Abstract
NetBookingLIMS is Internet-based software used for the total automation of workflow within core diagnostic or genetics laboratories. NetBookingLIMS organises all data incoming into the laboratory in a flat file and converts it to a relational database that can be used to generate auditing, billing, troubleshooting and usage reports. Also, it generates instrument compatible data and organises the day-to-day management of the laboratory in an efficient and structured manner. It is written using Perl5 and C and designed to run solely using an Internet browser. NetBookingLIMS software as well as a user manual and documentation are available at http://dcs.bbk.ac.uk/∼ahamo01/nblfrontpage.cgi.
INTRODUCTION
Pharmaceutical, commercial and academic scientific laboratories deal with enormous amounts of data. Whether this information is related to process, procedures, experimental design or analytical and clinical data, keeping it safe and organized - yet available to those who need it - is a formidable bioinformatics challenge. A core DNA sequencing and genetic analysis laboratory supports many research teams within an organisation and is responsible for the cloning and characterisation of genes. Usually a genetics laboratory employs many staff managed by one manager, has many instruments, supports many applications such as DNA sequencing, genotyping, fluorescent fingerprinting 1 and fluorescent mutation detection 2 and is used by many users (scientific researchers) with different requirements including high, medium and low throughput usage. Also it is difficult to carry out accurate auditing, billing and troubleshooting with such a complex working model. Thus, within such a laboratory there are many real world issues such as managerial and organisational problems and algorithmically complex problems such as samples scheduling and relational database administration and maintenance.
Several commercial genetics laboratory LIMS (Laboratory Information Management System) already exist, such as BioLIMS (http://www.appliedbiosystems.com/informatics/productpages/bl_body.html). However, they are difficult to integrate and use for supporting a multi-user and multi-tasking laboratory and they are not user friendly. NetBookingLIMS (Inter
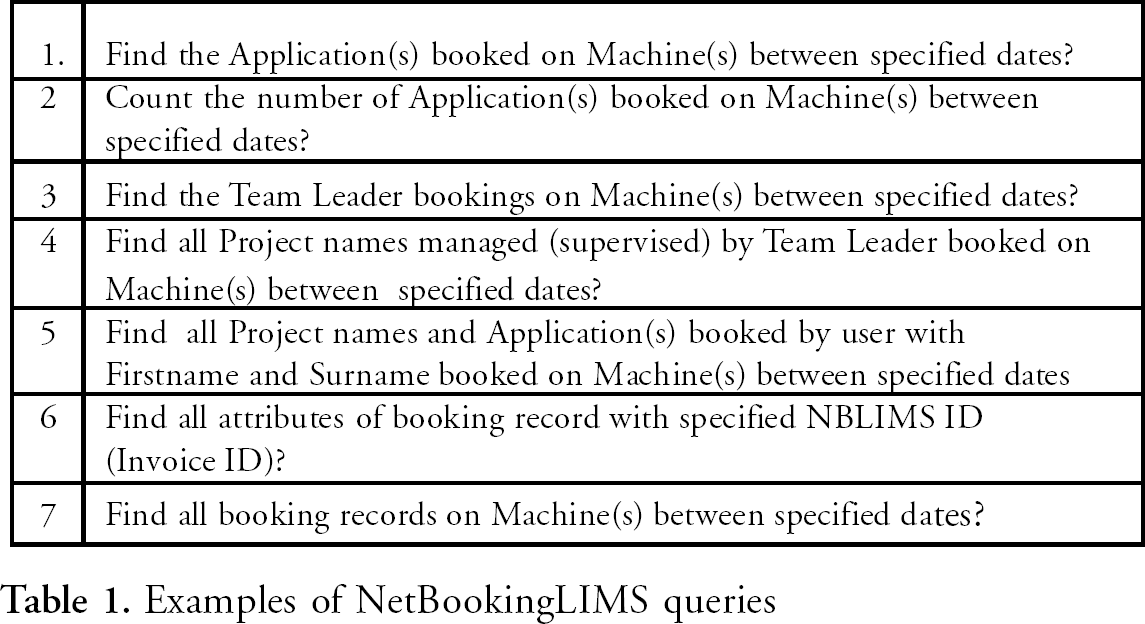
Examples of NetBookingLIMS queries
Invoice ID, Application Description, Instrument ID, Date of Run, User First Name, User Surname, Project Name, Team Leader, Email, Phone Number, Sample Sheet ID, Computer Results Sent to
where Invoice ID is the primary key, Sample Sheet ID is a foreign key that derives the lane number and the corresponding sample name booked on a run and Application Description defines the type of application to execute on the instrument.
