Abstract
High throughput robotic systems have been used since the 1990s to carry out biochemical assays in microtiter plates. However, before the application of such systems in industrial fermentation process development, some important specific demands should be taken into account. These are sufficient oxygen supply, optimal growth temperature, minimized sample evaporation, avoidance of contaminations, and simple but reliable process monitoring. A fully automated solution where all these aspects have been taken into account is presented. It is basically an arrangement of connected apparatuses (rail-mounted robotic arm, liquid handlers, sealer, shakers, fluorescence reader, etc.) that are integrated into a climatic chamber. Its readiness for use is demonstrated for the cultivation of Saccharomyces cerevisiae. This automation system permits to prepare, run, and monitor 768 aerobic micro-scale fermentations in parallel and without the need for any manual intervention. Although the applied quasi-continuous culture fluorescence measurement technique repeatedly requires a short interruption of the shaking process, this does not have a significant influence on the performance of cell culture experiments.
Keywords
Introduction
Modern biotechnological methods enable the rapid construction of mutant libraries containing thousands of genetic variants. Hence, the labor-intensive manual screening in shake flasks is no longer sufficient for such a huge number of samples, because one full-time employee is capable of performing only about 20 high-value experiments in parallel. In addition, it is generally known that automated processes can be superior to manual procedures regarding reliability, reproducibility, and measurement density. Microtiter plates recently have been considered as alternative shaken fermentation vessels, because these arrays of micro-scale reactors are easily handled by laboratory automation equipment.1,2 Such robotic systems have been used in the chemical and pharmaceutical industry for biochemical assay applications since the 1990s. The fermentation of microorganisms, however, requires some modifications to standard laboratory automation solutions to enable the aerobic cultivation of cells under appropriate biochemical engineering parameters: a high oxygen transfer rate is an important parameter during the development of aerobic biochemical processes, the incubation temperature has to be at the optimum of the cultivated species, the sample evaporation rate has to be minimized, airborne and cross-contaminations have to be avoided, and a noninvasive process monitoring technique is desirable. An implemented robotic system where all these aspects have been considered is shown in Figure 1.
Photograph of the fully automated high throughput fermentation system. A custom-arranged screening robot is placed inside an enclosure (length 4.9 m, width 2.3 m, height 2.2 m) providing temperature and humidity control. The centerpiece is a rail-mounted robotic arm (Sagian ORCA, Beckman-Coulter, Fullerton, CA) that transports the microtiter plates between the following apparatuses: barcode reader, lidding station, plate sealer, liquid handlers, shakers, optical readers, centrifuge, and deep freezer.
Robotic System
The solution is based on a Sagian Core System and the appropriate SAMI 3.3 software including a static scheduler (Beckman-Coulter, Fullerton, CA). Hardware from other vendors was integrated with custom-programmed drivers (University of Rostock, Germany). The system has been developed with a focus on the cultivation of Escherichia coli. 3 Nevertheless, it is also applicable for other organisms such as yeasts. Figure 2 shows a process flow scheme of a fully automated high throughput fermentation including preparation, cultivation, and monitoring of up to 768 parallel experiments. With a liquid handler, sixteen 48-well microtiter plates (nontreated cell culture plates; BD Biosciences Discovery Labware, Bedford, MA) are filled with 400 μL medium per well and inoculated from precultures. Next, the plates are sealed with two layers of a gas-permeable adhesive tape made of nonwoven rayon fibers (prototype, HJ Bioanalytik, Monchengladbach, Germany). This material does not reduce oxygen supply but reduces sample evaporation 4 and serves to avoid both airborne and well-to-well cross-contaminations. 3 Finally, the fermentation is started by shaking the plates at 1100 rpm on 16 orbital shaking machines (3 mm shaking diameter). These parameters were carefully chosen according to detailed biochemical engineering studies 5 and enable a high oxygen transfer rate of approximately 60 mmol L−1 h−1 The shakers had been custom modified with a cooling device before integration (Analytical Instrument Group, Rostock, Germany) to avoid additional heating of the microplates, and therefore, sample evaporation is significantly reduced during operation. 3 After defined time intervals (e.g., once per hour), the plates are transferred one after another to a fluorescence reader to monitor the fermentation processes by exciting the cells with UV light (370 nm) and measuring the nicotinamide adenine dinucleotide phosphate (NADPH)-dependent autofluorescence (460 nm) through the bottom of the transparent polystyrene microplate. This method has been described elsewhere in detail for E. coli, 6 but it can be adopted in certain cases for other organisms without any changes (e.g., for yeasts, as shown below). After the plate is read, it is immediately transferred back to its shaker. After several shaking and reading cycles (the exact number depends on the process under investigation), the process ends and the fermentation broths may optionally be investigated further by high throughput biochemical assays or may undergo an automatic sample preparation procedure (e.g., centrifugation, filtration, and freezing) for later offline analytics (e.g., chromatography or mass spectrometry).
