thereby mimicking blood flow in vitro. Finally, it is possible to mimic first-pass metabolism with this system by exploiting the continuous-flow feature. In well plates, this type of experiment involves placing two slices in one well; however, mutual interaction between slices can take place, and it is never clear which organ is responsible for any observed effect. In the microfluidic device, the two organ slices are compartmentalized, much as they are in the body, making it far easier to characterize interorgan interactions. The preservation of human organ tissue is crucial for performing these interorgan interaction studies with human tissue, as the probability of receiving surgical waste from two different organs on the same day is rather low. Future experiments are planned to study drug-induced toxicity in the microfluidic system.
In conclusion, this system has the potential to contribute to drug metabolism and toxicology studies of novel chemical entities. Although it will not fully replace the incubation of PCLS in well plates, the microfluidic system will provide important supplementary information at an early stage of drug development.
Footnotes
Acknowledgment
The work presented in this Feature Story has been supported in part by a grant from ZonMw. The authors thank Professor Dr. R. Porte and his colleagues of the Department of Hepatobiliary Surgery and Liver Transplantation, University Medical Center Groningen, The Netherlands, for providing the human tissue.
Competing Interests Statement: The authors certify that they have no relevant financial interests in this manuscript.