NetBookingLIMS is designed using SSADM (Structured System Analysis and Design Methodologies) and implemented as a 3-tiered architecture. Analysis includes a feasibility study which is carried out to determine the structure and relationship between various entities within the laboratory. An entity-relationship model (ERM) is produced as shown in Figure 2.
In order to determine the workflow path and identify problems with the management of the laboratory, a data flow diagram is constructed as shown in Figure 3.
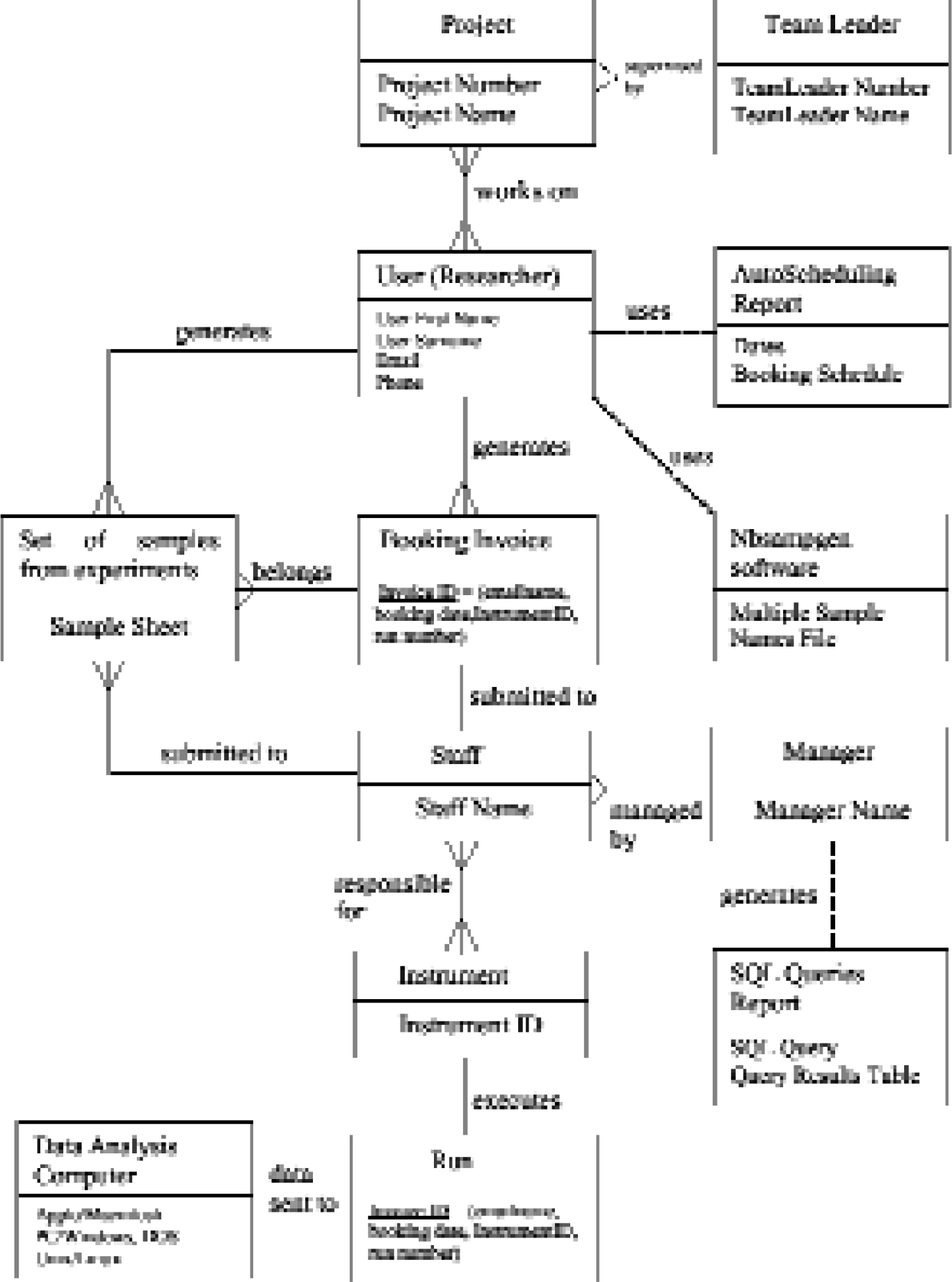
Entity-relationship model of a genetics core laboratory.