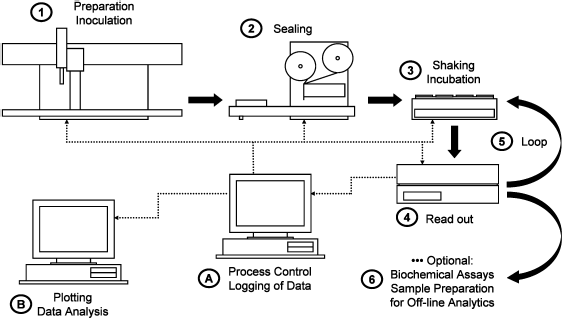
Process flow scheme of high throughput fermentation. (I) A liquid handler (Biomek 2000; Beckman-Coulter, Fullerton, CA) puts medium into the 48-well plates and inoculates the samples from precultures. (2) The plates are sealed with gas-permeable adhesive tape by a microplate sealer (RoboSeal; HJ Bioanalytik, Mönchengladbach, Germany). (3) Each of the 16 plates is shaken on its own shaker (H+P Labortechnik, Oberschleißheim, Germany). (4) After a certain period (e.g., every 60 min), the plates are read in a fluorescence reader (Fluostar; BMG Labtech, Offenburg, Germany). (5) The shaking and reading procedure is looped several times until the end of the processes. (6) Optionally, at the end of fermentation, the samples may be further analyzed by biochemical assays or prepared for additional offline analytics. (A) A process computer controls the whole procedure and logs the data from the fluorescence reader. (B) The data is finally analyzed with scientific graphing software at an office computer.
Experiments and Results
In the representative culture process described here (Fig. 3), Saccharomyces cerevisiae CEN.PK2-1C was cultivated in YPD medium (10 gL−1 yeast extract, 20 gL−1 peptone, 20 g L−1 dextrose; all from Sigma-Aldrich, St. Louis, MO) for 40 h at 30°C and 60% relative humidity. We mixed 99 parts medium with 1 part preculture to get inoculated medium. Four microtiter plates were used, and six wells of each plate were filled to enable six independant experiments in each plate. The shaking of a microtiter plate is interrupted for a period of almost 2 min during the transportation of the plate from the shaker to the reader, the measurement of the culture fluorescence inside the reader, and the transportation back to the shaker. As the complete robotic system is inside a climatic chamber, neither temperature nor humidity is influenced despite this procedure. The oxygen supply of the cells, however, is reduced due to nonshaking for the duration of the measurement procedure. 7 To examine whether this has a lasting influence on the physiology of the cells and on the fermentation characteristics, all four microplates were measured at different time intervals (0.5, 1, 2, and 4 h). The results in Figure 3 show that the different periods have no significant influence on the fluorescence curves during the fermentation. This is in accord with similar results shown elsewhere for E. coli. 3
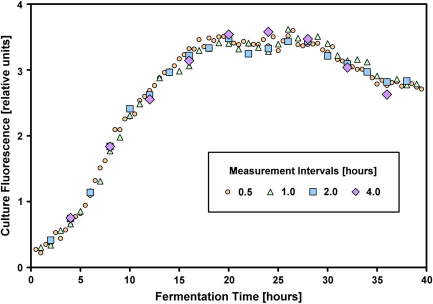
Results of some S. cerevisioe fermentations (details in the text). The NADPH-dependent culture fluorescence is plotted against the process time. Each data point represents the mean value of six independant experiments. The standard deviation was always below 5%.
Conclusion
The laboratory automation solution introduced here is capable of performing 768 parallel micro-scale fermentations under high oxygenation rates. It is fully automated and runs day and night. The quasi-continuous fermentation monitoring technique has little or no impact on the cell cultures, although the shaking of the microtiter plates has to be interrupted for the measurement procedure. The potential applications for this new robotic system are widespread. Among these, the function-driven analysis of metagenomic libraries, 8 the screening for better biocatalysts during directed evolution, 9 and the high throughput cultivation for metabolic flux analyses 10 are very promising examples.
Footnotes
Acknowledgments
This work was supported by the business unit Feed Additives of Degussa AG. The S. cerevisiae strain and preculture were kindly provided by Svenja Däubner, Thomas Hüller, and Daniel Börgel. H. F. Zimmermann wishes to thank Uwe Dingerdissen, Kerstin Thurow, Norbert Stoll, Jochen Büchs, and their coworkers for scientific and engineering support.