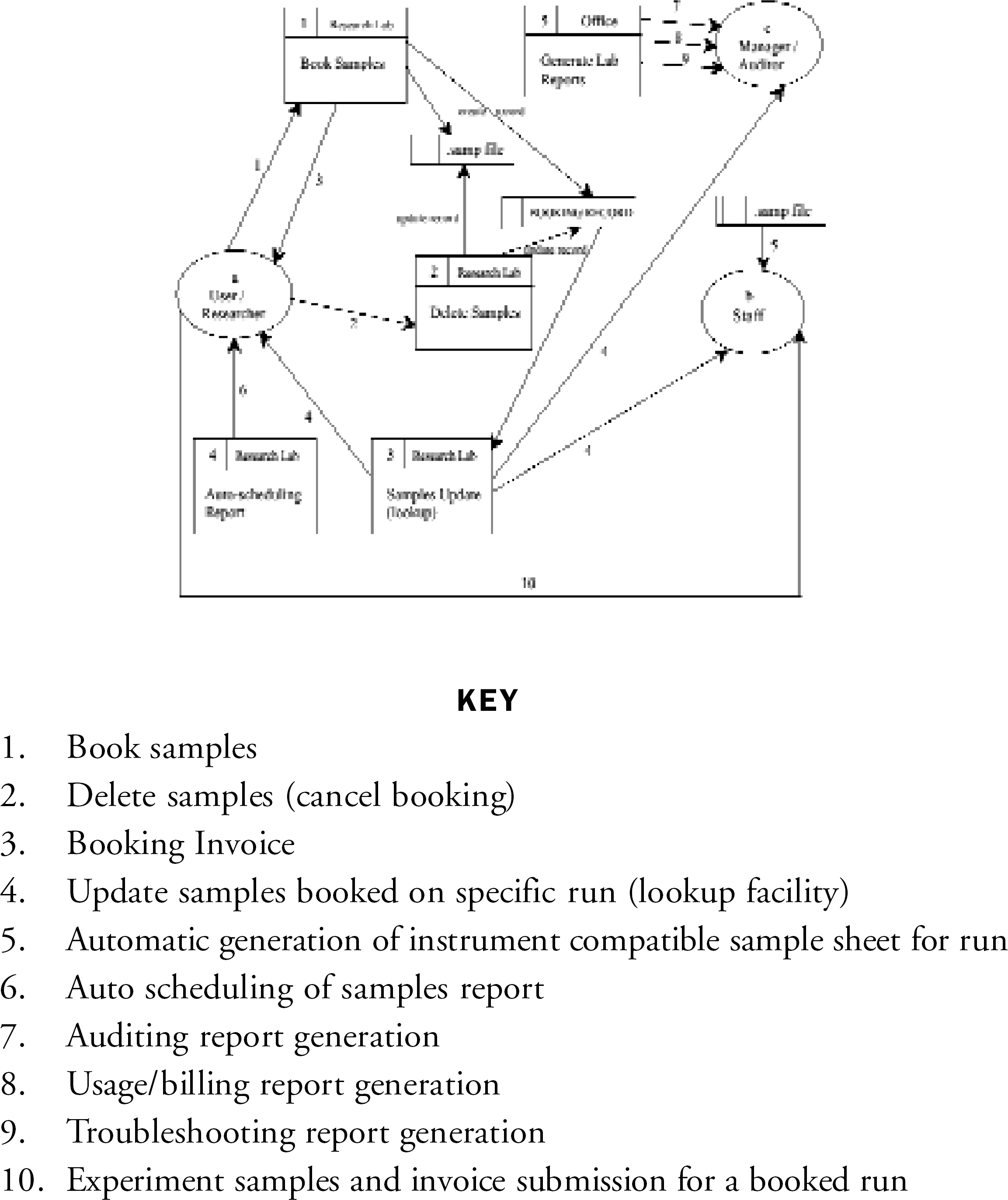
Data flow diagram showing the workflow within a genetics core laboratory.
Using the data flow diagram and entity relationship model as a basis, user requirements and specifications are determined and a 3-tier architectural design is conceived for the software. This consists of a front-end design which includes the data entry forms and the pages displayed to the end user via an Internet browser, a middle layer which is the main engine that takes the data from the front end and processes them according to the specification, and a backend layer which deals with how the data are stored. This includes the centralised ASCII flat file and its link to the relational database that is queried by the manager in order to obtain various types of management and troubleshooting reports. An overview of the architecture of the software is shown in Figure 4. Formal specification of the software was carried out using the formal specification language Z. 4 The main engine is implemented in Perl5 using CGI.pm. This generates the front end in HTML (Hyper Text Markup Language) that is displayed to the end user via an Internet browser and its central ASCII data store can be easily linked to a RDBMS (Relational Database Management System) such as Oracle (http://www.oracle.com), MySQL (http://www.mysql.com) or Sybase (http://www.sybase.com).
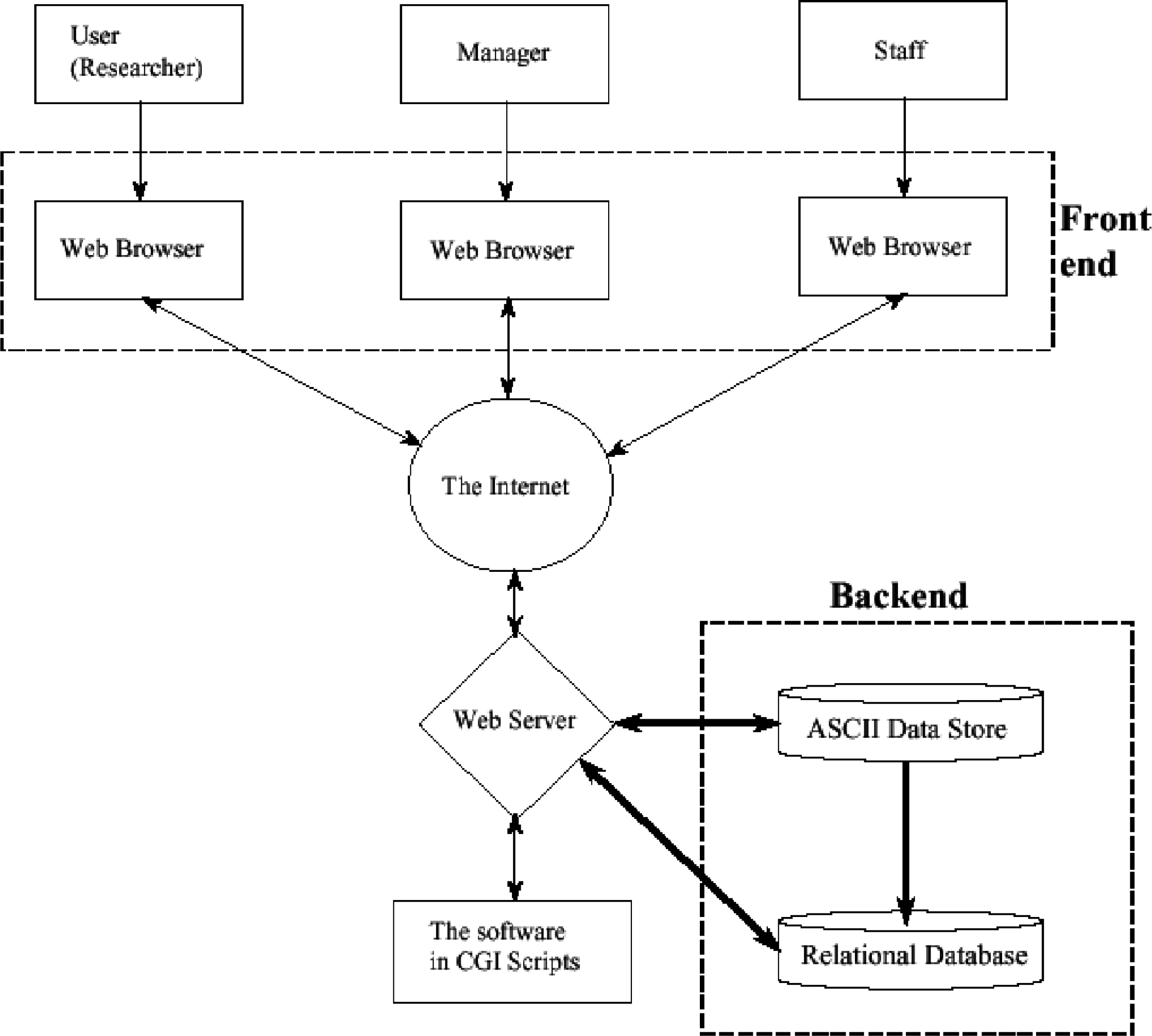
Overview of the architectural design of NetBookingLIMS software.
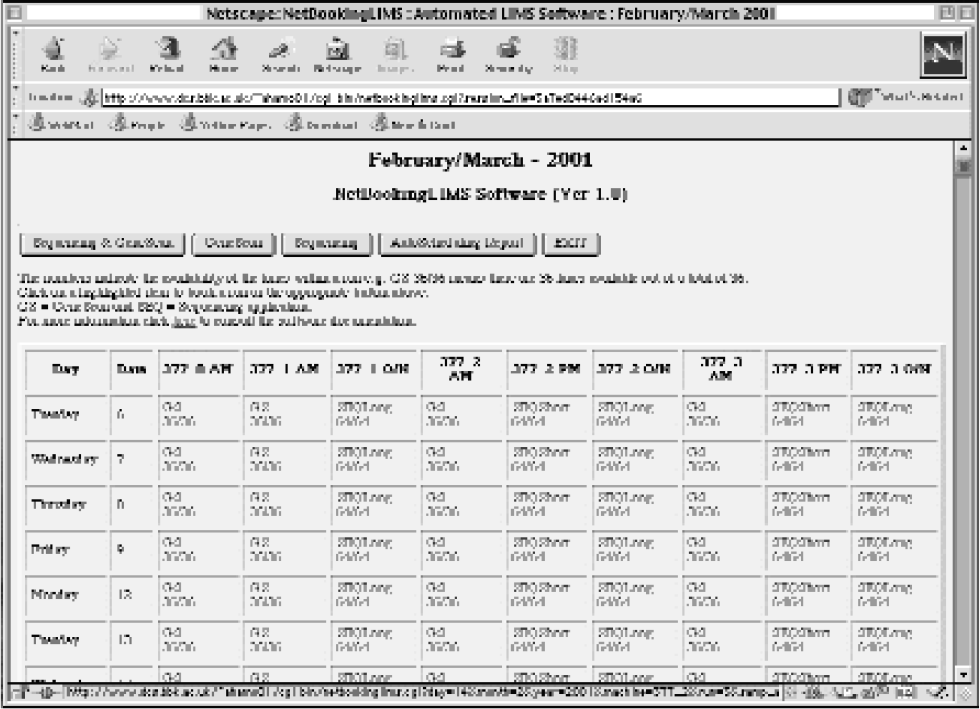
It has a user-friendly interface that automatically generates SQL queries and displays the output in table format together with the SQL query that generated it. The NetBookingLIMS software is designed and implemented to run via Internet browser, thus posing minimal requirements on the end user. Although it is Internet-based, it can be easily ported to run locally on an intranet web server and since it stores the records in a flat file, it can function without the need for RDMBS on the same web server where it is installed. It has been successfully used to automate and manage the workflow in many diagnostic and genetics core laboratories.
ACKNOWLEDGEMENTS
This work was funded by the Institute of Cancer Research supported by the Cancer Research Campaign.
